Abstract
Non-human primates are extensively used in neuroscience research as models of the human brain, with the rhesus macaque being a prominent example. We have previously introduced a set of tractography protocols (XTRACT) for reconstructing 42 corresponding white matter (WM) bundles in the human and the macaque brain and have shown cross-species comparisons using such bundles as WM landmarks. Our original XTRACT protocols were developed using the F99 macaque brain template. However, additional macaque template brains are becoming increasingly common. Here, we generalise the XTRACT tractography protocol definitions across five macaque brain templates, including the F99, D99, INIA, Yerkes and NMT. We demonstrate equivalence of such protocols in two ways: (a) Firstly by comparing the bodies of the tracts derived using protocols defined across the different templates considered, (b) Secondly by comparing the projection patterns of the reconstructed tracts across the different templates in two cross-species (human–macaque) comparison tasks. The results confirm similarity of all predictions regardless of the macaque brain template used, providing direct evidence for the generalisability of these tractography protocols across the five considered templates.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Non-human primates (NHPs) are extensively used in neuroscience research to explore the effects of evolution of the brain and its organisation, and as models of the human brain, with the rhesus macaque being a prominent example. This is possible because several neuroanatomical features and organisational principles are evolutionarily conserved across higher primates, including the presence of major white matter (WM) tracts (Markov et al. 2014; Petrides and Pandya 1999; Thiebaut de Schotten et al. 2012). For instance, several association pathways have been identified in humans, chimpanzees, and macaques, connecting similar broad brain areas, whilst differing in their branching patterns, as a result of ecological adaptation (connectivity specialisation) (Bryant et al. 2020; Eichert et al. 2020; Hecht et al. 2013; Petrides and Pandya 2009; Rilling et al. 2008). WM organisation is of particular functional interest as each functional brain subunit can be uniquely identified by its pattern of its extrinsic WM connections (Passingham et al. 2002).Therefore, mapping connectivity patterns in the NHP brain is of great interest, as it can provide insight into the human brain (Mars et al. 2018b).
Tracing (chemical, viral etc.) provides a gold standard (Jbabdi et al. 2015; Lehman et al. 2011) towards mapping brain connectivity, however, it is limited due to its invasiveness, and in its scalability, whole-brain applicability and translatability in humans. Magnetic resonance imaging (MRI) provides an alternative to invasive brain mapping, in both the human and NHP brain. Diffusion MRI (dMRI) specifically is a unique tool for identifying structural connections in the brain (Jbabdi et al. 2015). Various dMRI-based tractography methods have been developed to reliably extract defined sets of WM tracts. Such approaches typically use tractography protocols which consist of constraints/rules to guide the curve propagation process, reflecting underlying anatomy (Catani et al. 2002; Wakana et al. 2004). Protocols must be robust, consistent and reproducible whilst still reflecting the natural anatomical variation amongst individuals. This can be achieved in two ways, either manually prescribing subject-specific protocols (labour intensive especially for large cohorts; reduced consistency in protocol definition) or defining them in a standardised template space and mapping to the individual subjects in a consistent and automated fashion. The latter approach has proven to be rather powerful in the extraction of a considerable range of tracts, with a particular advantage when applied to large cohorts as it has allowed for the development of tract-specific atlases and the extraction of tract-specific features (Catani et al. 2012; de Schotten et al. 2011; Eichert et al. 2019; Makris et al. 2009; Takemura et al. 2017; Zhao et al. 2016).
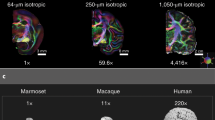
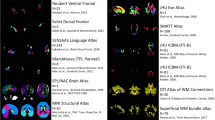
Building upon this approach, we previously devised tractography protocols for 42 WM tracts (FSL-XTRACT) that are standardised not only within species, but across very diverse brains, including humans and macaques (Mars et al. 2018b; Warrington et al. 2020), as well as adult and neonatal brains (Warrington et al. 2022) (Fig. 1). These protocols are defined in an analogous manner across brains and allow extraction of homologous WM bundles that exist across these different species, but are different in terms of their geometry, shape, exact location and projection patterns to grey matter. Subsequently, we have used these tracts as WM landmarks to anchor a common space and employed the uniqueness of cortical connection patterns to these tracts to probe areal specialisation. This has enabled us to quantitatively study divergences and similarities between brains with very different geometries (Mars et al. 2021), relying on connectivity patterns (instead of geometrically driven alignments), with higher relevance to brain function than allometric measurements (Eichert et al. 2020; Glasser et al. 2016; Mars et al. 2018a, b; Passingham et al. 2002; Van Essen et al. 2013; Xu et al. 2020).
Our original tractography protocols for the macaque brain were developed using the F99 macaque template, a choice motivated by the data availability and development around this template (Folloni et al. 2019; Mars et al. 2018b; Van Essen and Dierker 2007; Warrington et al. 2020). F99 is a hybrid surface-volume single-brain atlas containing a large set of associated reference data, including 15 areal partitioning schemes (with a probabilistic architectonic map), and connectivity data, as well as links to the CoCoMac and Markov connectivity databases (Markov et al. 2014; Paxinos et al. 2000; Stephan et al. 2001; Van Essen et al. 2012; Van Essen and Dierker 2007). However, a diverse set of alternative macaque brain templates have become available, either single-brain (such as D99; (Van Essen and Dierker 2007)), or population atlases (such as NMT (Seidlitz et al. 2018), Yerkes (Donahue et al. 2016), INIA19 (Rohlfing et al. 2012)), that are becoming more commonly used (e.g. for reporting chemical tracing experiments), providing their own sets of advantages and limitations. For instance, single-subject templates (F99, D99) tend to be of higher resolution but are more biased to the individual animal’s anatomy. On the contrary, multi-subject templates (INIA, Yerkes, NMT), are more representative of the group average, thus better for group comparisons, but can be limited in their relative neuroanatomical representation by the registration method (McLaren et al. 2009; Van Essen et al. 2001). Similarly, for in vivo templates (F99, NMT, INIA) the resolution and contrast are usually lower than for ex vivo templates (Yerkes, D99), with overall a single-subject ex vivo template having the highest contrast and resolution (D99) (Reveley et al. 2016; Seidlitz et al. 2018). Templates also differ in terms of the available associated data. For example, NMT has an extensive set of atlases both cortical and subcortical at various parcellation levels making it a useful template for atlas-based work (Seidlitz et al. 2018). Similarly, tracer studies often use the INIA (Howells et al. 2018) and NMT (Xu et al. 2022) templates (and less so the F99), thus making other templates more useful if data integration is in order. Therefore, depending on the data available and the questions at hand, a different template may be more or less appropriate, making the translation of our methods to multiple templates a valuable step.
