Abstract
DNA copy-number gains of chromosomes 8q, 13q, and 20q are frequently observed in gastric cancers. Moreover gain of chromosome 20q has been associated with lymph node metastasis. The aim of this study was to correlate DNA copy-number changes of individual genes on chromosomes 8q, 13q, and 20q in gastric adenocarcinomas to clinicopathological data. DNA isolated from 63 formalin-fixed and paraffin-embedded gastric adenocarcinoma tissue samples was analyzed by whole-genome microarray comparative genomic hybridization and by multiplex ligation-dependent probe amplification (MLPA), targeting 58 individual genes on chromosomes 8, 13, and 20. Using array comparative genomic hybridization, gains on 8q, 13q, and 20q were observed in 49 (77.8%), 25 (39.7%), and 49 (77.8%) gastric adenocarcinomas, respectively. Gain of chromosome 20q was significantly correlated with lymph node metastases (p = 0.05) and histological type (p = 0.02). MLPA revealed several genes to be frequently gained in DNA copy number. The oncogene c-myc on 8q was gained in 73% of the cancers, while FOXO1A and ATP7B on 13q were both gained in 28.6% of the cases. Multiple genes on chromosome 20q showed gains in more than 60% of the cancers. DNA copy-number gains of TNFRSF6B (20q13.3) and ZNF217 (20q13.2) were significantly associated with lymph node metastasis (p = 0.02) and histological type (p = 0.02), respectively. In summary, gains of chromosomes 8q, 13q, and 20q in gastric adenocarcinomas harbor DNA copy-number gains of known and putative oncogenes. ZNF217 and TNFRSF6B are associated with important clinicopathological variables, including lymph node status.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gastric cancer is a major health problem and ranks second as a cause of cancer death worldwide [1]. The only possible curative treatment is complete surgical resection. One of the hallmarks of solid cancers, including gastric cancer, is chromosomal instability leading to gains and losses of parts of, or even whole chromosomes [2]. The mechanisms underlying this instability phenotype in gastric cancers are still poorly understood. Patterns of DNA copy-number aberrations can be analyzed at high resolution by array comparative genomic hybridization (CGH). Gains of chromosomes 3q, 7p, 7q, 8q, 13q, 17q, and 20q and losses of chromosomes 4q, 5q, 6q, 9p, 17p, and 18q have been consistently described in gastric cancer studies using CGH or array CGH analysis [3–11]. In addition, gains of chromosomes 7q, 8q, 9q, 11q, 13q, and 20q and losses of chromosomes 4p, 5q, 6, 9p, 17p, and 18q can already be detected in gastric cancer precursor lesions with direct malignant potential, indicating these as early events in the pathogenesis of gastric cancer [12–16]. Results from array CGH studies have shown that patterns of chromosomal aberrations can be correlated with clinicopathological variables. In previous studies, we have shown different patterns of chromosomal instability to correlate with lymph node status and with age of onset of the disease [17, 18].
DNA copy-number gains of chromosomes 8, 13, and 20 have been described to play an important role in colorectal adenoma to carcinoma progression [19, 20]. Also in gastric adenomas and adenocarcinomas, gains of these chromosomes have frequently been detected, indicating the importance of genetic events on these chromosomes for gastric cancer pathogenesis. Moreover, gain of chromosome 20q has been described to correlate with lymph node metastasis in gastric cancer and poor clinical outcome in colorectal cancer [3, 21, 22]. The most widely used array CGH platforms, so far, use spotted bacterial artificial chromosomes (BACs), which means that the probe sequences on the arrays cover up to 1 kb of DNA. This approach is very suitable for detecting patterns of DNA copy-number changes but has limitations in pinpointing individual genes whose normal function is disrupted due to these chromosomal aberrations. Multiplex ligation-dependent probe amplification (MLPA) [23] allows to determine in a single experiment DNA copy-number ratios of up to 40 individual genes and can be used to zoom in on chromosomal areas that show aberrations with array CGH [24, 25]. The aim of the present study is a detailed analysis of DNA copy-number changes of individual genes within gained areas on chromosomes 8, 13, and 20 in gastric adenocarcinomas and to correlate gene specific alterations to clinicopathological data.
