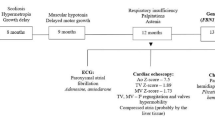Abstract
The cardinal phenotypic hallmarks of Marfan syndrome (MFS) include cardiac, ocular, and skeletal abnormalities. Since the clinical phenotype of MFS is highly heterogeneous, with certain symptoms appearing as children age, the diagnostic process and establishing a genotype-phenotype association in childhood MFS can be challenging. The lack of sufficient childhood studies also makes it difficult to interpret the subject. This study aims to evaluate the relationship between clinical symptoms used as diagnostic criteria and FBN1 variations in children with MFS. This study investigated the relationships between genotypes and phenotypes in 131 children suspected of having Marfan syndrome (MFS). Diagnosis of MFS was made according to the revised Ghent nosology. FBN1 variants were categorized based on exon regions, type of variant, and pathogenicity classes. These FBN1 variants were then correlated with the clinical manifestations including cardiovascular, ocular, facial, and skeletal abnormalities. Out of the children, 43 were diagnosed with MFS. FBN1 variant was identified in 32 (74.4%) of the MFS children. MFS diagnosis could not be made in five (15.6%) FBN1 variant-positive children. The most common cardinal finding is cardiac anomalies n = 38 (88.3%). The most common FBN1 pathogenic variant was c.1786 T > C/p.Cys596Arg n = 4 (12.5%). The distribution of pathogenic variants was as follows: 29 (90.6%) missense, 2 (6.3%) frameshift, and 1 (3.1%) nonsense. The numbers of AD and EL of the variant-positive children were 16 (50%) and 14 (43.7%), respectively. Ocular abnormalities were more common in children with FBN1-positive MFS (p = 0.009). There was no difference in the number of cardiac abnormalities between FBN1-positive and FBN1-negative MFS patients (p = 0.139).
Conclusion: This study examines the relationship between FBN1 variants and clinical features used as diagnostic criteria in MFS children. The findings emphasize the importance of long-term monitoring of heterogeneous clinical phenotypes and bioinformatic reanalysis in determining the genotype-phenotype relationship in children, as MFS symptoms can vary with age.
What is Known: • Marfan syndrome has highly variable phenotypic heterogeneity. • The genotype-phenotype relationship in childhood Marfan syndrome is not clear enough due to the variation in the time of onset of the findings. | |
What is New: • This article provides regional data for the field of research on genotype-phenotype relationships in childhood Marfan syndrome. • Long-term follow-up of clinical findings and bioinformatics reanalysis is an important requirement for a well-established genotype-phenotype relationship in childhood Marfan syndrome. |
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Marfan syndrome (MFS) is a type 1 fibrillinopathy, a connective tissue disease caused by FBN1 pathogenic variants [1, 2]. The main clinical findings involve the cardiovascular (CVS), ocular, and skeletal systems. Additionally, patients may exhibit cutaneous, respiratory, integumental, and nervous system findings [3]. MFS is an autosomal dominant disease, with approximately 25% of cases resulting from de novo variants [4, 5].
One of the most common manifestations is cardiovascular findings, which occur in 60–80% of cases and are the leading cause of morbidity and premature death [6]. The typical ocular finding is ectopia lentis (EL). However, the most distinct phenotype in MFS is characterized by skeletal system findings [7]. The diagnosis of MFS is based on the revised Ghent nosology [8]. The criteria for diagnosis included family history, positivity for FBN1 variants, aortic dilatation, presence of ectopic lens, and systemic findings.
Research on the genotype-phenotype relationship in Marfan syndrome plays a crucial role in establishing prognostic criteria for other potential systemic findings that can result in disability, particularly cardiac anomalies that can lead to premature death [9]. Although conflicting results have been reported in previous studies, it has been found that onset under 1 year of age, aortic involvement, diaphragmatic herniation, and neonatal exon region variants are associated with early death or a severe clinical course [10].