In this paper, we present the generalisation of the original F99 macaque XTRACT tractography protocol across five macaque brain templates (F99, D99, NMT, Yerkes, and INIA) to allow for greater flexibility in the questions these protocols can be used for. We took the approach of first aligning the existing protocols to new templates to provide a starting point and subsequently adjusting them in each space to ensure maximum equivalence across templates. This allowed us to also revise imperfections in the original F99 tractography protocols. We demonstrate robustness and strong similarity of tract reconstruction within individual animals using protocols from the five different template spaces in a number of ways, including: (a) correlations of the reconstructed tracts using each alternate template against those using the standard F99 template and also (b) agreement of within-bundle dMRI microstructure metrics. We further demonstrate equivalence in grey-to-white matter whole-brain connectivity patterns when using tracts reconstructed from different templates in two different tasks: (a) identifying homologue cortical regions between the human and the rhesus macaque and (b) predicting scalar whole-brain maps from humans to macaques. Results confirm similarity of all predictions regardless of the macaque brain template used, providing direct evidence for the generalisability of tractography protocols across the five macaque templates considered.
Methods
Our XTRACT tractography protocols comprise of regions of interest (ROIs) defined in F99 space that describe starting, exclusion, stopping, and waypoint criteria for 42 WM tracts. We transformed those to four new template spaces (Table 1), using the process summarised in Fig. 2. We first performed a non-linear registration between templates to obtain a starting point for transforming these protocols from F99 to the other four considered templates. Subsequently, we refined a number of them to ensure maximum generalisability. For example, a common challenge was induced left–right protocol asymmetry after registration in the new template spaces, which we had to manually adjust. Notably, we identified and adjusted F99 protocols that did not generalise well to other templates (for instance, comprising of small ROIs close to interfaces that were very sensitive to mis-alignment errors). The full list of tracts is shown in Table 2.
Specifically, we used the RheMAP approach (Klink and Sirmpilatze 2020) to calculate non-linear warp fields between the F99 space and the other four template spaces. These templates were based on both single-subject and group-averaged, in-vivo and ex-vivo anatomical (T1w) MRI data and included: D99 (Reveley et al. 2016), INIA (Rohlfing et al. 2012), NMTv2.0 (which will be referred to as NMT) (Seidlitz et al. 2018), Yerkes19 (YRK) (Donahue et al. 2016). RheMAP uses ANTs (Avants et al. 2010, 2011) to generate non-linear warp fields, which we converted to FSL format. These non-linear warp fields were then used to transform the F99-space protocol ROIs (for the 42 tracts) to each of the new template spaces (Table 2). This provided a good starting point for the protocols in the new template spaces. Where registration caused imperfections (such as in the case of the perigenual section of the cingulum bundle (CBP), the corticospinal tract (CST), and the anterior commissure (AC)) manual revisions of the protocols were performed in all five template spaces. The revisions involved modification of exclusion, seed and target ROIs, to achieve bilateral symmetry, as well as to avoid ROI overlap and eliminate mapping outside the brain. Thus, we achieved equivalent protocols in all template spaces. Twelve of the original 42 protocols required substantial adjustment.
Data
Macaque: we used ex vivo dMRI brain data of 6 rhesus macaques (4–16 years of age), publicly available from PRIME-DE (http://fcon_1000.projects.nitrc.org/indi/PRIME/oxford2.html) (Milham et al. 2018). Data were acquired using a 7T Agilent DirectDrive console, with a 2D diffusion-weighted spin-echo multi-shell (DW-SEMS) protocol (16 volumes acquired at b = 0 s/mm2, 128 volumes acquired at b = 4000 s/mm2), at 0.6 mm isotropic resolution.
Human: Minimally preprocessed dMRI data (Glasser et al. 2013; Sotiropoulos et al. 2013) from the young adult Human Connectome Project (HCP) (Van Essen et al. 2013) were used for the assessment of cross-species comparisons across the different macaque template spaces. Data from 50 unrelated subjects (age range, 22–35 years), (3T 1.25 mm isotropic resolution data, with 270 directions evenly distributed across b = 1000, 2000 and 3000 s/mm2 shells) were used.
Preprocessing, tractography and population white matter (WM) tract atlases
Animal-wise non-linear transformations to the F99 template space were obtained using FSL’s FNIRT (Andersson et al. 2007), using each animal’s FA maps. Animal-wise non-linear transformation fields to each of the other four template spaces were obtained by concatenating the animal-wise F99 warp fields and the warp fields between the F99 and every other template space (FSL format of RheMAP non-linear transforms described above) using the FSL package tools (Jenkinson et al. 2012).
Prior to tractography, for each animal, fibre orientations were estimated (up to three per voxel) using FSL’s BEDPOSTX (Hernández et al. 2013; Jbabdi et al. 2012). Using the template-specific protocols, probabilistic tractography was performed using FSL’s XTRACT (Warrington et al 2020). A curvature threshold of 80° was chosen, and subsidiary fibres were considered with a volume fraction greater than 1%. A maximum of 2000 streamlines steps were used, and a step size of 0.2 mm was chosen. The resultant distributions of streamline counts were normalised by the total number of valid streamlines generated (those not rejected by inclusion/exclusion mask criteria) and stored in each subject’s respective native space, representing the corresponding path distributions.
Tractography results of all animals per template were used to obtain a set of tract atlases per template, in the form of population percentage overlap. For each template space, the respective native-space normalised path distributions were transformed to the template space, binarised and averaged across animals. The resultant spatial maps describe the percentage of animals for which a given tract is present at a given voxel. These maps were used for qualitative comparison of the tractography output between F99 and each new template (Fig. 3).
KL divergence in our WM-anchored common space gives the (dis-)similarity between the connectivity patterns of cortical parcels across brains and allows for identification of corresponding cortical regions between the human and the macaque. A Minimisation of KL divergence maps to find the corresponding region in the macaque brain to human cortical ROIs (sensorimotor area shown here). B Use KL divergence maps to project the human myelin map to the macaque brain
Protocol assessment
The robustness of the protocols and the generalisability of our framework to the other template spaces were assessed.
Comparing tract similarity across templates
For each animal, we calculated, per WM tract, the correlation between the native-space path distribution obtained using the F99 protocol and the native-space path distribution obtained using the protocol for every other template (INIA, NMT, Yerkes, D99). To do so, each 3D path distribution was turned into a 1D vector and pairwise Pearson’s correlation between the corresponding vectors were computed. Then, we summarised the correlation between each template and F99 per WM tract as the mean ± std (standard deviation) across animals. Within-tract measures of microstructure — tract-wise fractional anisotropy (FA), tract-wise mean diffusivity (MD) — were also compared across templates, exploring whether the new protocols localise equivalent WM areas. To obtain such measures, path distributions for each tract were normalised and thresholded at 0.1% giving rise to a binary mask per tract and template. The median value of FA and MD within each of these masks was obtained.
