Abstract
Background
The importance of molecular diagnostics is increasingly emphasized in the 2021 WHO guidelines for gliomas. There is considerable variability in molecular features and prognosis among glioma patients with the same pathological WHO grade.
Methods
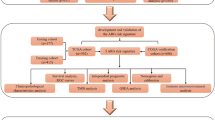
mRNA data and clinical information of human glioma patients were obtained from TCGA and CGGA databases, while expression profiles and TMZ resistance phenotypes of human glioma stem cells were acquired from the GEO database. Differentially expressed genes were identified across distinct WHO grades. Unsupervised clustering was performed on glioma patients based on DEG expression profiles. The Boruta algorithm was employed to identify feature genes for distinct molecular subtypes, and PCA was used to reduce the dimensionality of the feature gene expression data. Grade scores for each sample were calculated and correlated with patients' clinical molecular pathological features and immune microenvironment. Gene set enrichment analysis identified grade score-related functional pathways. Weighted gene co-expression network analysis identified grade score-associated biomarkers. The impact of the hub gene on malignant glioma behavior was validated through in vitro experiments, including CCK-8, EdU, colony formation, Transwell, wound healing, and immunofluorescence assays.
Results
A total of 672 and 687 samples were screened from TCGA and CGGA databases, respectively, along with 6 control, 24 low-grade, and 40 glioblastoma samples from our hospital. Two robust gene clusters were identified based on the expression profiles of 4,476 DEGs among grades 2, 3, and 4 tissues, revealing distinct prognoses. The grade scores exhibited significant heterogeneity across different WHO grade samples, representing diverse immune microenvironments. Grade scores served as independent risk factors for predicting patient prognosis, with higher sensitivity than traditional biomarkers. KIF20A, identified as a grade score-related biomarker, was independently associated with glioma prognosis. Exclusively expressed in tumor cells, KIF20A knockdown significantly inhibited tumor growth, invasion, and EMT biological behavior in glioma cells. Furthermore, KIF20A could serve as a biological marker for predicting patient response to TMZ treatment.
Conclusion
The grade scoring system enhances our understanding of the glioma tumor microenvironment. KIF20A, a novel biomarker for predicting TMZ treatment efficiency, influences malignant tumor behavior by affecting the EMT biological behavior of glioma cells.













Similar content being viewed by others
Data availability
Publicly available datasets were analyzed in this study. This data can be found below: https://www.cancer.gov/; http://www.cgga.org.cn/; http://tisch.comp-genomics.org/gallery/.
References
Babu D, Mudiraj A, Yadav N, Chandrashekhar YBVK, Panigrahi M, Prakash Babu P (2021) Rabeprazole has efficacy per se and reduces resistance to temozolomide in glioma via EMT inhibition. Cell Oncol (Dordrecht). https://doi.org/10.1007/s13402-021-00609-w
Barthel L, Hadamitzky M, Dammann P, Schedlowski M, Sure U, Thakur BK, Hetze S (2022) Glioma: molecular signature and crossroads with tumor microenvironment. Cancer Metastasis Rev. https://doi.org/10.1007/s10555-021-09997-9
Bindea G, Mlecnik B, Tosolini M, Kirilovsky A, Waldner M, Obenauf AC, Angell H, Fredriksen T, Lafontaine L, Berger A et al (2013) Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity. https://doi.org/10.1016/j.immuni.2013.10.003
Chen B, Khodadoust MS, Liu CL, Newman AM, Alizadeh AA (2018) Profiling Tumor Infiltrating Immune Cells with CIBERSORT. Methods Mol Biol (Clifton, NJ). https://doi.org/10.1007/978-1-4939-7493-1_12
Chen Y-H, Hueng D-Y, Tsai W-C (2018a) Proteolipid protein 2 overexpression indicates aggressive tumor behavior and adverse prognosis in human gliomas. Int J Mol Sci. https://doi.org/10.3390/ijms19113353
Cheng Y, Wang X, Xia Y (2021) Supervised t-distributed stochastic neighbor embedding for data visualization and classification. INFORMS J Comput. https://doi.org/10.1287/ijoc.2020.0961
