Abstract
Colorectal cancer (CRC) remains a highly prevalent gastrointestinal neoplasm, presenting significant prevalence and lethality rate. DEAD/H box RNA helicase 10 (DDX10) has been proposed as a potential oncogene in CRC, the specific action mechanism by which DDX10 modulates the aggressive biological cellular events in CRC remains implicitly elucidated, however. During this study, DDX10 expression was detected via RT-qPCR and Western blotting. Cell proliferation was estimated via EDU staining. TUNEL staining and Western blotting appraised cell apoptosis. Cell stemness was evaluated by sphere formation assay, RT-qPCR, Western blotting as well as immunofluorescence staining. Relevant assay kit examined aldehyde dehydrogenase (ALDH) activity. Western blotting and immunofluorescence staining also detected autophagy. DDX10 was hyper-expressed in CRC cells. Down-regulation of DDX10 hampered cell proliferation, aggravated the apoptosis while eliminated the ability to form spheroid cells in CRC. In addition, DDX10 deletion improved ATG10 expression and therefore activated autophagy in CRC cells. Consequently, ATG10 depletion or treatment with autophagy inhibitor 3-Methyladenine (3-MA) partially compensated the influences of DDX10 silencing on the proliferation, apoptosis and stemness of CRC cells. Accordingly, DDX10 deficiency may aggravate autophagy mediated by ATG10 to impede cell proliferation, stemness and facilitate cell apoptosis, hence blocking the progression of CRC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Colorectal cancer (CRC) represents a highly prevalent gastrointestinal neoplasm, rated as the second most deadly cancer following lung cancer (Siegel et al. 2020). It was estimated that in the United States, CRC contributed to over 53,000 deaths in 2020 (Siegel et al. 2020). The etiology of CRC begins as low-grade dysplasia, the slow development of which contributes to polyp generation in the colorectal lumen, finally progressing to a tumor with considerable size (Choi et al. 2015). Gender, obesity, a sedentary lifestyle, heavy consumption of alcohol and red meat, and smoking cigarettes are commonly believed as predisposing factors of CRC (Patel et al. 2022). Surgery with adjuvant chemotherapy and radiotherapy is the cornerstone of CRC therapy, the routine application of which fails to overcome tumor recurrence, metastasis as well as drug resistance, however (Simard et al. 2019; Fakih 2015; Ma et al. 2023). Accordingly, the identification of the latent mechanism of CRC may be conducive to the development of novel treatment regimens to increase the lifespan of patients with CRC.
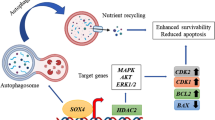
Autophagy is a self-cannibalization process in which biomacromolecules or damaged organelles are phagocytosed and encapsulated to sustain favorable survival conditions through providing cellular energy and protection under the circumstances of destructive environments, such as nutrient deficiency, drug toxicity, and hypoxia (Parzych and Klionsky 2014). Autophagy is implicated in diverse physiopathological processes, including oncogenesis, cellular development, apoptosis and survival (Klionsky et al. 2021). Autophagy-related gene 10 (ATG10) is known as an E2-like activating enzyme essential for the synthesis of the autophagosome complex (Hu and Reggiori 2022). Plenty studies have determined that ATG10 may act as a prognostic biomarker in CRC (Hu et al. 2022) and modulate the chemoresistance and metastasis of CRC cells (Sun et al. 2020; Jo et al. 2017).
DEAD/H box RNA helicases (DDX) are a group of proteins comprised of rapid amplified putative ATP-dependent RNA helicase that are related to tumor initiation and advancement (Zhang et al. 2021). Accumulated reports have well documented DEAD/H box RNA helicase 10 (DDX10) as a tumor promoter in lung adenocarcinoma (Liu et al. 2021), ovarian cancer (Gai et al. 2016) and osteosarcoma (Shi and Hao 2019). Zhou et al. have established that DDX10 mainly exerts oncogenic activities in CRC (Zhou et al. 2022). Recent literature has revealed that DDX10 can be degraded by promoting SQSTM1-mediated selective autophagy (Li et al. 2023). Importantly, ATG genes including ATG3, ATG7 and ATG10 were predicted to be negatively regulated by DDX10 by Biogrid database.