Comparing similarity of connectivity patterns across templates
Having extracted WM bundles across template spaces, we explored their equivalence in performing whole-brain comparisons between the human and macaque brains. Following the common space approach (Mars et al. 2018b), we extracted five versions of whole-brain grey-to-white matter connectivity patterns, one version corresponding to each of the considered macaque template spaces. For each of them, human and macaque comparisons were performed for two applications (scalar map prediction and homologue area identification, Fig. 3) and similarity of results across templates was evaluated. These comparisons allowed us to explore any differences between the proposed protocols in a complementary way to the previous pairwise similarities, which can capture the main body of a bundle, but less so any subtle deviations of the projections of these bundles.
To perform the above comparisons, we first built connectivity blueprints of the human and macaque brain (Mars et al. 2018b). These are whole brain (grey matter × WM tracts) matrices and we used the aforementioned 42 XTRACT tracts (Warrington et al. 2020). Specifically, tractography results for all 42 tracts were unwrapped in 1D, yielding a (whole brain × tracts) matrix. Whole-brain probabilistic tractography was performed to build a (cortex × whole brain) connectivity matrix, seeding 1000 streamlines from the white matter–grey matter boundary (WGB) and counting visitations in a whole-brain mask with the ventricles removed, down-sampled to 2 mm3. Connectivity blueprints (cortex × tracts) were then obtained by taking the product of this whole-brain connectivity matrix and the vectorised tract matrix. The columns of the resulting matrix represent the projection patterns of each tract on the WGB surface, whereas the rows represent the connectivity pattern of each of the WGB locations.
Surface data in F99 space were obtained following the approach described in Mars et al. (2018b). Briefly, a single set of macaque surfaces were derived using a set of high-quality structural data from one of the macaque subjects. The remaining macaque data were then non-linearly transformed to this space, and the surfaces were non-linearly transformed to the F99 standard space (Van Essen 2002). Prior to tractography, the surfaces were downsampled to approximately 10,000 vertices per hemisphere. Surfaces were only obtained in the F99 template space. For the other four template spaces, the template-specific tracts were non-linearly transformed in the F99 space and the above approach was followed to obtain connectivity blueprints using the F99-space surfaces. Following subject-wise construction of connectivity blueprints in each template space, we derived group-averaged blueprints for each template space, as well as one group-average blueprint for the human brain.
Once connectivity blueprints for the human and macaques are obtained, normalised rows of these matrices can be compared across species, as they correspond to patterns of connections of different grey matter locations to analogously defined WM tracts. We used this approach to identify the macaque homologues of two exemplar human brain areas (Fig. 3A) and explored the similarity when using tracts and blueprints obtained across the different macaque templates. We used the Glasser cortical atlas (Glasser et al. 2016) to define the human primary sensorimotor area (M1 + S1) and the medial temporal area (MT). For each of these areas, the average connectivity profile was extracted, by averaging the rows of the blueprint corresponding to locations within each of the areas, respectively. Subsequently, we looked for the vertices on the macaque cortical surface with the greatest similarity to the human connectivity profile for the same tracts. We used the symmetric Kullback–Leibler (KL) divergence (Kullback and Leibler 1951; Mars et al. 2018b) to assess similarities. KL divergence is zero for equivalent profiles and increases as they diverge. Hence, looking for the set of locations in the macaque with the minimum KL-divergence against the selected human brain areas allowed us to identify the macaque homologues of M1 + S1 and the MT. We compared the identified locations with reference regions obtained from the Paxinos macaque atlas (Paxinos et al. 2000) and we repeated this process independently for the five macaque template spaces. In all cases, we excluded the insula (Glasser atlas for the human and a combination of the M132 and MMF11 Markov atlases for the macaque brain (Glasser et al. 2016; Markov et al. 2014, 2011)), as it was poorly represented by the set of tracts reconstructed.
As a second step, we used the human-macaque similarities and divergences in the patterns of connectivity to our predefined set of tracts to project a scalar map (myelin maps (Glasser and Van Essen 2011)) from one species to the other (Fig. 3B). Effectively, we used connectivity patterns as a warp field and explored how tracts from the different macaque templates affect this transformation. To do so, for each of the macaque template spaces, we obtained a dis-similarity matrix to the human using the corresponding macaque and human blueprints. For each WGB location on the macaque brain we found the KL divergence across all WGB locations in the human. This KL divergence matrix was used to project the human myelin map (group average obtained from the HCP) to the macaque brain for each of the macaque templates (Fig. 3B), as described in (Mars et al. 2018b). Consistency of the myelin predictions across templates and with respect to the original F99 template, reinforces the generalisability of our framework to the other template spaces.
Results
We first assessed the robustness of tractography protocols that were directly registered from F99 to other template spaces. By assessing correlations of the resultant tracts between template spaces, we found that for 12 tracts the original F99 protocols did not generalise well when simply registered through non-linear warps (Fig. 4). We used large differences in correlation and qualitative inspection of the tracts as evidence that protocols needed further revision. We performed such revisions in two ways, both for the protocols in the new templates, but also for the original F99 protocols, when needed. Specifically, the original F99 protocols were modified for 6 tracts: CST (left), AR (right), AC, MDLF (right and left), CBT (right). In all tracts the seed, target and exclusion ROIs were modified, as detailed in Table 3.
Subject-wise correlation values between path distributions of tracts reconstructed using the F99-based protocols and each of the other template-based protocols. A subset of 12 (of the 42) tracts is shown comprising of the ones that required manual refinement following registration from the F99 template. For each tract and template, six samples are shown for “Original” (red asterisks, protocols are simply the F99 ones registered to new template space) and six for “Manually Adjusted” (blue circles, protocols have been refined in each template space), corresponding to the six macaque datasets we used. Adjustment was performed to improve tractography results and increase consistency across templates
This iterative process resulted in high agreement between within-animal tractography results when using the five different templates. Figure 5 demonstrates representative examples of tracts obtained from the same animal, when using the original F99 protocol registered to a new template (“Original”) vs the refined protocols (“Manually Adjusted”). Figures 4 and 7 highlight the improvement such revisions made across the six different animals and across the considered templates.
Examples of tracts obtained using protocols that were manually adjusted following the nonlinear registration from F99 to each other template space. A CBP and CST in NMT space, before (denoted as “Original”) and after adjustment of the registered protocol ROIs (from F99 space) (denoted as “Manually Adjusted”). B AR and CST in F99 space, before and following adjustment. Coronal sections are shown using radiological convention: R right, L left, A anterior, P posterior
Tract similarity across template spaces
Qualitatively, we observed high agreement between the path distributions generated using the standard F99 and the corresponding protocols in other template spaces across animals (Fig. 6). To quantify the agreement, we calculated the mean correlation ± std per tract across animals, between F99 and every other template. For all 42 WM tracts, median correlation was greater than 0.9 between F99 and every other template across animals. For each tract category (association, commissural, limbic, projection), the median agreement between the F99 and the in-vivo templates (INIA19, NMT) was ~ 0.99, as well as for the ex-vivo templates (D99, YRK) (Fig. 7). Complementary to the tract correlations, we also assessed within-tract DTI values (FA, MD). Both were consistent across all five templates considered, with the differences being mostly within 2% of the values obtained using the F99-based tractography protocols (Fig. 8).
