Abstract
Purpose
The current evaluation methods for tumor infiltrating lymphocytes (TILs), particularly CD8 + TILs, mainly rely on semiquantitative immunohistochemistry with high variability. We aimed to construct an individualized DNA methylation-based signature for CD8 + TILs (CD8 + MeTIL) that may characterize melanoma immune microenvironment and guide therapeutic selection.
Methods
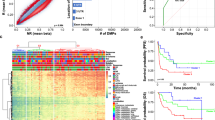
The transcriptome profiles and DNA methylation data of 457 melanoma patients from The Cancer Genome Atlas (TCGA) database were analyzed. Differential methylation analysis between groups with high and low CD8 + TILs was performed to select differentially methylated positions (DMPs) and define CD8 + MeTIL. The prognostic value of CD8 + MeTIL and its predictive value for immunotherapy response were investigated using multiple melanoma cohorts.
Results
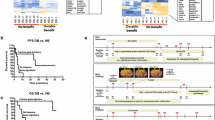
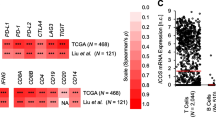
We successfully constructed the CD8 + MeTIL signature based on four DMPs. The survival analyses showed that higher CD8 + MeTIL score was associated with worse survival outcomes in TCGA-SKCM and GSE144487 cohorts. The ROC curve for the predictive analysis revealed that the survival prediction of CD8 + MeTIL score was superior compared with CD8 + TILs (CIBERSORT) and CD8B mRNA expression. Furthermore, we founded that tumors with higher CD8 + MeTIL score were marked with immunosuppressive characteristics, including low immune score and downregulated immune-related pathways. More importantly, the CD8 + MeTIL score showed a potential predictive value for the benefit from immunotherapy in two published cohorts. When combined CD8 + MeTIL with PD-L1 expression, the patient classification showed significantly different immunotherapy response rates and long-term survival outcomes.
Conclusions
The CD8 + MeTIL signature might be as a novel method to evaluate CD8 + TILs and guide immunotherapy approaches.







Similar content being viewed by others
Data availability
The data of TCGA-SKCM cohort were downloaded from the UCSC Xena database (http://xena.ucsc.edu/) and cbioPortal (http://www.cbioportal.org). The data of GSE144487 cohort were downloaded from the Gene Expression Omnibus (GEO) database (http://www.ncbi.nlm.nih.gov/geo/). The data of Jung19 (GSE119144 and GSE135222) and Cho20 (GSE126043 and GSE126044) were downloaded from the GEO database (http://www.ncbi.nlm.nih.gov/geo/). All patients’ data analyzed from published studies are ref-erenced to and publicly available accordingly.
References
Accomando WP, Wiencke JK, Houseman EA, Nelson HH, Kelsey KT (2014) Quantitative reconstruction of leukocyte subsets using DNA methylation. Genome Biol 15(3):R50. https://doi.org/10.1186/gb-2014-15-3-r50
Ayers M, Lunceford J, Nebozhyn M, Murphy E, Loboda A, Kaufman DR et al (2017) IFN-γ-related mRNA profile predicts clinical response to PD-1 blockade. J Clin Invest 127(8):2930–2940. https://doi.org/10.1172/JCI91190
Becht E, Giraldo NA, Lacroix L, Buttard B, Elarouci N, Petitprez F et al (2016) Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol 17(1):218. https://doi.org/10.1186/s13059-016-1070-5
Carlino MS, Larkin J, Long GV (2021) Immune checkpoint inhibitors in melanoma. Lancet 398(10304):1002–1014. https://doi.org/10.1016/S0140-6736(21)01206-X
Chen B, Khodadoust MS, Liu CL, Newman AM, Alizadeh AA (2018) Profiling tumor infiltrating immune cells with CIBERSORT. Methods Mol Biol 17(11):243–259. https://doi.org/10.1007/978-1-4939-7493-1_12
