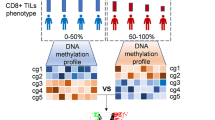
Abstract
Background
DNA methylation profiles have emerged as potential predictors of therapeutic response in various solid tumors.
Objective
This study aimed to analyze the DNA methylation profiles of patients with stage IV metastatic melanoma undergoing first-line immune checkpoint inhibitor treatment and evaluate their correlation with a radiological response according to immune-related Response Evaluation Criteria in Solid Tumors (iRECIST).
Methods
A total of 81 tissue samples from 71 patients with metastatic melanoma (27 female, 44 male) were included in this study. We utilized Illumina Methylation EPIC Beadchips to retrieve their genome-wide methylation profile by interrogating >850,000 CpG sites. Clustering based on the 500 most differentially methylated genes was conducted to identify distinct methylation patterns associated with immune checkpoint inhibitor response. Results were further aligned with an independent, previously published data set.
Results
The median progression-free survival was 8.5 months (range: 0–104.1 months), and the median overall survival was 30.6 months (range: 0–104.1 months). Objective responses were observed in 29 patients (40.8%). DNA methylation profiling revealed specific signatures that correlated with radiological response to immune checkpoint inhibitors. Three distinct clusters were identified based on the methylation patterns of the 500 most differentially methylated genes. Cluster 1 (12/12) and cluster 2 (12/24) exhibited a higher proportion of responders, while cluster 3 (39/45) predominantly consisted of non-responders. In the validation data set, responders also showed more frequent hypomethylation although differences in the data sets limit the interpretation.
Conclusions
These findings suggest that DNA methylation profiling of tumor tissues might serve as a predictive biomarker for immune checkpoint inhibitor response in patients with metastatic melanoma. Further validation studies are warranted to confirm the efficiency of DNA methylation profiling as a predictive tool in the context of immunotherapy for metastatic melanoma.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Immune checkpoint inhibitors have revolutionized the treatment of metastatic melanoma, exhibiting an objective response rate that ranges from 40% to 58%. Despite this success, there remains a significant gap in our understanding of response-associated biomarkers. |
However, the discovery of distinct methylation patterns linked to treatment response holds tremendous promise in guiding personalized therapeutic approaches and enhancing clinical outcomes for patients in this particular population observing similar trends in a validation cohort. |
Here, we identified a methylation signature with a high predictive performance, characterized by an 80% sensitivity, 81% specificity, and an area under the curve value of 0.829. These findings underscore its potential as a valuable additional tool in routine clinical practice and precision medicine. |
1 Introduction
Immune checkpoint inhibitors (ICIs) have revolutionized melanoma treatment. According to the EADO/EORTC/EDF and ESMO guidelines, patients with inoperable metastatic melanoma should receive first-line ICI therapy regardless of the mutation status because of prolonged long-term disease control with ICIs over targeted therapies [11, 12]. However, half of ICI-treated patients with melanoma do not benefit from treatment, underscoring the urgent need for reliable biomarkers.
Tumor mutational burden [8, 10, 30] and tumor neoantigen burden [25] are widely discussed biomarkers. However, analysis in a clinical routine is challenging, as protocols differ among papers, cut-off values are objects of discussion, and costs are rather high. Tumor tissue methylation is a well-established diagnostic biomarker for brain tumors and sarcomas [2, 22, 24]. Methylation analysis, however, including the required laboratory set-up, can be easily established, and thereby represents a promising method for biomarker evaluation in everyday clinical use. Alterations in DNA methylation are a key feature of tumorigenesis that result in dysregulated transcriptional gene activity. However, epigenetic changes in immune cells have also been reported to contribute to response and resistance to ICI-based therapies [6]. Indeed, tumor tissue methylation was previously shown to serve as a potential biomarker for ICI response in patients with sarcoma, head and neck cancer, lung cancer, and metastatic melanoma [7, 29, 39, 40]. Here, we report that DNA methylation profiling of treatment-naïve tumor samples independent of the tumor origin from patients with metastatic melanoma could be an additional useful tool for predicting response to ICIs and emphasizes that tissue-based DNA methylation profiling should be incorporated in future biomarker research studies.
2 Methods
2.1 Patients’ and Clinical Characteristics
A total of 71 patients (27 [38.0%] female and 44 [62.0%] male) with a median age of 65 years (range: 32–84 years) were retrospectively identified from the patient records of the Department of Dermatology, Medical University of Vienna, from 2015 to 2022. The inclusion criteria were as follows: (i) age ≥18 years; (ii) stage IVM1a–IVM1d metastatic melanoma according to the American Joint Committee on Cancer, 8th Edition [20]; (iii) sufficient formalin-fixed paraffin-embedded tumor tissue material prior to ICI initiation available; (iv) first-line therapy with ICIs including monotherapy with a programmed cell death protein 1 (PD-1) inhibitor (nivolumab, pembrolizumab), monotherapy with a cytotoxic T-lymphocyte associated protein 4 inhibitor (ipilimumab), or combination therapy of a PD-1 inhibitor (nivolumab) and cytotoxic T-lymphocyte associated protein 4 inhibitor (ipilimumab) and an ICI-based clinical trial study medication; (v) administration of a minimum of one cycle of ICI; (vi) radiologically measurable disease (lesion size ≥10×10 mm) prior to ICI therapy start; and (vii) availability of at least one radiological restaging under ICI therapy or the occurrence of death before the first performed radiological restaging. Patient and clinical characteristics, including age, sex, tumor mutation status (BRAF, NRAS, cKIT, or wild type), Eastern Cooperative Oncology Group (ECOG) performance status, baseline S100 and lactate dehydrogenase, and survival data (overall survival [OS], progression-free survival [PFS]) were obtained from patient records. Radiological assessment of response was defined according to the immune-based Response Evaluation Criteria in Solid Tumors (iRECIST) [38] and was performed by an independent radiologist blinded to patient outcomes. Objective response was determined by immune complete response or immune partial response, and disease control by immune complete response, immune partial response, or immune stable disease as best achieved response. Disease progression was defined as iPD and further subclassified as immune unconfirmed progressive disease and immune confirmed progressive disease. The date of immune unconfirmed progressive disease was classified as the timepoint of progression if no subsequent restaging was followed by a clear clinical progression or the occurrence of death. The date of the first radiologically confirmed progressive disease was defined as the immune confirmed progressive disease [38]. Patients with immune complete response and immune partial response were classified as responders, and patients with immune stable disease and iPD (immune unconfirmed progressive disease or immune confirmed progressive disease) were classified as non-responders. Patients who died due to early progression and received only one cycle of ICIs before the first restaging were classified as non-responders.