Thence, it is speculated that DDX10 may also participate in the process of CRC through mediating ATG-dependent autophagy, which is the objective of our present work.
Materials and methods
Cell lines, reagents and siRNA transfection
Normal human intestinal epithelial cell line NCM460 (SUNNCELL, Wuhan, China) was hatched in Roswell Park Memorial Institute (RPMI)-1640 medium (Welgene, Gyeongsan, Korea) and CRC cell lines (SW620, DLD-1 and LS174T) supplied by BeNa Culture Collection (Shanghai, China) were all subjected to Dulbecco’s modified Eagle medium (DMEM; Welgene, Gyeongsan, Korea). All mediums were appended with 10% fetal bovine serum (FBS; Bovogen Biologicals, Melbourne, VIC, Australia), along with 1% antibiotics (Welgene, Gyeongsan, Korea) in a water saturated atmosphere equipped with 5% CO2 at 37 °C. Besides, SW620 and DLD-1 cells were pretreated by 1 µM autophagy inhibitor 3-Methyladenine (3-MA; HARVEYBIO, Beijing, China) for 2 h (Huang et al. 2022), and then treated by EZ Trans Lipo transfection reagent (Life-iLab, Shanghai, China) for transfection of siRNAs targeting DDX10 (siRNA-DDX10-1/2) or/and ATG10 (siRNA-ATG10-1/2) as well as the relevant control siRNA (siRNA-NC) (Genomeditech, Shanghai, China).
5-Ethynyl-2’-deoxyuridine (EDU) staining
By means of iClick™ EdU Andy Fluor™ 488 Imaging Kit (GeneCopoeia, Inc.), cell proliferation was appraised in compliance with the indicate protocol. In brief, SW620 and DLD-1 cells (1 × 104 cells/well) prepared in 96-well plates received indicated treatment and transfection, followed by being mixed with 20 µM EDU working solution for 2 h. After being immobilized by 3.7% paraformaldehyde for 15 min and perforated with 0.5% Triton X-100 for 20 min at room temperature, the incorporated EdU was visualized by means of a click reaction using Alexa Fluor 488 azide for half an hour without exposure to light. Images were prepared for observation under a fluorescence microscope (Thermo Fisher Scientific) after the nuclear DNA was dyed by Hoechst 33,342.
Terminal-deoxynucleoitidyl transferase mediated nick end labeling (TUNEL)
Cell apoptosis was appraised applying TUNEL Apoptosis Assay kit (Shanghai Acmec Biochemical Co., Ltd) in agreement with the manufacturer’s instructions. In short, following indicated treatment and transfection, 4% paraformaldehyde was utilized to treat SW620 and DLD-1 cells for immobilization, followed by permeabilization with 0.1% Triton X-100. Subsequently, the cells were cultivated with TUNEL reaction reagent for 1 h and the nuclei were labeled with 10 mg/ml DAPI for 10 min. Finally, five random fields of view were selected to capture images using a fluorescence microscope (Thermo Fisher Scientific).
Sphere formation assay
SW620 and DLD-1 cells (1 × 103 cells per well) sowed into a 6-well ultralow attachment plate (Corning Glass) received indicated treatment and transfection and were incubated in DMEM deprived of serum comprising 20 ng/ml epidermal growth factor (EGF), 10 ng/ml human fibroblast growth factor basic (hFGFb) as well as 2% B27. Under a light microscope (Thermo Fisher Scientific), the formed spheres were calculated 10 days later.
Measurement of aldehyde dehydrogenase (ALDH) activity
Following the standard procedure of ALDH assay kit (Sigma-Aldrich), ALDH concentration was examined.
Immunofluorescence staining
After being immobilized by 4% paraformaldehyde and perforated with 0.1% Triton X-100 for 10 min, SW620 and DLD-1 cells were treated by 5% BSA blocking buffer and then cultivated with antibodies against octamer-binding transcription factor 4 (OCT4; cat. no. ab181557; 1/250; Abcam) overnight at 4 °C, prior to the addition of Alexa-Fluor 488-conjugated secondary antibody (cat. no. #S0018; 1/100; Affinity Biosciences) for another 1 h at room temperature on the following day. The fluorescence intensity was captured under the fluorescence microscope after the nuclear DNA was dyed by 1 mg/ml DAPI for 10 min.
