Abstract
Haemosporidian blood parasites are widely used in evolutionary ecological research when exploring the effects of parasites on different life-history traits of their bird hosts. However, their roles in bird migration are less studied. If these parasites deteriorate the body condition of the birds strongly, they might negatively affect the whole migration phenology and the survival of the birds as well. In our study, we tested the relationships between infection for parasite genera (Haemoproteus or Plasmodium), the three most frequent parasite lineages and body condition (body mass, fat deposit), and the timing of autumn migration in the European Robin (Erithacus rubecula). We found that mean body mass and fat scores did not differ between parasitized and non-parasitized individuals, but infected juveniles arrived later than their non-infected counterparts. The difference in the arrival time of parasitized and non-parasitized birds was greater in the case of Haemoproteus infections. However, when we analysed the effects of the distinct parasite lineages separately, we found that prevalence of parasite lineages correlated with the body mass, fat storage, and timing of autumn migration of the birds in a different direction. Our results therefore emphasize the importance of testing the impacts of the different parasites individually, because possible lineage-specific effects on bird condition during migration might exist.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Avian malaria and avian malaria-like parasites (Plasmodium and Haemoproteus spp.) are of special interest not only to veterinarians but also to ecologists. Because of the potential effects of these parasites on the whole life cycle of the birds, they have been in the focus of many previous studies (reviewed in Marzal 2012). However, it is difficult to explore the effects of avian malaria infections in wild bird populations. First, it is nearly impossible to detect individuals during the acute infection phase due to the inactivity of birds during this phase (Lapointe et al. 2012). So most knowledge on the effects of these parasites come from the chronic phase of infection or from experimental studies (e.g. Knowles et al. 2010; Palinauskas et al. 2018). Though detailed data about the mechanistic effects of these parasites are scarce (discussed later), some studies suggest that parasites negatively affect the physiological condition of the birds even in the chronic phase by destroying red blood cells. This results in reduced oxygen consumption rate and may manifest in slower flight distances of migratory birds (Valkiūnas 2005, but see Hahn et al. 2018). Furthermore, due to the increased oxidative stress, parasite infection may accelerate telomere shortening which ultimately affect the long-term survival of the host (Asghar et al. 2015).
It is important to note that there are more than 6000 lineages within the group of haemosporidians (Bensch et al. 2009) and their virulence potentially differs in different host species (Ilgūnas et al. 2019). Some lineages are host generalist infecting a broader range of host species, while some are host-specific (Waldenström et al. 2002; Dimitrov et al. 2010; Medeiros et al. 2014). Some studies found that generalist lineages reached lower intensities in their hosts than specialists did, because they are less adapted to certain host species but infected closely related host species more often than expected by chance (Hellgren et al. 2009). Generalist parasites might also be the most prevalent parasites in their compatible hosts (Hellgren et al. 2009; Drovetski et al. 2014) because these parasite species have strong immune evasion capabilities (Hellgren et al. 2007).
Different parasite lineages have different infectivity and pathogenicity on their avian hosts, leading also to various level of susceptibility of the hosts to these lineages (Dimitrov et al. 2015). Therefore, a number of studies have highlighted the importance of separately analysing the effects of different parasite lineages. Indeed, Ortego et al. (2008) found that male Lesser Kestrels (Falco naumanni) infected with a certain Plasmodium lineage (P-LK6) had less fledglings in their broods as the non-infected males, while in the Blue Tit (Cyanistes caeruleus), two closely related Plasmodium species (P. relictum and P. circumflexum) altered the mortality and recapture rate differently (Lachish et al. 2011).
Despite the relatively long known differences in the impacts and the transmission ability of these lineages, the exploration of how the effects of the lineages arise mechanistically has started recently. For instance, Hellgren et al. (2015) emphasized the importance of considering the origin of the lineages when studying the transmission and impacts of different avian malaria lineages. This is because they found that allelic variation in a specific parasite locus (MSP1, which is known to have a key role in the invasion of host red blood cells), might be linked to local differences in parasite virulence and host resistance.
Aželytė et al. (2022) found that during co-infection with two closely related lineages (P-GRW4 and P-SGS1; two lineages of the Plasmodium relictum), the lineage P-GRW4 started to disappear (and became undetectable with PCR methods) from the peripheral blood relatively fast when the intensity of P-SGS1 infection increased. Videvall et al. (2020) furthermore demonstrated that hosts infected with the highly virulent P-SGS1 showed a very high transcriptional response in genes which function primarily within the immune system or cell death regulation. However, the less virulent P-GRW4 caused much lower parasitaemia and just a minor transcriptome shift in the same direction as P-SGS1.
Similar differences were observed in the gene expression of the parasites, when Common Starlings (Sturnus vulgaris), i.e. a species with high tolerance for Plasmodium infection and a more susceptible species, the Common Crossbill (Loxia curvirostra) were experimentally infected with the same Plasmodium lineage. In the more sensitive host species, parasite expressed genes responsible for cell-invasion more intensely than in the parasite-tolerant host species. On the other hand, in the parasite-tolerant host species, parasite genes related to apoptosis or/and oxidative stress showed a higher expression level (Garcia-Longoria et al. 2020).