Group-average WM tracts from the final protocols using the five templates (F99, D99 INIA, YRK, NMT). Qualitative comparison between templates for all major tracts considered. For each tract, normalised path distributions were thresholded at 0.1% and binarised, and then averaged across the six animals. The above maps show group percentage averages of these distributions (colour codes depict from 30% to 100% of the group)
Correlation (mean ± std) across six subjects for each tract between F99-derived path distributions and those derived using each other template considered. The correlation values were used for template comparison and assessment of protocol robustness and generalisability. For each tract and template space, normalised path distributions were thresholded at 0.1% and the Pearson correlation against the F99 respective path distribution was obtained within a binary mask covering the F99 thresholded path distribution
Within-tract diffusion measures of microstructure — tract-wise fractional anisotropy (FA), tract-wise mean diffusivity (MD) — were used to assess protocol robustness and generalisability across templates, as more biologically relevant measures compared to correlation. A The median FA and MD values per WM tract across subjects for each template space. Values are comparable across templates. B Percent difference of FA and MD of each new template considered compared to F99. The majority of absolute differences are less than 2%
Similarity of connectivity patterns across templates
In addition to ensuring similarity in terms of the bodies of the tracts across templates, we also explored similarity in the projection patterns of these tracts. KL divergence was calculated between average human and macaque connectivity blueprints (for every template space) to obtain (dis-similarity matrices between the macaque and human connectivity patterns. We also calculated the KL divergence between macaque connectivity blueprints obtained using the F99 protocols and the protocols from the other template spaces. Figure 9 demonstrates that these divergences are very small (almost zero, median around 0.008 in all cases) compared, for instance, to macaque-human divergences which are on average two orders of magnitude higher (median 0.8). This provides evidence that projections patterns of individual tracts are very similar regardless of the protocols used.
Distributions of minimum KL divergence quantifying the connectivity-based (dis-)similarity between F99 and each other macaque template blueprint, as well as with the human blueprint. For each WGB location in F99 space, the divergence with the best matching vertex in other template spaces (i.e. minimum KL divergence) has been calculated. The boxplots represent the distribution of these minimum KL divergence values across the whole brain. The inset is the log10 of the same KLD values. The cross-species dis-similarity (i.e. between the human and the macaque) strongly surpasses the intra-species dis-similarity (i.e. amongst macaque templates)
We further explored whether any small differences in the projection patterns have an effect on downstream analyses. We performed two human-macaque comparisons on the basis of their connectivity patterns to the correspondingly defined WM tracts. Firstly, we identified the most similar cortical vertices from the macaque brain to the average connectivity pattern of two exemplar regions of the human brain, the primary sensorimotor area (M1 + S1) and the medial temporal area (MT), recapitulating previous work (Mars et al. 2018a, b; Warrington et al. 2022). Figure 10 demonstrates the corresponding KL-divergence values for both regions when using tracts reconstructed from protocols of the different macaque templates. We found high agreement in the cross-species cortical area mapping for the F99, and the other macaque template spaces considered, supporting the generalisability of the protocols to the new template spaces.
KL divergence maps quantifying the connectivity-based (dis-)similarity between two human regions (primary sensorimotor-left and MT-right) and the macaque brain, when using tracts defined by protocols across the different macaque templates. Macaque regions with the most similar connectivity patterns to the human regions (i.e. lowest KL divergence) are shown in purple. The macaque primary sensorimotor and MT areas from the Paxinos atlas are shown for reference (bottom row). In all template spaces, the resulting patterns closely resemble the expected location of the corresponding regions in the macaque, with regions of lowest divergence overlapping with the atlas-based ROIs for M1 + S1 and MT respectively
Secondly, whole-brain KL divergence matrices (i.e. matrices that depict for every vertex in the macaque brain the best agreement with the human ones) were used to project the human myelin map to the macaque brain for each of the macaque templates. Figure 11 demonstrates that the myelin prediction maps for each macaque brain were equivalent, each having a very similar correlation of ~ 0.84 to the measured (original) macaque myelin map. The pattern of the difference maps between predicted and measured is also equivalent against template spaces. With this step we do not only ensure generalisability of the tractography protocols across templates (agreement in the results across templates) but also the reliability of these results due to their high correlation to the measured myelin map.
Prediction of macaque myelin map from a measured (T1w/T2w ratio) human myelin map for each template space considered (left). The KL-divergence of connectivity patterns to the reconstructed WM tracts between human and macaque is used as a transformation field. The right column shows absolute difference between each myelin map prediction and a measured (T1w/T2w ratio) macaque myelin map
Taken together, these results support the robustness of the protocols to macaque template. As these new macaque tractography protocols are also correspondingly defined for the human brain, this expansion allows for greater flexibility and integration of the WM-based common space framework for probing brain differences and similarities across evolution and development (Mars et al. 2018b; Warrington et al. 2022).
Discussion
In this work, we presented generalised XTRACT tractography protocols across five different commonly used macaque brain template spaces, specifically F99, D99, INIA19, Yerkes, NMT. We aimed to define protocols for 42 WM tracts that provided equivalent tract reconstructions within the same animal, regardless of the template space used. We demonstrated such equivalence through a number of tests, ranging from similarity metrics that focussed on the body of the tracts to similarity tests of the projection patterns of each tract. As these tractography protocols are also correspondingly defined for the human brain (Warrington et al. 2020), this generalisability allows for greater flexibility and integration of our previously presented WM-based common space framework for probing brain differences and similarities across phylogeny (Mars et al. 2018b; Warrington et al. 2022). This also allows for easier integration of our WM connectivity data with other data types, thus harvesting the power of multimodal data in addressing more complex neuroanatomical questions. For example, in combination with histology or tracer data, which can be available in any of the other common template space, we can better target the biological underpinnings of the structural connectivity features we observe. Similarly, there can be great benefit from integrating structural with functional connectivity information to better elucidate the crosstalk between different brain regions, both within and across species (Garin et al. 2022; Zimmermann et al. 2018).
We used an iterative process involving the manual refinement of the original F99 protocol ROIs, the non-linear registration to every other template space and the subsequent manual refinement in each template space of protocols where the registration caused issues (such as ROI overlap, ROI expansion beyond the brain mask, ROI left–right asymmetry). This process resulted in reproducible tracking for all 42 tracts considered in all template spaces, with no further manual adjustment required by the user. Specifically, within-animal path distributions, produced using protocols based on macaque templates other than F99, were highly correlated (mean correlation ≥ 0.9) to those obtained using the standard F99 template. Furthermore, we looked at within-tract mean DTI measures (FA and MD) and found very high consistency across all five templates, indicating that all protocols resulted to almost identical tract localisation.