Reverse transcription-quantitative PCR (RT-qPCR)
With the aid of NucleoZOL® RNA Isolation Reagent (MACHEREY–NAGEL, Germany), total RNA was prepared from cells as described in the manufacturer’s manual, which was then reverse-transcripted into cDNA through SuperScript Reverse Transcriptase Kit (Vazyme, Nanjing, China). PCR reaction was implemented using LightCycler® 480 SYBR Mix (Roche Diagnostics Ltd., Shanghai, China). In terms of 2−ΔΔCq approach, DDX10, OCT4, Nanog and ATG10 expression were calibrated, with GAPDH as a normalizer.
Western blot
After isolated by RIPA buffer (BioRad, Hercules, CA, USA), quantified by BCA method (4 A Biotech, Beijing, China) and separated using gel electrophoresis, the proteins were loaded onto the PVDF membranes sealed in 5% BSA. The membranes were then processed with primary antibodies (Abcam) as well as HRP-linked secondary antibody (Abcam). By means of UltraSigna Supersensitive ECL Substrate Kit (4 A Biotech, Beijing, China), color development was conducted.
Statistics
Statistically significant differences were determined using One-way ANOVA as well as Tukey’s post hoc test, which was done with SPSS 22.0 software (SPSS, Inc., Chicago, IL, USA). P-values less than 0.05 were denoted significant in statistics.
Results
DDX10 demonstrates up-regulated expression in CRC cells
With the aim of illustrating the role of DDX10 in CRC, DDX10 expression in CRC cells was initially examined and the results from RT-qPCR and Western blotting hinted that DDX10 expression at both mRNA and protein level was conspicuously elevated in CRC cell lines (SW620, DLD-1 and LS174T) by contrast with in normal human intestinal epithelial cell line NCM460 (Fig. 1A-B). Consequently, SW620 and DLD-1 cells were selected for the follow-up experiments, in which the highest DDX10 expression was exhibited.
Deletion of DDX10 curtails the proliferation and promotes the apoptosis of CRC cells
Subsequently, functional experiments were carried out to specify the effects of DDX10 on the malignant process of CRC. Following the transduction of siRNA-DDX10-1/2 plasmids, DDX10 expression was discovered to be tremendously depleted in SW620 and DLD-1 cells (Fig. 2A-B). Moreover, siRNA-DDX10-1 was applied to the ensuing assays in view of the lower DDX10 expression in the siRNA-DDX10-1 group. As illustrated by EDU staining, the proliferation of SW620 and DLD-1 cells was retarded overtly after DDX10 was knocked down (Fig. 2C). Conversely, through TUNEL staining, it was noted that DDX10 deficiency resulted in the significant increase on the number of apoptotic cells (Fig. 2D). The data from Western blotting also manifested that when DDX10 was down-regulated, BCL2 expression was declined and Bax expression was improved in SW620 and DLD-1 cells (Fig. 2E).
Deletion of DDX10 curtails the proliferation and promotes the apoptosis of CRC cells. (A-B) Transfection efficacy of DDX10 interference plasmids. (C) EDU staining estimated cell proliferation. (D) TUNEL staining appraised cell apoptosis. (E) Examination of BCL2 and Bax expression by Western blotting. ***P < 0.001
Deletion of DDX10 inhibits the stemness of CRC cells
Cell stemness has been proposed as a culprit in the aggressive process of CRC. To comprehensively expound the influences of DDX10 on CRC, cell stemness was also appraised. The results from sphere formation assay delineated that after DDX10 was silenced, the sphere formation capacity of SW620 and DLD-1 cells was notably extenuated (Fig. 3A). Besides, ALDH concentration was depleted in SW620 and DLD-1 cells transfected with DDX10 interference plasmids (Fig. 3B). In addition, through RT-qPCR, Western blotting and immunofluorescence staining, OCT4 and Nanog expression were both discovered to be on a downward trend by shortage of DDX10 (Fig. 3C-E).