2.2 Genome-Wide DNA Methylation Analysis
Tumor tissue samples from patients with melanoma before to the first cycle of ICI therapy were collected retrospectively. A total of 81 tumor tissue samples from 71 patients were included in the final methylation analysis. These were categorized into four groups: 28 lymph node metastases (34.6%), 20 organ metastases (24.7%), 24 cutaneous metastases (29.6%), and nine primary tumors (11.1%) (Table 1). Of note, in a subset of patients, 8/71 (11.3%) had more than one treatment-naïve tumor sample from different organ sites available. In detail, one patient had a lymph node and an organ metastatic lesion (12.5%), two patients had a lymph node metastatic lesion and a primary tumor tissue (25%), three patients had one lymph node and one cutaneous metastatic lesion (37.5%), one patient had a cutaneous metastatic lesion and a primary tumor tissue (12.5%), and one patient had a lymph node metastatic lesion, two cutaneous metastatic lesions, and a primary tumor tissue (12.5%) [Table S1 of the Electronic Supplementary Material (ESM)]. The 5×10-μm slides were obtained from formalin-fixed paraffin-embedded tumor tissue after macrodissection with ≥70% tumor cells, as evaluated by an independent certified dermatopathologist blinded to the patients’ clinical outcomes. Genomic DNA was extracted and treated with sodium bisulfite following standard procedures, and genome-wide DNA methylation analyses were performed using Infinium MethylationEPIC BeadChip microarrays (Illumina, San Diego, CA, USA) as previously described [39]. Raw Microarray Data (.idat files) were loaded into R (version R 4.0.4, R Foundation for Statistical Computing, Vienna, Austria) using the RnBeads package (.idat files). Single nucleotide polymorphism-associated, non-specific/cross-hybridizing, and sex chromosome-specific probes were excluded from further analysis. The SWAN algorithm was applied for data normalization [26]. Calculation of differential methylation between groups was conducted using RnBeads implementation of the limma package as well as computation of a combined rank score, which depends on the difference in mean methylation levels of two groups, the mean methylation quotient, and statistical significance. For subsequent analyses, the top 500 differentially methylated CpG sites (DMPs) were selected. To evaluate the robustness of the results, we have used a validation cohort [29]. Analyses of gene ontology (GO) were conducted using the geometh function from the missMethyl package [33] and heatmaps were generated using ClustVis [27].
2.3 Statistics
Follow-up was calculated from the start date of ICI therapy until last contact or death, whatever occurred first. Descriptive statistics were used to describe patient characteristics, such as age, sex, and mutation status. For nominally scaled variables, absolute numbers and percentages were presented, and for metric variables, the mean, standard deviation, median, minimum, and maximum were applied. The Kaplan–Meier method was used to estimate PFS and OS from the first ICI application until the occurrence of disease progression or death. Patients without disease progression were censored on the date of their last visit. Kaplan–Meier curves were compared using the log-rank test and Cox’ proportional hazards models. To test the sensitivity-specificity trade-off, receiver operating characteristic classifier curve analyses were performed. The Kruskal–Wallis test and Mann–Whitney U test were used for comparisons between the groups. Statistical significance was determined by a p-value <0.05 (two-sided), multiple testing was corrected using the Bonferroni method. Statistical analyses were performed with Statistical Package for the Social Sciences (SPSS®) 23.0 software (SPSS Inc., Chicago, IL, USA) and standard R functions (version R 4.0.4; R Foundation for Statistical Computing, Vienna, Austria). The data cut-off date was 30 August, 2022.