The impacts of the parasite genera and lineages may also differ during migration. In a long-distant migrant species, the Great Reed Warbler (Acrocephalus arundinaceus), the onset of autumn migration was delayed with increasing intensity of Plasmodium or mixed-genus infection, while Haemoproteus infection did not affect migration timing (Emmenegger et al. 2021). Shurulinkov et al. (2012) found that body condition of Yellow Wagtails (Motacilla flava) infected with Haemoproteus motacillae during spring and autumn migration was worse than non-infected individuals but no such correlation was found in the case of Haemoproteus anthi infection. Furthermore, parasites might alter the landscape movements of infected migrants during the refuelling period (e.g. Hegemann et al. 2018; Eikenaar et al. 2020) and as a result blood parasite infections may slow down the whole migration process.
In our earlier work (Ágh et al. 2019) where we had no possibility to sequence parasite lineages and did not identify parasite genera either, we found that infected juvenile Robins arrived later during autumn migration; however, prevalence had no effects on the actual body condition of the individuals. Here, we sequenced and incorporated parasite lineage data to the data used in Ágh et al. (2019). We studied whether a relationship exists between infection status of the birds and their condition and timing of migration.
Material and methods
Data sampling
We captured Robins at the Ócsa Bird Ringing Station (Central Hungary: 47°17′ N, 19°12′ E) using mist nests and following the Actio Hungarica capturing protocol (for details, see Csörgő et al. 2016). Robins sporadically breed and overwinter at this study site and are regular and common passage migrants during spring and autumn. The birds included in our study arrived from different areas of Europe; however, their breeding origin cannot be determined based on morphological characteristics (reviewed in Harnos et al. 2018). In our analyses, we included blood samples only from birds that were most likely under migration (samples were collected between 20th August and 5th November 2016; N = 403). Based on a long-term ringing data series in Hungary, we can state with high confidence that after 20th of August, the percentage of the transmigrate Robins is markedly increasing. Therefore, we can define this date as the start of the autumn migration period. We choose 5th November to be the end of the sampling period because after that date, we mainly detect overwintering individuals (Gyurácz and Csörgő, 2009 in the Hungarian Bird Migration Atlas).
Capturing and DNA sampling was conducted periodically so that 4 sampling days were followed by a 3-day break period. Blood samples were collected into 96% ethanol and kept at − 20 °C until analysis. We determined the age [Njuvenile = 345 (hatched in the year of capture), Nadult = 58 (older); Demongin 2016], measured wing feather length (the flattened maximum wing chord) and body mass of the individuals (see Ágh et al. 2019 for more details). We also estimated the subcutaneous fat deposition by fat scores (the fat scores normally range from 0 to 8, see Kaiser 1993); however, the birds in this study had fat scores only between 0 and 3. Arrival date was defined as the day of first capture of the birds at the study site (Harnos et al. 2015).
Laboratory methods
The dataset used in this study was partially compiled by Ágh et al. (2019). In our current study, we added parasite lineage data to the previously used dataset where we had no possibility to sequence the lineages or to identify parasite genera and analysed the effects of the detected different lineages on their hosts. DNA was extracted with GeneAid Genomic DNA Mini Kit (Tissue) following the manufacturer’s protocol (Thermo Scientific™). Molecular sexing was performed to determine the sex of the birds and to check DNA quality (Suh et al. 2011). We excluded samples with unsuccessful amplification during molecular sexing (i.e. bad quality DNA samples, N = 7); therefore, we finally included 396 individuals (Njuvenile = 340 and Nadult = 56) in the analyses.
For the molecular detection of avian malaria and malaria-like parasites, we used a highly efficient nested polymerase chain reaction (PCR) method (Waldenström et al. 2004). To increase the sensitivity of the detection and to have enough DNA to the sequencing reaction, we used twice as much DNA as in our earlier study (Ágh et al. 2019). This resulted a higher overall prevalence (33.7%) in this study compared to Ágh et al. 2019 (14.9%). In all PCRs, both negative (ddH2O) and positive controls (samples that were previously confirmed to be infected) were included to control for possible contaminations and amplification failures during PCRs, respectively. However, neither negative controls nor positive controls ever showed contamination or amplification failures, respectively. To reduce the risk for false negatives, we screened negative samples twice for blood parasites.
