Abstract
Objective
Endoplasmic reticulum stress (ERS) and long non-coding RNAs (lncRNAs) are important in melanoma development and progression. This study aimed to explore the prognostic value of ERS-associated lncRNA profiles in cutaneous melanoma (CM).
Methods
The Cancer Genome Atlas (TCGA) provides the raw data of CM. GSEA website was used to obtain ERS-related genes, and mRNA and LncRNA co-expression network were used to obtain ERS-related lncRNAs. A Lasso regression analysis was used to identify a prognostic risk model for the composition of ERS-related lncRNAs. Patients were divided into high- and low-risk groups based on the model’s risk score. The researchers then compared the two groups’ survival rates, immune infiltration, chemotherapeutic drug sensitivity, and immune checkpoint gene expression.
Results
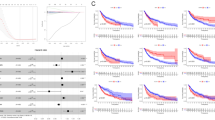
Thirty-nine ERS-related lncRNAs were discovered to be prognostic. A prognostic risk model made up of ten ERS-related lncRNAs was discovered. Patients in the low-risk group had a better prognosis than those in the high-risk group. An examination of tumor microenvironment revealed that risk scores correlated with immune cell infiltration in eight cases. Dacarbazine, paclitaxel, and cisplatin, three chemotherapy drugs, were more sensitive in the low-risk group than in the high-risk group.
Conclusion
This study identified a risk model of ten ERS-related lncRNAs that have significant prognostic value in CM and could help guide clinical treatment.









Similar content being viewed by others
Data availability
The datasets analyzed in this study can be found in this paper. Raw counts for RNA-seq transcriptome data and corresponding clinical data for skin cutaneous melanoma were extracted from TCGA database (https://portal.gdc.cancer.gov). ERS-related genes were extracted from GSEA database (https://www.gsea-msigdb.org/gsea/index.jsp).
References
Arslanbaeva LR, Santoro MM (2020) Adaptive redox homeostasis in cutaneous melanoma. Redox Biol 37:101753. https://doi.org/10.1016/j.redox.2020.101753
Atkins MB, Curiel-Lewandrowski C, Fisher DE et al (2021) The state of melanoma: emergent challenges and opportunities. Clin Cancer Res 27(10):2678–2697. https://doi.org/10.1158/1078-0432.CCR-20-4092
Bomar L, Senithilnathan A, Ahn C (2019) Systemic therapies for advanced melanoma. Dermatol Clin 37(4):409–423. https://doi.org/10.1016/j.det.2019.05.001
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA 68(6):394–424. https://doi.org/10.3322/caac.21492
Bremnes RM, Busund LT, Kilvær TL et al (2016) The role of tumor-infiltrating lymphocytes in development, progression, and prognosis of non-small cell lung cancer. J Thorac Oncol 11(6):789–800. https://doi.org/10.1016/j.jtho.2016.01.015
Chen Y, Chen L, Wang J, Tan J, Wang S (2021) Identification of six autophagy-related-lncRNA prognostic biomarkers in uveal melanoma. Dis Mark 2021:1–12. https://doi.org/10.1155/2021/2401617
Dillon LD, McPhee M, Davidson RS et al (2022) Expanded evidence that the 31-gene expression profile test provides clinical utility for melanoma management in a multicenter study. Curr Med Res Opin. https://doi.org/10.1080/03007995.2022.2033560
Dowling J, McGregor SP, Williford P (2019) Update on current treatment recommendations for primary cutaneous melanoma. Dermatol Clin 37(4):397–407. https://doi.org/10.1016/j.det.2019.06.001
Eggermont AMM, Chiarion-Sileni V, Grob JJ et al (2016) Prolonged survival in stage III melanoma with ipilimumab adjuvant therapy. N Engl J Med 375(19):1845–1855. https://doi.org/10.1056/NEJMoa1611299
Fan Y, Liang X, Yu D (2021) Low expression of endoplasmic reticulum stress-related gene SERP1 is associated with poor prognosis and immune infiltration in skin cutaneous melanoma. Aging 13(19):23036–23071. https://doi.org/10.18632/aging.203594
Gambichler T, Tsagoudis K, Kiecker F et al (2021) Prognostic significance of an 11-gene RNA assay in archival tissue of cutaneous melanoma stage I–III patients. Eur J Cancer 143:11–18. https://doi.org/10.1016/j.ejca.2020.10.016