3 Results
3.1 DNA Methylation Signatures Correlated with Response to ICI Therapy
To investigate whether DNA methylation signatures can predict the response to ICIs, we performed Infinium MethylationEPIC microarray analyses of tumor specimens collected prior to therapy onset. Eighty-one treatment-naïve tumor tissue samples derived from 71 patients with melanoma were included in the analysis, of which 29 (40.8%) were classified as responders and 42 (59.2%) were non-responders. After quality control and exclusion of 192,085 probes owing to cross-reactivity, unreliable measurements, and sex chromosome specificity, 674,810 probes remained for a further statistical analysis. Testing for methylation differences between responders and non-responders of all tumor samples was performed using RnBeads. This resulted in a total of 58,831 DMPs with a false discovery rate <0.05 (Fig. 1a). Differentially methylated CpG sites were predominantly located within intergenic regions, gene bodies, and transcriptional start sites (Fig. 1b). Based on the top 500 DMPs, three clusters were identified revealing that hypomethylation mainly corresponded to the response to ICIs. Cluster 1 only consisted of responders (12/12), cluster 2 was evenly distributed between responders and non-responders (12/24), and cluster 3 contained mainly non-responders (39/45) (Fig. 1c). Patients in cluster 1 (12/12) and cluster 2 (23/24) had predominantly an ECOG performance status of zero, in contrast to cluster 3, where 7/45 patients had an ECOG performance status ≥1. The baseline lactate dehydrogenase (LDH) levels of most patients in cluster 1 (11/12) and cluster 2 (18/24) had baseline LDH levels within the normal range, and patients in cluster 3 had a comparably high number of patients with baseline LDH levels ≥1 upper limit of normal (13/45). A further exploratory analysis revealed that non-responders compared with responders showed a trend towards a higher ECOG status ≥1 (p = 0.0887), albeit not significant, and no differences between non-responders and responders in baseline LDH (p = 0.6646) and S100 levels (p = 0.7500) were observed. The other clusters did not differ according to factors such as the BRAF mutation status immunotherapy (IO) regimen, sex, localization of the utilized tumor tissue, mutation status, and baseline LDH and S100 values (Fig. 1c).
DNA methylation-based prediction of response to immune checkpoint inhibitors in metastatic melanoma. A Scatter plot demonstrating the differences of DNA methylation signatures between responders (R) and non-responders (NR) to immunotherapy (IO). Red dots show differentially methylated CpG sites (DMPs). B The bar plot illustrates the locations of DMPs: intergenic, gene body, transcriptional start site (TSS), 5′ untranslated region (UTR), 3′ UTR, exon 1 and exon boundary. C Heatmap demonstrating three clusters based on the top 500 DMPs. Best overall response (iBOR); immune complete response (iCR); immune partial response (iPR); immune stable disease (iSD); immune progressive disease (iPD); immune unconfirmed progressive disease (iUPD); immune confirmed progressive disease (iCPD); including detailed information about IO scheme, sex, localization, mutation status (BRAF, cKIT, NRAS, wild type [wt]), Eastern Cooperative Oncology Group (ECOG) performance status, lactate dehydrogenase (LDH) and S100; not applicable (NA); beta values with a range from 0 (blue) to 1 (red) are shown. D Receiver operating characteristic (ROC) curve analysis presenting the sensitivity and specificity of DNA methylation signatures predicting the response to IO; area under the curve (AUC). E Kaplan–Meier curves demonstrating progression-free survival (PFS) and overall survival (OS) from patients with melanoma of each cluster indicated by different colors. Vertical lines display censored patients (statistical significance determined with alpha = 0.05, *p < 0.05, **p < 0.01, ***p < 0.001)
The predictive performance of the methylation signature was high with 80% sensitivity, 81% specificity, and an area under the curve of 0.829 (Fig. 1d). Progression-free survival and OS were significantly longer in patients from clusters 1 and 2 than in those from cluster 3 (p = 0.02 and p = 0.019, respectively) (Fig. 1e). Altogether, these results indicate that DNA methylation profiles in cutaneous metastatic melanoma correlate with the response to ICIs with high sensitivity and specificity.
3.2 Functional Characterization of Promotor Differentially Methylated Regions
A GO enrichment analysis including three sub-ontologies of biological processes (BP), cellular components (CC), and molecular functions (MF) of promotor differentially methylated regions analyzed with the missMethyl R package revealed that several pathways were significantly enriched. The most enriched pathways were related to olfactory pathways such as “detection of chemical stimulus involved in sensory perception of smell” with an enrichment false discovery rate of 6.13E−45, followed by “sensory perception of smell” and “detection of chemical stimulus involved in sensory perception” detected by GO:BP and “olfactory receptor activity” by GO:MF, which were recently demonstrated to be involved in the cell proliferation and migration processes of primary melanoma and melanoma metastasis [14]. Additionally, GO:BP and GO:MF revealed enrichments in “G protein-coupled receptor signaling pathways,” which were demonstrated to be involved in skin cancer development [32] and were shown to be engaged in melanogenesis including proliferation and migration [34]. Furthermore, GO:BP, GO:MF, and GO:CC displayed enrichments related to epidermal keratinocytes such as “keratinocyte differentiation,” “epidermis development,” “keratin filament,” and “intermediate filament cytoskeleton” (Table 2), indicative of an crosstalk between melanoma cells and the surrounding tumor microenvironment [21]. Finally, GO:BP enrichments related to B cells such as “humoral immune response” and “antimicrobial humoral immune response” were detected, which were demonstrated to promote a response to ICIs in metastatic melanoma [17].