To identify the different lineages, all samples with positive amplification were sequenced using the BigDye Terminator v3.1 cycle sequencing kit and sent to a capillary electrophoresis platform (Biological Research Centre, Hungary). Sequences were edited and aligned using the program BioEdit (Hall 1999) and identified to genus (Haemoproteus or Plasmodium) and lineage level by comparing sequence data with those of previously identified parasites reported in MalAvi database. Parasites with sequences differing by one nucleotide substitution were considered to represent evolutionary independent lineages (Bensch et al. 2004). We found 3 Haemoproteus and 12 Plasmodium lineages out of which 6 lineages were previously not reported (for GenBank accession numbers see Table 1). Out of these 15 lineages, there were only one frequent Haemoproteus (H-ROBIN1: Nadult = 13, Njuvenile = 39) and two Plasmodium lineages (P-LINN1: Nadult = 6, Njuvenile = 14; P-TURDUS1: Nadult = 6, Njuvenile = 28). Other lineages were present in the population in much less frequencies (prevalence ranged between 0.003 and 0.018). Therefore, the possible effects of the lineages on host physiology were analysed only for the frequent lineages (H-ROBIN1, P-LINN1, P-TURDUS1). Samples with multiple infections are difficult to identify to lineage level correctly due to the messy signal or the low signal intensity resulting in missing bases on the electropherogram. Therefore, mixed infections (only 4 cases) were omitted from the analyses. For prevalence data of the detected lineages, see Table 1.
Statistical methods
We calculated the prevalence of Haemoproteus and Plasmodium parasites and the most common lineages in all age and sex categories with Sterne methods (Sterne 1954) with Quantitative Parasitology 3.0 (Reiczigel and Rózsa 2005) and compared them with Fisher’s test.
To test for differences in the average body mass of infected and non-infected individuals, we used a linear model with multiple comparison tests and compared the means of infected (Plasmodium/Haemoproteus or infected with a given lineage) and non-infected individuals in each sex and age group combination. Independent variables were the age, sex, and wing length.
In the case of fat categories 2 and 3, the sample size was too low (N < 5) after separating individuals by infection status. Therefore, to reach the best statistical fit and well-interpretable biological effects, we categorized the birds into two fat categories, those with no visible fat (score = 0, N = 200) and with fat (score = 1, N = 196). In the first genus-specific analysis, we assessed the relationship between fat scores (0/1) and infection category (non-infected vs. Haemoproteus-infected/non-infected vs. Plasmodium-infected individuals) using a generalized linear model (GLM) with binomial distribution and logit link function (Venables and Ripley 2002). In the second lineage-specific analysis, we repeated the comparison among the most common parasite lineages (non-infected vs. infected with H-ROBIN1 or P-TURDUS1 or P-LINN1). As sex and age classes did not differ statistically in fat scores, these two factors were not used as dependent variables in the model.
To study the possible differences in the average arrival time of the individuals, we used general linear models. Independent variables were sex and infection categories (non-infected vs. Haemoproteus-infected/non-infected vs. Plasmodium-infected) and their interaction. In a separate model, we analysed the effects of the three most common lineages (non-infected vs. infected with H-ROBIN1 or P-TURDUS1 or P-LINN1) on arrival time; the other independent variable was the sex. Because adults and juveniles arrived in different migration waves and to eliminate the effects of the unbalanced sample sizes, we built separate models for adults and juveniles when analysing the effects of parasites on timing of migration. All analyses were performed using R version 3.4.2. (R Development Core Team R 2017). For GLM we used “stats” package; for multiple comparisons, we used “multcomp” package, which contains p-value correction too (Hothorn et al. 2008). For the calculation of the effect size, we used the “effectsize” package.
Results
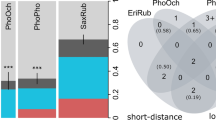
The overall prevalence of Haemoproteus and Plasmodium species was 13.6% (95% CI: 10.58–17.39%; N = 54) and 19.9% (95% CI: 16.27–24.22%; N = 79), respectively. When comparing the two genera, the prevalence did not differ between age (Haemoproteus: OR = 1.92, p = 0.069; Plasmodium: OR = 1.31, p = 0.390) or sex categories (Haemoproteus: OR = 0.90, p = 0.773; Plasmodium: OR = 0.74, p = 0.220). For more details on sex and age group interactions, see the supplementary material (Table S1).
There were no significant differences in the mean body mass between parasitized and non-parasitized individuals when compared non-infected individuals with Plasmodium-infected or Haemoproteus-infected individuals (F-value = 1.321, p = 0.270). However, P-TURDUS1-infected birds were heavier than non-infected individuals (estimate ± SE = 0.66 ± 0.224, t = 2.931, p = 0.004; Fig. 1; effect size: estimate [95% CI] = 0.52 [0.17, 0.86]). The interaction between age groups and different parasite lineages was non-significant; therefore, it was eliminated from the model (F-value = 0.807, p = 0.490).
Body mass in relation to the most common parasite lineages in all age and sex groups merged. Box plots show the median, lower and upper quartiles and the whiskers representing data within the 1.5 × interquartile range and were calculated from the data. The black dots represent the mean body mass in each category with SE, estimated from the final model
There was no significant difference in fat scores between Haemoproteus- or Plasmodium-infected and non-infected individuals (OR = 1.019, p = 0.797). However, we found that adult individuals infected with H-ROBIN1 had lower odds to have visible fat than non-infected adult individuals (OR = 0.084, CI = 0.005–0.423, z = − 2.387, p = 0.017; Fig. 2; effect size: − 2.55 [− 5.52, − 0.75]). While, juveniles infected with P-TURDUS1 had higher odds to have visible fat than their non-infected counterparts (OR = 2.500, CI = 1.142–6.033, z = 2.190, p = 0.029; Fig. 2; effect size: 0.95 [0.13, 1.87]). In this respect, individuals infected with P-LINN1, juveniles infected with H-ROBIN1, and adults infected with P-TURDUS1 did not differ significantly from the non-infected individuals in the same age category.
