Abstract
Background
Protocadherin 10 (PCDH 10), a member of the superfamily of protocadherins, is a Ca2+-dependent homophilic cell-cell adhesion molecule expressed on the surface of cell membranes. Protocadherin 10 plays a critical role in the central nervous system including in cell adhesion, formation and maintenance of neural circuits and synapses, regulation of actin assembly, cognitive function and tumor suppression. Additionally, Pcdh10 can serve as a non-invasive diagnostic and prognostic indicator for various cancers.
Methods
This paper collects and reviews relevant literature in Pubmed.
Conclusion
This review describes the latest research understanding the role of Pcdh10 in neurological disease and human cancer, highlighting the importance of scrutinizing its properties for the development of targeted therapies and identifying a need for further research to explore Pcdh10 functions in other pathways, cell types and human pathologies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cell–cell adhesion is a basic process in the morphogenesis of multicellular organisms. Originally described as cell adhesion molecules, cadherins play a crucial role in cell recognition, cell communication, morphogenesis, cytoskeletal organization, cell migration, and neural circuit formation (Flaherty and Maniatis 2020; Pancho et al. 2020). According to sequence similarities, cadherins can be divided into three subfamilies: classical cadherins, desmosomal cadherins, and protocadherins (Pcdhs) (Halbleib and Nelson 2006). Among these, Pcdhs are the largest and most diverse cadherin subfamily. Pcdhs are highly abundant in the developing brain, lungs and kidneys (Homayouni et al. 2001; Kim et al. 2007), being crucial for organ development and maintenance. Pcdhs are also involved in the establishment and function of specific cell–cell connections as well as in tumor development (Kahr et al. 2013). Based on the genomic organization, Pcdhs are further classified as clustered or non-clustered (Pancho et al. 2020).
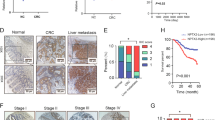
Pcdh10 is a non-clustered Pcdh (Light and Jontes 2017) that is initially highly expressed in CNS and is essential for neuronal development (Uemura et al. 2007). Pcdh10 has been identified as an autism-spectrum disorder gene (Morrow et al. 2008; Ferri et al. 2021; Hoshina et al. 2022). Additionally, Pcdh10 is a newly discovered tumor suppressor gene which is downregulated by hypermethylation or genetic deletion in various malignant tumors, and is linked to the occurrence, proliferation, invasion and metastasis of tumors (Zhong et al. 2013; Qiu et al. 2016; Yang et al. 2016, 2022). Importantly, tumor-associated Pcdh10 methylation status exhibit diagnostic and prognostic value for multiple human cancers, such as colorectal cancer, prostate cancer, cervial cancer, breast cancer, etc. (Lin et al. 2011; Jao et al. 2014; Deng et al. 2016; Liu et al. 2018b). Pcdh10 methylation does not occur in healthy tissues. However, the research of Pcdh10 is still in its early days, and there are many unknown biological characteristics and functions that have not yet been discovered.
In this review, we explore the role of Pcdh10 in neurological disease and human cancer, and provide further insight into the molecular mechanisms and disease-relationship that Pcdh10 controls.
The biological features of Pcdh10
Non-clustered PCDHs can be classified into three groups: δ1, δ2 and ε subgroup based on their structure and function (Kim et al. 2011). Most non-clustered Pcdhs have 6 or 7 extracellular cadherin repeats in the ectodomain, a transmembrane, and a cytoplasmic domain. Both Pcdhδ1 and Pcdhδ2 members contain conserved cytoplasmic motifs (CM1 and CM2) in their cytoplasmic domain, while Pcdhδ1 members have an additional protein phosphatase-1α binding domain (RRVTE, CM3).
Pcdh10, originally named OL-protocadherin, belongs to Pcdhδ2 and is located on chromosome 4 in humans and chromosome 3 in mice. It contains 6 extracellular cadherin repeats in the ectodomain, a transmembrane domain and a unique cytoplasmic domain (Hirano et al. 1999; Kim et al. 2011). Similar to other members of non-clustered Pcdhs, Pcdh10 mediates calcium-dependent cell–cell adhesion by homophilic binding through the extracellular cadherin domains, although this binding ability is generally weak (Hirano et al. 1999). Since cytoplasmic domains of non-clustered Pcdhs are distinct, they can act as major regulators via interacting with a variety of intracellular binding partners (Kim et al. 2011). A short isoform and a long isoform of Pcdh10 have been identified in human. Pcdh10 in mice contains a short isoform (iso1) and three long isoforms (iso2, iso3 and iso4), all of which only different at their carboxyterminal end of cytoplasmic domain (Kleinberger et al. 2022). Interestingly, all three long isoforms of mouse Pcdh10 contain several conserved motifs in their cytoplasmic domains (Kleinberger et al. 2022), suggesting that shared interacting partners are key for the basic functioning of Pcdh10 proteins. For example, mouse Pcdh10 interacts with Nck-associated protein 1 (Nap1), Sra-1/PIR121/cytoplasmic interacting FMR1 protein 2 (CYFIP2), Abl interactor 1 (Abi-1), hematopoietic stem/cell progenitor protein 300 (HSPC300) and WAVE1 to generate a Pcdh10-associated WAVE regulatory complex (Nakao et al. 2008). Overexpression of Pcdh10 recruits the WAVE regulatory complex at inter-axonal contact sites, which results in reorganization of F-actin and N-cadherin at these locations, and subsequently regulates cell migration of astrocytoma U251 cells (Fig. 1a). However, how Pcdh10/ Nap1/WAVE1 complex affect actin assembly needs to be further clarified.
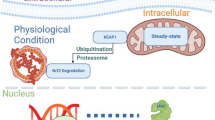
Mechanisms of Pcdh10 activity required to drive and maintain physiological and brain developmental functions. A Pcdh10 can bind Nap1, CYFIP2, Abi-1, HSPC300 and WAVE1 to form a Pcdh10-WAVE regulatory complex. By recruiting the WAVE regulatory complex to inter-axonal contact sites, Pcdh10 regulates F-actin organization and N-cadherin redistribution. Redistributed N-cadherin is unable to induce contact inhibition, leading to increased cell migration of glioblastoma cells. B Nuclear MEF2 activation initiates Mdm2 transcription, which results in ubiquitination of PSD-95. Pcdh10 then binds to ubiquitinated PSD-95 and links it to the proteasome for degradation, resulting in synapse elimination
PCDH10 in brain development
PCDH10 is detectable in nonneuronal tissue such as heart, kidney, lung and trachea (Wolverton and Lalande 2001), however its predominant expression is in the CNS (Kim et al. 2007), further indicating that Pcdh10 is critical for neural development. A high level of expression of Pcdh10 is detected in the striatum, piriform cortex, and preoptic region of the mouse brain at E13.5, but its expression in the globus pallidus is weak (Uemura et al. 2007). In all brain regions, Pcdh10 plays a key role in axon outgrowth and guidance. It has been shown, for instance, that Pcdh10 knockout mice do not form the cerebral peduncle, corticospinal tract, and striatonigral pathway, while corticofugal axons halting in the ventral telencephalon and thalamocortical axons fail to reach the internal capsule (Uemura et al. 2007). During embryonic and postnatal development, Pcdh10 is also expressed in the olfactory system, and variable levels of Pcdh10 are detected in olfactory sensory neurons and the diverse olfactory bulb glomeruli (Aoki et al. 2003; Williams et al. 2011). Pcdh10 expression is activity-dependent within olfactory sensory neurons, as reducing sensory odorant-evoked activity by naris occlusion or expression of an inactive form of cyclic nucleotide gated A2, reduces the expression of Pcdh10. Circuit formation is highly dependent on this regulation of expression, as the misexpression of Pcdh10 significantly impairs glomeruli formation in the olfactory system (Williams et al. 2011).
