Abstract
Purpose
Lung cancer has the highest morbidity and mortality among all cancer types. Reliable prognostic biomarkers are needed to identify high-risk patients apart from TNM system for precision medicine. The present study is designed to identify robust prognostic biomarkers in lung adenocarcinoma (LUAD) based on integration of multiple GEO datasets, The Cancer Genome Atlas (TCGA) database and Clinical Proteomic Tumor Analysis Consortium (CPTAC) database.
Methods
Four LUAD GEO datasets (GSE10072, GSE2514, GSE43458, and GSE32863) and TCGA database were implemented to analyze the differently expressed genes (DEGs). Gene ontology, KEGG pathway, and protein–protein interaction network (PPI) were conducted based on the above DEGs. Hub genes were selected based on connectivity degree in the PPI network. Expression analysis and Kaplan–Meier survival analysis were conducted in CPTAC lung adenocarcinomas cohort. Kaplan–Meier survival analysis and Cox proportional hazards regression were performed on these hub genes using TCGA and our own cohort.
Results
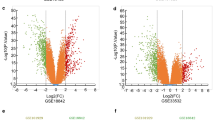
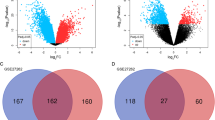
A total of 430 shared genes in all five datasets were identified as DEGs. Based on their PPI network, nine hub genes were selected and all of them were significantly associated with overall survival using GEPIA analysis. Two hub genes, TOP2A and UBE2C, were further combined and showed poorer prognosis in both TCGA dataset and our validated cohort. Analysis in CPTAC revealed that TOP2A and UBE2C were significantly highly expressed in tumor sample. Multivariable analysis suggested TOP2A and UBE2C as independent prognostic factors in LUAD.
Conclusion
Using data mining approach, we identified TOP2A and UBE2C as two robust prognostic factors in LUAD. We also demonstrated the TOP2A/UBE2C co-expression status in LUAD, and TOP2A/UBE2C co-expression correlated with poorer prognosis. More in-depth research is needed for transforming this result into clinical setting.













Similar content being viewed by others
References
Al-Kuraya K, Novotny H, Bavi P, Siraj AK, Uddin S, Ezzat A, Sanea NA, Al-Dayel F, Al-Mana H, Sheikh SS, Mirlacher M, Tapia C, Simon R, Sauter G, Terracciano L, Tornillo L (2007) HER2, TOP2A, CCND1, EGFR and C-MYC oncogene amplification in colorectal cancer. J Clin Pathol 60:768–772
Asamura H, Chansky K, Crowley J, Goldstraw P, Rusch VW, Vansteenkiste JF, Watanabe H, Wu YL, Zielinski M, Ball D, Rami-Porta R (2015) The International Association for the Study of Lung Cancer Lung Cancer Staging Project: Proposals for the Revision of the N Descriptors in the Forthcoming 8th Edition of the TNM Classification for Lung Cancer. J Thorac Oncol 10:1675–1684
Bui KT, Cooper WA, Kao S, Boyer M (2018) Targeted molecular treatments in non-small cell lung cancer: a clinical guide for oncologists. J Clin Med 7:192
Byron SA, Van Keuren-Jensen KR, Engelthaler DM, Carpten JD, Craig DW (2016) Translating RNA sequencing into clinical diagnostics: opportunities and challenges. Nat Rev Genet 17:257–271
Chang CC, Wang YR, Chen SF, Wu CC, Chan NL (2013) New insights into DNA-binding by type IIA topoisomerases. Curr Opin Struct Biol 23:125–133
Chen D, Maruschke M, Hakenberg O, Zimmermann W, Stief CG, Buchner A (2017) TOP2A, HELLS, ATAD2, and TET3 are novel prognostic markers in renal cell carcinoma. Urology 102:e261–e267
Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J (2016) Cancer statistics in China, 2015. CA Cancer J Clin 66:115–132
Chen SF, Huang NL, Lin JH, Wu CC, Wang YR, Yu YJ, Gilson MK, Chan NL (2018) Structural insights into the gating of DNA passage by the topoisomerase II DNA-gate. Nat Commun 9:3085
Chou C-P, Huang N-C, Jhuang S-J, Pan H-B, Peng N-J, Cheng J-T, Chen C-F, Chen J-J, Chang T-H (2014) Ubiquitin-conjugating enzyme UBE2C is highly expressed in breast microcalcification lesions. PLoS ONE 9:e93934
De Resende MF, Vieira S, Chinen LTD, Chiappelli F, da Fonseca FP, Guimarães GC, Soares FA, Neves I, Pagotty S, Pellionisz PA (2013) Prognostication of prostate cancer based on TOP2A protein and gene assessment: TOP2A in prostate cancer. J Transl Med 11:36
Deming PB, Cistulli CA, Zhao H, Graves PR, Piwnica-Worms H, Paules RS, Downes CS, Kaufmann WK (2001) The human decatenation checkpoint. Proc Natl Acad Sci USA 98:12044–12049
Eberhardt WE, Mitchell A, Crowley J, Kondo H, Kim YT, Turrisi A, Goldstraw P, Rami-Porta R (2015) The IASLC Lung Cancer Staging Project: Proposals for the Revision of the M Descriptors in the Forthcoming eighth edition of the TNM classification of lung cancer. J Thorac Oncol 10:1515–1522.