3.3 Tissue Origin Did Not Affect the DNA Methylation-Based Prediction of an ICI Response
Next, we performed differential methylation analyses of tissues derived from different organ sites. Based on the top 500 DMPs, lymph node metastasis-only samples revealed two clusters and clearly separated non-responders (17/18) in cluster 1 and responders (9/10) in cluster 2 (Fig. 2). Differential methylation analyses performed on different subgroups of patients (cutaneous metastases, primary tumors, and organ metastases) also identified two clusters containing mainly responders and non-responders (Fig. 3). In detail, an analysis of exclusively cutaneous metastases, based on 500 DMPs, identified two clusters, with the majority of responders in cluster 1 (9/10) and non-responders in cluster 2 (12/13) (Fig. 3a). A differential methylation analysis of only primary tumors, based on 500 DMPs, revealed a clear separation of non-responders (10/10) and responders (2/2); however, interpretation was limited by the low number of responders in this cohort (Fig. 3b). A differential analysis of the subset of organ metastases based on 500 mostly DMPs identified two clusters, with responders being mainly in cluster 1 (6/6) and non-responders preferentially in cluster 2 (10/12) (Fig. 3c). In a subset of patients, 8/71 (11.3%), more than one treatment-naïve tumor tissue sample was available (Table S1 of the ESM), and methylation profiling of all samples was performed accordingly. In 5/8 patients, the samples from different body sites clustered together (Fig. 4), whereas in 3/8 patients, an intra-patient different methylation pattern was observed. Irrespective, in-depth analysis per tissue methylation profiling showed that all samples correlated with response to ICI, irrespective of the tumor origin (Fig. 1c). These results support that tissues derived from different organ sites do not affect the DNA methylation-based prediction of ICI responses.
DNA methylation gene signatures of lymph node metastases in association with response to immune checkpoint inhibitors. Heatmap illustrates two main clusters based on the top 500 differentially methylated CpG sites: best overall response (iBOR); immune complete response (iCR); immune partial response (iPR); immune stable disease (iSD); immune progressive disease (iPD); immune unconfirmed progressive disease (iUPD); immune confirmed progressive disease (iCPD); including information about immunotherapy (IO scheme and sex. Beta values with a range from 0 (blue) to 1 (red) are shown
DNA methylation signatures of cutaneous metastases, primary tumors, and organ metastases in association with response to immune checkpoint inhibitors. Heatmaps of the top 500 differentially methylated CpG sites across tumor tissue samples derived from A cutaneous metastases, B primary tumors, and C organ metastases. Best overall response (iBOR); immune complete response (iCR); immune partial response (iPR); immune stable disease (iSD); immune progressive disease (iPD); immune unconfirmed progressive disease (iUPD); immune confirmed progressive disease (iCPD); including immunotherapy (IO) scheme and sex. Beta values with a range from 0 (blue) to 1 (red) are shown
DNA methylation-based prediction of response to immune checkpoint inhibitors of tumor tissues derived from different body sites. Heatmap illustrates the sample clustering of a subset of patients (n = 8) with more than one treatment-naïve tumor tissue sample derived from different body sites. Best overall response (iBOR); immune complete response (iCR); immune partial response (iPR); immune stable disease (iSD); immune progressive disease (iPD); immune unconfirmed progressive disease (iUPD); immune confirmed progressive disease (iCPD); including information about immunotherapy (IO) scheme, sex, localization, mutation status (BRAF, cKIT, NRAS, wild type [WT]), Eastern Cooperative Oncology Group (ECOG) performance status, lactate dehydrogenase (LDH) and S100. Beta values with a range from 0 (blue) to 1 (red) are shown
3.4 Methylation Signatures of an Independent Validation Data Set
We aligned our findings to an independent data set. This validation cohort consisted of 43 patients with tumor tissue samples, of which 26 patients were classified as responders and 17 as non-responders [29]. Based on the 500 differentially methylated regions from our cohort, we identified similar trends (p = 0.146). In detail, we identified three clusters, showing responders mainly cluster 2 (15/20, 75%) and non-responders in cluster 3 (5/8, 62.5%), with hypomethylation being more prevalent in responders (Fig. S1 of the ESM).
4 Discussion
This study demonstrates that DNA methylation signatures of treatment-naïve tumor samples, regardless of the tumor tissue origin, can predict responses to ICIs in patients with stage IV metastatic melanoma, and is warranted for further analysis in prospective cohorts. It has been shown that epigenetic changes can contribute to response and failure to ICIs and DNA methylation profiling has been proposed to be a useful tool to predict responses to ICIs in different solid tumor entities [39, 40, 42]. There is still a lack of data regarding DNA methylation profiling and responses to ICIs in the field of melanoma. Newell et al. and Filipinski et al. reported that DNA methylation might be useful for predicting the response to ICIs in metastatic melanoma [7, 29]. In contrast, these studies did not evaluate whether the tumor tissue origin affected the DNA methylation signature and prediction to ICIs, nor did they use exclusively treatment-naïve tumor tissues for analysis. We have utilized the study of Newell et al. as a validation cohort, because of the biggest similarity to our cohort, although differences certainly limited the usage as a validation dataset. The Newell et al. cohort differed in the radiological response assessment, using RECIST and not iRECIST criteria, and responders were classified as complete response, partial response, and stable disease (>6 months) and non-responders as stable disease (<6 months) and progressive disease. Furthermore, no detailed information on the origin of tumor tissue used for the methylation analysis was available. The whole study cohort’s distribution of tumor tissues (n = 71) mainly consisted of subcutaneous tissue (56%), lymph nodes (30%), brain metastases (9%), and primary tumors (4%). Sixty-six percent were treatment-naïve tumor tissue samples and 34% had an intervening systemic therapy before the start of ICI treatment. Nevertheless, we observed similar trends with hypomethylation being present mainly in responders. However, the heterogenous distribution of responders and non-responders observed in these two cohorts was not unexpected because of the above-mentioned variety in the cohorts. Therefore, tumor tissue characteristics alone might be insufficient to reliably predictive ICI responses and incorporating a multimodal concept including systemic inflammatory responses in combination with tissue-based parameters might be needed for a reproducible response prediction to ICIs in metastatic melanoma.