Establishing a genotype-phenotype relationship in children with MFS has several important limitations, and there are not enough studies on the subject [2]. Firstly, clinical manifestations are highly heterogeneous, with the clinical phenotype displaying highly variable expression. Phenotypic variability and penetrance can vary even among individuals within the same family [9]. Secondly, since the characteristic findings of the disease develop over time, diagnosing and predicting prognosis in pediatric patients can be challenging [11]. The timing of symptom onset and the specific distribution of manifestations in different tissues contribute to phenotypic variability [12]. Previous studies have reported that aortic anomalies, ectopic lens, and specific skeletal findings differ between individuals under and above the age of 18 [13, 14]. Thirdly, other disorders with symptoms similar to MFS, such as mitral valve prolapse syndrome or Shprintzen-Goldberg syndrome, can complicate the genotype-phenotype correlation due to fibrillinopathies caused by TGFβRII variants [15, 16]. Inconsistencies between the revised Ghent criteria developed for diagnostic purposes and FBN1 variant positivity can sometimes make it difficult to establish both a diagnosis and genotype-phenotype correlation [17].
The aim of this study is to investigate the genotype and phenotype characteristics used as diagnostic criteria for Marfan syndrome in Turkish children.
Materials and methods
This retrospective study was conducted at a single center and involved 131 children who underwent FBN1 screening due to suspicion of Marfan syndrome between 2016 and 2021.
One hundred seventy-three patients were referred from orthopedics, ophthalmology, cardiology, and pediatrics outpatient clinics with suspicion of MFS (Fig. 1). Clinical findings suggestive of MFS included tall stature (height > +2.5 SDS), family history, arachnodactyly, lens subluxation, and anomalies of the chest, face, and extremities. FBN1 screening was not performed in 42 patients with another specific diagnosis: familial tall stature (n = 34), isolated cardiovascular findings (n = 4), and skeletal system findings (n = 4). FBN1 screening was performed to 131 patients who presented with at least one MFS-related finding [18]. These findings included a family history, clinical signs, and symptoms suggestive of Marfan syndrome, such as a Marfanoid appearance (long limbs, long face, thumb sign, wrist sign), aortic dilatation, other specific cardiovascular findings, skeletal system abnormalities, or ectopia lentis. In addition, a total of 131 patients underwent FBN1 molecular analysis. FBN1 screening was conducted on children who could not be diagnosed with MFS according to the revised Ghent criteria but were strongly suspected of having MFS. These children were mostly those for whom specific findings were expected to develop with age. Additionally, FBN1 screening was scheduled for children whose height exceeded their midparental height. Several cases were referred from various departments (ophthalmology, orthopedics, cardiology, etc.) with an FBN1 analysis report. The researchers collected clinical signs, symptoms, and genotype characteristics. Out of the 43 patients diagnosed with MFS according to the Ghent nosology criteria, 16 were found to have the FBN1 variant. The diagnosis of MFS was excluded in 88 cases, with the FBN1 variant detected in 5 participants.
To describe clinical signs and symptoms of Marfan syndrome, annotations were sourced from the Human Phenotype Ontology, a standardized dictionary of phenotypic abnormalities in human diseases. These annotations covered a range of phenotypic abnormalities including growth (disproportionate tall stature, slender build), head and neck (long and narrow face, dental crowding, high narrow palate, downslanted palpebral fissures, open bite), eye (ectopia lentis, lens subluxation, lens luxation, glaucoma, retinal detachment, hypoplasia of the iris, increased axial length of the globe, flat cornea visual impairment, myopia), cardiovascular (ascending tubular aorta aneurysm, ascending aortic dissection, arterial dissection, mitral valve prolapse, tricuspid regurgitation, mitral regurgitation, bicuspid aorta, congestive heart failure), skeletal system (pectus carinatum, pectus excavatum, arachnodactyly, joint hypermobility, scoliosis, protrusio acetabuli, abnormal zygomatic bone morphology, dolichocephaly, retrognathia, micrognathia, kyphosis, limited elbow movement, osteoporosis, mitral valve calcification), and nervous system (dural ectasia). Marfanoid habitus was defined as a set of clinical signs and symptoms, with the most common features being disproportionate tall stature, long arms and legs, arachnodactyly, and joint hypermobility.