Haemoproteus- and Plasmodium-infected individuals arrived later compared to non-infected ones, but only in the case of juveniles (estimates for adults see Table S3). The differences were greater in the case of Haemoproteus infections than for Plasmodium. There was no difference in arrival time when comparing individuals infected with Haemoproteus vs. Plasmodium (Table 2). If sexes were analysed separately, the greatest difference in arrival time was between non-infected and Haemoproteus-infected females (Table 2).
Our lineage-specific analyses show that adults infected with P-TURDUS1 arrived marginally later than non-infected ones (estimate ± SE = 12.23 ± 6.197, t = 1.973, p = 0.054; Fig. 3; effect size: 0.85 [− 0.02, 1.71]); however, the sample size in the P-TURDUS1-infected group was very low (N = 6). On the other hand, juveniles infected with H-ROBIN1 (estimate ± SE = 10.37 ± 3.083, t = 3.363, p < 0.001; effect size: 0.57 [0.24, 0.90]) and P-TURDUS1 (estimate ± SE = 11.80 ± 3.567, t = 3.308, p < 0.001; effect size: 0.65 [0.26, 1.03]) arrived significantly later than non-infected ones (Fig. 3). The interaction between sex groups and parasite lineages was not significant; therefore, it was eliminated from the final model (F = 1.0291, p = 0.380).
Arrival date in relation to infections with the most common parasite lineages separately for adults (left) and juveniles (right). Box plots show the median, lower, and upper quartiles and the whiskers representing data within the 1.5 × interquartile range. The black dots represent the mean arrival time in each category
Discussion
Migratory birds cross different areas and habitats along their migratory routes and are exposed to a wide range of blood parasites (e.g. Waldenström et al. 2002). It was also hypothesized that they could potentially spread different parasite lineages to local resident species. However, later studies also showed that the number of migratory birds might not affect local parasite prevalence as prevalence was mostly influenced by macro-ecological patterns, like climatic differences among sites or vector communities (Shurulinkov and Ilieva 2009; Clark et al. 2016; Angeli et al. 2021).
Despite differences in the abundance and virulence of Haemoproteus and Plasmodium species (Scheuerlein and Ricklefs 2004; Astudillo et al. 2013), it is often difficult to detect any variation in the impacts of these parasites on the life history or morphological traits of the birds (e.g. Jenkins et al. 2015) because the effects of chronic infections are less pronounced (Valkiūnas 2005), and many times can be detected only on long term (Asghar et al. 2015). During experimental studies, the intensity of infection correlated with the rate of telomere loss (Asghar et al. 2015), affected the average haematocrit values, caused the blockage of blood vessels in the brain and resulted in higher mortality rate (Ilgūnas et al. 2019). However, to assess these questions, longitudinal data series or sometimes invasive sampling is needed.
The importance of analysing the effects of the different parasite lineages separately was supported by earlier studies. For instance in the House Martin (Delichon urbicum), the prevalence of the most common lineages showed opposite geographic patterns. The prevalence of Haemoproteus lineage (H-DELURB1) increased, while prevalence of Plasmodium relictum haplotypes decreased from North Africa to Europe. However, there were no lineage-specific differences in their impacts on body size, body condition, recapture rate, or survival of the individuals (van Rooyen et al. 2014). On the other hand, in other host and parasite species, differences were found among the lineages in the severity of the infection (Palinauskas et al. 2011), or in their effects on host fitness (Lachish et al. 2011; Shurulinkov et al. 2012).
In our study, the prevalence of two of the three most common lineages, P-TURDUS1 and H-ROBIN1 correlated with body condition, but in a different direction. Individuals infected with P-TURDUS1 was on average heavier than non-infected individuals, which was probably caused by the fact that P-TURDUS1-infected juveniles had a higher probability of having a fat deposit (having fat or not) than non-infected individuals. The interpretation of the results on P-TURDUS1 infection is complex. On one hand, it is possible that these individuals arrived at our study site in a better condition or we simply caught them at the end of their refuelling period. The fact that P-TURDUS1-infected individuals were captured later at our study site than non-infected individuals and individuals infected with H-ROBIN1 or P-LINN1 would suggest the latter.
On the other hand, adults infected with H-ROBIN1 had fat deposits less probably, but their mean body mass was similar to that of the non-infected individuals. It is possible that H-ROBIN1 infections caused a minimal delay in fat accumulation, but this was not manifested in differences in the body mass of the birds. However, it is important to note, that sample size for adult individuals was quite low, and the confidence intervals of the effect sizes were relatively wide (though the effect sizes were relatively high).