Giampietri C, Petrungaro S, Conti S, Facchiano A, Filippini A, Ziparo E (2015) Cancer microenvironment and endoplasmic reticulum stress response. Mediators Inflamm 2015:1–11. https://doi.org/10.1155/2015/417281
Guo JH, Yin SS, Liu H, Liu F, Gao FH (2021) Tumor microenvironment immune-related lncRNA signature for patients with melanoma. Ann Transl Med 9(10):857–857. https://doi.org/10.21037/atm-21-1794
Hersey P, Zhang XD (2008) Adaptation to ER stress as a driver of malignancy and resistance to therapy in human melanoma. Pigment Cell Melanoma Res 21(3):358–367. https://doi.org/10.1111/j.1755-148X.2008.00467.x
Hindupur SV, Schmid SC, Koch JA et al (2020) STAT3/5 inhibitors suppress proliferation in bladder cancer and enhance oncolytic adenovirus therapy. IJMS 21(3):1106. https://doi.org/10.3390/ijms21031106
Hinshaw DC, Shevde LA (2019) The tumor microenvironment innately modulates cancer progression. Cancer Res 79(18):4557–4566. https://doi.org/10.1158/0008-5472.CAN-18-3962
Kim J, Piao HL, Kim BJ et al (2018) Long noncoding RNA MALAT1 suppresses breast cancer metastasis. Nat Genet 50(12):1705–1715. https://doi.org/10.1038/s41588-018-0252-3
Kopp F, Mendell JT (2018) Functional classification and experimental dissection of long noncoding RNAs. Cell 172(3):393–407. https://doi.org/10.1016/j.cell.2018.01.011
Larkin J, Chiarion-Sileni V, Gonzalez R et al (2019) Five-Year survival with combined nivolumab and ipilimumab in advanced melanoma. N Engl J Med 381(16):1535–1546. https://doi.org/10.1056/NEJMoa1910836
Leonardi G, Falzone L, Salemi R et al (2018) Cutaneous melanoma: from pathogenesis to therapy (review). Int J Oncol. https://doi.org/10.3892/ijo.2018.4287
Li FW, Luo SK (2021) Identification and construction of a predictive immune-related lncRNA signature model for melanoma. IJGM 14:9227–9235. https://doi.org/10.2147/IJGM.S340025
Li Y, Shan Z, Liu C et al (2017) MicroRNA-294 promotes cellular proliferation and motility through the PI3K/AKT and JAK/STAT pathways by upregulation of NRAS in bladder cancer. Biochemistry Moscow 82(4):474–482. https://doi.org/10.1134/S0006297917040095
Li Z, Li Y, Zhong W, Huang P (2021a) m6A-related lncRNA to develop prognostic signature and predict the immune landscape in bladder cancer. J Oncol 2021:1–16. https://doi.org/10.1155/2021/7488188
Li X, Jin F, Li Y (2021b) A novel autophagy-related lncRNA prognostic risk model for breast cancer. J Cell Mol Med 25(1):4–14. https://doi.org/10.1111/jcmm.15980
Li Z, Wei J, Zheng H et al (2022) The new horizon of biomarker in melanoma patients: a study based on autophagy-related long non-coding RNA. Medicine (Baltimore) 101(1):e28553. https://doi.org/10.1097/MD.0000000000028553
Liao CL, Sun XY et al (2021) Identification and validation of tumor microenvironment-related lncRNA prognostic signature for uveal melanoma. Int J Ophthalmol 14(8):1151–1159. https://doi.org/10.18240/ijo.2021.08.03
Liu N, Liu Z, Liu X, Chen H (2019) Comprehensive analysis of a competing endogenous RNA network identifies seven-lncRNA signature as a prognostic biomarker for melanoma. Front Oncol 9:935. https://doi.org/10.3389/fonc.2019.00935
Lombard DB, Cierpicki T, Grembecka J (2019) Combined MAPK pathway and HDAC inhibition breaks melanoma. Cancer Discov 9(4):469–471. https://doi.org/10.1158/2159-8290.CD-19-0069
Peng YL, Wu ZS, Lu HM et al (2020) Prognostic significance of tumor-infiltrating immune cells in muscle-invasive bladder cancer. Am J Transl Res 12(10):6524–6536
Quail DF, Joyce JA (2013) Microenvironmental regulation of tumor progression and metastasis. Nat Med 19(11):1423–1437. https://doi.org/10.1038/nm.3394
Rather RA, Bhagat M, Singh SK (2020) Oncogenic BRAF, endoplasmic reticulum stress, and autophagy: crosstalk and therapeutic targets in cutaneous melanoma. Mutat Res/Rev Mutat Res 785:108321. https://doi.org/10.1016/j.mrrev.2020.108321
Ryabaya O, Prokofieva A, Khochenkov D, Abramov I, Zasedatelev A, Stepanova E (2018) Inhibition of endoplasmic reticulum stress-induced autophagy sensitizes melanoma cells to temozolomide treatment. Oncol Rep 40(1):385–394. https://doi.org/10.3892/or.2018.6430