Our study using exclusively treatment-naïve tissue samples adds to the current knowledge that there is no tissue-dependent influence on DNA methylation profiling or prediction of responses to ICIs. It has been reported that location-dependent, global DNA methylation patterns are shared between mucosal and cutaneous melanoma from primary sites, whereas uveal melanoma exhibits a different methylation signature [19]. Therefore, and because of the limited response rates of <10% to ICIs compared with cutaneous melanoma, we excluded uveal and mucosal melanomas from our analysis [1, 4, 36, 49]. Consequently, this further reduced potential batch effects when evaluating the DNA methylation signatures of different tissue sites, including primary tumors and affected body sites, by metastatic diseases of organs (including brain metastases), lymph nodes, and skin. To our knowledge, no such comparison regarding the DNA methylation signature and response to ICIs have been performed, and our results indicate that tissue samples from different sites may likewise predict the response to ICIs to a similar degree.
In addition, our pathway analysis provides further insights into the potentially differentially regulated pathways between responders and non-responders. Enrichments were detected in olfactory and G-protein coupled receptor signaling pathways, which were recently associated with migration and proliferation processes in melanoma [14, 34], as well as enrichments in the humoral response, which were shown to be supportive for ICI responses in metastatic melanoma [17].
Furthermore, our data imply that DNA methylation profiling has a higher sensitivity and specificity than established clinical markers such as LDH [47] and ECOG performance status [44]. An LDH ≥1 upper limit of normal is a well-established negative prognostic marker for reduced PFS and OS but has a low predictive value for a beneficial treatment outcome [47]. The ECOG performance status was initially established and used as a predictor of response and toxicity in patients receiving chemotherapy [31]. Regarding ICIs, a baseline ECOG performance status of zero has been associated with disease control [44], whereas other studies did not find a correlation between an ECOG score of zero and ICI-based survival benefit [48]. This confirms our findings that non-responders compared to responders showed a trend towards a higher ECOG status ≥1 (p = 0.0887), albeit not significant. However, we did not detect differences between non-responders and responders in baseline LDH (p = 0.6646) and S100 levels (p = 0.7500). We further observed that hypomethylation was associated with a beneficial response to ICIs. It was shown that global hypomethylation was associated with the expression of programmed death-ligand 1 (PD-L1) on melanoma cells in vitro [6, 15]. This is important because immunosuppressive receptors such as PD-L1 and PD-1 can inhibit the function of effector T cells [18]. However, the role of PD-L1 expression and response to ICIs in melanoma is discussed controversially and could not predict a positive clinical outcome [3, 28], Robert et al. 2015; [46], which led to the approval of PD-1 inhibitors for melanoma in metastatic [23, 45] and adjuvant [43] settings by the US Food and Drug Administration irrespective of the PD-L1 expression status of the tumor tissue.
Several limitations must be considered in the current analysis, such as the heterogeneity of the patient cohort in terms of patient characteristics and applied treatments. In comparison with other studies, we could demonstrate that tumor tissues of different origins can be used for DNA methylation profiling, which will potentially contribute to the clinical routine as a useful and easily applicable biomarker in the near future. Tissue-based DNA methylation profiling together with for example, systemic inflammatory signatures, should be addressed by future studies.
5 Conclusions
Our predictive DNA methylation signatures showed promising sensitivity and specificity; however, preferentially prospective validation cohorts are needed, possibly by using a combinational tissue-based and systemic inflammatory approach. Nevertheless, given the current lack of reliable and robust biomarkers for ICI responses in metastatic melanoma, the current study is of certain relevance. Indeed, methylation analysis is of high interest for everyday clinical use, as its analytical performance is robust, the costs are within a reasonable range, and a methylation analysis has already been established for diagnostic purposes in central nervous system malignancies [2, 24]. In summary, this study indicates that DNA methylation profiling is a useful tool for predicting the response to ICIs in patients with metastatic melanoma with high sensitivity and specificity, independent of the tissue origin.
References
Algazi AP, Tsai KK, Shoushtari AN, Munhoz RR, Eroglu Z, Piulats JM, et al. Clinical outcomes in metastatic uveal melanoma treated with PD-1 and PD-L1 antibodies. Cancer. 2016;122(21):3344–53. https://doi.org/10.1002/cncr.30258.
Capper D, Jones DTW, Sill M, Hovestadt V, Schrimpf D, Sturm D, et al. DNA methylation-based classification of central nervous system tumours. Nature. 2018;555(7697):469–74. https://doi.org/10.1038/nature26000.
Daud AI, Wolchok JD, Robert C, Hwu WJ, Weber JS, Ribas A, et al. Programmed death-ligand 1 expression and response to the anti-programmed death 1 antibody pembrolizumab in melanoma. J Clin Oncol. 2016;34(34):4102–9. https://doi.org/10.1200/jco.2016.67.2477.
Dimitriou F, Namikawa K, Reijers ILM, Buchbinder EI, Soon JA, Zaremba A, et al. Single-agent anti-PD-1 or combined with ipilimumab in patients with mucosal melanoma: an international, retrospective, cohort study. Ann Oncol. 2022;33(9):968–80. https://doi.org/10.1016/j.annonc.2022.06.004.
Elgundi Z, Papanicolaou M, Major G, Cox TR, Melrose J, Whitelock JM, et al. Cancer metastasis: the role of the extracellular matrix and the heparan sulfate proteoglycan perlecan. Front Oncol. 2019;9:1482. https://doi.org/10.3389/fonc.2019.01482.
Emran AA, Chatterjee A, Rodger EJ, Tiffen JC, Gallagher SJ, Eccles MR, et al. Targeting DNA methylation and EZH2 activity to overcome melanoma resistance to immunotherapy. Trends Immunol. 2019;40(4):328–44. https://doi.org/10.1016/j.it.2019.02.004.
