Abstract
Background and aim
Aberrant methylation of Ras association domain family 1, isoform A (RASSF1A), and short-stature homeobox gene 2 (SHOX2) promoters has been validated as a pair of valuable biomarkers for diagnosing early lung adenocarcinomas (LUADs). Epidermal growth factor receptor (EGFR) is the key driver mutation in lung carcinogenesis. This study aimed to investigate the aberrant promoter methylation of RASSF1A and SHOX2, and the genetic mutation of EGFR in 258 specimens of early LUADs.
Methods
We retrospectively selected 258 paraffin-embedded samples of pulmonary nodules measuring 2 cm or less in diameter and evaluated the diagnostic performance of individual biomarker assays and multiple panels between noninvasive (group 1) and invasive lesions (groups 2A and 2B). Then, we investigated the interaction between genetic and epigenetic alterations.
Results
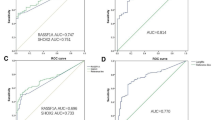
The degree of RASSF1A and SHOX2 promoter methylation and EGFR mutation was significantly higher in invasive lesions than in noninvasive lesions. The three biomarkers distinguished between noninvasive and invasive lesions with reliable sensitivity and specificity: 60.9% sensitivity [95% confidence interval (CI) 52.41–68.78] and 80.0% specificity (95% CI 72.14–86.07). The novel panel biomarkers could further discriminate among three invasive pathological subtypes (area under the curve value > 0.6). The distribution of RASSF1A methylation and EGFR mutation was considerably exclusive in early LUAD (P = 0.002).
Conclusion
DNA methylation of RASSF1A and SHOX2 is a pair of promising biomarkers, which may be used in combination with other driver alterations, such as EGFR mutation, to support the differential diagnosis of LUADs, especially for stage I.






Similar content being viewed by others
Data availability
The data utilized and analyzed in this study are available upon reasonable request to the corresponding author. Detailed data that might endanger participant consent and confidentiality are not published.
References
Aberle DR et al (2011) Reduced lung-cancer mortality with low-dose computed tomographic screening. N Engl J Med 365:395–409. https://doi.org/10.1056/NEJMoa1102873
Bao Y, Liu X, Liu Y, Wang S, Wu B (2019) Ras-association domain family 1 (RASSF1A) gene regulates progression, migration and invasion of bladder cancer. Surg Oncol 30:63–71. https://doi.org/10.1016/j.suronc.2019.05.009
Begum S et al (2011) An epigenetic marker panel for detection of lung cancer using cell-free serum DNA. Clin Cancer Res 17:4494–4503. https://doi.org/10.1158/1078-0432.Ccr-10-3436
Blandin Knight S, Crosbie PA, Balata H, Chudziak J, Hussell T, Dive C (2017) Progress and prospects of early detection in lung cancer. Open Biol. https://doi.org/10.1098/rsob.170070
Brocks D et al (2014) Intratumor DNA methylation heterogeneity reflects clonal evolution in aggressive prostate cancer. Cell Rep 8:798–806. https://doi.org/10.1016/j.celrep.2014.06.053
Buckingham L et al (2010) PTEN, RASSF1 and DAPK site-specific hypermethylation and outcome in surgically treated stage I and II nonsmall cell lung cancer patients. Int J Cancer 126:1630–1639. https://doi.org/10.1002/ijc.24896
Castro M et al (2010) Multiplexed methylation profiles of tumor suppressor genes and clinical outcome in lung cancer. J Transl Med 8:86. https://doi.org/10.1186/1479-5876-8-86
Cerfolio RJ et al (2010) The true false negative rates of esophageal and endobronchial ultrasound in the staging of mediastinal lymph nodes in patients with non-small cell lung cancer. Ann Thorac Surg 90:427–434. https://doi.org/10.1016/j.athoracsur.2010.04.062
Chen H et al (2019) Genomic and immune profiling of pre-invasive lung adenocarcinoma. Nat Commun 10:5472. https://doi.org/10.1038/s41467-019-13460-3
Cheung WK, Nguyen DX (2015) Lineage factors and differentiation states in lung cancer progression. Oncogene 34:5771–5780. https://doi.org/10.1038/onc.2015.85
Chung JH, Lee HJ, Kim BH, Cho NY, Kang GH (2011) DNA methylation profile during multistage progression of pulmonary adenocarcinomas. Virchows Arch 459:201–211. https://doi.org/10.1007/s00428-011-1079-9