Hegemann et al. (2018) found previously in short-distance migrants (including Robins) that the landscape movements of haemosporidian-infected individuals increased, and these infected individuals stayed ca. three times longer at their stopover site to refill their fat reserves compared to their non-infected conspecifics. Another study on the long-distant migrant Great Reed Warbler found that migration timing was delayed and migration distances were shortened for individuals that were highly parasitized with Plasmodium, or with both Plasmodium and Haemoproteus parasites. To compensate their delay, infected Great Reed Warblers migrated significantly less far and had shorter resting durations (Emmenegger et al. 2021). However, Hahn et al. (2018) did not find any effects of low parasitemia on phenotypic attributes in this species during the preparation for migration. These results suggest that infection status may affect the movement capacity and the migration phenology of migratory birds, but infection status may only weakly associate with the actual body condition of the individuals as was discussed by the review of Risely et al. (2018). The short or medium-distant migrants, like Robins, probably follow other migration strategies, and perhaps, impacts and consequences of infections on migration phenology of these birds may also be different.
However, if the findings of Emmenegger et al. (2021) are also valid in our case, this would explain why H-ROBIN1-infected juveniles arrived later at our study site than non-infected individuals. This would also imply that Haemoproteus infections could slow down the migration process also in Robins. The small sample size of H-ROBIN1-infected adults (N = 13) would, however, explain why we did not detect similar relationship also in this age group.
The delay in arrival time was greater in females. Females occupy a subordinate position also during migration (Campos et al. 2011) and as a result often feed in less favourable habitats (Catry et al. 2004). Together with the negative effects of the infection, this may cause a higher delay in the arrival time of females. Unfortunately, we had too few recapture data about the studied individuals [Nnon-infected = 43 (21 male, 22 female), Ninfected = 23 (13 male, 10 female)], so we were not able to statistically analyse whether differences exist in the preparation time and efficiency between males and females and Haemoproteus infected and non-infected individuals.
It is also possible that later arriving H-ROBIN1-infected individuals simply came from more distant geographic areas and as a consequence of their longer journey, they arrive with more depleted fat reserves. This would also explain why these individuals arrived later. In this case, we would predict the prevalence of H-ROBIN1 infection to be higher in more northern populations; however, there is no information about the prevalence of the different lineages in other Robin populations to compare with.
Our and previous results therefore emphasize, that studying the effects of haemosporidian parasites on their hosts is particularly important during migration or during the dispersal period (Martínez-de la Puente et al. 2010; López et al. 2013; Emmenegger et al. 2020; but see Santiago-Alarcon et al. 2013; Emmenegger et al. 2018). If the number of infected birds increases for some reason and these birds arrive at the stop-over or wintering site in suboptimal time, this may even cause local population decline on the long term.
In addition to the three generalist and widely distributed lineages, we found some other, previously not detected parasites, which might be specialized to Robins. However, these parasites were present in such a low prevalence (N12 rare lineages = 26; prevalence ranged between 0.003 and 0.018) that we were not able to analyse their effects on migration timing of the birds. We can only state that these rare parasites were detected mainly in the second half of the migration period (Fig. S1).
To summarize, individuals infected with haemosporidian parasites arrived later at the study site. The delay in arrival was more pronounced for Haemoproteus than for Plasmodium parasites. When focusing on individual lineages, we found that H-ROBIN1- and P-TURDUS1-infected juveniles arrived later than non-infected individuals. However, there were no differences between uninfected and Haemoproteus/Plasmodium-infected individuals in terms of fat storage or body mass. However, when we analysed the lineages individually, we found that TURDUS1-infected birds were heavier than non-infected individuals, adults infected with H-ROBIN1 had lower, while juveniles infected with P-TURDUS1 had higher probability to have visible fat than non-infected individuals.
Based on the magnitude of the effect sizes of the estimations and the biologically relevant differences found in timing of autumn migration of Robins, we stress the importance of studying the effects of haemosporidian parasites on migration phenology of this species. It is also important to note that we analysed only one migration period, and for some lineages, the sample sizes were so low that we can only carefully speculate the effects of the different lineages during autumn migration. To get a clearer picture on the effects of avian malaria during autumn migration and explore their effects on Robins on a broader scale, more data from different breeding populations and more detailed sampling of the migration populations (e.g. with radio-tracking, geolocators, see examples in Hegemann et al. 2018; Emmenegger et al. 2020) are needed.