DDX10 interference triggers ATG10-dependent autophagy
ATG genes including ATG3, ATG7 and ATG10 were predicted to be negatively regulated by DDX10 by Biogrid database. Via Western blotting, when DDX10 was down-regulated, ATG3, ATG7 and ATG10 expression were all raised, among which ATG10 was the most prominently altered (Fig. 4A). Correspondingly, ATG10 was chosen for the following assays. To verify whether the downstream mechanism of DDX10 was associated with ATG10, ATG10 expression was overtly lowered after transfection of siRNA-ATG10-1/2 (Fig. 4B-C). Furthermore, it turned out that LC3, Beclin1 expression were both increased and P62 expression was reduced by DDX10 absence in SW620 and DLD-1 cells, which were all reversed after ATG10 was depleted. Also, interference with ATG10 alone repressed LC3 expression and enhanced P62 expression in SW620 and DLD-1 cells (Fig. 4D-E).
DDX10 interference triggers ATG10-dependent autophagy. (A) Examination of ATG3, ATG7 and ATG10 expression by Western blotting. (B-C) Transfection efficacy of ATG10 interference plasmids. (D) SW620 and DLD-1 cells were co-transfected with siRNA-DDX10 and siRNA-ATG10 utilizing EZ Trans Lipo transfection reagent. Immunofluorescence staining examined LC3II expression. (E) SW620 and DLD-1 cells were co-transfected with siRNA-DDX10 and siRNA-ATG10 utilizing EZ Trans Lipo transfection reagent. Examination of LC3II/LC3I, P62 and Beclin1 expression by Western blotting. *P < 0.05, **P < 0.01, ***P < 0.001
ATG10-dependent autophagy mediated by DDX10 inhibition hampers the proliferation and aggravates the apoptosis of CRC cells
To prove that DDX10 might function in CRC through regulating ATG10-dependent autophagy, autophagy inhibitor 3-MA was applied and ATG10 was also silenced. From the results of EDU staining, it was observed that the weakened proliferative ability of SW620 and DLD-1 cells imposed by DDX10 reduction was strengthened by ATG10 interference or 3-MA (Fig. 5A). Moreover, when DDX10 was down-regulated, the apoptotic rate of SW620 and DLD-1 cells was elevated, BCL2 expression was lowered and Bax expression was promoted. After knockdown of ATG10 or treatment with 3-MA in DDX10-silencing SW620 and DLD-1 cells, the cell apoptotic rate was repressed again, coupled with the ascending BCL2 expression and the descending Bax expression (Fig. 5B-C).
ATG10-dependent autophagy mediated by DDX10 inhibition hampers the proliferation and aggravates the apoptosis of CRC cells. SW620 and DLD-1 cells were co-transfected with siRNA-DDX10 and siRNA-ATG10 utilizing EZ Trans Lipo transfection reagent or SW620 and DLD-1 cells were transfected with siRNA-DDX10 and treated by 3-MA. (A) EDU staining estimated cell proliferation. (B) TUNEL staining appraised cell apoptosis. (C) Examination of BCL2 and Bax expression by Western blotting. **P < 0.01, ***P < 0.001
ATG10-dependent autophagy mediated by DDX10 inhibition suppresses the stemness of CRC cells
As portrayed in Fig. 6A, sphere formation assay substantiated that the reduced size and number of spheres in DDX10-silencing SW620 and DLD-1 cells rose again by 3-MA treatment or deletion of ATG10 (Fig. 6A). 3-MA treatment or ATG10 reduction improved ALDH content again which was declined in SW620 and DLD-1 cells on account of DDX10 reduction (Fig. 6B). Similarly, the down-regulated OCT4 and Nanog expression attributed to DDX10 interruption in SW620 and DLD-1 cells were both up-regulated again after inactivation of autophagy or deficiency of ATG10 (Fig. 6C-E).
ATG10-dependent autophagy mediated by DDX10 inhibition suppresses the stemness of CRC cells. SW620 and DLD-1 cells were co-transfected with siRNA-DDX10 and siRNA-ATG10 utilizing EZ Trans Lipo transfection reagent or SW620 and DLD-1 cells were transfected with siRNA-DDX10 and treated by 3-MA. (A) Sphere formation assay evaluated cell stemness. (B) ALDH assay kit tested ALDH content. Examination of OCT4 and Nanog expression by (C) RT-qPCR and (D) Western blotting. (E) Immunofluorescence staining examined OCT4 expression. *P < 0.05, **P < 0.01, ***P < 0.001
Discussion
CRC is commonly termed as a heterogeneous disease, the tumorigenesis and deterioration of which involve a multitude of genetic and epigenetic aberrations. In the present work, DDX10 was found to be overexpressed in CRC cells and absence with DDX10 impaired the proliferation, stemness and exacerbated the apoptosis of CRC cells, the modulatory mechanism of which might be driven by activation of autophagy mediated by ATG10.