Neurons of the lateral and basolateral amygdala in mice heterozygous for Pcdh10 also have more filopodia and dendritic spines and (Schoch et al. 2017), and Pcdh10 appears to be necessary for the elimination of hippocampal and cortical synapses (Tsai et al. 2012). Pcdh10 acts as the downstream molecule of myocyte enhancer factor 2 (MEF2), which initiates the transcription of Murine double minute 2 (Mdm2). Post-synaptic scaffolding protein 95 (PSD-95), the synaptic scaffolding protein, was ubiquitinated by Mdm2 in response to MEF2 activation. Pcdh10 then binds PSD-95 and links it to the proteasome for ubiquitination and degradation, leading to synapse elimination (Fig. 1b). These data imply that Pcdh10 is an important player in dendritogenesis, axon development and synaptogenesis.
Pcdh10 in neurological disease
Autism spectrum disorders (ASD, also known as autism) is a highly genetically heterogeneous neurodevelopmental disorder characterized by impaired social communications. ASD commonly co-presents with other neurological conditions, such as epilepsy, intellectual disability or bipolar disorders. Several studies have identified Pcdh10 act as an autism associated gene (Morrow et al. 2008; Bucan et al. 2009).
It has been reported that the pathophysiology of autism is highly related with homozygous deletion of Pcdh10 in families with ASD (Morrow et al. 2008). Patients with homozygous Pcdh10 deletion exhibit disrupted elimination of activity-dependent excitatory synapses, as a result of altered ubiquitination and degradation pathways (Tsai et al. 2012). In line with the study, Hoshina and colleague showed that Pcdh10 deletion in mice display mild impairment in their social recognition and communication responses, suggesting that Pcdh10 mutations may cause ASD-related symptoms (Hoshina et al. 2022).
Heterozygous male mice of this mutant strain display sociability defects (Schoch et al. 2017) and altered γ oscillations (Port et al. 2017), which are known to be crucial for fear memory retrieval (Bocchio et al. 2017). Accordingly, a recent study showed that juvenile and adult Pcdh10-heterozygous mice displayed an increase in immature dendritic spine density, reduced NMDAR expression, altered γ synchronization of the basolateral amygdala, and disrupted fear conditioning behaviours (Ferri et al. 2021). Interestingly, Pcdh10+/– females showed deficits only as adults in the cued fear memory, which might relate to hormonal changes. However, the mechanism underlying the Pcdh10 knockdown-induced behavioral differences between sexes is unclear.
In addition to ASD, several recent studies have implicated human Pcdh10 in other neurological conditions, such as familial amyloidotic polyneuropathy (FAP), obsessive–compulsive disorder (OCD), major depression (MD) and schizophrenia (Fromer et al. 2014; Goncalves et al. 2016; Qin et al. 2016; Bodea et al. 2017; Tang et al. 2019). A large-scale single nucleotide polymorphism genotyping data on chromosome 4 suggested that Pcdh10 is one of the susceptibility genes of schizophrenia and bipolar disorder (Tang et al. 2019). Similarly, 428 methylated genes, including Pcdh10, have been linked to early-onset major depression in an epigenome-wide association study of 75 monozygotic twin pairs (Roberson-Nay et al. 2020). As aforementioned, deletion of Pcdh10 in mice does not affect the growth of striatal and nigrostriatal axons, but rather leads to defects in development of excitatory synapses in the dorsal basolateral nucleus of the amygdala, reduces anxiety, and causes fear and stress in ASD, OCD and MD (Hoshina et al. 2022). These results imply a strong association between Pcdh10 and relevant psychiatric disorders, including ASD, OCD and MD, and also suggest Pcdh10 as a potential target for designing anxiolytics.
Role of Pcdh10 in cancers
Numerous studies have reported that Pcdh10 acts as a tumor suppressor in a wide range of human tumors. Although the Pcdh10 gene is widely expressed in normal tissues, it is silenced or decreased in malignant tumors.
Pcdh10 in colorectal cancer
Colorectal cancer (CRC) is one of the most common types of malignant tumors. Pcdh10 is well-known as a tumor suppressor in colorectal carcinogenesis, invasion and metastasis (Zhong et al. 2017).
Several studies have reported that aberrant CpG methylation of the Pcdh10 promoter is observed in 43–85% of colorectal cancer tissues, indicating that downregulated Pcdh10 caused by methylation is a common feature of colorectal carcinogenesis (Yu et al. 2010; Silva et al. 2013; Zhong et al. 2013). A recent microarray analysis has reported that Pcdh10 expression is lost in more than half of patients with CRC (Skuja et al. 2019), raising the possibility that genetic deletion could be another mechanism for Pcdh10 inactivation in CRC. Mutations in BRAF genes, which are linked to dysregulated DNA methylation (Tanaka et al. 2010), has been found in about 10% of CRC cases (Caputo et al. 2019). Dobre and colleagues further demonstrated BRAF positive cases have higher Pcdh10 methylation levels than BRAF negative cases (Dobre et al. 2021). Taken together, Pcdh10 genetic modification and epigenetic inactivation play critical roles in the development of CRC.
In support of a role for Pcdh10 as a potential antioncogene, re-expressing Pcdh10 in colorectal cancer RKO cells leads to G1 cell cycle arrest without affecting apoptosis (Zhong et al. 2013). More specifically, Pcdh10 could inhibit cell proliferation and survival by modulating p53/p21/Rb and Bcl-2 pathways in CRC cells (Jao et al. 2021). Meanwhile, it also suppressed epithelial‐mesenchymal transition (EMT), a cellular biological process that promotes cancer cells to migrate, and stemness in CRC by negatively affecting the EGFR/AKT/GSK3β/β‐catenin signaling pathway (Jao et al. 2021).