References
Ágh N, Piross IS, Majoros G et al (2019) Malaria infection status of European Robins seems to associate with timing of autumn migration but not with actual condition. Parasitology 146:814–820. https://doi.org/10.1017/S0031182018002184
Asghar M, Hasselquist D, Hansson B et al (2015) Hidden costs of infection: chronic malaria accelerates telomere degradation and senescence in wild birds. Science 347:436–438. https://doi.org/10.1126/science.1261121
Astudillo VG, Hernández SM, Kistler WM et al (2013) Spatial, temporal, molecular, and intraspecific differences of haemoparasite infection and relevant selected physiological parameters of wild birds in Georgia, USA. Int J Parasitol Parasites Wildl 2:178–189. https://doi.org/10.1016/j.ijppaw.2013.04.005
Aželytė J, Platonova E, Bensch S et al (2022) A comparative analysis of the dynamics of Plasmodium relictum (GRW4) development in the blood during single and co-infections. Acta Trop 226:106247. https://doi.org/10.1016/j.actatropica.2021.106247
Bensch S, Pé Rez-Tris J, Waldenstr J, Hellgren O (2004) Linkage between nuclear and mitochondrial DNA sequences in avian malaria parasites: multiple cases of cryptic speciation? Evolution (n y) 58:1617–1621. https://doi.org/10.1111/j.0014-3820.2004.tb01742.x
Bensch S, Hellgren O, Pérez-Tris J (2009) MalAvi: a public database of malaria parasites and related haemosporidians in avian hosts based on mitochondrial cytochrome b lineages. Mol Ecol Resour 9:1353–1358. https://doi.org/10.1111/j.1755-0998.2009.02692.x
Campos AR, Catry P, Ramos J, Robalo J (2011) Competition among European Robins Erithacus rubecula in the winter quarters: sex is the best predictor of priority of access to experimental food resources. Ornis Fenn 88:226–233
Catry P, Campos A, Almada V, Cresswell W (2004) Winter segregation of migrant European robins Erithacus rubecula in relation to sex, age and size. J Avian Biol 35:204–209. https://doi.org/10.1111/j.0908-8857.2004.03266.x
Clark NJ, Clegg SM, Klaassen M (2016) Migration strategy and pathogen risk: non-breeding distribution drives malaria prevalence in migratory waders. Oikos 125:1358–1368. https://doi.org/10.1111/oik.03220
Csörgő T, Harnos A, Rózsa L, Karcza Zs, Fehérvári P (2016) Detailed description of the Ócsa Bird Ringing Station, Hungary. Ornis Hungarica 24(2):91– 108. https://doi.org/10.1515/orhu-2016-0018
de Angeli DD, Filion A, Fecchio A et al (2021) Migrant birds disperse haemosporidian parasites and affect their transmission in avian communities. Oikos 130:979–988. https://doi.org/10.1111/oik.08199
Demongin L (2016) European Robin (Erithacus rubecula). In: Identification guide to birds in the hand. Beauregard-Vernon. pp 243–244.
Dimitrov D, Zehtindjiev P, Bensch S (2010) Genetic diversity of avian blood parasites in SE Europe: cytochrome b lineages of the genera Plasmodium and Haemoproteus (Haemosporida) from Bulgaria. Acta Parasitol 55:201–209. https://doi.org/10.2478/s11686-010-0029-z
Dimitrov D, Palinauskas V, Iezhova TA et al (2015) Plasmodium spp.: an experimental study on vertebrate host susceptibility to avian malaria. Exp Parasitol 148:1–16. https://doi.org/10.1016/j.exppara.2014.11.005
Drovetski SV, Aghayan SA, Mata VA et al (2014) Does the niche breadth or trade-off hypothesis explain the abundance-occupancy relationship in avian Haemosporidia? Mol Ecol 23:3322–3329. https://doi.org/10.1111/mec.12744
Eikenaar C, Hessler S, Hegemann A (2020) Migrating birds rapidly increase constitutive immune function during stopover. R Soc Open Sci 7:192031. https://doi.org/10.1098/rsos.192031
Emmenegger T, Bensch S, Hahn S et al (2020) Effects of blood parasite infections on spatiotemporal migration patterns and activity budgets in a long-distance migratory passerine. Ecol Evol 11:753–762. https://doi.org/10.1002/ece3.7030
Emmenegger T, Bensch S, Hahn S et al (2021) Effects of blood parasite infections on spatiotemporal migration patterns and activity budgets in a long-distance migratory passerine. Ecol Evol 11:753–762. https://doi.org/10.1002/ece3.7030
Emmenegger, T, Bauer, S, Hahn, S, Müller, SB, Spina, F, Jenni, L (2018) Blood parasites prevalence of migrating passerines increases over the spring passage period. J Zool 306(1):23–27. https://doi.org/10.1111/jzo.12565
Garcia-Longoria L, Palinauskas V, Ilgūnas M et al (2020) Differential gene expression of Plasmodium homocircumflexum (lineage pCOLL4) across two experimentally infected passerine bird species. Genomics 112:2857–2865. https://doi.org/10.1016/j.ygeno.2020.03.025
Gyurácz J, Csörgő T (2009) European Robin. In: Csörgő T, Karcza Zs, Halmos G, Magyar G, Gyurácz J, Szép T, Bankovics A, Schmidt A, Schmidt E (eds) Hungarian bird Migration Atlas. Kossuth Kiadó, Budapest, Hungary, pp 440–442
