Abstract
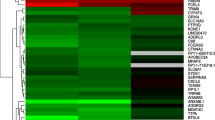
Lung adenocarcinoma (LUAD) shows heterogeneous morphological features and the stepwise progression from adenocarcinoma in situ to minimally invasive adenocarcinoma to invasive LUAD. Although multiple genetic alterations have been linked to the progression, the differences between the gene expression profiles of non-invasive lesions (non-ILs) and adjacent histologically normal lung (aNL) tissues within invasive LUAD have not been investigated. Herein, we analyzed differentially expressed genes (DEGs) specific to early-stage carcinogenesis in LUAD. Invasive LUAD tissue samples containing both non-ILs and aNL tissues were obtained from seven patients with pathological stage I LUAD, and each component was subjected to microdissection. Gene expression profiles of each component were determined using targeted RNA-sequencing. In total, 2536 DEGs, including 863 upregulated and 1673 downregulated genes, were identified in non-ILs. In non-ILs, the expression of SLC44A5, a choline transporter-like protein-coding gene, was significantly upregulated, and that of TMEM100, a gene encoding a transmembrane protein, was significantly downregulated. Reportedly, SLC44A5 plays an important role in the development and progression of hepatocellular carcinoma, whereas TMEM100 functions as a tumor suppressor in non-small cell lung cancer. Gene set enrichment analysis showed that DEGs in non-ILs were negatively enriched in cell death and immune response. Immunohistochemical analysis revealed that increased SLC44A5 expression and decreased TMEM100 expression were maintained in ILs. A protein–protein interaction (PPI) network analysis identified several upregulated and downregulated hub genes with high degrees in non-ILs. In conclusion, several new DEGs and key PPI network hub genes were identified in non-ILs, contributing to understanding of early-stage carcinogenesis in LUAD.





Similar content being viewed by others
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249. https://doi.org/10.3322/caac.21660
Travis WD, Brambilla E, Bruke AP, Marx A, Nicholson AG (2015) WHO classification of tumors of the lung, pleura, thymus, and heart, 4th edn. IARC Press France, Lyon, pp 26–30
Hu X, Fujimoto J, Ying L et al (2019) Multi-region exome sequencing reveals genomic evolution from preneoplasia to lung adenocarcinoma. Nat Commun 10:2978. https://doi.org/10.1038/s41467-019-10877-8
Pao W, Hutchinson KE (2012) Chipping away at the lung cancer genome. Nat Med 18:349–351. https://doi.org/10.1038/nm.2697
The Cancer Genome Atlas Research Network (2014) Comprehensive molecular profiling of lung adenocarcinoma. Nature 511:543–550. https://doi.org/10.1038/nature13385
Desai TJ, Brownfield DG, Krasnow MA (2014) Alveolar progenitor and stem cells in lung development, renewal and cancer. Nature 507:190–194. https://doi.org/10.1038/nature12930
Ferone G, Lee MC, Sage J, Berns A (2020) Cells of origin of lung cancers: lessons from mouse studies. Genes Dev 34:1017–1032. https://doi.org/10.1101/gad.338228.120
Murphy SJ, Wigle DA, Lima JF et al (2014) Genomic rearrangements define lineage relationships between adjacent lepidic and invasive components in lung adenocarcinoma. Cancer Res 74:3157–3167. https://doi.org/10.1158/0008-5472.can-13-1727
Vinayanuwattikun C, Le Calvez-Kelm F, Abedi-Ardekani B et al (2016) Elucidating genomic characteristics of lung cancer progression from in situ to invasive adenocarcinoma. Sci Rep 6:31628. https://doi.org/10.1038/srep31628
Nishimura T, Nakamura H, Tan KT et al (2020) A proteogenomic profile of early lung adenocarcinomas by protein co-expression network and genomic alteration analysis. Sci Rep 10:13604. https://doi.org/10.1038/s41598-020-70578-x
Qian J, Zhao S, Zou Y, Rahman SMJ, Senosain MF, Stricker T, Chen H, Powell CA, Borczuk AC, Massion PP (2020) Genomic understandings of tumor behavior in in situ and early lung adenocarcinoma. Am J Respir Crit Care Med 201:697–706. https://doi.org/10.1164/rccm.201902-0294oc
Chen H, Carrot-Zhang J, Zhao Y et al (2019) Genomic and immune profiling of pre-invasive lung adenocarcinoma. Nat Commun 10:5472. https://doi.org/10.1038/s41467-019-13460-3