Do QA, Su P-H, Chen C-W, Wang H-C, Lee Y-X, Weng Y-C, Chen L-Y, Hsu Y-H, Lai H-C (2023) DNA methylation of window of implantation genes in cervical secretions predicts ongoing pregnancy in infertility treatment. Int J Mol Sci. https://doi.org/10.3390/ijms24065598
Duan J, Huang W, Shi H (2016) Positive expression of KIF20A indicates poor prognosis of glioma patients. Onco Targets Ther. https://doi.org/10.2147/OTT.S115974
Geeleher P, Cox N, Huang RS (2014) pRRophetic: an R package for prediction of clinical chemotherapeutic response from tumor gene expression levels. PLoS ONE. https://doi.org/10.1371/journal.pone.0107468
Horbinski C, Berger T, Packer RJ, Wen PY (2022) Clinical implications of the 2021 edition of the WHO classification of central nervous system tumours. Nat Rev Neurol. https://doi.org/10.1038/s41582-022-00679-w
Iasonos A, Schrag D, Raj GV, Panageas KS (2008) How to build and interpret a nomogram for cancer prognosis. J Clin Oncol. https://doi.org/10.1200/JCO.2007.12.9791
Imai K, Hirata S, Irie A, Senju S, Ikuta Y, Yokomine K, Harao M, Inoue M, Tomita Y, Tsunoda T et al (2011) Identification of HLA-A2-restricted CTL epitopes of a novel tumour-associated antigen, KIF20A, overexpressed in pancreatic cancer. Br J Cancer. https://doi.org/10.1038/sj.bjc.6606052
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. https://doi.org/10.1186/1471-2105-9-559
Lee YS, Lee YS (2020) Molecular characteristics of meningiomas. J Pathol Transl Med. https://doi.org/10.4132/jptm.2019.11.05
Liang B, Zhou Y, Jiao J, Xu L, Yan Y, Wu Q, Tong X, Yan H (2022) Integrated analysis of transcriptome data revealed AURKA and KIF20A as critical genes in medulloblastoma progression. Front Oncol. https://doi.org/10.3389/fonc.2022.875521
Liu B, Su J, Fan B, Ni X, Jin T (2023) High expression of KIF20A in bladder cancer as a potential prognostic target for poor survival of renal cell carcinoma. Medicine. https://doi.org/10.1097/MD.0000000000032667
Louis DN, Perry A, Wesseling P, Brat DJ, Cree IA, Figarella-Branger D, Hawkins C, Ng HK, Pfister SM, Reifenberger G et al (2021) The 2021 WHO classification of tumors of the central nervous system: a summary. Neuro Oncol. https://doi.org/10.1093/neuonc/noab106
Maurya NS, Kushwah S, Kushwaha S, Chawade A, Mani A (2023) Prognostic model development for classification of colorectal adenocarcinoma by using machine learning model based on feature selection technique boruta. Sci Rep. https://doi.org/10.1038/s41598-023-33327-4
Nakamura M, Takano A, Thang PM, Tsevegjav B, Zhu M, Yokose T, Yamashita T, Miyagi Y, Daigo Y (2020) Characterization of KIF20A as a prognostic biomarker and therapeutic target for different subtypes of breast cancer. Int J Oncol. https://doi.org/10.3892/ijo.2020.5060
Nicholson JG, Fine HA (2021) Diffuse glioma heterogeneity and its therapeutic implications. Cancer Discov. https://doi.org/10.1158/2159-8290.CD-20-1474
Obuchowski NA, Bullen JA (2018) Receiver operating characteristic (ROC) curves: review of methods with applications in diagnostic medicine. Phys Med Biol. https://doi.org/10.1088/1361-6560/aab4b1
Picca A, Finocchiaro G (2022) Deciphering diffuse glioma immune microenvironment as a key to improving immunotherapy results. Curr Opin Oncol. https://doi.org/10.1097/CCO.0000000000000895
Picca A, Berzero G, Sanson M (2018) Current therapeutic approaches to diffuse grade II and III gliomas. Ther Adv Neurol Disord. https://doi.org/10.1177/1756285617752039
Reifenberger G, Wirsching H-G, Knobbe-Thomsen CB, Weller M (2017) Advances in the molecular genetics of gliomas—implications for classification and therapy. Nat Rev Clin Oncol. https://doi.org/10.1038/nrclinonc.2016.204
Ren X, Chen X, Ji Y, Li L, Li Y, Qin C, Fang K (2020) Upregulation of KIF20A promotes tumor proliferation and invasion in renal clear cell carcinoma and is associated with adverse clinical outcome. Aging. https://doi.org/10.18632/aging.202153
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. https://doi.org/10.1093/nar/gkv007
Saito K, Ohta S, Kawakami Y, Yoshida K, Toda M (2017) Functional analysis of KIF20A, a potential immunotherapeutic target for glioma. J Neurooncol. https://doi.org/10.1007/s11060-016-2360-1
Song Q, Zhou R, Shu F, Fu W (2022) Cuproptosis scoring system to predict the clinical outcome and immune response in bladder cancer. Front Immunol. https://doi.org/10.3389/fimmu.2022.958368