DDX dysregulation has been reported to engage in splicing, mRNA nuclear output, and mRNA degradation to translation and alter the expression of certain oncogenes or anti-oncogenes to impact the behaviors of cancer cells (Kouyama et al. 2019; Fuller-Pace 2013). A great body of studies have claimed that DDX chiefly plays cancerous roles in CRC. For instance, DDX54 may acts as an activator of p65 and AKT signaling to facilitate the proliferation and mobility of CRC cells (Yu et al. 2021). DDX39B contributes to the metastasis of CRC via upregulating FUT3 expression and stimulating the TGFβ/SMAD2 signaling (He et al. 2021). As a member of DDX family, DDX10 expression is increased in CRC tissues and may splice RPL35 to aggravate the proliferation and metastasis of CRC cells (Zhou et al. 2022). Consistently, our research firstly presented that DDX10 expression was raised in CRC cells relative to normal human intestinal epithelial cell line and interference with DDX10 notably halted the proliferation of CRC cells. Liu et al. have held the view that inhibition of DDX10 hinders the proliferation while accelerates the apoptosis of lung adenocarcinoma cells (Liu et al. 2021). Expectedly, our experimental findings delineated that when DDX10 was depleted, the apoptotic ability of CRC cells was robustly intensified, accompanied with the elevated pro-apoptotic Bax expression and the lowered anti-apoptotic BCL2 expression. As aforementioned above, DDX10 might accelerate the occurrence of CRC.
CSCs are a subpopulation of cancer cells featured by the self-renewal and differentiation capacities similar to stem cells. Numerous studies have broadly clarified that cancer stemness represents a major source of the tumorigenesis, metastasis, recurrence and treatment resistance of CRC (Du et al. 2022; Ebrahimi et al. 2023). OCT4 and Nanog have been regarded to sustain the pluripotency of stem cells, thence being viewed as crucial regulators of stem cells (Hatina et al. 2022). ALDH is also known as a highly robust CSC marker across the vast majority of cancer types (Toledo-Guzmán et al. 2019). Here, DDX10 down-regulation was confirmed to diminish the stem-like properties of CRC cells, which was also evidenced by the declined ALDH activity, OCT4 and Nanog expression.
Interestingly, the data from Biogrid database predicated that DDX10 had a negative correlation with ATG3, ATG7 and ATG10. Our investigation further validated that ATG3, ATG7 and ATG10 expression were all enhanced in CRC cells after DDX10 was silenced. In particular, ATG10 was the most notably up-regulated. As an autophagic E2-like enzyme, ATG10 can assemble ATG12 and regulate the transformation of LC3-I to LC3-II by binding with ATG7, which is dispensable for the formation of autophagosome (Kaiser et al. 2013). P62/SQSTM1 is a multifunctional adapter protein which can interact with polyubiquitinated proteins through the ubiquitin-associated domain, further participating in the autophagic degradation of ubiquitinated substrates (Lamark et al. 2017). Beclin1 is a central protein in autophagy that triggers a cascade of proteins involved in autophagosome formation (Prerna and Dubey 2022). It is worth mentioning that DDX10 can be degraded by the activation of selective autophagy mediated by P62 (Li et al. 2023). The current experimental results uncovered that DDX10 knockdown resulted in the increase on LC3II, Beclin1 expression as well as the decrease on P62 expression in CRC cells, which were all reversed after ATG10 was depleted, highlighting that DDX10 deficiency might lead to the activation of autophagy through up-regulating ATG10 in CRC cells. Plenty studies have documented that autophagy is closely associated with the advancement of CRC and targeting autophagy may be considered as a potential therapeutic approach for CRC (Zhou et al. 2016; Zhang et al. 2023; Gil et al. 2016). Recent evidence has supported that ascending LC3B and p62 expression predicts favorable overall survival of CRC patients, whereas Beclin-1 exerts the opposite impacts (Hu et al. 2022). Specifically, ATG10 has also been determined as a biomarker for predicting the outcome of CRC patients (Hu et al. 2022) and to be implicated in the drug resistance and metastasis of CRC cells (Sun et al. 2020; Jo et al. 2017). Our study demonstrated that interference with ATG10 alone repressed LC3 expression and enhanced P62 expression in CRC cells and further testified that the impacts of DDX10 silencing on the proliferation, apoptosis as well as stemness of CRC cells were all partially abrogated following ATG10 deletion or treatment with autophagy inhibitor 3-MA.