In addition to tumor inhibition, methylation of Pcdh10 may serve as a non-invasive biomarker for CRC diagnosis as Pcdh10 methylation that is present in tissues could be detected in serum/plasma (Danese et al. 2013). Pcdh10 methylation detected in plasma increases with increasing methylation rate in tumor tissues only in early CRC (stage I/II). Additionally, allelic loss of Pcdh10 was ascertained in primary CRC tumors, and highly related with tumor progression and distant metastasis, suggesting that its allelic loss predicts an adverse prognosis (Jao et al. 2014). Moreover, patients receiving adjuvant treatment with no methylation in Pcdh10, SPARC and UCHL1, had longer disease-free rates and overall survival rates than those with hypermethylation (Heitzer et al. 2014). In contrast, unmethylated genes were related to shorter survival in surveillance group. These findings suggest that promoter methylation status of Pcdh10, SPARC and UCHL1 provide a suitable tool for predicting prognosis of stage II colorectal cancer patients.
Pcdh10 in tumors of the female reproductive system
Tumors in the female genital tract represent a leading cause of morbidity and mortality among women worldwide. Cervical and endometrial cancers are two very different diseases, having differing pathogenesis and treatments. However, PCDH10 promoter hypermethylation is a frequent hallmark observed during the progression of cervical and endometrial cancers, as previously reported (Narayan et al. 2009; Wang et al. 2009; Zhao et al. 2014; Bhat et al. 2017).
According to GEO2R analysis, Pcdh10 is downregulated and is likely to be one of the most significant genes in tumor differentiation in endometrial cancer (Liu et al. 2018a). Endometrial cancer is the most common gynecologic malignant cancer and about 80% of these cancers are endometrial endometrioid carcinomas (EEC). Pcdh10 is repressed in EEC cells due to its promoter CpG hypermethylation. A novel PCDH10-Wnt/β-catenin-MALAT1 regulatory axis that contributes to ECC development and progression, delays tumor growth and induces cell apoptosis (Zhao et al. 2019). Yang and colleagues also identified DEPDC1 as a downstream mediator of Pcdh10, and they further demonstrated that Pcdh10 suppress cell proliferation and induce apoptosis through DEPDC1-caspase signaling in EEC cell lines (HEC-1-A and KLE) (Yang et al. 2016). In the future, it would be interesting to investigate the clinical significance of Pcdh10 and MALAT1/DEPDC1. Moreover, a recent study has reported that low expression of Pcdh10 is associated with high Enhancer of Zeste Homolog 2 (EZH2) expression and Histone H3 (H3K27me3) enrichment in the tissue of endometriosis patients (Xiaolei et al. 2022). Silencing EZH2 by siRNA reduced H3K27me3 enrichment and increased PCDH10 expression, resulting in decreased invasion and migration of endometrial stromal cells and providing a target for the treatment of endometriosis patients (Xiaolei et al. 2022).
Similarly, Pcdh10 is also inactivated epigenetically in 75% cervical cell lines (Ying et al. 2006). In cervical Hela cells, knockdown of HOTAIR lncRNAs inhibits the Wnt/β-catenin signaling cascade by decreasing promoter methylation of Pcdh10, demonstrating the potential mechanism of how Pcdh10 reguates the progression of cervical cancer (Salmeron-Barcenas et al. 2019). Notably, analysis of Pcdh10 in cervical scrapings is superior to the Human Papillomavirus (HPV) test, implying its potential function as a specific diagnostic biomarker (Lin et al. 2011). Collectively, these findings demonstrate the potential role of Pcdh10 in inducing the development of different tumors of the female genital tract.
Pcdh10 in gastric cancer
Gastric cancer (GC) is the third most common fatal form of cancer around the globe and the detailed mechanism underlying gastric carcinogenesis remains unclear. Pcdh10 expression is silenced or down-regulated in gastric cancer cells and tissues (Yu et al. 2009; Li et al. 2012b), suggesting it may act as a tumor suppressor in GC. Re-expression Pcdh10 in MKN45 gastric cancer cells inhibited tumor growth, cell proliferation and invasion, induced cell apoptosis, and also increased the expression of pro-apoptotic genes including Fas, Caspase8, Jun, and CDKN1A; the anti-proliferation gene FGFR; and the anti-invasion gene HTATIP2 (Yu et al. 2009). Another study showed Pcdh10 overexpression in gastric cancer cell lines (MNK74, 7901 and AGS) suppressed cell proliferation but had no effect on cell apoptosis (Li et al. 2012b). Further investigations are required to fully understand the function of Pcdh10 in regulating apoptosis in gastric cancer.
Numerous studies have indicated that aberrant methylation of Pcdh10 could be used as a non-invasive biomarker to facilitate diagnosis and prognostic guidance for gastric cancer patients (Deng et al. 2014; Hou et al. 2015; Schneider et al. 2015; Pimson et al. 2016). For example, using MSP qPCR method, Pimson and colleagues demonstrated that Pcdh10 promoter methylation was detected in 94.06% of plasma DNA from gastric cancer patients whereas it was found in only 2.97% of matched controls, serving as a reliable non-invasive diagnostic indicator for GC (Pimson et al. 2016). In terms of prognosis prediction, Pcdh10 promoter methylation at CpG site was found in 91.92% in GC tissues (Deng et al. 2014). GC patients with 5 or more methylated CpG sites of PCDH10 promoter were dramatically related to poorer survival rates. Meanwhile, using multivariate survival analysis, the authors demonstrated methylation of combined CpG sites (− 115, − 108, − 13, and + 3) was an independent predictor, with overall survival, of gastric cancer patients postoperatively (Deng et al. 2014). Multiple studies have also confirmed this finding (Hou et al. 2015; Schneider et al. 2015). Therefore, Pcdh10 methylated at CpG sites has significant clinical applicability for GC prognosis evaluation.
Pcdh10 in pancreatic cancer
Pancreatic cancer (PC) is one of the most lethal diseases worldwide (Kamisawa et al. 2016). To date, surgical resection is the best choice for treatment of PC, however, the recurrence rate of patients who undergo resection remains very high (Ilic and Ilic 2016). Therefore, the identification of new predictive biomarkers and exploration of the pathogenesis is crucial for the development of novel therapeutics for management of PC.
Previous study identified that Pcdh10 expression is silenced by methylation in pancreatic cancer cell lines, and re-expression of Pcdh10 prevents the malignant biological process of PC cells (Qiu et al. 2016). An earlier study analyzed Pcdh10 promoter methylation in pancreatic tumor samples, but high-resolution melting analysis failed to detect a significant association between Pcdh10 promoter methylation status and tumor-staging (Yu et al. 2010). Recently, high methylation levels of Pcdh10 were found to correlate with worse progression-free survival rates instead of the overall survival, suggesting that Pcdh10 methylation status predicts poor prognosis in patients with pancreatic ductal adenocarcinomas (Curia et al. 2019).
In terms of anti-tumor effects, Pcdh10 overexpression can prevent the proliferation, migration, invasion ability of pancreatic cancer cells and trigger apoptosis by activating the AKT pathway (Qiu et al. 2016). Meanwhile, Pcdh10 can interact with human telomerase reverse transcriptase (hTERT) to reduce telomerase activity, hence mediating the inhibitory effect of PC phenotype (Zhou et al. 2015). Zhang and colleagues demonstrated that the Pcdh10 gene could generate circular RNA of Pcdh10 (circPcdh10) in PC tissue, indicating a worse prognosis (Zhang et al. 2021).