The researchers gathered data on family history, cardiac and ocular findings, and skeletal findings. They also documented measurements of height, weight, and body mass index. Aortic diameter Z-scores were extracted from echocardiography records. Aortic diameters were calculated using an automated software created by Cantinotti et al. [19]. The program captured measurements such as aortic valve annulus (mm), sinus of Valsalva (mm), sino-tubular junction (mm), and ascending aorta (mm) from the echocardiography results. The revised Ghent nosology was utilized to categorize phenotypes [18]. Systemic findings were employed as diagnostic criteria for cases with a score of ≥ 7. Marfan syndrome (MFS) was diagnosed based on the outlined criteria in the Ghent nosology.
Molecular analysis
The NGS (next-generation sequencing) analysis method was utilized to analyze the FBN1 gene. Primers covering the specified exons of the gene were employed. DNA was extracted from blood samples collected in EDTA tubes. The DNA concentration was measured using a Qubit 3.0 device (ThermoScientific, USA) and adjusted to a final concentration of 2 nanograms per microliter. Initial PCR steps (library preparation) were conducted with the obtained DNA samples and primer pairs specifically designed for sequencing the FBN1 gene, not for complete panel sequencing. The library fragments acted as templates for synthesizing new DNA fragments. Following the library preparation phase, the manufacturer’s kit was utilized for the emulsion PCR steps, which were carried out on the Illumina MiSeq device (USA) according to their protocols. The digital recording of the introduction of nucleotides into the growing DNA helix took place. The sample mixtures were processed using the relevant kits on the Illumina MiSeq device and loaded onto the appropriate chips to initiate the sequencing stage.
Variant classification
FBN1 variants were identified as exon regions that exhibited a strong genotype-phenotype correlation based on previous data. Consequently, the exon regions were categorized into three groups: N-terminus (exon 1–21), middle region (exon 22–42), and C-terminus (exon 43–65). The relationship between genotypes in these regions and the manifestations in the patients was assessed.
The pathogenicity classification of variants was determined in accordance with the 2015 American College of Medical Genetics (ACMG) standards and guidelines [20]: Class 1 (benign), Class 2 (likely benign), Class 3 VUS (variant of unknown significance), Class 4 (likely pathogenic), and Class 5 (pathogenic). Specific guidelines for FBN1 were utilized to determine variant pathogenicity based on ACMG [21]. The pathogenicity of the variants was compared to the phenotypic abnormalities.
Bioinformatics analyses were conducted using various databases (Encode, ClinVar, OMIM, COSMIC, PubMed, HGMD, PGMD, Ensemble, Mutation Taster, Varsome). The variants’ domains were identified using Prosit (http://www.expasy.org/prosite/) and Smart (https://www.smartgene.com/). The MutationTaster analysis program was employed for in silico analysis.
The cases were clinically divided into two groups based on FBN1 positivity and cardiac findings: FBN1 positive and negative groups, and cardiac findings positive and negative groups.
Statistics
Quantitative variables were defined as mean ± standard deviation, while qualitative variables were expressed as percentages. Group comparisons were conducted using the chi-square (χ2) test, with the Fisher test used if the chi-square test was unsuccessful. The distribution was tested using the Kolmogorov-Smirnov test. The Student t test was used for normally distributed variables, and the Mann–Whitney U test was used for non-normally distributed variables. Statistical analyses were conducted using SPSS v21 (IBM Corporation, New York, USA).
Results
Out of the 131 participants screened for FBN1, 72 (55%) were male. Table 1 displays the anthropometric and clinical characteristics of the participants (n = 131) who were screened for FBN1. It was found that CVS, ocular, and facial findings were significantly higher in the FBN1 positive group (p < 0.001, p < 0.001, and p = 0.007), while no significant difference was observed between the groups in terms of the frequency of skeletal findings (p = 0.134). The most common manifestation was CVS findings (n = 61) (48%). Additional CVS findings can be found in Table 1.
The mean age at diagnosis for cases with AD and EL was as follows: 12.5 ± 5.43 and 12.96 ± 4.79, respectively. Figure 2 shows the percentage distribution of clinical manifestations by age groups.