Conclusion
All in all, this study revealed the anti-tumor role of DDX10 interference in CRC. The anti-proliferation, anti-stemness and pro-apoptosis activities of DDX10 inhibition might be associated with the activation of autophagy mediated by up-regulation of ATG10. Therefore, targeting autophagy dependent on DDX10 might be valued as a potential novel strategy to treat CRC.
Data availability
No datasets were generated or analysed during the current study.
References
Choi CH, Ignjatovic-Wilson A, Askari A, Lee GH, Warusavitarne J, Moorghen M et al (2015) Low-grade dysplasia in ulcerative colitis: risk factors for developing high-grade dysplasia or colorectal cancer. Am J Gastroenterol 110(10):1461–1471 quiz 72
Du L, Cheng Q, Zheng H, Liu J, Liu L, Chen Q (2022) Targeting stemness of cancer stem cells to fight colorectal cancers. Semin Cancer Biol 82:150–161
Ebrahimi N, Afshinpour M, Fakhr SS, Kalkhoran PG, Shadman-Manesh V, Adelian S et al (2023) Cancer stem cells in colorectal cancer: signaling pathways involved in stemness and therapy resistance. Crit Rev Oncol Hematol 182:103920
Fakih MG (2015) Metastatic colorectal cancer: current state and future directions. J Clin Oncol 33(16):1809–1824
Fuller-Pace FV (2013) DEAD box RNA helicase functions in cancer. RNA Biol 10(1):121–132
Gai M, Bo Q, Qi L (2016) Epigenetic down-regulated DDX10 promotes cell proliferation through Akt/NF-κB pathway in ovarian cancer. Biochem Biophys Res Commun 469(4):1000–1005
Gil J, Pesz KA, Sąsiadek MM (2016) May autophagy be a novel biomarker and antitumor target in colorectal cancer? Biomark Med 10(10):1081–1094
Hatina J, Kripnerová M, Houdek Z, Pešta M, Tichánek F (2022) Pluripotency stemness and Cancer: more questions than answers. Adv Exp Med Biol 1376:77–100
He C, Li A, Lai Q, Ding J, Yan Q, Liu S et al (2021) The DDX39B/FUT3/TGFβR-I axis promotes tumor metastasis and EMT in colorectal cancer. Cell Death Dis 12(1):74
Hu Y, Reggiori F (2022) Molecular regulation of autophagosome formation. Biochem Soc Trans 50(1):55–69
Hu D, Huo Y, Xue Y, Feng H, Sun W, Wang H et al (2022) Clinical application of autophagy proteins as prognostic biomarkers in colorectal cancer: a meta-analysis. Future Oncol 18(31):3537–3549
Huang Z, Gan S, Zhuang X, Chen Y, Lu L, Wang Y et al (2022) Artesunate inhibits the cell growth in Colorectal Cancer by promoting ROS-Dependent cell senescence and autophagy. Cells. ;11(16)
Jo YK, Roh SA, Lee H, Park NY, Choi ES, Oh JH et al (2017) Polypyrimidine tract-binding protein 1-mediated down-regulation of ATG10 facilitates metastasis of colorectal cancer cells. Cancer Lett 385:21–27
Kaiser SE, Qiu Y, Coats JE, Mao K, Klionsky DJ, Schulman BA (2013) Structures of Atg7-Atg3 and Atg7-Atg10 reveal noncanonical mechanisms of E2 recruitment by the autophagy E1. Autophagy 9(5):778–780
Klionsky DJ, Petroni G, Amaravadi RK, Baehrecke EH, Ballabio A, Boya P et al (2021) Autophagy in major human diseases. EMBO J 40(19):e108863
Kouyama Y, Masuda T, Fujii A, Ogawa Y, Sato K, Tobo T et al (2019) Oncogenic splicing abnormalities induced by DEAD-Box helicase 56 amplification in colorectal cancer. Cancer Sci 110(10):3132–3144
Lamark T, Svenning S, Johansen T (2017) Regulation of selective autophagy: the p62/SQSTM1 paradigm. Essays Biochem 61(6):609–624
