Abstract
Background
Dysregulation of Long Non-coding RNAs (lncRNAs) emerges to be a hallmark of cancers. Metastatic prostate cancer and localized disease that recurs after treatment are clinical challenges, it remains unclear how lncRNA plays a role in those processes.
Methods
From previous RNA-Seq data on 65 prostate cancer and adjacent normal tissues. We identified a novel lncRNA ENST00000503625 down-regulated in prostate cancer and correlated with tumor progression characteristics. Public datasets were examined for associations between ENST00000503625 expression and clinical parameters and prognoses. Subsequently, we constructed and externally validated a nomogram for predicting biochemical recurrence (BCR). Finally, in vitro experiments were carried out to determine how ENST00000503625 functions biologically in prostate cancer.
Results
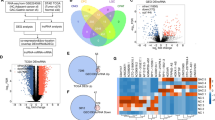
Low ENST00000503625 in tumor was associated with poor clinical features and prognoses. TCGA pan-cancer analysis found that ENST00000503625 was deregulated in a variety of tumors and correlated with overall survival, disease-specific survival, and progression-free survival. The nomogram for predicting BCR was constructed using TCGA data, which exhibited excellent accuracy in external validation with Chinese Prostate Cancer Genome and Epigenome Atlas data. Gene Ontology and KEGG pathway analysis found that genes related to ENST00000503625 were enriched in multiple tumor progression related pathways. When ENST00000503625 was knocked down in vitro, the epithelial-mesenchymal transition was induced, by which cancer cells migrated and invaded more readily.
Conclusion
Our data suggested that ENST00000503625 may serve as a potential prognostic marker or a therapeutic target for prostate cancer metastases.






Similar content being viewed by others
Data availability
Data supporting the findings of this study are available on the official websites of TCGA, GEO, CPGEA, and GTEx.
References
Batista PJ, Chang HY (2013) Long noncoding RNAs: cellular address codes in development and disease. Cell 152:1298–1307. https://doi.org/10.1016/j.cell.2013.02.012
Bhan A, Soleimani M, Mandal SS (2017) Long noncoding RNA and cancer: a new paradigm. Cancer Res 77:3965–3981. https://doi.org/10.1158/0008-5472.CAN-16-2634
Bilusic M, Einstein DJ, Karzai FH et al (2020) The potential role for immunotherapy in biochemically recurrent prostate cancer. Urol Clin North Am 47:457–467. https://doi.org/10.1016/j.ucl.2020.07.004
Bridges MC, Daulagala, AC, Kourtidis, A (2021) Lnccation: lncrna localization and function. J Cell Biol 220: e202009045. https://doi.org/10.1083/jcb.202009045
Cooperberg MR, Pasta DJ, Elkin EP et al (2005) The University of California, san Francisco cancer of the prostate risk assessment score: a straightforward and reliable preoperative predictor of disease recurrence after radical prostatectomy. J Urol 173:1938–1942. https://doi.org/10.1097/01.ju.0000158155.33890.e7
Denham JW, Steigler A, Wilcox C et al (2008) Time to biochemical failure and prostate-specific antigen doubling time as surrogates for prostate cancer- specific mortality: evidence from the TROG 96.01 randomised controlled trial. Lancet Oncol 9:1058–1068. https://doi.org/10.1016/S14702045(08)70236-5
Derrien T, Johnson R, Bussotti G et al (2012) The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res 22:1775–1789. https://doi.org/10.1101/gr.132159.111
Hessels D, Schalken JA (2009) The use of PCA3 in the diagnosis of prostate cancer. Nat Rev Urol 6:255–261. https://doi.org/10.1038/nrurol.2009.40
Jathar S, Kumar V, Srivastava J, Tripathi V (2017) Technological developments in lncRNA biology. Adv Exp Med Biol 1008:283–323. https://doi.org/10.1007/978-981-10-5203-3_10
Kidd SG, Carm KT, Bogaard M et al (2021) High expression of SCHLAP1 in primary prostate cancer is an independent predictor of biochemical recurrence, despite substantial heterogeneity. Neoplasia 23:634–641. https://doi.org/10.1016/j.neo.2021.05.012
Li X, Wu Z, Fu X, Han W (2013) Long noncoding RNAs: insights from biological features and functions to diseases. Med Res Rev 33:517–553. https://doi.org/10.1002/med.21254
Lim YWS, Xiang X, Garg M et al (2021) The double-edged sword of H19 lncRNA: Insights into cancer therapy. Cancer Lett 500:253–262. https://doi.org/10.1016/j.canlet.2020.11.006
Mehra R, Udager AM, Ahearn TU et al (2016) Overexpression of the long non-coding RNA SChLAP1 independently predicts lethal prostate cancer. Euro Urol 70:549–552. https://doi.org/10.1016/j.eururo.2015.12.003
Pound CR, Partin AW, Eisenberger MA, Chan DW, Pearson JD, Walsh PC (1999) Natural history of progression after PSA elevation following radical prostatectomy. JAMA 281:1591–1597. https://doi.org/10.1001/jama.281.17.1591