Hahn S, Bauer S, Dimitrov D et al (2018) Low intensity blood parasite infections do not reduce the aerobic performance of migratory birds. Proc R Soc B Biol Sci 285:20172307. https://doi.org/10.1098/rspb.2017.2307
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. In Nucleic acids symposium series (41(41):95–98). [London]: Information Retrieval Ltd., c1979–c2000
Harnos A, Fehérvári P, Csörgő T (2015) Hitchhikers’ guide to analysing bird ringing data. Ornis Hungarica 23:163–188. https://doi.org/10.1515/orhu-2015-0018
Harnos A, Ágh N, Fehérvári P et al (2018) Exploratory analyses of migration timing and morphometrics of the European Robin (Erithacus rubecula). Ornis Hungarica 26:124–148. https://doi.org/10.1515/orhu-2018-0009
Hegemann A, Alcalde Abril P, Muheim R et al (2018) Immune function and blood parasite infections impact stopover ecology in passerine birds. Oecologia 188:1011–1024. https://doi.org/10.1007/s00442-018-4291-3
Hellgren O, Waldenström J, Peréz-Tris J et al (2007) Detecting shifts of transmission areas in avian blood parasites - a phylogenetic approach. Mol Ecol 16:1281–1290. https://doi.org/10.1111/j.1365-294X.2007.03227.x
Hellgren O, Pérez-Tris J, Bensch S (2009) A jack-of-all-trades and still a master of some: prevalence and host range in avian malaria and related blood parasites. Ecology 90:2840–2849. https://doi.org/10.1890/08-1059.1
Hellgren O, Atkinson CT, Bensch S et al (2015) Global phylogeography of the avian malaria pathogen Plasmodium relictum based on MSP1 allelic diversity. Ecography (cop) 38:842–850. https://doi.org/10.1111/ecog.01158
Hothorn T, Bretz F, Westfall P (2008) Simultaneous inference in general parametric models. Biometrical J 50:346–363
Ilgūnas M, Bukauskaitė D, Palinauskas V et al (2019) Patterns of Plasmodium homocircumflexum virulence in experimentally infected passerine birds. Malar J 18:174. https://doi.org/10.1186/s12936-019-2810-2
Jenkins T, Delhaye J, Christe P (2015) Testing local adaptation in a natural great tit-malaria system: an experimental Approach. PLoS ONE 10:e0141391. https://doi.org/10.1371/journal.pone.0141391
Kaiser A (1993) A new multi-category classification of subcutaneous fat deposits of songbird. J F Ornithol 64:246–255
Knowles SCL, Palinauskas V, Sheldon BC (2010) Chronic malaria infections increase family inequalities and reduce parental fitness: experimental evidence from a wild bird population. J Evol Biol 23:557–569. https://doi.org/10.1111/j.1420-9101.2009.01920.x
Lachish S, Knowles SCL, Alves R et al (2011) Fitness effects of endemic malaria infections in a wild bird population: the importance of ecological structure. J Anim Ecol 80:1196–1206. https://doi.org/10.1111/j.1365-2656.2011.01836.x
Lapointe DA, Atkinson CT, Samuel MD (2012) Ecology and conservation biology of avian malaria. Ann N Y Acad Sci 1249:211–226. https://doi.org/10.1111/j.1749-6632.2011.06431.x
López G, Muñoz J, Soriguer R, Figuerola J (2013) Increased endoparasite infection in late-arriving individuals of a Trans-Saharan passerine migrant bird. PLoS ONE 8(4):e61236. https://doi.org/10.1371/journal.pone.0061236
Martínez-de la Puente J, Merino S, Tomás G et al (2010) The blood parasite Haemoproteus reduces survival in a wild bird: a medication experiment. Biol Lett 6:663–665. https://doi.org/10.1098/rsbl.2010.0046
Marzal A (2012) Recent advances in studies on avian malaria parasites. In: Okwa OO (ed) Malaria Parasites. In Tech, Rijeka, Croatia, pp 135–158
Medeiros MCI, Ellis VA, Ricklefs RE (2014) Specialized avian Haemosporida trade reduced host breadth for increased prevalence. J Evol Biol 27:2520–2528. https://doi.org/10.1111/jeb.12514
Ortego J, Cordero PJ, Aparicio JM, Calabuig G (2008) Consequences of chronic infections with three different avian malaria lineages on reproductive performance of Lesser Kestrels (Falco naumanni). J Ornithol 149:337–343. https://doi.org/10.1007/s10336-008-0287-9
Palinauskas V, Valkiūnas G, Bolshakov CV, Bensch S (2011) Plasmodium relictum (lineage SGS1) and Plasmodium ashfordi (lineage GRW2): the effects of the co-infection on experimentally infected passerine birds. Exp Parasitol 127:527–533. https://doi.org/10.1016/j.exppara.2010.10.007
Palinauskas V, Žiegytė R, Šengaut J, Bernotienė R (2018) Different paths – the same virulence: experimental study on avian single and co-infections with Plasmodium relictum and Plasmodium elongatum. Int J Parasitol 48:1089–1096. https://doi.org/10.1016/j.ijpara.2018.08.003
R Development Core Team R (2017) A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Reiczigel J, Rózsa L (2005) Quantitative Parasitology 3.0 Budapest, Hungary
Risely A, Klaassen M, Hoye BJ (2018) Migratory animals feel the cost of getting sick: a meta-analysis across species. J Anim Ecol 87:301–314. https://doi.org/10.1111/1365-2656.12766