Schmitz SU, Grote P, Herrmann BG (2016) Mechanisms of long noncoding RNA function in development and disease. Cell Mol Life Sci 73(13):2491–2509. https://doi.org/10.1007/s00018-016-2174-5
Slack FJ, Chinnaiyan AM (2019) The role of non-coding RNAs in oncology. Cell 179(5):1033–1055. https://doi.org/10.1016/j.cell.2019.10.017
Turner N, Ware O, Bosenberg M (2018) Genetics of metastasis: melanoma and other cancers. Clin Exp Metastasis 35(5–6):379–391. https://doi.org/10.1007/s10585-018-9893-y
Wang H, Yang L, Wang D, Zhang Q, Zhang L (2017) Pro-tumor activities of macrophages in the progression of melanoma. Hum Vaccin Immunother 13(7):1556–1562. https://doi.org/10.1080/21645515.2017.1312043
Weber J, Mandala M, Del Vecchio M et al (2017) Adjuvant nivolumab versus ipilimumab in resected stage III or IV melanoma. N Engl J Med 377(19):1824–1835. https://doi.org/10.1056/NEJMoa1709030
Wolchok JD, Chiarion-Sileni V, Gonzalez R et al (2017) Overall survival with combined nivolumab and ipilimumab in advanced melanoma. N Engl J Med 377(14):1345–1356. https://doi.org/10.1056/NEJMoa1709684
Wu Z, Zhu K, Liu Q et al (2020) Profiles of immune infiltration in bladder cancer and its clinical significance: an integrative genomic analysis. Int J Med Sci 17(6):762–772. https://doi.org/10.7150/ijms.42151
Xiao B, Liu L, Li A et al (2021) Identification and validation of immune-related lncRNA prognostic signatures for melanoma. Immun Inflamm Dis 9(3):1044–1054. https://doi.org/10.1002/iid3.468
Xue L, Wu P, Zhao X et al (2021) Using immune-related lncRNA signature for prognosis and response to immunotherapy in cutaneous melanoma. IJGM 14:6463–6475. https://doi.org/10.2147/IJGM.S335266
Yavartanoo M, Yi GS (2021) Development and validation of tumor immunogenicity based gene signature for skin cancer risk stratification. IJMS 22(21):12025. https://doi.org/10.3390/ijms222112025
Zhang E, He X, Zhang C et al (2018) A novel long noncoding RNA HOXC-AS3 mediates tumorigenesis of gastric cancer by binding to YBX1. Genome Biol 19(1):154. https://doi.org/10.1186/s13059-018-1523-0
(2015) Combined nivolumab and ipilimumab or monotherapy in untreated melanoma. N Engl J Med 373(13):1270–1271. https://doi.org/10.1056/NEJMc1509660
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
Preparation of the manuscript: A-aL. Grammar modification: FL and ML. Interpretation or analysis of data: A-aL, YZ, and DX. Revision for important intellectual content: M-yY.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Ethical approval
Our study was approved by the Medical Ethics Committee of the Second Affiliated Hospital of Nanchang University.
Consent for publication
Our manuscript does not contains any individual data in any form.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
432_2022_4086_MOESM1_ESM.tif
Supplementary file1 A–L The infiltrating levels of B cells memory, dendritic cells, Macrophages M0, Macrophages M1, Macrophages M2, Mast cells activated, NK cells resting, Plasma cells, T cells CD4 memory activated, T cells CD8, T cells follicular helper and T cells regulatory in cluster 1/2 subtypes (TIF 10800 kb)
432_2022_4086_MOESM2_ESM.tif
Supplementary file2 Validation of the prognosis of risk scores in different groups of CM patients, including age (A, B) and gender (C, D) (TIF 10800 kb)
432_2022_4086_MOESM3_ESM.tif
Supplementary file3 Validation of the prognosis of risk scores in different groups of CM patients, including clinical stage (A) and TNM stage (B–D) (TIF 10800 kb)
432_2022_4086_MOESM4_ESM.tif
Supplementary file4 The low- and high-risk groups displaying different endoplasmic reticulum stress (ERS) statuses. A–C Principal component analysis (PCA) between low- and high-risk groups based on the whole-genome, ERS-related encoding genes, and the risk model of 10 ERS-related lncRNA expression profiles (TIF 10800 kb)
Rights and permissions
About this article
Cite this article
Li, Aa., Li, F., Lan, M. et al. A novel endoplasmic reticulum stress-related lncRNA prognostic risk model for cutaneous melanoma. J Cancer Res Clin Oncol 148, 3227–3241 (2022). https://doi.org/10.1007/s00432-022-04086-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-022-04086-y