Cucuruz B et al (2012) MAGE qPCR improves the sensitivity and accuracy of EBUS-TBNA for the detection of lymphatic cancer spread. J Thorac Oncol 7:690–697. https://doi.org/10.1097/JTO.0b013e31824294de
Darwiche K et al (2013) Assessment of SHOX2 methylation in EBUS-TBNA specimen improves accuracy in lung cancer staging. Ann Oncol 24:2866–2870. https://doi.org/10.1093/annonc/mdt365
Dochez V, Caillon H, Vaucel E, Dimet J, Winer N, Ducarme G (2019) Biomarkers and algorithms for diagnosis of ovarian cancer: CA125, HE4, RMI and ROMA, a review. J Ovarian Res 12:28. https://doi.org/10.1186/s13048-019-0503-7
Dubois F et al (2016) RASSF1A suppresses the invasion and metastatic potential of human non-small cell lung cancer cells by inhibiting YAP activation through the GEF-H1/RhoB pathway. Cancer Res 76:1627–1640. https://doi.org/10.1158/0008-5472.Can-15-1008
Esteller M, Herman JG (2002) Cancer as an epigenetic disease: DNA methylation and chromatin alterations in human tumours. J Pathol 196:1–7. https://doi.org/10.1002/path.1024
Gao H et al (2022) The diagnostic potential of SHOX2 and RASSF1A DNA methylation in early lung adenocarcinoma. Front Oncol. https://doi.org/10.3389/fonc.2022.849024
Grawenda AM, O’Neill E (2015) Clinical utility of RASSF1A methylation in human malignancies. Br J Cancer 113:372–381. https://doi.org/10.1038/bjc.2015.221
Hawes SE et al (2010) DNA hypermethylation of tumors from non-small cell lung cancer (NSCLC) patients is associated with gender and histologic type. Lung Cancer 69:172–179. https://doi.org/10.1016/j.lungcan.2009.11.002
Heller G et al (2013) Genome-wide CpG island methylation analyses in non-small cell lung cancer patients. Carcinogenesis 34:513–521. https://doi.org/10.1093/carcin/bgs363
Hu X et al (2019) Multi-region exome sequencing reveals genomic evolution from preneoplasia to lung adenocarcinoma. Nat Commun 10:2978. https://doi.org/10.1038/s41467-019-10877-8
Hu X et al (2021) Evolution of DNA methylome from precancerous lesions to invasive lung adenocarcinomas. Nat Commun 12:687. https://doi.org/10.1038/s41467-021-20907-z
Hua X et al (2020) Genetic and epigenetic intratumor heterogeneity impacts prognosis of lung adenocarcinoma. Nat Commun 11:2459. https://doi.org/10.1038/s41467-020-16295-5
Hulbert A et al (2017) Early detection of lung cancer using DNA promoter hypermethylation in plasma and sputum. Clin Cancer Res 23:1998–2005. https://doi.org/10.1158/1078-0432.Ccr-16-1371
Izumchenko E et al (2015) Targeted sequencing reveals clonal genetic changes in the progression of early lung neoplasms and paired circulating DNA. Nat Commun 6:8258. https://doi.org/10.1038/ncomms9258
Kakinuma R et al (2016) Natural history of pulmonary subsolid nodules: a prospective multicenter study. J Thorac Oncol 11:1012–1028. https://doi.org/10.1016/j.jtho.2016.04.006
Karemaker ID, Vermeulen M (2018) Single-cell DNA methylation profiling: technologies and biological applications. Trends Biotechnol 36:952–965. https://doi.org/10.1016/j.tibtech.2018.04.002
Klutstein M, Nejman D, Greenfield R, Cedar H (2016) DNA methylation in cancer and aging. Cancer Res 76:3446–3450. https://doi.org/10.1158/0008-5472.Can-15-3278
Kneip C et al (2011) SHOX2 DNA methylation is a biomarker for the diagnosis of lung cancer in plasma. J Thorac Oncol 6:1632–1638. https://doi.org/10.1097/JTO.0b013e318220ef9a
Ko E et al (2013) Association of RASSF1A and p63 with poor recurrence-free survival in node-negative stage I-II non-small cell lung cancer. Clin Cancer Res 19:1204–1212. https://doi.org/10.1158/1078-0432.Ccr-12-2848
Kobayashi Y, Mitsudomi T, Sakao Y, Yatabe Y (2015) Genetic features of pulmonary adenocarcinoma presenting with ground-glass nodules: the differences between nodules with and without growth. Ann Oncol 26:156–161. https://doi.org/10.1093/annonc/mdu505
Kodama K et al (2001) Prognostic value of ground-glass opacity found in small lung adenocarcinoma on high-resolution CT scanning. Lung Cancer 33:17–25. https://doi.org/10.1016/s0169-5002(01)00185-4
Li N, Zeng Y, Huang J (2020) Signaling pathways and clinical application of RASSF1A and SHOX2 in lung cancer. J Cancer Res Clin Oncol 146:1379–1393. https://doi.org/10.1007/s00432-020-03188-9