Filipski K, Scherer M, Zeiner KN, Bucher A, Kleemann J, Jurmeister P, et al. DNA methylation-based prediction of response to immune checkpoint inhibition in metastatic melanoma. J Immunother Cancer. 2021;9(7): e002226. https://doi.org/10.1136/jitc-2020-002226.
Forschner A, Battke F, Hadaschik D, Schulze M, Weißgraeber S, Han CT, et al. Tumor mutation burden and circulating tumor DNA in combined CTLA-4 and PD-1 antibody therapy in metastatic melanoma: results of a prospective biomarker study. J Immunother Cancer. 2019;7(1):180. https://doi.org/10.1186/s40425-019-0659-0.
Gallo S, Vitacolonna A, Crepaldi T. NMDA receptor and its emerging role in cancer. Int J Mol Sci. 2023;24(3):2540. https://doi.org/10.3390/ijms24032540.
Gao Y, Yang C, He N, Zhao G, Wang J, Yang Y. Integration of the tumor mutational burden and tumor heterogeneity identify an immunological subtype of melanoma with favorable survival. Front Oncol. 2020;10: 571545. https://doi.org/10.3389/fonc.2020.571545.
Garbe C, Amaral T, Peris K, Hauschild A, Arenberger P, Basset-Seguin N, et al. European consensus-based interdisciplinary guideline for melanoma. Part 2: treatment: update 2022. Eur J Cancer. 2022;170:256–84. https://doi.org/10.1016/j.ejca.2022.04.018.
Garbe C, Amaral T, Peris K, Hauschild A, Arenberger P, Bastholt L, et al. European consensus-based interdisciplinary guideline for melanoma. Part 1: diagnostics: update 2019. Eur J Cancer. 2020;126:141–58. https://doi.org/10.1016/j.ejca.2019.11.014.
Ge SX, Jung D, Yao R. ShinyGO: a graphical gene-set enrichment tool for animals and plants. Bioinformatics. 2019;36(8):2628–9. https://doi.org/10.1093/bioinformatics/btz931.
Gelis L, Jovancevic N, Bechara FG, Neuhaus EM, Hatt H. Functional expression of olfactory receptors in human primary melanoma and melanoma metastasis. Exp Dermatol. 2017;26(7):569–76. https://doi.org/10.1111/exd.13316.
Gowrishankar K, Gunatilake D, Gallagher SJ, Tiffen J, Rizos H, Hersey P. Inducible but not constitutive expression of PD-L1 in human melanoma cells is dependent on activation of NF-κB. PLoS ONE. 2015;10(4): e0123410. https://doi.org/10.1371/journal.pone.0123410.
He Y, Liu T, Dai S, Xu Z, Wang L, Luo F. Tumor-associated extracellular matrix: how to be a potential aide to anti-tumor immunotherapy? Front Cell Dev Biol. 2021;9: 739161. https://doi.org/10.3389/fcell.2021.739161.
Helmink BA, Reddy SM, Gao J, Zhang S, Basar R, Thakur R, et al. B cells and tertiary lymphoid structures promote immunotherapy response. Nature. 2020;577(7791):549–55. https://doi.org/10.1038/s41586-019-1922-8.
Jiang X, Wang J, Deng X, Xiong F, Ge J, Xiang B, et al. Role of the tumor microenvironment in PD-L1/PD-1-mediated tumor immune escape. Mol Cancer. 2019;18(1):10. https://doi.org/10.1186/s12943-018-0928-4.
Jurmeister P, Wrede N, Hoffmann I, Vollbrecht C, Heim D, Hummel M, et al. Mucosal melanomas of different anatomic sites share a common global DNA methylation profile with cutaneous melanoma but show location-dependent patterns of genetic and epigenetic alterations. J Pathol. 2022;256(1):61–70. https://doi.org/10.1002/path.5808.
Keung EZ, Gershenwald JE. The eighth edition American Joint Committee on Cancer (AJCC) melanoma staging system: implications for melanoma treatment and care. Expert Rev Anticancer Ther. 2018;18(8):75–84. https://doi.org/10.1080/14737140.2018.1489246.
Kodet O, Lacina L, Krejčí E, Dvořánková B, Grim M, Štork J, et al. Melanoma cells influence the differentiation pattern of human epidermal keratinocytes. Mol Cancer. 2015;14(1):1. https://doi.org/10.1186/1476-4598-14-1.
Koelsche C, Schrimpf D, Stichel D, Sill M, Sahm F, Reuss DE, et al. Sarcoma classification by DNA methylation profiling. Nat Commun. 2021;12(1):498. https://doi.org/10.1038/s41467-020-20603-4.
Larkin J, Chiarion-Sileni V, Gonzalez R, Grob JJ, Cowey CL, Lao CD, et al. Combined nivolumab and ipilimumab or monotherapy in untreated melanoma. N Engl J Med. 2015;373(1):23–34. https://doi.org/10.1056/NEJMoa1504030.
Louis DN, Perry A, Wesseling P, Brat DJ, Cree IA, Figarella-Branger D, et al. The 2021 WHO classification of tumors of the central nervous system: a summary. Neuro Oncol. 2021;23(8):1231–51. https://doi.org/10.1093/neuonc/noab106.
Lu T, Wang S, Xu L, Zhou Q, Singla N, Gao J, et al. Tumor neoantigenicity assessment with CSiN score incorporates clonality and immunogenicity to predict immunotherapy outcomes. Sci Immunol. 2020;5(44):eaaz3199. https://doi.org/10.1126/sciimmunol.aaz3199.