Li J, Zhou Y, Zhao W, Liu J, Ullah R, Fang P et al (2023) Porcine reproductive and respiratory syndrome virus degrades DDX10 via SQSTM1/p62-dependent selective autophagy to antagonize its antiviral activity. Autophagy 19(8):2257–2274
Liu C, Tang J, Duan X, Du Y, Wang X, Cui Y (2021) DDX10 promotes human lung carcinoma proliferation by U3 small nucleolar ribonucleoprotein IMP4. Thorac Cancer 12(12):1873–1880
Ma SC, Zhang JQ, Yan TH, Miao MX, Cao YM, Cao YB et al (2023) Novel strategies to reverse chemoresistance in colorectal cancer. Cancer Med 12(10):11073–11096
Parzych KR, Klionsky DJ (2014) An overview of autophagy: morphology, mechanism, and regulation. Antioxid Redox Signal 20(3):460–473
Patel SG, Karlitz JJ, Yen T, Lieu CH, Boland CR (2022) The rising tide of early-onset colorectal cancer: a comprehensive review of epidemiology, clinical features, biology, risk factors, prevention, and early detection. Lancet Gastroenterol Hepatol 7(3):262–274
Prerna K, Dubey VK (2022) Beclin1-mediated interplay between autophagy and apoptosis: New understanding. Int J Biol Macromol 204:258–273
Shi JH, Hao YJ (2019) DDX10 overexpression predicts worse prognosis in osteosarcoma and its deletion prohibits cell activities modulated by MAPK pathway. Biochem Biophys Res Commun 510(4):525–529
Siegel RL, Miller KD, Goding Sauer A, Fedewa SA, Butterly LF, Anderson JC et al (2020) Colorectal cancer statistics, 2020. CA Cancer J Clin 70(3):145–164
Simard J, Kamath S, Kircher S (2019) Survivorship Guidance for patients with colorectal Cancer. Curr Treat Options Oncol 20(5):38
Sun W, Li J, Zhou L, Han J, Liu R, Zhang H et al (2020) The c-Myc/miR-27b-3p/ATG10 regulatory axis regulates chemoresistance in colorectal cancer. Theranostics 10(5):1981–1996
Toledo-Guzmán ME, Hernández MI, Gómez-Gallegos ÁA, Ortiz-Sánchez E (2019) ALDH as a stem cell marker in solid tumors. Curr Stem Cell Res Ther 14(5):375–388
Yu Y, Wang JL, Meng LL, Hu CT, Yan ZW, He ZP et al (2021) DDX54 plays a cancerous role through activating P65 and AKT Signaling Pathway in Colorectal Cancer. Front Oncol 11:650360
Zhang L, Li X, DEAD-Box RNA (2021) Helicases in cell cycle control and clinical therapy. Cells. ;10(6)
Zhang Y, Li H, Lv L, Lu K, Li H, Zhang W et al (2023) Autophagy: dual roles and perspective for clinical treatment of colorectal cancer. Biochimie 206:49–60
Zhou H, Yuan M, Yu Q, Zhou X, Min W, Gao D (2016) Autophagy regulation and its role in gastric cancer and colorectal cancer. Cancer Biomark 17(1):1–10
Zhou X, Liu Z, He T, Zhang C, Jiang M, Jin Y et al (2022) DDX10 promotes the proliferation and metastasis of colorectal cancer cells via splicing RPL35. Cancer Cell Int 22(1):58
Funding
The research received support from Project ZR2022QH391 supported by Shandong Provincial Natural Science Foundation.
Author information
Authors and Affiliations
Contributions
Kai Wang and Hong Jiang conceived the study. Kai Wang and Hao Zhan drafted and revised the manuscript. Song Fan and Shicheng Chu analyzed the data and searched the literature. Kai Wang, Hao Zhan, Song Fan, Shicheng Chu and Hongli Xu performed the experiments. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, K., Zhan, H., Fan, S. et al. ATG10-dependent autophagy is required for DDX10 to regulate cell proliferation, apoptosis and stemness in colorectal cancer. J Cancer Res Clin Oncol 150, 386 (2024). https://doi.org/10.1007/s00432-024-05910-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00432-024-05910-3