Farcomeni A (2008) A review of modern multiple hypothesis testing, with particular attention to the false discovery proportion. Stat Methods Med Res 17:347–388
Gao Z, Shi R, Yuan K, Wang Y (2016) Expression and prognostic value of E2F activators in NSCLC and subtypes: a research based on bioinformatics analysis. Tumour Biol 37:14979–14987
Goldstraw P, Chansky K, Crowley J, Rami-Porta R, Asamura H, Eberhardt WE, Nicholson AG, Groome P, Mitchell A, Bolejack V (2016) The IASLC lung cancer staging project: proposals for revision of the TNM stage groupings in the forthcoming (eighth) edition of the TNM classification for lung cancer. J Thorac Oncol 11:39–51
Hao Z, Zhang H, Cowell J (2012) Ubiquitin-conjugating enzyme UBE2C: molecular biology, role in tumorigenesis, and potential as a biomarker. Tumor Biol 33:723–730
Hershko A (1997) Roles of ubiquitin-mediated proteolysis in cell cycle control. Curr Opin Cell Biol 9:788–799
Huang WT, Yang X, He RQ, Ma J, Hu XH, Mo WJ, Chen G (2019) Overexpressed BSG related to the progression of lung adenocarcinoma with high-throughput data-mining, immunohistochemistry, in vitro validation and in silico investigation. Am J Transl Res 11:4835–4850
Ieta K, Ojima E, Tanaka F, Nakamura Y, Haraguchi N, Mimori K, Inoue H, Kuwano H, Mori M (2007) Identification of overexpressed genes in hepatocellular carcinoma, with special reference to ubiquitin-conjugating enzyme E2C gene expression. Int J Cancer 121:33–38
Jamal-Hanjani M, Wilson GA, McGranahan N, Birkbak NJ, Watkins TBK, Veeriah S, Shafi S, Johnson DH, Mitter R, Rosenthal R, Salm M, Horswell S, Escudero M, Matthews N, Rowan A, Chambers T, Moore DA, Turajlic S, Xu H, Lee SM, Forster MD, Ahmad T, Hiley CT, Abbosh C, Falzon M, Borg E, Marafioti T, Lawrence D, Hayward M, Kolvekar S, Panagiotopoulos N, Janes SM, Thakrar R, Ahmed A, Blackhall F, Summers Y, Shah R, Joseph L, Quinn AM, Crosbie PA, Naidu B, Middleton G, Langman G, Trotter S, Nicolson M, Remmen H, Kerr K, Chetty M, Gomersall L, Fennell DA, Nakas A, Rathinam S, Anand G, Khan S, Russell P, Ezhil V, Ismail B, Irvin-Sellers M, Prakash V, Lester JF, Kornaszewska M, Attanoos R, Adams H, Davies H, Dentro S, Taniere P, O'Sullivan B, Lowe HL, Hartley JA, Iles N, Bell H, Ngai Y, Shaw JA, Herrero J, Szallasi Z, Schwarz RF, Stewart A, Quezada SA, Le Quesne J, Van Loo P, Dive C, Hackshaw A, Swanton C (2017) Tracking the evolution of non-small-cell lung cancer. N Engl J Med 376:2109–2121
Kalemkerian GP, Narula N, Kennedy EB, Biermann WA, Donington J, Leighl NB, Lew M, Pantelas J, Ramalingam SS, Reck M, Saqi A, Simoff M, Singh N, Sundaram B (2018) Molecular testing guideline for the selection of patients with lung cancer for treatment with targeted tyrosine kinase inhibitors: American Society of Clinical Oncology Endorsement of the College of American Pathologists/International Association for the Study of Lung Cancer/Association for Molecular Pathology Clinical Practice Guideline Update. J Clin Oncol 36:911–919
Kang X, Song C, Du X, Zhang C, Liu Y, Liang L, He J, Lamb K, Shen WH, Yin Y (2015) PTEN stabilizes TOP2A and regulates the DNA decatenation. Sci Rep 5:17873
Kotera M, Okuno Y, Hattori M, Goto S, Kanehisa M (2004) Computational assignment of the EC numbers for genomic-scale analysis of enzymatic reactions. J Am Chem Soc 126:16487–16498
Kruglyak KM, Lin E, Ong FS (2016) Next-generation sequencing and applications to the diagnosis and treatment of lung cancer. Adv Exp Med Biol 890:123–136