Kato Y, Nakamura H, Tojo H et al (2015) A proteomic profiling of laser-microdissected lung adenocarcinoma cells of early lepidic-types. Clin Transl Med 4:64. https://doi.org/10.1186/s40169-015-0064-3
Kidokoro Y, Sakabe T, Haruki T, Kadonaga T, Nosaka K, Nakamura H, Umekita Y (2020) Gene expression profiling by targeted RNA sequencing in pathological stage I lung adenocarcinoma with a solid component. Lung Cancer 147:56–63. https://doi.org/10.1016/j.lungcan.2020.06.035
Goldstraw P, Chansky K, Crowley J, Rami-Porta R, Asamura H, Eberhardt WE, Nicholson AG, Groome P, Mitchell A, Bolejack V (2016) The IASLC lung cancer staging project: TNM classification for lung cancer. J Thorac Oncol 11:39–51. https://doi.org/10.1016/j.jtho.2015.09.009
Subramanian A, Tamayo P, Mootha VK et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A 102:15545–15550. https://doi.org/10.1073/pnas.0506580102
Mootha VK, Lindgren CM, Eriksson KF et al (2003) PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet 34:267–273. https://doi.org/10.1038/ng1180
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. https://doi.org/10.1101/gr.1239303
Assenov Y, Ramírez F, Schelhorn SE, Lengauer T, Albrecht M (2008) Albrecht, computing topological parameters of biological networks. Bioinformatics 24:282–284. https://doi.org/10.1093/bioinformatics/btm554
Basnet H, Tian L, Ganesh K, Huang YH, Macalinao DG, Brogi E, Finley LW, Massagué J (2019) Flura-seq identifies organ-specific metabolic adaptations during early metastatic colonization. Elife 8:e43627. https://doi.org/10.7554/elife.43627
Pyo KH, Kim JH, Lee JM, Kim SE, Cho JS, Lim SM, Cho BC (2019) Promising preclinical platform for evaluation of immuno-oncology drugs using Hu-PBL-NSG lung cancer models. Lung Cancer 127:112–121. https://doi.org/10.1016/j.lungcan.2018.11.035
Szklarczyk D, Morris JH, Cook H et al (2017) The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res 45:D362–D368. https://doi.org/10.1093/nar/gkw937
Chen YJ, Roumeliotis TI, Chang YH et al (2020) Proteogenomics of non-smoking lung cancer in East Asia delineates molecular signatures of pathogenesis and progression. Cell 182:226-244.e217. https://doi.org/10.1016/j.cell.2020.06.012
Gillette MA, Satpathy S, Cao S et al (2020) Proteogenomic characterization reveals therapeutic vulnerabilities in lung adenocarcinoma. Cell 182:200-225.e235. https://doi.org/10.1016/j.cell.2020.06.013
Inazu M (2014) Choline transporter-like proteins CTLs/SLC44 family as a novel molecular target for cancer therapy. Biopharm Drug Dispos 35:431–449. https://doi.org/10.1002/bdd.1892
Peng GZ, Ye QF, Wang R, Li MX, Yang ZX (2016) Knockdown by shRNA identifies SLC44A5 as a potential therapeutic target in hepatocellular carcinoma. Mol Med Rep 13:4845–4852. https://doi.org/10.3892/mmr.2016.5136
Moon EH, Kim YS, Seo J, Lee S, Lee YJ, Oh SP (2015) Essential role for TMEM100 in vascular integrity but limited contributions to the pathogenesis of hereditary haemorrhagic telangiectasis. Cardiovasc Res 105:353–360. https://doi.org/10.1093/cvr/cvu260
Frullanti E, Colombo F, Falvella FS et al (2012) Association of lung adenocarcinoma clinical stage with gene expression pattern in noninvolved lung tissue. Int J Cancer 131:E643–E648. https://doi.org/10.1002/ijc.27426
Han Z, Wang T, Han S et al (2017) Low-expression of TMEM100 is associated with poor prognosis in non-small cell lung cancer. Am. J. Trans. Res 9:2567–2578. http://www.ncbi.nlm.nih.gov/pmc/articles/pmc5446537/
He Q, Dong Y, Zhu Y, Ding Z, Zhang X, Wang Z, Ai R, He Y (2021) TMEM100 induces cell death in non-small cell lung cancer via the activation of autophagy and apoptosis. Oncol Rep 45:63. https://doi.org/10.3892/or.2021.8014
Zhuang J, Huang Y, Zheng W, Yang S, Zhu G, Wang J, Lin X, Ye J (2020) TMEM100 expression suppresses metastasis and enhances sensitivity to chemotherapy in gastric cancer. Biol Chem 401:285–296. https://doi.org/10.1515/hsz-2019-0161
Chen Y, Hu R, Li X, Shi Z, Tian H, Feng J, Yu S (2020) B7–H4 and HHLA2, members of B7 family, are aberrantly expressed in EGFR mutated lung adenocarcinoma. Pathol Res Pract 216:153134. https://doi.org/10.1016/j.prp.2020.153134