Cho JW, Hong MH, Ha SJ, Kim YJ, Cho BC, Lee I et al (2020) Genome-wide identification of differentially methylated promoters and enhancers associated with response to anti-PD-1 therapy in non-small cell lung cancer. Exp Mol Med 52(9):1550–1563. https://doi.org/10.1038/s12276-020-00493-8
Curti BD, Faries MB (2021) Recent advances in the treatment of melanoma. N Engl J Med 384(23):2229–2240. https://doi.org/10.1056/NEJMra2034861
Doi A, Park IH, Wen B, Murakami P, Aryee MJ, Irizarry R et al (2009) Differential methylation of tissue- and cancer-specific CpG island shores distinguishes human induced pluripotent stem cells, embryonic stem cells and fibroblasts. Nat Genet 41(12):1350–1353. https://doi.org/10.1038/ng.471
Eggermont A, Blank CU, Mandala M, Long GV, Atkinson V, Dalle S et al (2018) Adjuvant pembrolizumab versus placebo in resected stage III melanoma. N Engl J Med 378(19):1789–1801. https://doi.org/10.1056/NEJMoa1802357
Farlik M, Halbritter F, Müller F, Choudry FA, Ebert P, Klughammer J et al (2016) DNA methylation dynamics of human hematopoietic stem cell differentiation. Cell Stem Cell 19(6):808–822. https://doi.org/10.1016/j.stem.2016.10.019
Fecher LA, Cummings SD, Keefe MJ, Alani RM (2007) Toward a molecular classification of melanoma. J Clin Oncol 25(12):1606–1620. https://doi.org/10.1200/JCO.2006.06.0442
Fu C, Jiang A (2018) Dendritic cells and CD8 T Cell immunity in tumor microenvironment. Front Immunol 9:3059. https://doi.org/10.3389/fimmu.2018.03059
Galon J, Costes A, Sanchez-Cabo F, Kirilovsky A, Mlecnik B, Lagorce-Pagès C et al (2006) Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science 313(5795):1960–1964. https://doi.org/10.1126/science.1129139
Grasso CS, Tsoi J, Onyshchenko M, Abril-Rodriguez G, Ross-Macdonald P, Wind-Rotolo M et al (2021) Conserved interferon-γ signaling drives clinical response to immune checkpoint blockade therapy in melanoma. Cancer Cell 39(1):122. https://doi.org/10.1016/j.ccell.2020.11.015
Hamid O, Robert C, Daud A, Hodi FS, Hwu WJ, Kefford R et al (2013) Safety and tumor responses with lambrolizumab (anti-PD-1) in melanoma. N Engl J Med 369(2):134–144. https://doi.org/10.1056/NEJMoa1305133
Horn L, Spigel DR, Vokes EE, Holgado E, Ready N, Steins M et al (2017) Nivolumab versus docetaxel in previously treated patients with advanced non-small-cell lung cancer: two-year outcomes from two randomized, open-label, phase III trials (CheckMate 017 and CheckMate 057). J Clin Oncol 35(35):3924–3933. https://doi.org/10.1200/JCO.2017.74.3062
Hou P, Bao S, Fan D, Yan C, Su J, Qu J et al (2021) Machine learning-based integrative analysis of methylome and transcriptome identifies novel prognostic DNA methylation signature in uveal melanoma. Brief Bioinform. https://doi.org/10.1093/bib/bbaa371
Houseman EA, Accomando WP, Koestler DC, Christensen BC, Marsit CJ, Nelson HH et al (2012) DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinform 13:86. https://doi.org/10.1186/1471-2105-13-86
Irizarry RA, Ladd-Acosta C, Wen B, Wu Z, Montano C, Onyango P et al (2009) The human colon cancer methylome shows similar hypo- and hypermethylation at conserved tissue-specific CpG island shores. Nat Genet 41(2):178–186. https://doi.org/10.1038/ng.298
Jeschke J, Collignon E, Fuks F (2015) DNA methylome profiling beyond promoters—taking an epigenetic snapshot of the breast tumor microenvironment. FEBS J 282(9):1801–1814. https://doi.org/10.1111/febs.13125
Jeschke J, Bizet M, Desmedt C, Calonne E, Dedeurwaerder S, Garaud S et al (2017) DNA methylation-based immune response signature improves patient diagnosis in multiple cancers. J Clin Invest 127(8):3090–3102. https://doi.org/10.1172/JCI91095