Connectivity blueprints were subsequently used to explore whether using protocols from other than F99 templates causes changes in the connectivity patterns. We used connectivity patterns to reconstruct WM tracts to perform two human-macaque comparative experiments: one identifying homologue areas between human and macaque brains using similarity in connectivity patterns for localising functionally similar regions across species; and one using connectivity patterns to obtain a transformation for mapping cortical maps across two species. Equivalence in reconstructed tracts across macaque template spaces would result into equivalence of results in these two experiments. Indeed, results were consistent across all five templates and in agreement with previously published results (Mars et al. 2018b; Warrington et al. 2022). These complementary analyses indicated that the more subtle projection patterns of the reconstructed tracts are also equivalent across templates.
Although we have extended XTRACT protocols to five commonly used macaque template spaces, we recognise that there are still more templates available, so it would be of great interest and utility to further generalise our framework to even more template spaces (Van Essen 2002). It would also be beneficial to similarly extend to other NHP species and templates, with a natural extension being on the protocols for which we already have a first version, including the chimpanzee (Bryant et al. 2020), the gibbon (Bryant et al. 2023), the gorilla (Roumazeilles et al. 2020) and the marmoset (Schaeffer et al. 2017). This will allow for even greater flexibility in the questions our framework can be used to address. This work would not have been possible without the dedication of many in the NHP community to data collection, standardisation, documentation, and sharing (Milham et al. 2020, 2022). It is only through open science and sharing of our data and expertise that we can overcome shared problems and push the boundaries of the field. We aim with our work, including the one presented here, to contribute to this goal moving forward.
Data availability
Macaque data used in this study are publicly available from PRIME-DE (http://fcon_1000.projects.nitrc.org/indi/PRIME/oxford2.html) (Milham et al. 2018). Human data are publicly available from the young adult Human Connectome Project (HCP) (https://www.humanconnectome.org/study/hcp-young-adult). Tractography protocols and template-specific tract averages for all five template spaces are available on (https://github.com/SPMIC-UoN/XTRACT_Macaque_MultiTemplate) and will be incorporated into FSL-XTRACT (https://fsl.fmrib.ox.ac.uk/fsl/fslwiki/XTRACT).
References
Andersson JLR, Jenkinson M, Smith SM (2007) Non-linear optimisation (FMRIB Technical Report TR07JA1)
Avants BB, Yushkevich P, Pluta J, Minkoff D, Korczykowski M, Detre J, Gee JC (2010) The optimal template effect in hippocampus studies of diseased populations. Neuroimage 49:2457–2466. https://doi.org/10.1016/j.neuroimage.2009.09.062
Avants BB, Tustison NJ, Song G, Cook PA, Klein A, Gee JC (2011) A reproducible evaluation of ANTs similarity metric performance in brain image registration. Neuroimage 54:2033–2044. https://doi.org/10.1016/j.neuroimage.2010.09.025
Bryant KL, Li L, Eichert N, Mars RB (2020) A comprehensive atlas of white matter tracts in the chimpanzee. PLoS Biol 18:e3000971. https://doi.org/10.1371/journal.pbio.3000971
Bryant KL, Manger PR, Bertelsen MF, Khrapitchev AA, Sallet J, Benn RA, Mars RB (2023) A map of white matter tracts in a lesser ape, the lar gibbon. Brain Struct Funct. https://doi.org/10.1007/s00429-023-02709-9
Catani M, Howard RJ, Pajevic S, Jones DK (2002) Virtual in vivo interactive dissection of white matter fasciculi in the human brain. Neuroimage 17:77–94. https://doi.org/10.1006/nimg.2002.1136
Catani M, Dell’Acqua F, Vergani F, Malik F, Hodge H, Roy P, Valabregue R, Thiebaut de Schotten M (2012) Short frontal lobe connections of the human brain. Cortex 48:273–291. https://doi.org/10.1016/j.cortex.2011.12.001
de Schotten MT, Dell’Acqua F, Forkel SJ, Simmons A, Vergani F, Murphy DGM, Catani M (2011) A lateralized brain network for visuospatial attention. Nat Neurosci 14:1245–1246. https://doi.org/10.1038/nn.2905
Donahue CJ, Sotiropoulos SN, Jbabdi S, Hernandez-Fernandez M, Behrens TE, Dyrby TB, Coalson T, Kennedy H, Knoblauch K, Van Essen DC, Glasser MF (2016) Using diffusion tractography to predict cortical connection strength and distance: a quantitative comparison with tracers in the monkey. J Neurosci 36:6758–6770. https://doi.org/10.1523/JNEUROSCI.0493-16.2016
Eichert N, Verhagen L, Folloni D, Jbabdi S, Khrapitchev AA, Sibson NR, Mantini D, Sallet J, Mars RB (2019) What is special about the human arcuate fasciculus? Lateralization, projections, and expansion. Cortex 118:107–115. https://doi.org/10.1016/j.cortex.2018.05.005
Eichert N, Robinson EC, Bryant KL, Jbabdi S, Jenkinson M, Li L, Krug K, Watkins KE, Mars RB (2020) Cross-species cortical alignment identifies different types of anatomical reorganization in the primate temporal lobe. Elife 9:e53232. https://doi.org/10.7554/eLife.53232
Folloni D, Sallet J, Khrapitchev AA, Sibson N, Verhagen L, Mars RB (2019) Dichotomous organization of amygdala/temporal-prefrontal bundles in both humans and monkeys. Elife 8:e47175. https://doi.org/10.7554/eLife.47175
Garin CM, Hori Y, Everling S, Whitlow CT, Calabro FJ, Luna B, Froesel M, Gacoin M, Ben Hamed S, Dhenain M, Constantinidis C (2022) An evolutionary gap in primate default mode network organization. Cell Rep 39:110669. https://doi.org/10.1016/j.celrep.2022.110669