Santiago-Alarcon D, Mettler R, Segelbacher G, Schaefer HM (2013) Haemosporidian parasitism in the blackcap Sylvia atricapilla in relation to spring arrival and body condition. J Avian Biol 44:521–530. https://doi.org/10.1111/j.1600-048X.2013.00181.x
Scheuerlein A, Ricklefs RE (2004) Prevalence of blood parasites in European passeriform birds. Proc R Soc B Biol Sci 271:1363–1370. https://doi.org/10.1098/rspb.2004.2726
Shurulinkov P, Ilieva M (2009) Spatial and temporal differences in the blood parasite fauna of passerine birds during the spring migration in Bulgaria. Parasitol Res 104:1453–1458. https://doi.org/10.1007/s00436-009-1349-5
Shurulinkov P, Chakarov N, Daskalova G (2012) Blood parasites, body condition, and wing length in two subspecies of yellow wagtail (Motacilla flava) during migration. Parasitol Res 110:2043–2051. https://doi.org/10.1007/s00436-011-2733-5
Sterne TE (1954) Some remarks on confidence or fiducial limits. Biometrika 41:275–278
Suh A, Kriegs JO, Brosius J, Schmitz J (2011) Retroposon insertions and the chronology of avian sex chromosome evolution. Mol Biol Evol 28:2993–2997. https://doi.org/10.1093/molbev/msr147
Valkiūnas G (2005) Avian malaria parasites and other Haemosporidia. CRC Press, London, New York, Washington
van Rooyen J, Jenkins T, Lahlah N, Christe P (2014) North-African house martins endure greater haemosporidian infection than their European counterparts. J Avian Biol 45:450–456. https://doi.org/10.1111/jav.00408
Venables WN, Ripley BD (2002) Generalized linear models. Springer, New York, New York, NY
Videvall E, Palinauskas V, Valkiūnas G, Hellgren O (2020) Host transcriptional responses to high- and low-virulent avian malaria parasites. Am Nat 195:1079–1084. https://doi.org/10.1086/708530
Waldenström J, Bensch S, Kiboi S et al (2002) Cross-species infection of blood parasites between resident and migratory songbirds in Africa. Mol Ecol 11:1545–1554. https://doi.org/10.1046/j.1365-294X.2002.01523.x
Waldenström J, Bensch S, Hasselquist D, Ostman O (2004) A new nested polymerase chain reaction method very efficient in detecting Plasmodium and Haemoproteus infections from avian blood. J Parasitol 90:191–194. https://doi.org/10.1645/GE-3221RN
Acknowledgements
The authors express their gratitude for the work of the volunteers at the Ócsa Bird Ringing Station, with special thanks for Imre Sándor Piross, Ármin Csipak, and Krisztián Barna. We are grateful for the members of Molecular Ecology Research Group of the University of Veterinary Medicine who helped in the laboratory work and Gábor Majoros, Alexandra Juhász, Lajos Rózsa, and Károly Erdélyi for their helpful comments in the study and methods development.
Funding
Open access funding provided by University of Pannonia. This work was supported by the Hungarian Ministry of Human Capacities (National Talent Program, grant numbers NTP-NFTÖ–16–0493) to NÁ, the National Scientific Research Fund of Hungary (OTKA under Grant No. 108571) to NÁ, the National Research, Development and Innovation Office (Grant no. K132490 to NÁ and PD124043, FK127917 to ES), and by the New National Excellence Program by the Ministry of Innovation and Technology (ÚNKP-19–4-ELTE-460) and the János Bolyai research scholarship from the Hungarian Academy of Sciences (BO/00163/22) to ES.
Author information
Authors and Affiliations
Contributions
NÁ and ES designed the study. NÁ collected the field data; molecular identification was conducted by NÁ and ES. Statistical analyses were conducted by NÁ. All authors interpreted the data and wrote the manuscript.
Corresponding author
Ethics declarations
Ethical standards
All international, national, and institutional guidelines for the care and use of animals were followed. Research was permitted by the Middle-Danube-Valley Inspectorate for Environmental Protection, Nature Conservation and Water Management (under Registration Number KTF: 27251–1/2014). Blood sampling was performed by N.Á. (Certificate Registration Number 6/2015, issued by the Institutional Animal Welfare Committee of the University of Veterinary Medicine Budapest).
Conflict of interest
The authors declare no competing interests.
Additional information
Section Editor: Elizabeth Marie Warburton
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Data
The dataset analysed for this study are available on MendeleyData (https://doi.org/10.17632/jm4t6ygxrj.1).
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ágh, N., Csörgő, T. & Szöllősi, E. Delay in arrival: lineage-specific influence of haemosporidians on autumn migration of European robins. Parasitol Res 121, 2831–2840 (2022). https://doi.org/10.1007/s00436-022-07621-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-022-07621-5