Maksimovic J, Gordon L, Oshlack A. SWAN: subset-quantile within array normalization for illumina infinium human methylation 450 bead chips. Genome Biol. 2012;13(6):R44. https://doi.org/10.1186/gb-2012-13-6-r44.
Metsalu T, Vilo J. ClustVis: a web tool for visualizing clustering of multivariate data using principal component analysis and heatmap. Nucleic Acids Res. 2015;43(W1):W566–70. https://doi.org/10.1093/nar/gkv468.
Morrison C, Pabla S, Conroy JM, Nesline MK, Glenn ST, Dressman D, et al. Predicting response to checkpoint inhibitors in melanoma beyond PD-L1 and mutational burden. J Immunother Cancer. 2018;6(1):32. https://doi.org/10.1186/s40425-018-0344-8.
Newell F, Pires da Silva I, Johansson PA, Menzies AM, Wilmott JS, Addala V, et al. Multiomic profiling of checkpoint inhibitor-treated melanoma: identifying predictors of response and resistance, and markers of biological discordance. Cancer Cell. 2022;40(1):88–102. https://doi.org/10.1016/j.ccell.2021.11.012.
Ning B, Liu Y, Wang M, Li Y, Xu T, Wei Y. The predictive value of tumor mutation burden on clinical efficacy of immune checkpoint inhibitors in melanoma: a systematic review and meta-analysis. Front Pharmacol. 2022;13: 748674. https://doi.org/10.3389/fphar.2022.748674.
Pater JL, Loeb M. Nonanatomic prognostic factors in carcinoma of the lung: a multivariate analysis. Cancer. 1982;50(2):326–31. https://doi.org/10.1002/1097-0142(19820715)50:2%3c326::aid-cncr2820500227%3e3.0.co;2-g.
Pedro MP, Lund K, Iglesias-Bartolome R. The landscape of GPCR signaling in the regulation of epidermal stem cell fate and skin homeostasis. Stem Cells. 2020;38(12):520-31. https://doi.org/10.1002/stem.3273.
Phipson B, Maksimovic J, Oshlack A. missMethyl: an R package for analyzing data from Illumina’s human methylation 450 platform. Bioinformatics. 2016;32(2):286–8. https://doi.org/10.1093/bioinformatics/btv560.
Raymond JH, Aktary Z, Larue L, Delmas V. Targeting GPCRs and their signaling as a therapeutic option in melanoma. Cancers (Basel). 2022;14(3):706. https://doi.org/10.3390/cancers14030706.
Rober C, Schachter J, Long GV, Arance A, Grob JJ, Mortier L, et al. Pembrolizumab versus ipilimumab in advanced melanoma. N Engl J Med. 2015;372(26):2521–32. https://doi.org/10.1056/NEJMoa1503093.
Rodrigues M, Koning L, Coupland SE, Jochemsen AG, Marais R, Stern MH, et al. So close, yet so far: discrepancies between uveal and other melanomas. A position paper from UM Cure 2020. Cancers (Basel). 2019;11(7):1032. https://doi.org/10.3390/cancers11071032.
Rømer AMA, Thorseth ML, Madsen DH. Immune modulatory properties of collagen in cancer. Front Immunol. 2021;12: 791453. https://doi.org/10.3389/fimmu.2021.791453.
Seymour L, Bogaerts J, Perrone A, Ford R, Schwartz LH, Mandrekar S, et al. iRECIST: guidelines for response criteria for use in trials testing immunotherapeutics. Lancet Oncol. 2017;18(3):e143–52. https://doi.org/10.1016/s1470-2045(17)30074-8.
Starzer AM, Berghoff AS, Hamacher R, Tomasich E, Feldmann K, Hatziioannou T, et al. Tumor DNA methylation profiles correlate with response to anti-PD-1 immune checkpoint inhibitor monotherapy in sarcoma patients. J Immunother Cancer. 2021;9(3): e001458. https://doi.org/10.1136/jitc-2020-001458.
Starzer AM, Heller G, Tomasich E, Melchardt T, Feldmann K, Hatziioannou T, et al. DNA methylation profiles differ in responders versus non-responders to anti-PD-1 immune checkpoint inhibitors in patients with advanced and metastatic head and neck squamous cell carcinoma. J Immunother Cancer. 2022;10(3): e003420. https://doi.org/10.1136/jitc-2021-003420.
Thomas D, Rathinavel AK, Radhakrishnan P. Altered glycosylation in cancer: a promising target for biomarkers and therapeutics. Biochim Biophys Acta Rev Cancer. 2021;1875(1): 188464. https://doi.org/10.1016/j.bbcan.2020.188464.
Wang P, Xu G, Gao E, Xu Y, Liang L, Jiang G, et al. Identification of prognostic DNA methylation signatures in lung adenocarcinoma. Oxid Med Cell Longev. 2022;2022:8802303. https://doi.org/10.1155/2022/8802303.
Weber J, Mandala M, Del Vecchio M, Gogas HJ, Arance AM, Cowey CL, et al. Adjuvant nivolumab versus ipilimumab in resected stage III or IV melanoma. N Engl J Med. 2017;377(19):1824–35. https://doi.org/10.1056/NEJMoa1709030.
Wells L, Cerniglia M, Hall S, Jost AC, Britt G. Treatment of metastatic disease with immune checkpoint inhibitors nivolumab and pembrolizumab: effect of performance status on clinical outcomes. J Immunother Precis Oncol. 2022;5(2):37–42. https://doi.org/10.36401/jipo-22-3.