Tables 2 and 3 provide clinical phenotypes associated with the modified Ghent criteria. Out of all the participants, 43 children (32.82%) were diagnosed with Marfan syndrome (MFS). The FBN1 variant was detected in 32 patients (74.4%) diagnosed with MFS in children. Out of the 32 FBN1 variants, 27 (63%) were diagnosed with MFS, while the remaining 5 were still being followed up for a possible diagnosis of MFS. Among the patients with manifestations related to MFS, 5 patients were FBN1 positive (3.8%), while 83 (63.35%) were FBN1 negative according to the Ghent criteria.
There were 16 cases (12.21%) diagnosed with MFS without an FBN1 variant (Table 4). Table 3 displays the anthropometric and clinical characteristics of patients diagnosed with MFS (n = 43) in relation to FBN1 positivity. While there were no differences between the FBN1 positive and negative groups in terms of CVS, skeletal, and facial findings (p = 1, p = 0.832, and p = 0.188), ocular findings were more frequently observed in the FBN1 positive group (p = 0.009). Other findings are detailed in Table 4.
Out of all cases (n = 131), 61 (46.56%) exhibited cardiovascular manifestations (Table 5). Table 4 illustrates the relationship between aortic root dilatation and other clinical features in children diagnosed with Marfan syndrome. There were no differences in the frequency of accompanying ocular, skeletal, and facial findings between the groups with and without aortic root dilatation.
The distribution of FBN1 variants based on three exon regions was as follows (Table 6). Table 6 displays the distribution of pathogenic variants and phenotypic abnormalities based on exon regions. The numbers of all variants were as follows: missense n = 29 (90.6%), frameshift n = 2 (6.4%), and nonsense n = 1 (3%). All cases with Cys-missense variants had aortic dilatation. The distribution of FBN1 variant domains was as follows: EGF-like n = 15 (46.8%), cbEGF-like n = 9 (28%), and TGFb n = 8 (25.2%).
The frequency of the most common FBN1 pathogenic variants was determined as follows: c.1786 T > C/p.Cys596Arg (exon 15) (n = 4, 12.5%) and c.7964C > T/p.Ala2655Val (exon 64) (n = 4, 12.5%), and c.4450A > G/p.N1484D (exon 36) (n = 2, 6.25%) (Table 7). Table 7 also shows the association of pathogenic variants with skeletal and facial abnormalities.
Discussion
This study evaluates the genotype-phenotype association in children with Marfan syndrome. There have been few published reports on this subject in children to date [22, 23]. It contributes to the limited number of pediatric reports that include genotype-phenotype associations in Marfan syndrome. This study revealed that the most common cardinal finding in children with MFS is cardiac anomalies. The Ghent criteria present diagnostic challenges in children, and there was no difference in the frequency of cardiac findings between FBN1-positive and FBN1-negative children. Ocular findings were more common in children with FBN1-positive MFS. Additionally, although the study did not have a large enough sample size to establish a well-defined genotype-phenotype relationship, it provides data for clinical phenotypes associated with some exonic regions/domains.
Previous studies have reported varying rates of FBN1 variant positivity in cases screened with suspicion of MFS. In a study screening for suspected MFS, the FBN1 variant was identified in 131 (40%) of 327 pediatric patients [11]. Out of the cases, 202 (61.17%) were diagnosed with Marfan syndrome (MFS) using the revised Ghent criteria. Among these cases, 39% had a clinical diagnosis. The average age at which genetic screening was conducted was 7.8 ± 5.4 years. In our study, an FBN1 variant was identified in 32 out of 131 participants (24.4%). The detection rate of FBN1 variants appears to be relatively low. Although the rate reported is acceptable compared to other studies (40% in Stark et al.), the data are not directly comparable [23]. The referenced study mainly reported (likely) pathogenic variants, while our study included all variants in the calculations. Additionally, the number of children diagnosed with the Ghent criteria was lower (35.5%) compared to the previous study. In another study conducted in adults at a different center, FBN1 variant positivity (n = 51) was found to be 67.1% in 76 patients who underwent genetic screening [23]. This difference seems to be due to variances in patient selection. In our study, another reason for the low FBN1 positivity rate may be that screening indications were broadened in children presenting with tall stature compared to their midparental height. Based on the data from the literature, it is clear that selecting the suitable cases for genetic screening is crucial. Out of the cases, 16 (37.2%) were diagnosed based on the Ghent criteria without detecting any FBN1 variant. As mentioned above, low FBN1 positivity has been reported in children diagnosed with MFS in previous studies. There could be several reasons for this low FBN1 positivity. These patients may have one of the MFS-like fibrillinopathies, but unfortunately, they were not screened for these diseases. Another reason could be the diagnostic challenges posed by the Ghent criteria in children, or there may be differences in the definition of clinical phenotypes that underlie the Ghent criteria [11, 17, 24].