Pcdh10 in other cancers
The deletion of Pcdh10 has been reported in various human tumors. In addition to the aforementioned CRC, GC, PC, cervical and endometrial cancers, Pcdh10 loss has been observed in non-small-cell lung cancer (NSCLC; (Tang et al. 2012), nasopharyngeal and esophageal cancer (Ying et al. 2006), bladder cancer (Lin et al. 2012, 2013), hepatocellular carcinoma (Fang et al. 2013; Bing et al. 2018), multiple myeloma (Li et al. 2012a), lymphoid malignancies (Narayan et al. 2013), medulloblastoma (Bertrand et al. 2011), breast cancer (Liu et al. 2018b), and prostate cancer (Li et al. 2011), implying that Pcdh10 plays an oncosuppressor role in tumors.
In support of a role for Pcdh10 as a tumor suppressor, restoration of Pcdh10 in hepatocellular carcinoma cell lines inhibits proliferation and induces cell apoptosis via suppressing PI3K/Akt signaling pathway (Ye et al. 2017). In multiple myeloma cells, rescue of Pcdh10 expression induces apoptosis by impeding the the NF-κB pathway (Li et al. 2014), and suppresses cell proliferation via the negative modulation of Wnt/β-catenin/BCL-9 signaling (Xu et al. 2015). In oncogenic KRAS-mutated NSCLC mouse model, KRAS mutation increases the expression of Miz1, which in turn suppresses Pcdh10, leading to enhanced cell proliferation and promotion of lung tumorigenesis (Yang et al. 2022). Further evidence that Pcdh10 acts as an oncosuppressor derives from the observations that downregulated Pcdh10 expression caused by methylation predicted poor prognosis in patients with hepatocellular carcinoma (Bing et al. 2018), breast cancer (Liu et al. 2018b; Xu et al. 2021), prostate cancer (Wang et al. 2014; Deng et al. 2016), and non-small-cell lung cancer (Harada et al. 2015). These data also indicated that Pcdh10 methylation was a potential prognostic biomarker for those human cancers.
It should be noted that Pcdh10 might act as a tumor oncogene in gliomas, as it is essential for the proliferation and tumorigenicity of human glioblastoma cell lines GB2 and GB16 (Echizen et al. 2014). In human astrocytoma cell (U251), the cytoplasmic domain of Pcdh10 can interact with Nap1 and recruit the WAVE complex, and this interaction promotes adhesion and motility at the cell junctions to facilitate migration (Nakao et al. 2008). However, Pcdh10 signaling has the opposite effect in medulloblastoma cells, where Pcdh10 expression is decreased due to DNA hypermethylation and histone modification, but its restoration impedes migration (Bertrand et al. 2011). Similarly, after treatment with cytochalasin H in U87MG malignant human glioma cells, the proliferation is inhibited along with upregulated Pcdh10 expression (Heidarzadeh et al. 2019). It is not yet clear how Pcdh10 can promote tumorigenicity under one circumstance and impede it under another.
It is interesting to note that the methylation status of Pcdh10 may be able to predict the response of lymphomas to doxorubicin (Narayan et al. 2013), a common chemotherapeutic drug used to treat a variety of human cancers. Both B-cell (100%) and T-cell (79%) acute lymphoblastic leukemia frequently exhibit Pcdh10 promoter hypermethylation. Non-Hodgkin lymphoma (NHL) cell lines with down-regulated Pcdh10 expression were less sensitive to leukemia specific drugs including dexamathasone and methotrexate, while T-cell and B-cell lymphoma cell lines with Pcdh10 methylation or down-regulated expression showed doxorubicin resistance, providing new evidence for the selection of treatment plans (Narayan et al. 2013). Meanwhile, Pcdh10 could be a potential target gene for establishing epigenetic therapies in lymphomas. Imatinib is a molecular target drug used to treat chronic myeloid leukemia. In imatinib-resistant K562 leukemia cell line (KR cells), silencing of hBEX1 can repress imatinib-induced apoptosis (Ding et al. 2009). Gain expression of hBex1 enhanced PCDH10 expression and partially restored sensitivity to imatinib, implying a novel hBex1/PCDH10 pathway which contributes to drug resistance. However, the mechanism of the involvement of Pcdh10 in apoptosis has not been examined (Table 1).
Potential epigenetic therapies targeting Pcdh10
An increasing number of studies have demonstrated that Pcdh10 plays an important role in cancer. Therefore, exploring therapeutic strategies targeting Pcdh10 may be of great importance in the management of several types of tumors.
Therapeutic strategies that target Pcdh10 may be relevant to CRC. Zhou et al. demonstrated that hsa_circ_0001666 functions as a tumor suppressor by directly binding miR‐576‐5p and lessening its inhibitory effect on the target gene Pcdh10, thereby inhibiting cell proliferation, metastasis, EMT progression and stemness as well as triggering apoptosis of CRC cells (Zhou et al. 2021). Notably, hsa_circ_0001666 can also suppress Wnt/β‐catenin signaling, a well‐known cancer‐promoting pathway, via promoting PCDH10 expression.
As previously mentioned, HOTAIR lncRNA acts as the upstream regulator of Pcdh10 in cervical hela cells (Salmeron-Barcenas et al. 2019). The expression of multiple mRNAs, including MAGI2, AJAP1, SOX17, PCDH10, and TET1, was downregulated by HOTAIR knockdown, which also reduced the activity of the Wnt/-catenin signaling pathway. Similarly, HOTAIR interacted with miR-148 and DNMT1, promoting the methylation of PCDH10, and bringing about oncogenic changes in GC (Seo et al. 2021). Moreover, canonical oncogenic lncRNA MALAT1 can bind EZH2 to counteract PCDH10 by inducing the methylation of its promoter, resulting in an increase in GC cell migration and invasion (Qi et al. 2016).
CircPcdh10 promotes tumor progression of pancreatic cancer by increasing hTERT expression through interacting with miR-338-3p (Zhang et al. 2021). Further experiments confirmed that there was a targeted regulatory association between CircPcdh10 and miR-338-3p/hTERT; the inhibitory effects of circPCDH10 depletion on the viability, proliferation, invasion, and migration of PC cells were significantly abolished by treating with miR-338-3p inhibitor and hTERT. Similarly, a recent study revealed an oncogenic transcription factor FOXM1 which activated expression of miR-552, and further inhibited downstream target genes including Pcdh10, DACH1 and SMAD, which in turn promoted tumor progression and resulted in poor prognosis in PC patients (Wang et al. 2021). However, the in-depth molecular mechanisms underlying these conditions require further elucidation. In general, these results highlight the potentiality of targeting Pcdh10 gene in human cancers (Fig. 2; Table2).