Li N et al (2021) Analysis of the prognostic value and gene expression mechanism of SHOX2 in lung adenocarcinoma. Front Mol Biosci. https://doi.org/10.3389/fmolb.2021.688274
Liu Y, Gao W, Siegfried JM, Weissfeld JL, Luketich JD, Keohavong P (2007) Promoter methylation of RASSF1A and DAPK and mutations of K-ras, p53, and EGFR in lung tumors from smokers and never-smokers. BMC Cancer 7:74. https://doi.org/10.1186/1471-2407-7-74
Mari-Alexandre J et al (2017) Translating cancer epigenomics into the clinic: focus on lung cancer. Transl Res 189:76–92. https://doi.org/10.1016/j.trsl.2017.05.008
Matsuguma H et al (2013) Characteristics of subsolid pulmonary nodules showing growth during follow-up with CT scanning. Chest 143:436–443. https://doi.org/10.1378/chest.11-3306
Mazor T et al (2015) DNA methylation and somatic mutations converge on the cell cycle and define similar evolutionary histories in brain tumors. Cancer Cell 28:307–317. https://doi.org/10.1016/j.ccell.2015.07.012
Myong NH (2003) Thyroid transcription factor-1 (TTF-1) expression in human lung carcinomas: its prognostic implication and relationship with wxpressions of p53 and Ki-67 proteins. J Korean Med Sci 18:494–500. https://doi.org/10.3346/jkms.2003.18.4.494
Nguyen QN et al (2019) 1-Genetic and epigenetic alterations of the EGFR and mutually independent association with BRCA1, MGMT, and RASSF1A methylations in Vietnamese lung adenocarcinomas. Pathol Res Pract 215:885–892. https://doi.org/10.1016/j.prp.2019.01.032
Osarogiagbon RU et al (2019) Early-stage NSCLC: advances in thoracic oncology 2018. J Thorac Oncol 14:968–978. https://doi.org/10.1016/j.jtho.2019.02.029
Pao W et al (2004) EGF receptor gene mutations are common in lung cancers from “never smokers” and are associated with sensitivity of tumors to gefitinib and erlotinib. Proc Natl Acad Sci U S A 101:13306–13311. https://doi.org/10.1073/pnas.0405220101
Patz EF et al (2014) Overdiagnosis in low-dose computed tomography screening for lung cancer. JAMA Intern Med 174:269–274. https://doi.org/10.1001/jamainternmed.2013.12738
Ram RR, Mendiratta S, Bodemann BO, Torres MJ, Eskiocak U, White MA (2014) RASSF1A inactivation unleashes a tumor suppressor/oncogene cascade with context-dependent consequences on cell cycle progression. Mol Cell Biol 34:2350–2358. https://doi.org/10.1128/mcb.01506-13
Remy-Jardin M, Edme JL, Boulenguez C, Remy J, Mastora I, Sobaszek A (2002) Longitudinal follow-up study of smoker’s lung with thin-section CT in correlation with pulmonary function tests. Radiology 222:261–270. https://doi.org/10.1148/radiol.2221001154
Ren M, Wang C, Sheng D, Shi Y, Jin M, Xu S (2017) Methylation analysis of SHOX2 and RASSF1A in bronchoalveolar lavage fluid for early lung cancer diagnosis. Ann Diagn Pathol 27:57–61. https://doi.org/10.1016/j.anndiagpath.2017.01.007
Roy D, Tiirikainen M (2020) Diagnostic power of DNA methylation classifiers for early detection of cancer. Trends Cancer 6:78–81. https://doi.org/10.1016/j.trecan.2019.12.006
Sandoval J, Peiró-Chova L, Pallardó FV, García-Giménez JL (2013) Epigenetic biomarkers in laboratory diagnostics: emerging approaches and opportunities. Expert Rev Mol Diagn 13:457–471. https://doi.org/10.1586/erm.13.37
Schmidt B et al (2010) SHOX2 DNA methylation is a biomarker for the diagnosis of lung cancer based on bronchial aspirates. BMC Cancer 10:600. https://doi.org/10.1186/1471-2407-10-600
Schmidt ML, Hobbing KR, Donninger H, Clark GJ (2018) RASSF1A deficiency enhances RAS-driven lung tumorigenesis. Cancer Res 78:2614–2623. https://doi.org/10.1158/0008-5472.Can-17-2466
Selamat SA et al (2011) DNA methylation changes in atypical adenomatous hyperplasia, adenocarcinoma in situ, and lung adenocarcinoma. PLoS ONE. https://doi.org/10.1371/journal.pone.0021443
Shi J et al (2020) Performance evaluation of SHOX2 and RASSF1A methylation for the aid in diagnosis of lung cancer based on the analysis of FFPE specimen. Front Oncol. https://doi.org/10.3389/fonc.2020.565780
Shigematsu H et al (2005) Clinical and biological features associated with epidermal growth factor receptor gene mutations in lung cancers. J Natl Cancer Inst 97:339–346. https://doi.org/10.1093/jnci/dji055
Shopland DR (1995) Tobacco use and its contribution to early cancer mortality with a special emphasis on cigarette smoking. Environ Health Perspect 103(Suppl 8):131–142. https://doi.org/10.1289/ehp.95103s8131