On the other hand, while an FBN1 variant was detected in 5 cases, they were not diagnosed according to the Ghent criteria. This suggests that the presentation of MFS with characteristic clinical findings that develop with age, and the use of Ghent criteria in childhood, may lead to diagnostic inconsistencies. It is stated that the Ghent criteria are not reliable in children [24]. These findings should be followed up on, as specific changes will occur with age. This needs to be considered when evaluating the Ghent criteria in the childhood age group. Similarly, the age-dependent appearance of these findings also leads to differences in genotype-phenotype correlation. In a study involving the adult age group, the rate of patients diagnosed with MFS using the Ghent criteria was reported as 93.3% [16]. This high rate could be due to the adult age of the subjects, with a mean age of 41.33 ± 3.77. Therefore, cross-sectional studies in childhood may not fully reveal the genotype-phenotype relationship [24].
Previous studies have reported an association between the severity of cardiac findings and frameshift and nonsense variant types [25, 26]. However, in our study, we did not find any association with cardiac or other clinical manifestations. This lack of difference may be due to the relatively small number of cases in our cohort. Additionally, the age of the cases may also contribute to this discrepancy. The presence of cardiac findings and their impact on life expectancy, as well as the rate of progression of aortic dilatation, may vary with advancing age.
The variant p.Pro1148Ala c.3442 C > G was reported as benign in the ClinVar database. Cardiac findings were detected in all 3 cases with this variant, and EL was also present in one case. In silico analysis also classified it as benign strong. This variant is a frequently observed polymorphism in previous studies [27]. In our study, these 3 cases were diagnosed as MFS cases according to the Ghent criteria. Unfortunately, parental segregation analysis could not be performed for the same variant due to a possible presence of another variant in the case. Comprehensive bioinformatic re-analyses were planned for these 3 cases with the delegation, but the parents did not cooperate. Additionally, regulations of social security institutions posed an obstacle to this. Reanalysis of exon data should be standard practice for patients. For these cases, the reverse phenotyping approach may provide a potential breakthrough [28]. Today, reverse phenotyping appears to be a useful approach that will change our conventional understanding of establishing the genotype-phenotype relationship. While previous approaches have focused on assigning phenotypes to individuals with genetic disorders, the reverse phenotyping approach involves recruiting individuals with a certain genotype and analyzing hypotheses for possible predicted phenotypes. Recognizing phenotypes from genotypes offers a rational tool for comprehensive insight into the molecular classification of diseases.
When evaluating the concordance between genotype and phenotype based on pathogenicity classes according to ACMG, although a well-established phenotypic association with some exon regions and pathogenic variants is known in patients with MFS, there are also incompatibilities in some cases [2]. In the current study, the number of patients who participated is insufficient to definitively state that there is no well-established correlation between clinical findings and genotypes based on pathogenicity classes and exon regions in children with Marfan syndrome. Further research with a larger sample size is necessary to establish a strong genotype-phenotype relationship in children with MFS.