Wang Z, Wang Y, Yang T, Xing H, Wang Y, Gao L, Guo X, Xing B, Wang Y, Ma W (2021) Machine learning revealed stemness features and a novel stemness-based classification with appealing implications in discriminating the prognosis, immunotherapy and temozolomide responses of 906 glioblastoma patients. Brief Bioinform. https://doi.org/10.1093/bib/bbab032
Wilkerson MD, Hayes DN (2010) ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics (oxford, Engl). https://doi.org/10.1093/bioinformatics/btq170
Yang C, Zhang Y, Lin S, Liu Y, Li W (2021) Suppressing the KIF20A/NUAK1/Nrf2/GPX4 signaling pathway induces ferroptosis and enhances the sensitivity of colorectal cancer to oxaliplatin. Aging. https://doi.org/10.18632/aging.202774
Yang K, Wu Z, Zhang H, Zhang N, Wu W, Wang Z, Dai Z, Zhang X, Zhang L, Peng Y et al (2022) Glioma targeted therapy: insight into future of molecular approaches. Mol Cancer. https://doi.org/10.1186/s12943-022-01513-z
Yi M, Nissley DV, McCormick F, Stephens RM (2020) ssGSEA score-based Ras dependency indexes derived from gene expression data reveal potential Ras addiction mechanisms with possible clinical implications. Sci Rep. https://doi.org/10.1038/s41598-020-66986-8
Zhang Z, Reinikainen J, Adeleke KA, Pieterse ME, Groothuis-Oudshoorn CGM (2018) Time-varying covariates and coefficients in Cox regression models. Ann Transl Med. https://doi.org/10.21037/atm.2018.02.12
Zhao L, Li Y, Zhu J, Sun N, Song N, Xing Y, Huang H, Zhao J (2019) Chlorotoxin peptide-functionalized polyethylenimine-entrapped gold nanoparticles for glioma SPECT/CT imaging and radionuclide therapy. J Nanobiotechnol. https://doi.org/10.1186/s12951-019-0462-6
Acknowledgements
We gratefully acknowledge The Cancer Genome Atlas pilot project, Chinese Glioma Genome Atlas and Tumor Immune Single-cell Hub 2, which made the genomic data and clinical data available.
Funding
This work was funded by the Beijing Municipal Natural Science Foundation (7202150) and the National High Level Hospital Clinical Research Funding (2022-PUMCH-A-019) for Yu Wang, and by the National High Level Hospital Clinical Research Funding (2022-PUMCH-B-113), the Tsinghua University-Peking Union Medical College Hospital Initiative Scientific Research Program (2019ZLH101) and the Beijing Municipal Natural Science Foundation (19JCZDJC64200[Z]) for Wenbin Ma.
Author information
Authors and Affiliations
Contributions
LGY and ST contributed to the conception and design of this study. YNW, WM and YW contributed to the analysis and interpretation of data. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval and consent to participate
Ethics approval was obtained from the ethics committee at Peking Union Medical College Hospital (2022-PUMCH-B-113). Informed consent was obtained from all subjects and/or their legal guardians. All experiments were performed in accordance with relevant guidelines and regulations.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
432_2023_4898_MOESM1_ESM.tif
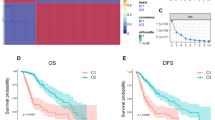
Figure S1. Prognosis and clinical features of grade-related gene clusters in CGGA-gliomas. (A) Consensus clustering matrix for glioma samples with k = 2. (B) CDF of consensus clustering for k = 2 to k = 9. (C) Relative changes in the area under the CDF curve by cluster number. (D) Principal component analysis (PCA) of glioma samples with K=2. (E) Survival analysis of clusters A and B in CGGA datasets. (F) Heatmap illustrating clinicopathological features of the two gene clusters (TIF 5147 KB)
432_2023_4898_MOESM2_ESM.tif
Figure S2. Expression and prognostic significance of hub genes in gliomas in the CGGA database. (A) The mRNA expression of hub genes among different WHO grade gliomas in CGGA cohort. (B) Forest plot of univariate and multivariate Cox regression analysis (TIF 2252 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ye, L., Tong, S., Wang, Y. et al. Grade scoring system reveals distinct molecular subtypes and identifies KIF20A as a novel biomarker for predicting temozolomide treatment efficiency in gliomas. J Cancer Res Clin Oncol 149, 9857–9876 (2023). https://doi.org/10.1007/s00432-023-04898-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-04898-6