Conclusion
The functions of Pcdh10, its regulatory targets and the role it plays in human pathologies, remain largely unexplored. Pcdh10 is considered to play important roles in brain development and is implicated in human neurological disorders like autism, obsessive–compulsive disorder, major depression and schizophrenia. Pcdh10 has also been implicated in a range of human cancers, acting as a tumor suppressor and playing key roles in regulating tumor growth, invasion and metastasis. In contrast, Pcdh10 has also been shown to be an oncogene for the tumorigenesis of glioblastoma. Further research is required to fully elucidate the role of Pcdh10 in neurological conditions and different types of the cancers, and determine whether Pcdh10 is implicated in any other human condition. In addition, aberrant methylations of Pcdh10 have been recognized as a non-invasive biomarker for tumor diagnosis and prognosis. Though the involvement of Pcdh10 in the pathogenesis of neural diseases and human cancers has been recently established, our understanding of the molecular functions and related signaling pathways involved is limited. Moreover, the genes targeting Pcdh10 function and the relevant molecular mechanisms involved also remain to be investigated but could provide further insights into therapeutic strategies that could be developed for the treatment of Pcdh10-regulated conditions. Our review focuses on the known conditions where Pcdh10 is disrupted and its potential as a cancer biomarker, however given the various pathways regulated by Pcdh10, further research is likely to determine its role in other conditions and further explore its potential as a non-invasive biomarker of disease.
References
Aoki E, Kimura R, Suzuki ST, Hirano S (2003) Distribution of OL-protocadherin protein in correlation with specific neural compartments and local circuits in the postnatal mouse brain. Neuroscience 117(3):593–614. https://doi.org/10.1016/s0306-4522(02)00944-2
Bertrand KC, Mack SC, Northcott PA, Garzia L, Dubuc A, Pfister SM et al (2011) PCDH10 is a candidate tumour suppressor gene in medulloblastoma. Childs Nerv Syst 27(8):1243–1249. https://doi.org/10.1007/s00381-011-1486-x
Bhat S, Kabekkodu SP, Varghese VK, Chakrabarty S, Mallya SP, Rotti H et al (2017) Aberrant gene-specific DNA methylation signature analysis in cervical cancer. Tumour Biol 39(3):1010428317694573. https://doi.org/10.1177/1010428317694573
Bing Y, Tian M, Li G, Jiang B, Ma Z, Li L et al (2018) Down-regulated of PCDH10 predicts poor prognosis in hepatocellular carcinoma patients. Medicine (Baltimore) 97(35):e12055. https://doi.org/10.1097/MD.0000000000012055
Bocchio M, Nabavi S, Capogna M (2017) Synaptic plasticity, engrams, and network oscillations in amygdala circuits for storage and retrieval of emotional memories. Neuron 94(4):731–743. https://doi.org/10.1016/j.neuron.2017.03.022
Bodea CA, Middleton FA, Melhem NM, Klei L, Song Y, Tiobech J et al (2017) Analysis of shared haplotypes amongst palauans maps loci for psychotic disorders to 4q28 and 5q23-q31. Mol Neuropsychiatry 2(4):173–184. https://doi.org/10.1159/000450726
Bucan M, Abrahams BS, Wang K, Glessner JT, Herman EI, Sonnenblick LI et al (2009) Genome-wide analyses of exonic copy number variants in a family-based study point to novel autism susceptibility genes. PLoS Genet 5(6):e1000536. https://doi.org/10.1371/journal.pgen.1000536
Caputo F, Santini C, Bardasi C, Cerma K, Casadei-Gardini A, Spallanzani A et al (2019) BRAF-mutated colorectal cancer: clinical and molecular insights. Int J Mol Sci. https://doi.org/10.3390/ijms20215369
Curia MC, Fantini F, Lattanzio R, Tavano F, Di Mola F, Piantelli M et al (2019) High methylation levels of PCDH10 predict poor prognosis in patients with pancreatic ductal adenocarcinoma. BMC Cancer 19(1):452. https://doi.org/10.1186/s12885-019-5616-2
Danese E, Minicozzi AM, Benati M, Montagnana M, Paviati E, Salvagno GL et al (2013) Epigenetic alteration: new insights moving from tissue to plasma - the example of PCDH10 promoter methylation in colorectal cancer. Br J Cancer 109(3):807–813. https://doi.org/10.1038/bjc.2013.351
Deng J, Liang H, Ying G, Dong Q, Zhang L, Yu J et al (2014) Clinical significance of the methylated cytosine-phosphate-guanine sites of protocadherin-10 promoter for evaluating the prognosis of gastric cancer. J Am Coll Surg 219(5):904–913. https://doi.org/10.1016/j.jamcollsurg.2014.06.014
Deng QK, Lei YG, Lin YL, Ma JG, Li WP (2016) Prognostic value of protocadherin10 (PCDH10) methylation in serum of prostate cancer patients. Med Sci Monit 22:516–521. https://doi.org/10.12659/msm.897179
Ding K, Su Y, Pang L, Lu Q, Wang Z, Zhang S et al (2009) Inhibition of apoptosis by downregulation of hBex1, a novel mechanism, contributes to the chemoresistance of Bcr/Abl+ leukemic cells. Carcinogenesis 30(1):35–42. https://doi.org/10.1093/carcin/bgn251
Dobre M, Salvi A, Pelisenco IA, Vasilescu F, De Petro G, Herlea V et al (2021) Crosstalk between DNA methylation and gene mutations in colorectal cancer. Front Oncol 11:697409. https://doi.org/10.3389/fonc.2021.697409
Echizen K, Nakada M, Hayashi T, Sabit H, Furuta T, Nakai M et al (2014) PCDH10 is required for the tumorigenicity of glioblastoma cells. Biochem Biophys Res Commun 444(1):13–18. https://doi.org/10.1016/j.bbrc.2013.12.138
Fang S, Huang SF, Cao J, Wen YA, Zhang LP, Ren GS (2013) Silencing of PCDH10 in hepatocellular carcinoma via de novo DNA methylation independent of HBV infection or HBX expression. Clin Exp Med 13(2):127–134. https://doi.org/10.1007/s10238-012-0182-9
Ferri SL, Dow HC, Schoch H, Lee JY, Brodkin ES, Abel T (2021) Age- and sex-specific fear conditioning deficits in mice lacking Pcdh10, an autism associated gene. Neurobiol Learn Mem 178:107364. https://doi.org/10.1016/j.nlm.2020.107364