Cheng H, Borczuk A, Janakiram M, Ren X, Lin J, Assal A, Halmos B, Perez-Soler R, Zang X (2018) Wide expression and significance of alternative immune checkpoint molecules, B7x and HHLA2, in PD-L1-Negative Human Lung Cancers. Clin Cancer Res 24:1954–1964. https://doi.org/10.1158/1078-0432.Ccr-17-2924
Cheng H, Janakiram M, Borczuk A, Lin J, Qiu W, Liu H, Chinai JM, Halmos B, Perez-Soler R, Zang X (2017) HHLA2, a new immune checkpoint member of the B7 family, is widely expressed in human lung cancer and associated with EGFR mutational status. Clin Cancer Res 23:825–832. https://doi.org/10.1158/1078-0432.Ccr-15-3071
Zhao R, Chinai JM, Buhl S et al (2013) HHLA2 is a member of the B7 family and inhibits human CD4 and CD8 T-cell function. Proc Natl Acad Sci 110:9879–9884. https://doi.org/10.1073/pnas.1303524110
Kai Y, Amatya VJ, Kushitani K, Kambara T, Suzuki R, Tsutani Y, Miyata Y, Okada M, Takeshima Y (2019) Mucin 21 is a novel, negative immunohistochemical marker for epithelioid mesothelioma for its differentiation from lung adenocarcinoma. Histopathology 74:545–554. https://doi.org/10.1111/his.13775
Kataoka T, Okudela K, Nakashima Y et al (2019) Unique expression profiles of mucin proteins in interstitial pneumonia-associated lung adenocarcinomas. Histol Histopathol 34:1243–1254. https://doi.org/10.14670/hh-18-114
Yoshimoto T, Matsubara D, Soda M et al (2019) Mucin 21 is a key molecule involved in the incohesive growth pattern in lung adenocarcinoma. Cancer Sci 110:3006–3011. https://doi.org/10.1111/cas.14129
Sakamoto H, Shimizu J, Horio Y, Ueda R, Takahashi T, Mitsudomi T, Yatabe Y (2007) Disproportionate representation of KRAS gene mutation in atypical adenomatous hyperplasia, but even distribution of EGFR gene mutation from preinvasive to invasive adenocarcinomas. J Pathol 212:287–294. https://doi.org/10.1002/path.2165
Yoshida Y, Shibata T, Kokubu A, Tsuta K, Matsuno Y, Kanai Y, Asamura H, Tsuchiya R, Hirohashi S (2005) Mutations of the epidermal growth factor receptor gene in atypical adenomatous hyperplasia and bronchioloalveolar carcinoma of the lung. Lung Cancer 50:1–8. https://doi.org/10.1016/j.lungcan.2005.04.012
Soh J, Toyooka S, Ichihara S et al (2008) Sequential molecular changes during multistage pathogenesis of small peripheral adenocarcinomas of the lung. J Thorac Oncol 3:340–347. https://doi.org/10.1097/jto.0b013e318168d20a
Sivakumar S, Lucas FAS, McDowell TL et al (2017) Genomic landscape of atypical adenomatous hyperplasia reveals divergent modes to lung adenocarcinoma. Cancer Res 77:6119–6130. https://doi.org/10.1158/0008-5472.can-17-1605
Acknowledgements
The authors are grateful to Professor Eiji Nanba, Kaori Adachi, and Masachika Kai for technical assistance with the NGS experiment and Shoji Yashima and Yuko Urakami for pathological specimen preparation.
Funding
This work was supported by JSPS KAKENHI [Grant Number JP19K18214].
Author information
Authors and Affiliations
Contributions
T. Kadonaga: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Visualization, Writing—original draft. T. Sakabe: Conceptualization, Data curation, Investigation, Methodology, Project administration, Supervision, Visualization, Writing—original draft. Y. Kidokoro: Funding acquisition, Methodology, Visualization, Writing—review and editing. T. Haruki: Data curation, Resources, Writing—review and editing. K. Nosaka: Data curation, Resources, Writing—review and editing. H. Nakamura: Funding acquisition, Resources, Writing—review and editing. Y. Umekita: Conceptualization, Funding acquisition, Methodology, Project administration, Supervision, Writing—original draft.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Written informed consent for the use of clinical data in research was obtained from all patients, and the study was approved by the Ethical Review Board of Tottori University, Japan (approval number: 19A165, December 4, 2019).
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kadonaga, T., Sakabe, T., Kidokoro, Y. et al. Gene expression profiling using targeted RNA-sequencing to elucidate the progression from histologically normal lung tissues to non-invasive lesions in invasive lung adenocarcinoma. Virchows Arch 480, 831–841 (2022). https://doi.org/10.1007/s00428-021-03250-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-021-03250-y