Abstract
Purpose
Bevacizumab improves survival in patients with metastatic colorectal cancer (mCRC) under chemotherapy, but few predictive markers have been identified.
Methods
To investigate chemosensitive single nucleotide polymorphisms (SNPs) of mCRC, we performed exome sequencing and RNA sequencing in 19 patients. A clinical association analysis was performed with the other 116 patients who had received chemotherapy to bevacizumab regimens. In vivo biodistribution studies and [18F]FDG-PET imaging were performed on mice bearing human colorectal cancer (HCT116 and SW480) xenografts after injection of bevacizumab with 5-FU, leucovorin, and irinotecan (FOLFIRI).
Results
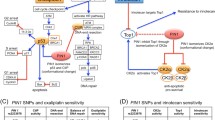
PPP1R15A rs557806 showed the most significant association with FRB-driven tumor IR in exome sequencing and the highest correlation (r = 0.74) with drug responses in RNA sequencing. Patients homozygous for the reference alleles (GG) of PPP1R15A rs557806 exhibited greater disease control rate and a tendency toward greater objective response rate (ORR) than those with homozygous or heterozygous substitution alleles (GC and CC; P = 0.027 and 0.073, respectively). In xenografted mice, HCT116 clones transfected with the G allele at PPP1R15A rs557806 were more sensitive to bevacizumab regimens than those with the C allele. Tumor volume of xenografts with the G allele was significantly lower than that of xenografts with the C allele (P = 0.004, day 13). [18F]FDG uptake decreased to 75 % in HCT116 xenograft-bearing mice with the G allele, whereas [18F]FDG uptake was 42 % in mice xenografts with the C allele (P = 0.032). ANXA11 rs1049550, a predictive biomarker of SNP described in our previous study, was validated using the xenograft model. Tumor volume and [18F]FDG uptake analyses showed that tumors in the SW480 xenografts expressing the substitution allele (T) at ANXA11 rs1049550 were more susceptible to FOLFIRI plus bevacizumab-induced suppression than those expressing the reference allele (C) (P = 0.001 and 0.026, respectively).
Conclusion
ANXA11 rs1049550 and PPP1R15A rs557806 may improve the identification of mCRC patients sensitive to bevacizumab regimens, and further validation is required in large cohorts.




Similar content being viewed by others
References
Ashktorab H, Daremipouran M, Devaney J et al (2015) Identification of novel mutations by exome sequencing in African American colorectal cancer patients. Cancer 121:34–42. doi:10.1002/cncr.28922
Celik B, Yalcin AD, Bisgin A, Dimitrakopoulou-Strauss A, Kargi A, Strauss LG (2013) Level of TNF-related apoptosis-inducing-ligand and CXCL8 correlated with 2-[18F]Fluoro-2-deoxy-d-glucose uptake in anti-VEGF treated colon cancers. Med Sci Monit 19:875–882. doi:10.12659/msm.889605
Cifola I, Pietrelli A, Consolandi C et al (2013) Comprehensive genomic characterization of cutaneous malignant melanoma cell lines derived from metastatic lesions by whole-exome sequencing and SNP array profiling. PLoS ONE 8:e63597. doi:10.1371/journal.pone.0063597
De Angelis PM, Svendsrud DH, Kravik KL, Stokke T (2006) Cellular response to 5-fluorouracil (5-FU) in 5-FU-resistant colon cancer cell lines during treatment and recovery. Mol Cancer 5:20. doi:10.1186/1476-4598-5-20
De Mattia E, Cecchin E, Toffoli G (2015) Pharmacogenomics of intrinsic and acquired pharmacoresistance in colorectal cancer: toward targeted personalized therapy. Drug Resist Updat 20:39–70. doi:10.1016/j.drup.2015.05.003
Fang LT, Lee S, Choi H et al (2014) Comprehensive genomic analyses of a metastatic colon cancer to the lung by whole exome sequencing and gene expression analysis. Int J Oncol 44:211–221. doi:10.3892/ijo.2013.2150
Hegde PS, Jubb AM, Chen D et al (2013) Predictive impact of circulating vascular endothelial growth factor in four phase III trials evaluating bevacizumab. Clin Cancer Res 19:929–937. doi:10.1158/1078-0432.ccr-12-2535
Hollander MC, Sheikh MS, Yu K et al (2001) Activation of Gadd34 by diverse apoptotic signals and suppression of its growth inhibitory effects by apoptotic inhibitors. Int J Cancer 96:22–31