Li B, Cui Y, Diehn M, Li R (2017) Development and validation of an individualized immune prognostic signature in early-stage nonsquamous non-small cell lung cancer. JAMA Oncol 3:1529–1537
McGranahan N, Swanton C (2017) Clonal heterogeneity and tumor evolution: past, present, and the future. Cell 168:613–628
Meyerson M, Gabriel S, Getz G (2010) Advances in understanding cancer genomes through second-generation sequencing. Nat Rev Genet 11:685–696
Nitiss JL (2009) DNA topoisomerase II and its growing repertoire of biological functions. Nat Rev Cancer 9:327–337
Olsen KE, Knudsen H, Rasmussen BB, Balslev E, Knoop A, Ejlertsen B, Nielsen KV, Schönau A, Overgaard J (2004) Amplification of HER2 and TOP2A and deletion of TOP2A genes in breast cancer investigated by new FISH probes. Acta Oncol 43:35–42
Osmani L, Askin F, Gabrielson E, Li QK (2018) Current WHO guidelines and the critical role of immunohistochemical markers in the subclassification of non-small cell lung carcinoma (NSCLC): moving from targeted therapy to immunotherapy. Semin Cancer Biol 52:103–109
Ozsolak F, Milos PM (2011) RNA sequencing: advances, challenges and opportunities. Nat Rev Genet 12:87–98
Panvichian R, Tantiwetrueangdet A, Angkathunyakul N, Leelaudomlipi S (2015) TOP2A amplification and overexpression in hepatocellular carcinoma tissues. BioMed Res Int 2015:381602
Passiglia F, Bronte G, Castiglia M, Listi A, Calo V, Toia F, Cicero G, Fanale D, Rizzo S, Bazan V, Russo A (2015) Prognostic and predictive biomarkers for targeted therapy in NSCLC: for whom the bell tolls? Expert Opin Biol Ther 15:1553–1566
Pickart CM (2001) Mechanisms underlying ubiquitination. Annu Rev Biochem 70:503–533
Rape M, Kirschner MW (2004) Autonomous regulation of the anaphase-promoting complex couples mitosis to S-phase entry. Nature 432:588
Relli V, Trerotola M, Guerra E, Alberti S (2019) Abandoning the notion of non-small cell lung cancer. Trends Mol Med 25:585–594
Rolfo C, Caglevic C, Santarpia M, Araujo A, Giovannetti E, Gallardo CD, Pauwels P, Mahave M (2017) Immunotherapy in NSCLC: a promising and revolutionary weapon. Adv Exp Med Biol 995:97–125
Rybarczyk-Kasiuchnicz A, Ramlau R (2018) Current views on molecularly targeted therapy for lung cancer—a review of literature from the last five years. Kardiochirurgia i torakochirurgia polska 15:119–124
Shedden K, Taylor JM, Enkemann SA, Tsao MS, Yeatman TJ, Gerald WL, Eschrich S, Jurisica I, Giordano TJ, Misek DE, Chang AC, Zhu CQ, Strumpf D, Hanash S, Shepherd FA, Ding K, Seymour L, Naoki K, Pennell N, Weir B, Verhaak R, Ladd-Acosta C, Golub T, Gruidl M, Sharma A, Szoke J, Zakowski M, Rusch V, Kris M, Viale A, Motoi N, Travis W, Conley B, Seshan VE, Meyerson M, Kuick R, Dobbin KK, Lively T, Jacobson JW, Beer DG (2008) Gene expression-based survival prediction in lung adenocarcinoma: a multi-site, blinded validation study. Nat Med 14:822–827
Shen Z, Jiang X, Zeng C, Zheng S, Luo B, Zeng Y, Ding R, Jiang H, He Q, Guo J (2013) High expression of ubiquitin-conjugating enzyme 2C (UBE2C) correlates with nasopharyngeal carcinoma progression. BMC Cancer 13:192
Siegel RL, Miller KD, Jemal A (2015) Cancer statistics, 2015. CA A Cancer J Clin 65:5–29
Siegel RL, Miller KD, Jemal A (2018) Cancer statistics, 2018. CA Cancer J Clin 68:7–30
Simon R, Atefy R, Wagner U, Forster T, Fijan A, Bruderer J, Wilber K, Mihatsch MJ, Gasser T, Sauter G (2003) HER-2 and TOP2A coamplification in urinary bladder cancer. Int J Cancer 107:764–772
Stark R, Grzelak M, Hadfield J (2019) RNA sequencing: the teenage years. Nat Rev Genet