Glasser MF, Van Essen DC (2011) Mapping human cortical areas in vivo based on myelin content as revealed by T1- and T2-weighted MRI. J Neurosci 31:11597–11616. https://doi.org/10.1523/JNEUROSCI.2180-11.2011
Glasser MF, Sotiropoulos SN, Wilson JA, Coalson TS, Fischl B, Andersson JL, Xu J, Jbabdi S, Webster M, Polimeni JR, Van Essen DC, Jenkinson M (2013) The minimal preprocessing pipelines for the Human Connectome Project. Neuroimage 80:105–124. https://doi.org/10.1016/j.neuroimage.2013.04.127
Glasser MF, Coalson TS, Robinson EC, Hacker CD, Harwell J, Yacoub E, Ugurbil K, Andersson J, Beckmann CF, Jenkinson M, Smith SM, Van Essen DC (2016) A multi-modal parcellation of human cerebral cortex. Nature 536:171–178. https://doi.org/10.1038/nature18933
Hecht EE, Gutman DA, Preuss TM, Sanchez MM, Parr LA, Rilling JK (2013) Process versus product in social learning: comparative diffusion tensor imaging of neural systems for action execution-observation matching in macaques, chimpanzees, and humans. Cereb Cortex 23:1014–1024. https://doi.org/10.1093/cercor/bhs097
Hernández M, Guerrero GD, Cecilia JM, García JM, Inuggi A, Jbabdi S, Behrens TEJ, Sotiropoulos SN (2013) Accelerating fibre orientation estimation from diffusion weighted magnetic resonance imaging using GPUs. PLoS ONE 8:e61892. https://doi.org/10.1371/journal.pone.0061892
Howells H, Thiebaut de Schotten M, Dell’Acqua F, Beyh A, Zappalà G, Leslie A, Simmons A, Murphy DG, Catani M (2018) Frontoparietal tracts linked to lateralized hand preference and manual specialization. Cereb Cortex 28:2482–2494. https://doi.org/10.1093/cercor/bhy040
Jbabdi S, Sotiropoulos SN, Savio AM, Graña M, Behrens TEJ (2012) Model-based analysis of multishell diffusion MR data for tractography: how to get over fitting problems. Magn Reson Med 68:1846–1855. https://doi.org/10.1002/mrm.24204
Jbabdi S, Sotiropoulos SN, Haber SN, Van Essen DC, Behrens TE (2015) Measuring macroscopic brain connections in vivo. Nat Neurosci 18:1546–1555. https://doi.org/10.1038/nn.4134
Jenkinson M, Beckmann CF, Behrens TEJ, Woolrich MW, Smith SM (2012) FSL. Neuroimage 62:782–790. https://doi.org/10.1016/j.neuroimage.2011.09.015
Klink C, Sirmpilatze N (2020) PRIME-RE/RheMAP: RheMAP v1.3 “Corona Edition”
Kullback S, Leibler RA (1951) On information and sufficiency. Ann Math Stat 22:79–86. https://doi.org/10.1214/aoms/1177729694
Lehman JF, Greenberg BD, McIntyre CC, Rasmussen SA, Haber SN (2011) Rules ventral prefrontal cortical axons use to reach their targets: implications for diffusion tensor imaging tractography and deep brain stimulation for psychiatric illness. J Neurosci 31:10392–10402. https://doi.org/10.1523/JNEUROSCI.0595-11.2011
Makris N, Papadimitriou GM, Kaiser JR, Sorg S, Kennedy DN, Pandya DN (2009) Delineation of the middle longitudinal fascicle in humans: a quantitative, in vivo, DT-MRI study. Cereb Cortex 19:777–785. https://doi.org/10.1093/cercor/bhn124
Markov NT, Misery P, Falchier A, Lamy C, Vezoli J, Quilodran R, Gariel MA, Giroud P, Ercsey-Ravasz M, Pilaz LJ, Huissoud C, Barone P, Dehay C, Toroczkai Z, Van Essen DC, Kennedy H, Knoblauch K (2011) Weight consistency specifies regularities of macaque cortical networks. Cereb Cortex 21:1254–1272. https://doi.org/10.1093/cercor/bhq201
Markov NT, Ercsey-Ravasz MM, Ribeiro Gomes AR, Lamy C, Magrou L, Vezoli J, Misery P, Falchier A, Quilodran R, Gariel MA, Sallet J, Gamanut R, Huissoud C, Clavagnier S, Giroud P, Sappey-Marinier D, Barone P, Dehay C, Toroczkai Z, Knoblauch K, Van Essen DC, Kennedy H (2014) A weighted and directed interareal connectivity matrix for macaque cerebral cortex. Cereb Cortex 24:17–36. https://doi.org/10.1093/cercor/bhs270
Mars RB, Passingham RE, Jbabdi S (2018a) Connectivity fingerprints: from areal descriptions to abstract spaces. Trends Cogn Sci 22:1026–1037. https://doi.org/10.1016/j.tics.2018.08.009
Mars RB, Sotiropoulos SN, Passingham RE, Sallet J, Verhagen L, Khrapitchev AA, Sibson N, Jbabdi S (2018b) Whole brain comparative anatomy using connectivity blueprints. Elife 7:1–15. https://doi.org/10.7554/eLife.35237
Mars RB, Jbabdi S, Rushworth MFS (2021) A common space approach to comparative neuroscience. Annu Rev Neurosci 44:69–86. https://doi.org/10.1146/annurev-neuro-100220-025942
McLaren DG, Kosmatka KJ, Oakes TR, Kroenke CD, Kohama SG, Matochik JA, Ingram DK, Johnson SC (2009) A population-average MRI-based atlas collection of the rhesus macaque. Neuroimage 45:52–59. https://doi.org/10.1016/j.neuroimage.2008.10.058
Milham MP, Ai L, Koo B, Xu T, Amiez C, Balezeau F, Baxter MG, Blezer ELA, Brochier T, Chen A, Croxson PL, Damatac CG, Dehaene S, Everling S, Fair DA, Fleysher L, Freiwald W, Froudist-Walsh S, Griffiths TD, Guedj C, Hadj-Bouziane F, Ben Hamed S, Harel N, Hiba B, Jarraya B, Jung B, Kastner S, Klink PC, Kwok SC, Laland KN, Leopold DA, Lindenfors P, Mars RB, Menon RS, Messinger A, Meunier M, Mok K, Morrison JH, Nacef J, Nagy J, Rios MO, Petkov CI, Pinsk M, Poirier C, Procyk E, Rajimehr R, Reader SM, Roelfsema PR, Rudko DA, Rushworth MFS, Russ BE, Sallet J, Schmid MC, Schwiedrzik CM, Seidlitz J, Sein J, Shmuel A, Sullivan EL, Ungerleider L, Thiele A, Todorov OS, Tsao D, Wang Z, Wilson CRE, Yacoub E, Ye FQ, Zarco W, Zhou Y, Margulies DS, Schroeder CE (2018) An open resource for non-human primate imaging. Neuron 100:61–74.e2. https://doi.org/10.1016/j.neuron.2018.08.039