Flaherty E, Maniatis T (2020) The role of clustered protocadherins in neurodevelopment and neuropsychiatric diseases. Curr Opin Genet Dev 65:144–150. https://doi.org/10.1016/j.gde.2020.05.041
Fromer M, Pocklington AJ, Kavanagh DH, Williams HJ, Dwyer S, Gormley P et al (2014) De novo mutations in schizophrenia implicate synaptic networks. Nature 506(7487):179–184. https://doi.org/10.1038/nature12929
Goncalves NP, Martins D, Saraiva MJ (2016) Overexpression of protocadherin-10 in transthyretin-related familial amyloidotic polyneuropathy. Am J Pathol 186(7):1913–1924. https://doi.org/10.1016/j.ajpath.2016.02.020
Halbleib JM, Nelson WJ (2006) Cadherins in development: cell adhesion, sorting, and tissue morphogenesis. Genes Dev 20(23):3199–3214. https://doi.org/10.1101/gad.1486806
Harada H, Miyamoto K, Yamashita Y, Taniyama K, Mihara K, Nishimura M et al (2015) Prognostic signature of protocadherin 10 methylation in curatively resected pathological stage I non-small-cell lung cancer. Cancer Med 4(10):1536–1546. https://doi.org/10.1002/cam4.507
Heidarzadeh S, Motalleb GH, Zorriehzahra MJ (2019) Evaluation of tumor regulatory genes and apoptotic pathways in the cytotoxic effect of cytochalasin H on malignant human glioma cell line (U87MG). Cell J 21(1):62–69. https://doi.org/10.22074/cellj.2019.5948
Heitzer E, Artl M, Filipits M, Resel M, Graf R, Weissenbacher B et al (2014) Differential survival trends of stage II colorectal cancer patients relate to promoter methylation status of PCDH10, SPARC, and UCHL1. Mod Pathol 27(6):906–915. https://doi.org/10.1038/modpathol.2013.204
Hirano S, Yan Q, Suzuki ST (1999) Expression of a novel protocadherin, OL-protocadherin, in a subset of functional systems of the developing mouse brain. J Neurosci 19(3):995–1005. https://doi.org/10.1523/JNEUROSCI.19-03-00995.1999
Homayouni R, Rice DS, Curran T (2001) Disabled-1 interacts with a novel developmentally regulated protocadherin. Biochem Biophys Res Commun 289(2):539–547. https://doi.org/10.1006/bbrc.2001.5998
Hoshina N, Johnson-Venkatesh EM, Rally VR, Sant J, Hoshina M, Seiglie MP et al (2022) ASD/OCD-linked protocadherin-10 regulates synapse, but not axon, development in the amygdala and contributes to fear- and anxiety-related behaviors. J Neurosci 42(21):4250–4266. https://doi.org/10.1523/JNEUROSCI.1843-21.2022
Hou YC, Deng JY, Zhang RP, Xie XM, Cui JL, Wu WP et al (2015) Evaluating the clinical feasibility: The direct bisulfite genomic sequencing for examination of methylated status of protocadherin10 (PCDH10) promoter to predict the prognosis of gastric cancer. Cancer Biomark 15(5):567–573. https://doi.org/10.3233/CBM-150496
Ilic M, Ilic I (2016) Epidemiology of pancreatic cancer. World J Gastroenterol 22(44):9694–9705. https://doi.org/10.3748/wjg.v22.i44.9694
Jao TM, Tsai MH, Lio HY, Weng WT, Chen CC, Tzeng ST et al (2014) Protocadherin 10 suppresses tumorigenesis and metastasis in colorectal cancer and its genetic loss predicts adverse prognosis. Int J Cancer 135(11):2593–2603. https://doi.org/10.1002/ijc.28899
Jao TM, Fang WH, Ciou SC, Yu SL, Hung YL, Weng WT et al (2021) PCDH10 exerts tumor-suppressor functions through modulation of EGFR/AKT axis in colorectal cancer. Cancer Lett 499:290–300. https://doi.org/10.1016/j.canlet.2020.11.017
Kahr I, Vandepoele K, van Roy F (2013) Delta-protocadherins in health and disease. Prog Mol Biol Transl Sci 116:169–192. https://doi.org/10.1016/B978-0-12-394311-8.00008-X
Kamisawa T, Wood LD, Itoi T, Takaori K (2016) Pancreatic cancer. Lancet 388(10039):73–85. https://doi.org/10.1016/S0140-6736(16)00141-0
Kim SY, Chung HS, Sun W, Kim H (2007) Spatiotemporal expression pattern of non-clustered protocadherin family members in the developing rat brain. Neuroscience 147(4):996–1021. https://doi.org/10.1016/j.neuroscience.2007.03.052
Kim SY, Yasuda S, Tanaka H, Yamagata K, Kim H (2011) Non-clustered protocadherin. Cell Adh Migr 5(2):97–105. https://doi.org/10.4161/cam.5.2.14374
Kleinberger I, Sanders E, Staes K, Van Troys M, Hirano S, Hochepied T et al (2022) Innovative mouse models for the tumor suppressor activity of Protocadherin-10 isoforms. BMC Cancer 22(1):451. https://doi.org/10.1186/s12885-022-09381-y
Li Z, Li W, Xie J, Wang Y, Tang A, Li X et al (2011) Epigenetic inactivation of PCDH10 in human prostate cancer cell lines. Cell Biol Int 35(7):671–676. https://doi.org/10.1042/CBI20100568
Li Y, Yang ZS, Song JJ, Liu Q, Chen JB (2012a) Protocadherin-10 is involved in angiogenesis and methylation correlated with multiple myeloma. Int J Mol Med 29(4):704–710. https://doi.org/10.3892/ijmm.2012.880
Li Z, Chim JC, Yang M, Ye J, Wong BC, Qiao L (2012b) Role of PCDH10 and its hypermethylation in human gastric cancer. Biochim Biophys Acta 1823(2):298–305. https://doi.org/10.1016/j.bbamcr.2011.11.011
Li Z, Yang Z, Peng X, Li Y, Liu Q, Chen J (2014) Nuclear factor-kappaB is involved in the protocadherin-10-mediated pro-apoptotic effect in multiple myeloma. Mol Med Rep 10(2):832–838. https://doi.org/10.3892/mmr.2014.2285
Light SEW, Jontes JD (2017) Delta-protocadherins: organizers of neural circuit assembly. Semin Cell Dev Biol 69:83–90. https://doi.org/10.1016/j.semcdb.2017.07.037
Lin CJ, Lai HC, Wang KH, Hsiung CA, Liu HW, Ding DC et al (2011) Testing for methylated PCDH10 or WT1 is superior to the HPV test in detecting severe neoplasms (CIN3 or greater) in the triage of ASC-US smear results. Am J Obstet Gynecol 204(1):21 e21-27. https://doi.org/10.1016/j.ajog.2010.07.036
Lin YL, Li ZG, He ZK, Guan TY, Ma JG (2012) Clinical and prognostic significance of protocadherin-10 (PCDH10) promoter methylation in bladder cancer. J Int Med Res 40(6):2117–2123. https://doi.org/10.1177/030006051204000609
Lin YL, Li ZG, Guan TY (2013) The clinical significance of PCDH10 promoter methylation in patients with bladder transitional cell carcinoma. Urol Int 90(2):219–224. https://doi.org/10.1159/000345053