Limitations
There are several limitations to this study. A comprehensive molecular panel analysis was not conducted to determine if patients diagnosed with MFS clinically had genetic variants associated with similar disorders such as Loeys-Dietz syndrome or isolated ectopia lentis. Only FBN1 sequencing analysis was performed, making it impossible to distinguish between conditions like isolated ectopia lentis or MFS. The fact that neither a panel nor CNV-sequencing (or software equivalent, e.g., ExomeDepth) was used is a significant limitation, since patients with exon deletions or Loeys-Dietz syndrome could have been included in the FBN1 negative group. This is especially important in the patient who presented with an aortic dissection and in the two patients with aortic dilatation and EL, in whom a pathogenic variant was not found. Additionally, the study was unable to test the tissue-specific expression of FBN1 with different transcripts, which could be a potential cause of phenotypic variability [29]. Another limitation is the inability to perform CNV sequencing or other techniques to detect larger insertions/deletions in the FBN1 gene. Similarly, the retrospective nature of the study constituted another limitation. Parental segregation analyses could not be performed when necessary. Of the patients with a family history, only 3 parents had previous FBN1 analysis results, and they had the same pathogenic variant as the children with MFS. Unfortunately, pathogenic FBN1 variants detected with NGS could not be validated by Sanger sequencing in this genotype-phenotype incompatibility. Ultimately, this is a retrospective study, and genotype-phenotype discrepancies may result from differences in study designs, ranging from patient selection criteria to the definition of clinical phenotypes.
Conclusion
This study presents data on diagnostic clinical findings of childhood MFS and associated FBN1 variants. It indicates that the Ghent diagnostic criteria, which include clinical findings and FBN1 variant features, have some diagnostic difficulties in childhood MFS. It also emphasizes the importance of performing bioinformatic reanalysis to evaluate the genotype-phenotype relationship due to the changing nature of clinical phenotypes with age.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Sakai LY, Keene DR, Renard M, De Backer J (2016) FBN1: the disease-causing gene for Marfan syndrome and other genetic disorders. Gene 591(1):279–291. https://doi.org/10.1016/j.gene.2016.07.033. Epub 2016 Jul 18. PMID: 27437668; PMCID: PMC6639799
Chen ZX, Jia WN, Jiang YX (2022) Genotype-phenotype correlations of marfan syndrome and related fibrillinopathies: phenomenon and molecular relevance. Front Genet 16(13):943083. https://doi.org/10.3389/fgene.2022.943083. PMID: 36176293; PMCID: PMC9514320
Groth KA, Hove H, Kyhl K, Folkestad L, Gaustadnes M, Vejlstrup N et al (2015) Prevalence, incidence, and age at diagnosis in Marfan Syndrome. Orphanet J Rare Dis 2(10):153. https://doi.org/10.1186/s13023-015-0369-8. PMID: 26631233; PMCID: PMC4668669
Chiu HH, Wu MH, Chen HC, Kao FY, Huang SK (2014) Epidemiological profile of Marfan syndrome in a general population: a national database study. Mayo Clin Proc 89(1):34–42. https://doi.org/10.1016/j.mayocp.2013.08.022. PMID: 24388020
Collod-Béroud G, Le Bourdelles S, Ades L, Ala-Kokko L, Booms P, Boxer M et al (2003) Update of the UMD-FBN1 mutation database and creation of an FBN1 polymorphism database. Hum Mutat 22(3):199–208. https://doi.org/10.1002/humu.10249. PMID: 12938084
Chung BH, Lam ST, Tong TM, Li SY, Lun KS, Chan DH et al (2009) Identification of novel FBN1 and TGFBR2 mutations in 65 probands with Marfan syndrome or Marfan-like phenotypes. Am J Med Genet A 149A(7):1452–1459. https://doi.org/10.1002/ajmg.a.32918. PMID: 19533785