Jung H, Kim HS, Kim JY, Sun JM, Ahn JS, Ahn MJ et al (2019) DNA methylation loss promotes immune evasion of tumours with high mutation and copy number load. Nat Commun 10(1):4278. https://doi.org/10.1038/s41467-019-12159-9
Kandoth C, McLellan MD, Vandin F, Ye K, Niu B, Lu C et al (2013) Mutational landscape and significance across 12 major cancer types. Nature 502(7471):333–339. https://doi.org/10.1038/nature12634
Kim TK, Vandsemb EN, Herbst RS, Chen L (2022) Adaptive immune resistance at the tumour site: mechanisms and therapeutic opportunities. Nat Rev Drug Discov 21(7):529–540. https://doi.org/10.1038/s41573-022-00493-5
Koestler DC, Christensen B, Karagas MR, Marsit CJ, Langevin SM, Kelsey KT et al (2013) Blood-based profiles of DNA methylation predict the underlying distribution of cell types: a validation analysis. Epigenetics 8(8):816–826. https://doi.org/10.4161/epi.25430
Lin H, Wei S, Hurt EM, Green MD, Zhao L, Vatan L et al (2018) Host expression of PD-L1 determines efficacy of PD-L1 pathway blockade-mediated tumor regression. J Clin Invest 128(2):805–815. https://doi.org/10.1172/JCI96113
Mao L, Qi Z, Zhang L, Guo J, Si L (2021) Immunotherapy in acral and mucosal melanoma: current status and future directions. Front Immunol. https://doi.org/10.3389/fimmu.2021.680407
Pagès F, Mlecnik B, Marliot F, Bindea G, Ou FS, Bifulco C et al (2018) International validation of the consensus Immunoscore for the classification of colon cancer: a prognostic and accuracy study. Lancet 391(10135):2128–2139. https://doi.org/10.1016/S0140-6736(18)30789-X
Pan JH, Zhou H, Cooper L, Huang JL, Zhu SB, Zhao XX et al (2019) LAYN is a prognostic biomarker and correlated with immune infiltrates in gastric and colon cancers. Front Immunol 10:6. https://doi.org/10.3389/fimmu.2019.00006
Postow MA, Chesney J, Pavlick AC, Robert C, Grossmann K, McDermott D et al (2015) Nivolumab and ipilimumab versus ipilimumab in untreated melanoma. N Engl J Med 372(21):2006–2017. https://doi.org/10.1056/NEJMoa1414428
Ribas A, Puzanov I, Dummer R, Schadendorf D, Hamid O, Robert C et al (2015) Pembrolizumab versus investigator-choice chemotherapy for ipilimumab-refractory melanoma (KEYNOTE-002): a randomised, controlled, phase 2 trial. Lancet Oncol 16(8):908–918. https://doi.org/10.1016/S1470-2045(15)00083-2
Robert C, Long GV, Brady B, Dutriaux C, Maio M, Mortier L et al (2015) Nivolumab in previously untreated melanoma without BRAF mutation. N Engl J Med 372(4):320–330. https://doi.org/10.1056/NEJMoa1412082
Robert C, Ribas A, Schachter J, Arance A, Grob JJ, Mortier L et al (2019) Pembrolizumab versus ipilimumab in advanced melanoma (KEYNOTE-006): post-hoc 5-year results from an open-label, multicentre, randomised, controlled, phase 3 study. Lancet Oncol 20(9):1239–1251. https://doi.org/10.1016/S1470-2045(19)30388-2
Rozeman EA, Hoefsmit EP, Reijers I, Saw R, Versluis JM, Krijgsman O et al (2021) Survival and biomarker analyses from the OpACIN-neo and OpACIN neoadjuvant immunotherapy trials in stage III melanoma. Nat Med 27(2):256–263. https://doi.org/10.1038/s41591-020-01211-7
Salgado R, Denkert C, Demaria S, Sirtaine N, Klauschen F, Pruneri G et al (2015) The evaluation of tumor-infiltrating lymphocytes (TILs) in breast cancer: recommendations by an International TILs working group 2014. Ann Oncol 26(2):259–271. https://doi.org/10.1093/annonc/mdu450
Salhab A, Nordström K, Gasparoni G, Kattler K, Ebert P, Ramirez F et al (2018) A comprehensive analysis of 195 DNA methylomes reveals shared and cell-specific features of partially methylated domains. Genome Biol 19(1):150. https://doi.org/10.1186/s13059-018-1510-5