Su X, Wan Y, Xie L, Lin X, Zhao H, Ju X, Fang A (2019) Expression of SUMO1P3 compared with SUMO1 is an independent predictor of patient outcome in lung adenocarcinoma. Med Sci Monit 25:6691–6701
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z (2017) GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res 45:W98–w102
Tedesco D, Zhang J, Trinh L, Lalehzadeh G, Meisner R, Yamaguchi KD, Ruderman DL, Dinter H, Zajchowski DA (2007) The ubiquitin-conjugating enzyme E2-EPF is overexpressed in primary breast cancer and modulates sensitivity to topoisomerase II inhibition. Neoplasia (New York, NY) 9:601
Thakur MK, Gadgeel SM (2016) Predictive and prognostic biomarkers in non-small cell lung cancer. Semin Respir Crit Care Med 37:760–770
Travis WD, Asamura H, Bankier AA, Beasley MB, Detterbeck F, Flieder DB, Goo JM, MacMahon H, Naidich D, Nicholson AG, Powell CA, Prokop M, Rami-Porta R, Rusch V, van Schil P, Yatabe Y (2016) The IASLC lung cancer staging project: proposals for coding T categories for subsolid nodules and assessment of tumor size in part-solid tumors in the forthcoming eighth edition of the TNM classification of lung cancer. J Thorac Oncol 11:1204–1223
Turajlic S, Sottoriva A, Graham T, Swanton C (2019) Resolving genetic heterogeneity in cancer. Nat Rev Genet 20:404–416
Vos SM, Tretter EM, Schmidt BH, Berger JM (2011) All tangled up: how cells direct, manage and exploit topoisomerase function. Nat Rev Mol Cell Biol 12:827–841
Wang JC (2002) Cellular roles of DNA topoisomerases: a molecular perspective. Nat Rev Mol Cell Biol 3:430–440
Wang H, Gu R, Tian F, Liu Y, Fan W, Xue G, Cai L, Xing Y (2019) PHLPP2 as a novel metastatic and prognostic biomarker in non-small cell lung cancer patients. Thorac Cancer 10:2124–2132
Wistuba II, Behrens C, Lombardi F, Wagner S, Fujimoto J, Raso MG, Spaggiari L, Galetta D, Riley R, Hughes E, Reid J, Sangale Z, Swisher SG, Kalhor N, Moran CA, Gutin A, Lanchbury JS, Barberis M, Kim ES (2013) Validation of a proliferation-based expression signature as prognostic marker in early stage lung adenocarcinoma. Clin Cancer Res 19:6261–6271
Yu G, Wang LG, Han Y, He QY (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16:284–287
Zheng H, Zhan Y, Zhang Y, Liu S, Lu J, Yang Y, Wen Q, Fan S (2019) Elevated expression of G3BP1 associates with YB1 and p-AKT and predicts poor prognosis in nonsmall cell lung cancer patients after surgical resection. Cancer Med 8:6894–6903
Acknowledgements
We thank all the staff in the Department of Thoracic Surgery for their support during the study.
Funding
This study was supported by the National Key R&D Program of China (2017YFC1311000), the CAMS Initiative for Innovative Medicine (2017-I2M-1-005, 2019-I2M-2-002), and the Institutional Fundamental Research Funds (2018PT32033).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Ethic approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the ethics committee of National Cancer Center/Cancer Hospital, Chinese Academy of Medical Sciences and Peking Union Medical College and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Guo, W., Sun, S., Guo, L. et al. Elevated TOP2A and UBE2C expressions correlate with poor prognosis in patients with surgically resected lung adenocarcinoma: a study based on immunohistochemical analysis and bioinformatics. J Cancer Res Clin Oncol 146, 821–841 (2020). https://doi.org/10.1007/s00432-020-03147-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-020-03147-4