Becerra-Muñoz VM, Gómez-Doblas JJ, Porras-Martín C, Such-Martínez M, Crespo-Leiro MG, Barriales-Villa R et al (2018) The importance of genotype-phenotype correlation in the clinical management of Marfan syndrome. Orphanet J Rare Dis 13(1):16. https://doi.org/10.1186/s13023-017-0754-6. PMID: 29357934; PMCID: PMC5778633
Faivre L, Collod-Beroud G, Loeys BL, Child A, Binquet C, Gautier E et al (2007) Effect of mutation type and location on clinical outcome in 1,013 probands with Marfan syndrome or related phenotypes and FBN1 mutations: an international study. Am J Hum Genet 81(3):454–466. https://doi.org/10.1086/520125. Epub 2007 Jul 25. PMID: 17701892; PMCID: PMC1950837
Marelli S, Micaglio E, Taurino J, Salvi P, Rurali E, Perrucci GL et al (2023) Marfan Syndrome: enhanced diagnostic tools and follow-up management strategies. Diagnostics (Basel) 13(13):2284. https://doi.org/10.3390/diagnostics13132284. PMID: 37443678; PMCID: PMC10340634
Stheneur C, Faivre L, Collod-Béroud G, Gautier E, Binquet C, Bonithon-Kopp C, Claustres M et al (2011) Prognosis factors in probands with an FBN1 mutation diagnosed before the age of 1 year. Pediatr Res 69(3):265–270. https://doi.org/10.1203/PDR.0b013e3182097219. PMID: 21135753
Stark VC, Hensen F, Kutsche K, Kortüm F, Olfe J, Wiegand P et al (2020) Genotype-phenotype correlation in children: the impact of FBN1 variants on pediatric Marfan care. Genes (Basel) 11(7):799. https://doi.org/10.3390/genes11070799. PMID: 32679894; PMCID: PMC7397236
Dietz HC (2015) Potential phenotype-genotype correlation in Marfan syndrome: when less is more? Circ Cardiovasc Genet 8(2):256–260. https://doi.org/10.1161/CIRCGENETICS.115.001040. PMID: 25901038
Seo GH, Kim YM, Kang E, Kim GH, Seo EJ, Lee BH et al (2018) The phenotypic heterogeneity of patients with Marfan-related disorders and their variant spectrums. Medicine (Baltimore) 97(20):e10767. https://doi.org/10.1097/MD.0000000000010767. PMID: 29768367; PMCID: PMC5976283
Hannuksela M, Lundqvist S, Carlberg B (2006) Thoracic aorta—dilated or not? Scand Cardiovasc J 40:175–178. https://doi.org/10.1080/14017430600565999
Becerra-Muñoz VM, Gómez-Doblas JJ, Porras-Martín C, Such-Martínez M, Crespo-Leiro MG, Barriales-Villa R et al (2018) The importance of genotype-phenotype correlation in the clinical management of Marfan syndrome. Orphanet J Rare Dis 13(1):16. https://doi.org/10.1186/s13023-017-0754-6. PMID: 29357934; PMCID: PMC5778633
Coelho SG, Almeida AG (2020) Marfan syndrome revisited: from genetics to the clinic. Rev Port Cardiol (Engl Ed). 39(4):215-226 English, Portuguese. https://doi.org/10.1016/j.repc.2019.09.008. Epub 2020 May 18. PMID: 32439107
Baudhuin L, Kotzer K, Lagerstedt S (2015) Decreased frequency of FBN1 missense variants in Ghent criteria-positive Marfan syndrome and characterization of novel FBN1 variants. J Hum Genet 60:241–252. https://doi.org/10.1038/jhg.2015.10
Loeys BL, Dietz HC, Braverman AC, Callewaert BL, De Backer J, Devereux RB et al (2010) The revised Ghent nosology for the Marfan syndrome. J Med Genet 47(7):476–485. https://doi.org/10.1136/jmg.2009.072785. PMID: 20591885
Cantinotti M, Giordano R, Scalese M, Murzi B, Assanta N, Spadoni I et al (2017) Nomograms for two-dimensional echocardiography derived valvular and arterial dimensions in Caucasian children. J Cardiol 69(1):208–215. https://doi.org/10.1016/j.jjcc.2016.03.010. Epub 2016 Apr 24 PMID: 27118699
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, Laboratory Quality Assurance Committee ACMG et al (2015) Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 17(5):405–424. https://doi.org/10.1038/gim.2015.30. PMID: 25741868; PMCID: PMC4544753
De Backer J (2022) ClinGen FBN1 expert panel specifications to the ACMG/AMP variant interpretation guidelines version 1. https://www.clinicalgenome.org/affiliation/50046/docs/assertion-criteria. Accessed 4 Jan 2022