Prensner JR, Iyer MK, Balbin OA et al (2011) Transcriptome sequencing across a prostate cancer cohort identifies PCAT-1, an unannotated lincRNA implicated in disease progression. Nat Biotechnol 29:742–749. https://doi.org/10.1038/nbt.1914
Prensner JR, Iyer MK, Sahu A et al (2013) The long noncoding RNA SChLAP1 promotes aggressive prostate cancer and antagonizes the SWI/SNF complex. Nat Genet 45:1392–1398. https://doi.org/10.1038/ng.2771
Prensner JR, Zhao S, Erho N et al (2014) RNA biomarkers associated with metastatic progression in prostate cancer: a multi-institutional high-throughput analysis of SChLAP1. Lancet Oncol 15:1469–1480. https://doi.org/10.1016/S1470-2045(14)71113-1
Ren S, Wang F, Shen J et al (2013) Long non-coding RNA metastasis associated in lung adenocarcinoma transcript 1 derived miniRNA as a novel plasma-based biomarker for diagnosing prostate cancer. Eur J Cancer 49:2949–2959. https://doi.org/10.1016/j.ejca.2013.04.026
Ren S, Wei GH, Liu D et al (2018) Whole-genome and transcriptome sequencing of prostate cancer identify new genetic alterations driving disease progression. Eur Urol 73:322–339. https://doi.org/10.1016/j.eururo.2017.08.027
Roehl KA, Han M, Ramos CG, Antenor JAV, Catalona WJ (2004) Cancer progression and survival rates following anatomical radical retropubic prostatectomy in 3,478 consecutive patients: long-term results. J Urol 172:910–914. https://doi.org/10.1097/01.ju.0000134888.22332.bb
Shi X, Zhang W, Nian X et al (2020) The previously uncharacterized lncRNA APP promotes prostate cancer progression by acting as a competing endogenous RNA. Int J Cancer 146:475–486. https://doi.org/10.1002/ijc.32422
Siegel RL, Miller KD, Fuchs HE, Jemal A (2022) Cancer statistics, 2022. CA Cancer J Clin 72:7–33. https://doi.org/10.3322/caac.21708
Singh N, Ramnarine VR, Song JH et al (2021) The long noncoding RNA H19 regulates tumor plasticity in neuroendocrine prostate cancer. Nat Commun 12:1–20. https://doi.org/10.1038/s41467-021-26901-9
Sung H, Ferlay J, Siegel RL et al (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249. https://doi.org/10.3322/caac.21660
Tilki D, Preisser F, Graefen M, Huland H, Pompe RS (2019) External validation of the European Association of Urology biochemical recurrence risk groups to predict metastasis and mortality after radical prostatectomy in a European cohort. Eur Urol 75:896–900. https://doi.org/10.1016/j.eururo.2019.03.016
Van den Broeck T, van den Bergh R, Arfi N et al (2019) Prognostic value of biochemical recurrence following treatment with curative intent for prostate cancer: a systematic review. Eur Urol 75:967–987. https://doi.org/10.1016/j.eururo.2018.10.011
Wen S, Wei Y, Zen C, Xiong W, Niu Y, Zhao Y (2020) Long non-coding RNA NEAT1 promotes bone metastasis of prostate cancer through N6-methyladenosine. Mol Cancer 19:1–18. https://doi.org/10.1186/s12943-020-01293-4
Zhang W, Shi X, Chen R et al (2020) Novel long non-coding RNA lncAMPC promotes metastasis and immunosuppression in prostate cancer by stimulating LIF/LIFR expression. Mol Ther 28:2473–2487. https://doi.org/10.1016/j.ymthe.2020.06.013
Zhao L, Wang J, Li Y et al (2021) NONCODEV6:an updated database dedicated to long non-coding RNA annotation in both animals and plants. Nucleic Acids Res 49:D165–D171. https://doi.org/10.1093/nar/gkaa1046
Funding
This work was supported by Natural Science Foundation of Chongqing (cstc2018jcyjAX0666), National Natural Science Foundation of China (81902600), and National Natural Science Foundation of China (81800624).
Author information
Authors and Affiliations
Contributions
YL, SR, and LW conceived and designed the study. ZF and JL contributed to analysis and interpretation of data. XS, WZ, and LQ contributed to technical support. YL, ZF, and SG wrote the paper. YS and SR provided experimental samples and techniques. LW revised the manuscript. The study was supervised by SR and LW.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Ethics approval
The use of human tissue samples in the study was approved by the Clinical Research Ethics Committee of Changhai Hospital. All subjects in the study provided informed consent. The patients have given their written informed consent to publish this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, Y., Fang, Z., Ge, S. et al. Long non-coding RNA ENST00000503625 is a potential prognostic biomarker and metastasis suppressor gene in prostate cancer. J Cancer Res Clin Oncol 149, 7305–7317 (2023). https://doi.org/10.1007/s00432-023-04676-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-04676-4