Milham M, Petkov CI, Margulies DS, Schroeder CE, Basso MA, Belin P, Fair DA, Fox A, Kastner S, Mars RB, Messinger A, Poirier C, Vanduffel W, Van Essen DC, Alvand A, Becker Y, Ben Hamed S, Benn A, Bodin C, Boretius S, Cagna B, Coulon O, El-Gohary SH, Evrard H, Forkel SJ, Friedrich P, Froudist-Walsh S, Garza-Villarreal EA, Gao Y, Gozzi A, Grigis A, Hartig R, Hayashi T, Heuer K, Howells H, Ardesch DJ, Jarraya B, Jarrett W, Jedema HP, Kagan I, Kelly C, Kennedy H, Klink PC, Kwok SC, Leech R, Liu X, Madan C, Madushanka W, Majka P, Mallon A-M, Marche K, Meguerditchian A, Menon RS, Merchant H, Mitchell A, Nenning K-H, Nikolaidis A, Ortiz-Rios M, Pagani M, Pareek V, Prescott M, Procyk E, Rajimehr R, Rautu I-S, Raz A, Roe AW, Rossi-Pool R, Roumazeilles L, Sakai T, Sallet J, García-Saldivar P, Sato C, Sawiak S, Schiffer M, Schwiedrzik CM, Seidlitz J, Sein J, Shen Z, Shmuel A, Silva AC, Simone L, Sirmpilatze N, Sliwa J, Smallwood J, Tasserie J, Thiebaut De Schotten M, Toro R, Trapeau R, Uhrig L, Vezoli J, Wang Z, Wells S, Williams B, Xu T, Xu AG, Yacoub E, Zhan M, Ai L, Amiez C, Balezeau F, Baxter MG, Blezer ELA, Brochier T, Chen A, Croxson PL, Damatac CG, Dehaene S, Everling S, Fleysher L, Freiwald W, Griffiths TD, Guedj C, Hadj-Bouziane F, Harel N, Hiba B, Jung B, Koo B, Laland KN, Leopold DA, Lindenfors P, Meunier M, Mok K, Morrison JH, Nacef J, Nagy J, Pinsk M, Reader SM, Roelfsema PR, Rudko DA, Rushworth MFS, Russ BE, Schmid MC, Sullivan EL, Thiele A, Todorov OS, Tsao D, Ungerleider L, Wilson CRE, Ye FQ, Zarco W, Zhou Y (2020) Accelerating the evolution of nonhuman primate neuroimaging. Neuron 105:600–603. https://doi.org/10.1016/j.neuron.2019.12.023
Milham M, Petkov C, Belin P, Ben Hamed S, Evrard H, Fair D, Fox A, Froudist-Walsh S, Hayashi T, Kastner S, Klink C, Majka P, Mars R, Messinger A, Poirier C, Schroeder C, Shmuel A, Silva AC, Vanduffel W, Van Essen DC, Wang Z, Roe AW, Wilke M, Xu T, Aarabi MH, Adolphs R, Ahuja A, Alvand A, Amiez C, Autio J, Azadi R, Baeg E, Bai R, Bao P, Basso M, Behel AK, Bennett Y, Bernhardt B, Biswal B, Boopathy S, Boretius S, Borra E, Boshra R, Buffalo E, Cao L, Cavanaugh J, Celine A, Chavez G, Chen LM, Chen X, Cheng L, Chouinard-Decorte F, Clavagnier S, Cléry J, Colcombe SJ, Conway B, Cordeau M, Coulon O, Cui Y, Dadarwal R, Dahnke R, Desrochers T, Deying L, Dougherty K, Doyle H, Drzewiecki CM, Duyck M, Arachchi WE, Elorette C, Essamlali A, Evans A, Fajardo A, Figueroa H, Franco A, Freches G, Frey S, Friedrich P, Fujimoto A, Fukunaga M, Gacoin M, Gallardo G, Gao L, Gao Y, Garside D, Garza-Villarreal EA, Gaudet-Trafit M, Gerbella M, Giavasis S, Glen D, Ribeiro Gomes AR, Torrecilla SG, Gozzi A, Gulli R, Haber S, Hadj-Bouziane F, Fujimoto SH, Hawrylycz M, He Q, He Y, Heuer K, Hiba B, Hoffstaedter F, Hong S-J, Hori Y, Hou Y, Howard A, De La Iglesia-Vaya M, Ikeda T, Jankovic-Rapan L, Jaramillo J, Jedema HP, Jin H, Jiang M, Jung B, Kagan I, Kahn I, Kiar G, Kikuchi Y, Kilavik B, Kimura N, Klatzmann U, Kwok SC, Lai H-Y, Lamberton F, Lehman J, Li P, Li X, Li X, Liang Z, Liston C, Little R, Liu C, Liu N, Liu X, Liu X, Lu H, Loh KK, Madan C, Magrou L, Margulies D, Mathilda F, Mejia S, Meng Y, Menon R, Meunier D, Mitchell AJ, Mitchell A, Murphy A, Mvula T, Ortiz-Rios M, Ortuzar Martinez DE, Pagani M, Palomero-Gallagher N, Pareek V, Perkins P, Ponce F, Postans M, Pouget P, Qian M, Ramirez JB, Raven E, Restrepo I, Rima S, Rockland K, Rodriguez NY, Roger E, Hortelano ER, Rosa M, Rossi A, Rudebeck P, Russ B, Sakai T, Saleem KS, Sallet J, Sawiak S, Schaeffer D, Schwiedrzik CM, Seidlitz J, Sein J, Sharma J, Shen K, Sheng W, Shi NS, Shim WM, Simone L, Sirmpilatze N, Sivan V, Song X, Tanenbaum A, Tasserie J, Taylor P, Tian X, Toro R, Trambaiolli L, Upright N, Vezoli J, Vickery S, Villalon J, Wang X, Wang Y, Weiss AR, Wilson C, Wong T-Y, Woo C-W, Wu B, Xiao D, Xu AG, Xu D, Xufeng Z, Yacoub E, Ye N, Ying Z, Yokoyama C, Yu X, Yue S, Yuheng L, Yumeng X, Zaldivar D, Zhang S, Zhao Y, Zuo Z (2022) Toward next-generation primate neuroscience: a collaboration-based strategic plan for integrative neuroimaging. Neuron 110:16–20. https://doi.org/10.1016/j.neuron.2021.10.015
Passingham RE, Stephan KE, Kötter R (2002) The anatomical basis of functional localization in the cortex. Nat Rev Neurosci 3:606–616. https://doi.org/10.1038/nrn893
Paxinos G, Huang X, Toga A (2000) The Rhesus monkey brain in stereotaxic coordinates. Academic Press, San Diego
Petrides M, Pandya DN (1999) Dorsolateral prefrontal cortex: comparative cytoarchitectonic analysis in the human and the macaque brain and corticocortical connection patterns. Eur J Neurosci 11:1011–1036. https://doi.org/10.1046/j.1460-9568.1999.00518.x
Petrides M, Pandya DN (2009) Distinct parietal and temporal pathways to the homologues of Broca’s area in the monkey. PLoS Biol 7:e1000170. https://doi.org/10.1371/journal.pbio.1000170
Reveley C, Gruslys A, Ye FQ, Glen D, Samaha J, Russ BE, Saad Z, Seth AK, Leopold DA, Saleem KS (2016) Three-dimensional digital template atlas of the macaque brain. Cereb Cortex 27(9):4463–4477. https://doi.org/10.1093/cercor/bhw248
Rilling JK, Glasser MF, Preuss TM, Ma X, Zhao T, Hu X, Behrens TEJ (2008) The evolution of the arcuate fasciculus revealed with comparative DTI. Nat Neurosci 11:426–428. https://doi.org/10.1038/nn2072
Rohlfing T, Kroenke CD, Sullivan EV, Dubach MF, Bowden DM, Grant KA, Pfefferbaum A (2012) The INIA19 template and NeuroMaps atlas for primate brain image parcellation and spatial normalization. Front Neuroinform 6:27. https://doi.org/10.3389/fninf.2012.00027
Roumazeilles L, Eichert N, Bryant KL, Folloni D, Sallet J, Vijayakumar S, Foxley S, Tendler BC, Jbabdi S, Reveley C, Verhagen L, Dershowitz LB, Guthrie M, Flach E, Miller KL, Mars RB (2020) Longitudinal connections and the organization of the temporal cortex in macaques, great apes, and humans. PLoS Biol 18:e3000810. https://doi.org/10.1371/journal.pbio.3000810