Siegel RL, Miller KD, Jemal A (2018) Cancer statistics, 2018. CA Cancer J Clin. https://doi.org/10.3322/caac.21442
Sung H et al (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249. https://doi.org/10.3322/caac.21660
Suzuki K, Asamura H, Kusumoto M, Kondo H, Tsuchiya R (2002) “Early” peripheral lung cancer: prognostic significance of ground glass opacity on thin-section computed tomographic scan. Ann Thorac Surg 74:1635–1639. https://doi.org/10.1016/s0003-4975(02)03895-x
Toyooka S et al (2003) Smoke exposure, histologic type and geography-related differences in the methylation profiles of non-small cell lung cancer. Int J Cancer 103:153–160. https://doi.org/10.1002/ijc.10787
Toyooka S et al (2006) Mutational and epigenetic evidence for independent pathways for lung adenocarcinomas arising in smokers and never smokers. Cancer Res 66:1371–1375. https://doi.org/10.1158/0008-5472.CAN-05-2625
Tsao MS et al (2005) Erlotinib in lung cancer: molecular and clinical predictors of outcome. N Engl J Med 353:133–144. https://doi.org/10.1056/NEJMoa050736
Vaissière T et al (2009) Quantitative analysis of DNA methylation profiles in lung cancer identifies aberrant DNA methylation of specific genes and its association with gender and cancer risk factors. Cancer Res 69:243–252. https://doi.org/10.1158/0008-5472.Can-08-2489
Vizoso M et al (2015) Aberrant DNA methylation in non-small cell lung cancer-associated fibroblasts. Carcinogenesis 36:1453–1463. https://doi.org/10.1093/carcin/bgv146
Wang XW et al (2018) CT features differentiating pre- and minimally invasive from invasive adenocarcinoma appearing as mixed ground-glass nodules: mass is a potential imaging biomarker. Clin Radiol 73:549–554. https://doi.org/10.1016/j.crad.2018.01.017
Wei H, Fang N, Guo L, Wu Z, Zhou Q (2015) Meta-analysis of the association between RASSF1A gene promoter methylation and non-small cell lung cancer. Zhongguo Fei Ai Za Zhi 18:443–450. https://doi.org/10.3779/j.issn.1009-3419.2015.07.09
Weiss G, Schlegel A, Kottwitz D, König T, Tetzner R (2017) Validation of the SHOX2/PTGER4 DNA methylation marker panel for plasma-based discrimination between patients with malignant and nonmalignant lung disease. J Thorac Oncol 12:77–84. https://doi.org/10.1016/j.jtho.2016.08.123
Yatabe Y, Borczuk AC, Powell CA (2011) Do all lung adenocarcinomas follow a stepwise progression? Lung Cancer 74:7–11. https://doi.org/10.1016/j.lungcan.2011.05.021
Zhang C et al (2017) DNA methylation analysis of the SHOX2 and RASSF1A panel in bronchoalveolar lavage fluid for lung cancer diagnosis. J Cancer 8:3585–3591. https://doi.org/10.7150/jca.21368
Zhang C et al (2019) Genomic landscape and immune microenvironment features of preinvasive and early invasive lung adenocarcinoma. J Thorac Oncol 14:1912–1923. https://doi.org/10.1016/j.jtho.2019.07.031
Zhao QT et al (2015) Diagnostic value of SHOX2 DNA methylation in lung cancer: a meta-analysis. Onco Targets Ther 8:3433–3439. https://doi.org/10.2147/ott.S94300
Acknowledgements
The authors thank the pathology department of The Nanfang Hospital of Southern Medical University for providing excellent technical support for this study.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-proft sectors.
Author information
Authors and Affiliations
Contributions
Jie Lin conceived and provided the research ideas for this paper. Xiangyu Ji collected samples and performed data analysis and wrote the original manuscript. Hong Li and Hui-Hui Chen are responsible for performing the experiments and evaluating the results. Xiangyu Ji# and Hong Li# contributed equally to this work and share first authorship. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interests
No grants or conflicts of interest supported this effort.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ji, XY., Li, H., Chen, HH. et al. Diagnostic performance of RASSF1A and SHOX2 methylation combined with EGFR mutations for differentiation between small pulmonary nodules. J Cancer Res Clin Oncol 149, 8557–8571 (2023). https://doi.org/10.1007/s00432-023-04745-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-04745-8