Liu L, Chen F, Xiu A, Du B, Ai H, Xie W (2018a) Identification of key candidate genes and pathways in endometrial cancer by integrated bioinformatical analysis. Asian Pac J Cancer Prev 19(4):969–975. https://doi.org/10.22034/APJCP.2018.19.4.969
Liu W, Wu J, Shi G, Yue X, Liu D, Zhang Q (2018b) Aberrant promoter methylation of PCDH10 as a potential diagnostic and prognostic biomarker for patients with breast cancer. Oncol Lett 16(4):4462–4470. https://doi.org/10.3892/ol.2018.9214
Morrow EM, Yoo SY, Flavell SW, Kim TK, Lin Y, Hill RS et al (2008) Identifying autism loci and genes by tracing recent shared ancestry. Science 321(5886):218–223. https://doi.org/10.1126/science.1157657
Nakao S, Platek A, Hirano S, Takeichi M (2008) Contact-dependent promotion of cell migration by the OL-protocadherin-Nap1 interaction. J Cell Biol 182(2):395–410. https://doi.org/10.1083/jcb.200802069
Narayan G, Scotto L, Neelakantan V, Kottoor SH, Wong AH, Loke SL et al (2009) Protocadherin PCDH10, involved in tumor progression, is a frequent and early target of promoter hypermethylation in cervical cancer. Genes Chromosomes Cancer 48(11):983–992. https://doi.org/10.1002/gcc.20703
Narayan G, Xie D, Freddy AJ, Ishdorj G, Do C, Satwani P et al (2013) PCDH10 promoter hypermethylation is frequent in most histologic subtypes of mature lymphoid malignancies and occurs early in lymphomagenesis. Genes Chromosomes Cancer 52(11):1030–1041. https://doi.org/10.1002/gcc.22098
Pancho A, Aerts T, Mitsogiannis MD, Seuntjens E (2020) Protocadherins at the crossroad of signaling pathways. Front Mol Neurosci 13:117. https://doi.org/10.3389/fnmol.2020.00117
Pimson C, Ekalaksananan T, Pientong C, Promthet S, Putthanachote N, Suwanrungruang K et al (2016) Aberrant methylation of PCDH10 and RASSF1A genes in blood samples for non-invasive diagnosis and prognostic assessment of gastric cancer. PeerJ 4:e2112. https://doi.org/10.7717/peerj.2112
Port RG, Gajewski C, Krizman E, Dow HC, Hirano S, Brodkin ES et al (2017) Protocadherin 10 alters gamma oscillations, amino acid levels, and their coupling; baclofen partially restores these oscillatory deficits. Neurobiol Dis 108:324–338. https://doi.org/10.1016/j.nbd.2017.08.013
Qi Y, Ooi HS, Wu J, Chen J, Zhang X, Tan S et al (2016) MALAT1 long ncRNA promotes gastric cancer metastasis by suppressing PCDH10. Oncotarget 7(11):12693–12703. https://doi.org/10.18632/oncotarget.7281
Qin H, Samuels JF, Wang Y, Zhu Y, Grados MA, Riddle MA et al (2016) Whole-genome association analysis of treatment response in obsessive-compulsive disorder. Mol Psychiatry 21(2):270–276. https://doi.org/10.1038/mp.2015.32
Qiu C, Bu X, Jiang Z (2016) Protocadherin-10 acts as a tumor suppressor gene, and is frequently downregulated by promoter methylation in pancreatic cancer cells. Oncol Rep 36(1):383–389. https://doi.org/10.3892/or.2016.4793
Roberson-Nay R, Lapato DM, Wolen AR, Lancaster EE, Webb BT, Verhulst B et al (2020) An epigenome-wide association study of early-onset major depression in monozygotic twins. Transl Psychiatry 10(1):301. https://doi.org/10.1038/s41398-020-00984-2
Salmeron-Barcenas EG, Illades-Aguiar B, Del Moral-Hernandez O, Ortega-Soto A, Hernandez-Sotelo D (2019) HOTAIR knockdown decreased the activity Wnt/beta-catenin signaling pathway and increased the mRNA levels of its negative regulators in hela cells. Cell Physiol Biochem 53(6):948–960. https://doi.org/10.33594/000000188
Schneider BG, Mera R, Piazuelo MB, Bravo JC, Zabaleta J, Delgado AG et al (2015) DNA methylation predicts progression of human gastric lesions. Cancer Epidemiol Biomark Prev 24(10):1607–1613. https://doi.org/10.1158/1055-9965.EPI-15-0388
Schoch H, Kreibich AS, Ferri SL, White RS, Bohorquez D, Banerjee A et al (2017) Sociability deficits and altered amygdala circuits in mice lacking Pcdh10, an autism associated gene. Biol Psychiatry 81(3):193–202. https://doi.org/10.1016/j.biopsych.2016.06.008
Seo SI, Yoon JH, Byun HJ, Lee SK (2021) HOTAIR induces methylation of PCDH10, a tumor suppressor gene, by regulating DNMT1 and sponging with miR-148b in gastric adenocarcinoma. Yonsei Med J 62(2):118–128. https://doi.org/10.3349/ymj.2021.62.2.118
Silva TD, Vidigal VM, Felipe AV, Jacqueline Miranda DEL, Neto RA, Saad SS et al (2013) DNA methylation as an epigenetic biomarker in colorectal cancer. Oncol Lett 6(6):1687–1692. https://doi.org/10.3892/ol.2013.1606
Skuja E, Butane D, Nakazawa-Miklasevica M, Daneberga Z, Purkalne G, Miklasevics E (2019) Deletions in metastatic colorectal cancer with chromothripsis. Exp Oncol 41(4):323–327. https://doi.org/10.32471/exp-oncology.2312-8852.vol-41-no-4.13841
Tanaka N, Huttenhower C, Nosho K, Baba Y, Shima K, Quackenbush J et al (2010) Novel application of structural equation modeling to correlation structure analysis of CpG island methylation in colorectal cancer. Am J Pathol 177(6):2731–2740. https://doi.org/10.2353/ajpath.2010.100361
Tang X, Yin X, Xiang T, Li H, Li F, Chen L et al (2012) Protocadherin 10 is frequently downregulated by promoter methylation and functions as a tumor suppressor gene in non-small cell lung cancer. Cancer Biomark 12(1):11–19. https://doi.org/10.3233/CBM-2012-00280
Tang J, Chen X, Cai B, Chen G (2019) A logical relationship for schizophrenia, bipolar, and major depressive disorder. Part 4: evidence from chromosome 4 high-density association screen. J Comp Neurol 527(2):392–405. https://doi.org/10.1002/cne.24543
Tsai NP, Wilkerson JR, Guo W, Maksimova MA, DeMartino GN, Cowan CW et al (2012) Multiple autism-linked genes mediate synapse elimination via proteasomal degradation of a synaptic scaffold PSD-95. Cell 151(7):1581–1594. https://doi.org/10.1016/j.cell.2012.11.040