Stueve TR, Marconett CN, Zhou B, Borok Z, Laird-Offringa IA (2016) The importance of detailed epigenomic profiling of different cell types within organs. Epigenomics 8(6):817–829. https://doi.org/10.2217/epi-2016-0005
Sun R, Limkin EJ, Vakalopoulou M, Dercle L, Champiat S, Han SR et al (2018) A radiomics approach to assess tumour-infiltrating CD8 cells and response to anti-PD-1 or anti-PD-L1 immunotherapy: an imaging biomarker, retrospective multicohort study. Lancet Oncol 19(9):1180–1191. https://doi.org/10.1016/S1470-2045(18)30413-3
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A et al (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71(3):209–249. https://doi.org/10.3322/caac.21660
Sunshine JC, Nguyen PL, Kaunitz GJ, Cottrell TR, Berry S, Esandrio J et al (2017) PD-L1 Expression in Melanoma: a quantitative immunohistochemical antibody comparison. Clin Cancer Res 23(16):4938–4944. https://doi.org/10.1158/1078-0432.CCR-16-1821
Taube JM, Anders RA, Young GD, Xu H, Sharma R, McMiller TL et al (2012) Colocalization of inflammatory response with B7–h1 expression in human melanocytic lesions supports an adaptive resistance mechanism of immune escape. Sci Transl Med 4(127):127ra37. https://doi.org/10.1126/scitranslmed.3003689
Topalian SL, Sznol M, McDermott DF, Kluger HM, Carvajal RD, Sharfman WH et al (2014) Survival, durable tumor remission, and long-term safety in patients with advanced melanoma receiving nivolumab. J Clin Oncol 32(10):1020–1030. https://doi.org/10.1200/JCO.2013.53.0105
Tsao H, Atkins MB, Sober AJ (2004) Management of cutaneous melanoma. N Engl J Med 351(10):998–1012. https://doi.org/10.1056/NEJMra041245
Xiao M, Liang X, Yan Z, Chen J, Zhu Y, Xie Y et al (2022) A DNA-methylation-driven genes based prognostic signature reveals immune microenvironment in pancreatic cancer. Front Immunol. https://doi.org/10.3389/fimmu.2022.803962
Zhou W, Dinh HQ, Ramjan Z, Weisenberger DJ, Nicolet CM, Shen H et al (2018) DNA methylation loss in late-replicating domains is linked to mitotic cell division. Nat Genet 50(4):591–602. https://doi.org/10.1038/s41588-018-0073-4
Zito Marino F, Ascierto PA, Rossi G, Staibano S, Montella M, Russo D et al (2017) Are tumor-infiltrating lymphocytes protagonists or background actors in patient selection for cancer immunotherapy. Expert Opin Biol Ther 17(6):735–746. https://doi.org/10.1080/14712598.2017.1309387
Zou Q, Wang X, Ren D, Hu B, Tang G, Zhang Y et al (2021) DNA methylation-based signature of CD8+ tumor-infiltrating lymphocytes enables evaluation of immune response and prognosis in colorectal cancer. J Immunother Cancer. https://doi.org/10.1136/jitc-2021-002671
Funding
This work was supported by grants from National Natural Science Foundation of China (82002906 and 81902789) and CSCO-Roche Cancer Research Fund 2019 (Y-Roche2019/2-0028).
Author information
Authors and Affiliations
Contributions
SC and JY contributed to the study conception and design. Material preparation, data collection and analysis were performed by JY, XW, and YZ. The first draft of the manuscript was written by JY and XW. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All authors have no conflicts of interest to declare.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yan, J., Wu, X., Zhu, Y. et al. Genome-wide DNA methylation profile analysis identifies an individualized predictive signature for melanoma immune response. J Cancer Res Clin Oncol 149, 343–356 (2023). https://doi.org/10.1007/s00432-022-04566-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-022-04566-1