Jubb AM, Harris AL (2010) Biomarkers to predict the clinical efficacy of bevacizumab in cancer. Lancet Oncol 11:1172–1183. doi:10.1016/s1470-2045(10)70232-1
Kim JC, Kim SY, Cho DH et al (2011) Novel chemosensitive single-nucleotide polymorphism markers to targeted regimens in metastatic colorectal cancer. Clin Cancer Res 17:1200–1209. doi:10.1158/1078-0432.ccr-10-1907
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL (2013a) TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14:R36. doi:10.1186/gb-2013-14-4-r36
Kim JC, Ha YJ, Roh SA et al (2013b) Feasibility of proposed single-nucleotide polymorphisms as predictive markers for targeted regimens in metastatic colorectal cancer. Br J Cancer 108:1862–1869. doi:10.1038/bjc.2013.163
Koboldt DC, Zhang Q, Larson DE et al (2012) VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res 22:568–576. doi:10.1101/gr.129684.111
Koutras AK, Antonacopoulou AG, Eleftheraki AG et al (2012) Vascular endothelial growth factor polymorphisms and clinical outcome in colorectal cancer patients treated with irinotecan-based chemotherapy and bevacizumab. Pharmacogenomics J 12:468–475. doi:10.1038/tpj.2011.37
Lambrechts D, Lenz HJ, de Haas S, Carmeliet P, Scherer SJ (2013) Markers of response for the antiangiogenic agent bevacizumab. J Clin Oncol 31:1219–1230. doi:10.1200/jco.2012.46.2762
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25:1754–1760. doi:10.1093/bioinformatics/btp324
Liu X, Jian X, Boerwinkle E (2013) dbNSFP v2.0: a database of human non-synonymous SNVs and their functional predictions and annotations. Hum Mutat 34:E2393–E2402. doi:10.1002/humu.22376
Luo HY, Xu RH (2014) Predictive and prognostic biomarkers with therapeutic targets in advanced colorectal cancer. World J Gastroenterol 20:3858–3874. doi:10.3748/wjg.v20.i14.3858
McKenna A, Hanna M, Banks E et al (2010) The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303. doi:10.1101/gr.107524.110
Mertens J, De Bruyne S, Van Damme N et al (2013) Standardized added metabolic activity (SAM) IN (1)(8)F-FDG PET assessment of treatment response in colorectal liver metastases. Eur J Nucl Med Mol Imaging 40:1214–1222. doi:10.1007/s00259-013-2421-z
Meyerson M, Gabriel S, Getz G (2010) Advances in understanding cancer genomes through second-generation sequencing. Nat Rev Genet 11:685–696. doi:10.1038/nrg2841
Pavlidis ET, Pavlidis TE (2013) Role of bevacizumab in colorectal cancer growth and its adverse effects: a review. World J Gastroenterol 19:5051–5060. doi:10.3748/wjg.v19.i31.5051
Purcell S, Neale B, Todd-Brown K et al (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575. doi:10.1086/519795
Therasse P, Arbuck SG, Eisenhauer EA et al (2000) New guidelines to evaluate the response to treatment in solid tumors. European Organization for Research and Treatment of Cancer, National Cancer Institute of the United States, National Cancer Institute of Canada. J Natl Cancer Inst 92:205–216. doi:10.1093/jnci/92.3.205
Acknowledgments
This work was supported in part by Grants (to J.C. Kim) from the Korea Research Foundation (2013R1A2A1A03070986), Ministry of Science, ICT, and Future Planning, and the Korea Health 21 R&D Project (HI06C0868 and HI13C1750), Ministry of Health, Welfare, and Family Affairs, Republic of Korea.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
None.
Ethical approval
The study protocol was approved by the Institutional Review Board for Human Genetic and Genomic Research (Registration No. 2014-0150). All participants provided their written informed consent. This study was also reviewed and approved by the Institutional Animal Care and Use Committee of the Asan Institute for Life Sciences, Asan Medical Center, Seoul, Korea (Registration No. 2014-03-038).
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Roh, S.A., Park, I.J., Yoon, Y.S. et al. Feasibility of novel PPP1R15A and proposed ANXA11 single nucleotide polymorphisms as predictive markers for bevacizumab regimen in metastatic colorectal cancer. J Cancer Res Clin Oncol 142, 1705–1714 (2016). https://doi.org/10.1007/s00432-016-2177-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-016-2177-5