Uemura M, Nakao S, Suzuki ST, Takeichi M, Hirano S (2007) OL-protocadherin is essential for growth of striatal axons and thalamocortical projections. Nat Neurosci 10(9):1151–1159. https://doi.org/10.1038/nn1960
Wang KH, Liu HW, Lin SR, Ding DC, Chu TY (2009) Field methylation silencing of the protocadherin 10 gene in cervical carcinogenesis as a potential specific diagnostic test from cervical scrapings. Cancer Sci 100(11):2175–2180. https://doi.org/10.1111/j.1349-7006.2009.01285.x
Wang L, Xie PG, Lin YL, Ma JG, Li WP (2014) Aberrant methylation of PCDH10 predicts worse biochemical recurrence-free survival in patients with prostate cancer after radical prostatectomy. Med Sci Monit 20:1363–1368. https://doi.org/10.12659/MSM.891241
Wang X, Dou N, Wang J, Zhang Y, Li Y, Gao Y (2021) FOXM1-induced miR-552 expression contributes to pancreatic cancer progression by targeting multiple tumor suppressor genes. Int J Biol Sci 17(4):915–925. https://doi.org/10.7150/ijbs.56733
Williams EO, Sickles HM, Dooley AL, Palumbos S, Bisogni AJ, Lin DM (2011) Delta protocadherin 10 is regulated by activity in the mouse main olfactory system. Front Neural Circ 5:9. https://doi.org/10.3389/fncir.2011.00009
Wolverton T, Lalande M (2001) Identification and characterization of three members of a novel subclass of protocadherins. Genomics 76(1–3):66–72. https://doi.org/10.1006/geno.2001.6592
Xiaolei T, Jiang M, Yang N, Jing Z (2022) Effects of EZH2 on invasion and migration of endometrial stromal cells in endometriosis patients by regulating PCDH10 gene H3K27 methylation. Altern Ther Health Med 29:42–49
Xu Y, Yang Z, Yuan H, Li Z, Li Y, Liu Q et al (2015) PCDH10 inhibits cell proliferation of multiple myeloma via the negative regulation of the Wnt/beta-catenin/BCL-9 signaling pathway. Oncol Rep 34(2):747–754. https://doi.org/10.3892/or.2015.4056
Xu M, Liu C, Pu L, Lai J, Li J, Ning Q et al (2021) Systemic analysis of the expression levels and prognosis of breast cancer-related cadherins. Exp Biol Med (maywood) 246(15):1706–1720. https://doi.org/10.1177/15353702211010417
Yang Y, Jiang Y, Jiang M, Zhang J, Yang B, She Y et al (2016) Protocadherin 10 inhibits cell proliferation and induces apoptosis via regulation of DEP domain containing 1 in endometrial endometrioid carcinoma. Exp Mol Pathol 100(2):344–352. https://doi.org/10.1016/j.yexmp.2016.03.002
Yang J, Hou C, Wang H, Perez EA, Do-Umehara HC, Dong H et al (2022) Miz1 promotes KRAS-driven lung tumorigenesis by repressing the protocadherin Pcdh10. Cancer Lett 555:216025. https://doi.org/10.1016/j.canlet.2022.216025
Ye M, Li J, Gong J (2017) PCDH10 gene inhibits cell proliferation and induces cell apoptosis by inhibiting the PI3K/Akt signaling pathway in hepatocellular carcinoma cells. Oncol Rep 37(6):3167–3174. https://doi.org/10.3892/or.2017.5630
Ying J, Li H, Seng TJ, Langford C, Srivastava G, Tsao SW et al (2006) Functional epigenetics identifies a protocadherin PCDH10 as a candidate tumor suppressor for nasopharyngeal, esophageal and multiple other carcinomas with frequent methylation. Oncogene 25(7):1070–1080. https://doi.org/10.1038/sj.onc.1209154
Yu J, Cheng YY, Tao Q, Cheung KF, Lam CN, Geng H et al (2009) Methylation of protocadherin 10, a novel tumor suppressor, is associated with poor prognosis in patients with gastric cancer. Gastroenterology 136(2):640-651 e641. https://doi.org/10.1053/j.gastro.2008.10.050
Yu B, Yang H, Zhang C, Wu Q, Shao Y, Zhang J et al (2010) High-resolution melting analysis of PCDH10 methylation levels in gastric, colorectal and pancreatic cancers. Neoplasma 57(3):247–252. https://doi.org/10.4149/neo_2010_03_247
Zhang S, Qiu M, Gao S, Tian T (2021) Circular RNA PCDH10 regulates the tumorigenesis of pancreatic cancer through the miR-338-3p/hTERT axis. Am J Transl Res 13(4):2181–2197
Zhao Y, Yang Y, Trovik J, Sun K, Zhou L, Jiang P et al (2014) A novel wnt regulatory axis in endometrioid endometrial cancer. Cancer Res 74(18):5103–5117. https://doi.org/10.1158/0008-5472.CAN-14-0427
Zhao Y, Yang Y, Trovik J, Sun K, Zhou L, Jiang P et al (2019) Novel PCDH10-Wnt-MALAT1 regulatory axis in endometrioid endometrial adenocarcinoma. Hong Kong Med J 25(Suppl 7(5)):17–22
Zhong X, Zhu Y, Mao J, Zhang J, Zheng S (2013) Frequent epigenetic silencing of PCDH10 by methylation in human colorectal cancer. J Cancer Res Clin Oncol 139(3):485–490. https://doi.org/10.1007/s00432-012-1353-5
Zhong X, Shen H, Mao J, Zhang J, Han W (2017) Epigenetic silencing of protocadherin 10 in colorectal cancer. Oncol Lett 13(4):2449–2453. https://doi.org/10.3892/ol.2017.5733
Zhou LN, Hua X, Deng WQ, Wu QN, Mei H, Chen B (2015) PCDH10 interacts with hTERT and negatively regulates telomerase activity. Medicine (Baltimore) 94(50):e2230. https://doi.org/10.1097/MD.0000000000002230
Zhou J, Wang L, Sun Q, Chen R, Zhang C, Yang P et al (2021) Hsa_circ_0001666 suppresses the progression of colorectal cancer through the miR-576–5p/PCDH10 axis. Clin Transl Med 11(11):e565. https://doi.org/10.1002/ctm2.565
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant no. 82200668) and Natural Science Foundation of the Anhui Education Department (KJ2021A0281).
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions.
Author information
Authors and Affiliations
Contributions
YZ drafted the manuscript. MP revised and finalized the review. XL provided supervision and obtained the funding.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This manuscript is a review article and does not involve a research protocol requiring approval by the relevant institutional review board or ethics committee.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhen, Y., Pavez, M. & Li, X. The role of Pcdh10 in neurological disease and cancer. J Cancer Res Clin Oncol 149, 8153–8164 (2023). https://doi.org/10.1007/s00432-023-04743-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-04743-w