Gezdirici A, Teralı K, Gülec EY, Bornaun H, Dogan M, Eröz R (2021) An integrated clinical and molecular study of a cohort of Turkish patients with Marfan syndrome harboring known and novel FBN1 variants. J Hum Genet 66(7):647–657. https://doi.org/10.1038/s10038-021-00899-w. Epub 2021 Jan 22 PMID: 33483584
Stark VC, Hensen F, Kutsche K, Kortüm F, Olfe J, Wiegand P et al (2020) Genotype-phenotype correlation in children: the ımpact of FBN1 variants on pediatric Marfan care. Genes (Basel) 11(7):799. https://doi.org/10.3390/genes11070799. PMID: 32679894; PMCID: PMC7397236
Dean J (2007) Marfan syndrome: clinical diagnosis and management. Eur J Hum Genet 15:724–733. https://doi.org/10.1038/sj.ejhg.5201851
Xu S, Li L, Fu Y, Wang X, Sun H, Wang J et al (2020) (2020) Increased frequency of FBN1 frameshift and nonsense mutations in Marfan syndrome patients with aortic dissection. Mol Genet Genomic Med 8(1):e1041. https://doi.org/10.1002/mgg3.1041. Epub 2019 Dec 12. PMID: 31830381; PMCID: PMC6978253
Stengl R, Ágg B, Pólos M, Mátyás G, Szabó G, Merkely B et al (2021) Potential predictors of severe cardiovascular involvement in Marfan syndrome: the emphasized role of genotype-phenotype correlations in improving risk stratification-a literature review. Orphanet J Rare Dis 16(1):245. https://doi.org/10.1186/s13023-021-01882-6. PMID: 34059089; PMCID: PMC8165977
Frédéric MY, Lalande M, Boileau C, Hamroun D, Claustres M, Béroud C, Collod-Béroud G (2009) UMD-predictor, a new prediction tool for nucleotide substitution pathogenicity – application to four genes: FBN1, FBN2, TGFBR1, and TGFBR2. Hum Mutat 30(6):952–959. https://doi.org/10.1002/humu.20970. PMID: 19370756
Wilczewski CM, Obasohan J, Paschall JE, Zhang S, Singh S, Maxwell GL, Similuk M, Wolfsberg TG, Turner C, Biesecker LG, Katz AE (2023) Genotype first: clinical genomics research through a reverse phenotyping approach. Am J Hum Genet 110(1):3–12. https://doi.org/10.1016/j.ajhg.2022.12.004. PMID: 36608682; PMCID: PMC9892776
Benarroch L, Aubart M, Gross MS, Arnaud P, Hanna N, Jondeau G et al (2019) Reference expression profile of three FBN1 transcript ısoforms and their association with clinical variability in Marfan syndrome. Genes (Basel) 10(2):128. https://doi.org/10.3390/genes10020128. PMID: 30754709; PMCID: PMC6409622
Acknowledgements
We would like to thank Mertcan Karaoglan for his contribution to the fluency of the manuscript in English.
Funding
Open access funding provided by the Scientific and Technological Research Council of Türkiye (TÜBİTAK).
Author information
Authors and Affiliations
Contributions
Karaoglan M: designed, collected data, interpreted analysis and wrote, Nacarkahya G: analyzed genetic data and performed molecular analysis, Aytaç Emel, collected data, Keskin M, designed and collected data.
Corresponding author
Ethics declarations
Ethical approval
The authors declare adherence to international ethical standards. All the procedures carried out in the research with the participation of humans were in compliance with the ethical standards of the National Research Ethics Committee and with the Helsinki Declaration of 1964 and its subsequent changes or with comparable ethics standards. The study was approved by the Gaziantep University Clinical Research Ethics Committee.
Consent to participate
Informed voluntary consent was obtained from every participant of the study".
Conflict of interest
The authors declare no competing interests.
Additional information
Communicated by Peter de Winter
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Karaoglan, M., Nacarkahya, G., Aytac, E.H. et al. Genotype and clinical phenotype of children with Marfan syndrome in Southeastern Anatolia. Eur J Pediatr 183, 3219–3232 (2024). https://doi.org/10.1007/s00431-024-05579-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00431-024-05579-3