Schaeffer DJ, Adam R, Gilbert KM, Gati JS, Li AX, Menon RS, Everling S (2017) Diffusion-weighted tractography in the common marmoset monkey at 9.4T. J Neurophysiol 118:1344–1354. https://doi.org/10.1152/jn.00259.2017
Seidlitz J, Sponheim C, Glen D, Ye FQ, Saleem KS, Leopold DA, Ungerleider L, Messinger A (2018) A population MRI brain template and analysis tools for the macaque. Neuroimage 170:121–131. https://doi.org/10.1016/j.neuroimage.2017.04.063
Sotiropoulos SN, Jbabdi S, Xu J, Andersson JL, Moeller S, Auerbach EJ, Glasser MF, Hernandez M, Sapiro G, Jenkinson M, Feinberg DA, Yacoub E, Lenglet C, Van Essen DC, Ugurbil K, Behrens TEJ (2013) Advances in diffusion MRI acquisition and processing in the Human Connectome Project. Neuroimage 80:125–143. https://doi.org/10.1016/j.neuroimage.2013.05.057
Stephan KE, Kamper L, Bozkurt A, Burns GAPC, Young MP, Kötter R (2001) Advanced database methodology for the collation of connectivity data on the macaque brain (CoCoMac). Philos Trans R Soc Lond B Biol Sci 356:1159–1186. https://doi.org/10.1098/rstb.2001.0908
Takemura H, Pestilli F, Weiner KS, Keliris GA, Landi SM, Sliwa J, Ye FQ, Barnett MA, Leopold DA, Freiwald WA, Logothetis NK, Wandell BA (2017) Occipital white matter tracts in human and macaque. Cereb Cortex 27:3346–3359. https://doi.org/10.1093/cercor/bhx070
Thiebaut de Schotten M, Dell’Acqua F, Valabregue R, Catani M (2012) Monkey to human comparative anatomy of the frontal lobe association tracts. Cortex 48:82–96. https://doi.org/10.1016/j.cortex.2011.10.001
Van Essen DC (2002) Surface-based atlases of cerebellar cortex in the human, macaque, and mouse. Ann N Y Acad Sci 978:468–479. https://doi.org/10.1111/j.1749-6632.2002.tb07588.x
Van Essen DC, Dierker DL (2007) Surface-based and probabilistic atlases of primate cerebral cortex. Neuron 56:209–225. https://doi.org/10.1016/j.neuron.2007.10.015
Van Essen DC, Lewis JW, Drury HA, Hadjikhani N, Tootell RBH, Bakircioglu M, Miller MI (2001) Mapping visual cortex in monkeys and humans using surface-based atlases. Vision Res 41:1359–1378. https://doi.org/10.1016/S0042-6989(01)00045-1
Van Essen DC, Glasser MF, Dierker DL, Harwell J (2012) Cortical parcellations of the macaque monkey analyzed on surface-based atlases. Cereb Cortex 22:2227–2240. https://doi.org/10.1093/cercor/bhr290
Van Essen DC, Smith SM, Barch DM, Behrens TEJ, Yacoub E, Ugurbil K (2013) The WU-Minn Human Connectome Project: an overview. Neuroimage 80:62–79. https://doi.org/10.1016/j.neuroimage.2013.05.041
Wakana S, Jiang H, Nagae-Poetscher LM, van Zijl PCM, Mori S (2004) Fiber tract–based atlas of human white matter anatomy. Radiology 230:77–87. https://doi.org/10.1148/radiol.2301021640
Warrington S, Bryant KL, Khrapitchev AA, Sallet J, Charquero-Ballester M, Douaud G, Jbabdi S, Mars RB, Sotiropoulos SN (2020) XTRACT—standardised protocols for automated tractography in the human and macaque brain. Neuroimage 217:116923. https://doi.org/10.1016/j.neuroimage.2020.116923
Warrington S, Thompson E, Bastiani M, Dubois J, Baxter L, Slater R, Jbabdi S, Mars RB, Sotiropoulos SN (2022) Concurrent mapping of brain ontogeny and phylogeny within a common space: standardized tractography and applications. Sci Adv 8:abq2022. https://doi.org/10.1126/sciadv.abq2022
Xu T, Nenning K-H, Schwartz E, Hong S-J, Vogelstein JT, Goulas A, Fair DA, Schroeder CE, Margulies DS, Smallwood J, Milham MP, Langs G (2020) Cross-species functional alignment reveals evolutionary hierarchy within the connectome. Neuroimage 223:117346. https://doi.org/10.1016/j.neuroimage.2020.117346
Xu R, Bichot NP, Takahashi A, Desimone R (2022) The cortical connectome of primate lateral prefrontal cortex. Neuron 110:312–327.e7. https://doi.org/10.1016/j.neuron.2021.10.018
Zhao J, Thiebaut de Schotten M, Altarelli I, Dubois J, Ramus F (2016) Altered hemispheric lateralization of white matter pathways in developmental dyslexia: evidence from spherical deconvolution tractography. Cortex 76:51–62. https://doi.org/10.1016/j.cortex.2015.12.004
Zimmermann J, Griffiths JD, McIntosh AR (2018) Unique mapping of structural and functional connectivity on cognition. J Neurosci 38:9658–9667. https://doi.org/10.1523/JNEUROSCI.0900-18.2018
Acknowledgements
SA, SW and SNS acknowledge funding from the European Research Council (ERC Consolidator—101000969 to SNS). The Wellcome Centre for Integrative Neuroimaging (SJ, KLB, and RM) is supported by the Wellcome Trust [203139/Z/16/Z to SJ, KLB, RM; [221933/Z/20/Z] and [215573/Z/19/Z] to SJ].
Author information
Authors and Affiliations
Contributions
All authors contributed significantly to this work from different and, in cases, overlapping roles. The specific contributions of each are as follows. SA: conceptualisation, methodology, software, data curation, formal analysis, investigation, visualisation, writing—original draft, writing—review and editing, SW: methodology, software, data curation, formal analysis, writing—original draft, writing—review and editing, KLB: resources, methodology, writing—review and editing, SP: resources, writing—review and editing, SJ: methodology, resources, writing—review and editing, RM: methodology, formal analysis, writing—review and editing, SNS: conceptualisation, methodology, formal analysis, resources, writing—original draft, writing—review and editing, supervision, funding acquisition.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Assimopoulos, S., Warrington, S., Bryant, K.L. et al. Generalising XTRACT tractography protocols across common macaque brain templates. Brain Struct Funct (2024). https://doi.org/10.1007/s00429-024-02760-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00429-024-02760-0