Wolchok JD, Chiarion-Sileni V, Gonzalez R, Rutkowski P, Grob JJ, Cowey CL, et al. Overall survival with combined nivolumab and ipilimumab in advanced melanoma. N Engl J Med. 2017;377(14):1345–56. https://doi.org/10.1056/NEJMoa1709684.
Wolchok JD, Kluger H, Callahan MK, Postow MA, Rizvi NA, Lesokhin AM, et al. Nivolumab plus ipilimumab in advanced melanoma. N Engl J Med. 2013;369(2):122–33. https://doi.org/10.1056/NEJMoa1302369.
Xu J, Zhao J, Wang J, Sun C, Zhu X. Prognostic value of lactate dehydrogenase for melanoma patients receiving anti-PD-1/PD-L1 therapy: a meta-analysis. Medicine (Baltimore). 2021;100(14): e25318. https://doi.org/10.1097/md.0000000000025318.
Yang F, Markovic SN, Molina JR, Halfdanarson TR, Pagliaro LC, Chintakuntlawar AV, et al. Association of sex, age, and eastern cooperative oncology group performance status with survival benefit of cancer immunotherapy in randomized clinical trials: a systematic review and meta-analysis. JAMA Netw Open. 2020;3(8): e2012534. https://doi.org/10.1001/jamanetworkopen.2020.12534.
Zimmer L, Vaubel J, Mohr P, Hauschild A, Utikal J, Simon J, et al. Phase II DeCOG-study of ipilimumab in pretreated and treatment-naïve patients with metastatic uveal melanoma. PLoS ONE. 2015;10(3): e0118564. https://doi.org/10.1371/journal.pone.0118564.
Acknowledgments
We thank the Department of Pathology, Medical University of Vienna for providing tissue samples for the DNA methylation profiling analysis.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
Open access funding provided by Medical University of Vienna. This study was supported by the Christian Doppler Research Association, the Austrian Federal Ministry for Digital and Economic Affairs, and the National Foundation for Research, Technology, and Development.
Conflicts of Interest
Julia Maria Ressler received honoraria for lectures from Bristol-Myers Squibb, Roche, Amgen, and Novartis and travel support from Sanofi, Roche, and Bristol-Myers Squibb. Angelika Martina Starzer received honoraria for lectures from AstraZeneca and travel and congress support from PharmaMar, MSD, and Lilly. Christoph Höller reports speaker honoraria from Amgen, BMS, MSD, Novartis, and Roche, participation in advisory boards for Amgen, Astra Zeneca, BMS, Incyte, MSD, Novartis, Pierre Fabre, and Roche, and research funding by Amgen to the institution. Matthias Preusser has received honoraria for lectures, consultation, and advisory board participation from the following for-profit companies: Bayer, Bristol-Myers Squibb, Novartis, Gerson Lehrman Group (GLG), CMC Contrast, GlaxoSmithKline, Mundipharma, Roche, BMJ Journals, MedMedia, Astra Zeneca, AbbVie, Lilly, Medahead, Daiichi Sankyo, Sanofi, Merck Sharp & Dome, Tocagen, Adastra, Gan & Lee Pharmaceuticals, and Servier. Anna Sophie Berghoff has research support from Daiichi Sankyo and Roche, honoraria for lectures, consultation, and advisory board participation from Roche Bristol-Meyers Squibb, Merck, Daiichi Sankyo, AstraZeneca, and CeCaVa, and travel support from Roche, Amgen, and AbbVie. Philipp Tschandl received honoraria from Silverchair, unrestricted grants for education projects from Lilly, and honoraria for lectures from AbbVie, Lilly, FotoFinder, and Novartis. Erwin Tomasich, Teresa Hatziioannou, Helmut Ringl, Gerwin Heller, Rita Silmbrod, Lynn Gottmann, and Nina Zila have no conflicts of interest that are directly relevant to the content of this article.
Ethics Approval
This study was conducted in accordance with the Declaration of Helsinki and approved by the Institutional Ethics committee of the Medical University of Vienna (Number 1312/2021).
Consent to Participate
Written informed consent from the study participants was obtained.
Consent for Publication
Not applicable.
Availability of Data and Material
Raw data of DNA methylation analysis are publicly available at the Gene Expression Omnibus database respiratory under the accession number GSE235122. Further data will be shared upon reasonable request to the corresponding author. The relevant data set of the study is included in the article or uploaded as supplementary information. Additional data will be shared upon request by the authors.
Code Availability
Not applicable.
Authors’ Contributions
JMR: conceptualization, resources, data curation, formal analysis, validation, writing of the original draft. ET: formal analysis, validation, writing. TH: formal analysis, validation. HR: radiological assessment, validation. GW: bioinformatical analysis, visualization, validation, writing. RS: data curation, formal analysis. LG: data curation, formal analysis. AMS: data curation, formal analysis. NZ data curation, formal analysis. PT: resources, formal analysis. CH: resources, validation, supervision. MP: conceptualization, resources, validation, writing, supervision. ASB: conceptualization, resources, formal analysis, validation, writing. All authors read and approved the final manuscript.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Ressler, J.M., Tomasich, E., Hatziioannou, T. et al. DNA Methylation Signatures Correlate with Response to Immune Checkpoint Inhibitors in Metastatic Melanoma. Targ Oncol 19, 263–275 (2024). https://doi.org/10.1007/s11523-024-01041-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11523-024-01041-4