Materials and methods
Materials and DNA isolation
DNA of 63 formalin-fixed and paraffin-embedded (FFPE) gastric adenocarcinomas of which 53 were obtained from Leeds (Leeds University Hospital, UK) and 10 were obtained from the Dutch D1/D2 trial [26], was isolated as previously described [17] using a commercial available DNA isolation kit (QIAmp DNA microkit, Qiagen, Hilden, Germany). Briefly, areas of at least 70% tumor cells were demarcated on a 4-µm haematoxylin- and eosin-stained tissue section. Adjacent serial sections of 10 µm were cut, and after deparaffination, the tumor tissue was macrodissected using a needle. After an overnight incubation with sodium thiocyanate (1 M) at 37°C, followed by proteinase K treatment, DNA was extracted. DNA concentrations were measured on a Nanodrop ND-1000 spectrophotometer (Isogen, IJsselstein, The Netherlands), and DNA quality was assessed by isothermal amplification as previously described [27]. Only DNA of excellent, good, and intermediate quality was used for further analysis.
DNA labeling and array CGH)
DNA labeling and array CGH were essentially performed as previously described [17, 28]. In short, tumor and normal DNA were differentially labeled using random priming (Bioprime DNA Labeling System; Invitrogen, Breda, The Netherlands) and hybridized on a BAC array containing approximately 5,000 clones printed in triplicate, consisting of the Sanger BAC clone set with an average resolution along the whole genome of 1.0 Mb (http://www.ensembl.org/Homo_sapiens/cytoview), the OncoBac set (http://informa.bio.caltech.edu/Bac_onc.html), containing approximately 600 clones corresponding to 200 cancer-related genes, and selected clones of interest obtained from the Children’s Hospital Oakland Research Institute (CHORI) to fill any gaps larger than 1 Mb on chromosome 6 and to have full-coverage contigs of regions on chromosomes 8, 11, 13, and 20. All clones were printed on Codelink™ slides (Amersham BioSciences, Roosendaal, The Netherlands) at a concentration of 1 μg/μl in 150-mM sodium phosphate, pH 8.5, using an OmniGrid 100 microarrayer (Genomic Solutions, Ann Arbor, MI, USA) equipped with SMP3 pins (TeleChem International, Sunnyvale, CA, USA), and processed according to the manufacturer’s protocol.
After hybridization, images of the arrays were acquired by scanning (Microaray scanner G2505B; Agilent technologies, Palo Alto, USA), and spot analysis and quality control were automatically performed using BlueFuse 3.4 software (BlueGnome, Cambridge, UK). When the BlueFuse quality flag was below one or the confidence value was below 0.1, spots were excluded from further analysis. The log2 tumor to normal fluorescence ratio was calculated for each spot and normalized against the mode of the ratios of all autosomes. The package CGH call was used for data segmentation and defining copy-number gains and losses of each clone in the array CGH profile [29].
Multiplex ligation-dependent probe amplification
MLPA was performed, as previously described, [24] using two different probe mixes. One probemix contained 38 probes representing 31 different genes on chromosome 20 and 10 control probes located on chromosomes 2, 3, 4, 5, 12, and 16. The second probemix contained 11 probes on chromosome 8, 12 probes on chromosome 13, 16 probes on chromosome 20 representing 14 different genes, and 8 control probes located on chromosomes 2, 4, 12, and 16. Some genes on chromosome 20 are present in both probe mixes, leaving 35 different genes on chromosome 20 by combining these two probe mixes. DNA of the cell line HT29, showing a gain on chromosomes 8, 13, and 20, was used as positive control. A human pool of DNA isolated from blood of 36 healthy individuals and a pool of DNA isolated of 30 normal gastric and colon mucosa, spleen, liver, and kidney tissue samples (FFPE), were used as normal controls.
Of each sample, approximately 100 ng of DNA in a volume of 5 µl was denaturated at 98°C for 5 min. A mixture of 1.5-µl salsa probes (1–4 fmol of each short synthetic probe oligonucleotide and each phage M13-derived long probe oligonucleotide in TE (10 mM Tris-HCl, pH 8.2, 1 mM ethylenediamine tetraacetic acid (EDTA))) and 1.5 µl of MLPA buffer (1.5 M KCl, 300 mM Tris-HCl, pH 8.5, 1 mM EDTA) was added. The mixture was heated for 1 min at 95°C followed by 16 h of incubation at 60°C to allow the MLPA hemipobes to hybridize. Next, 32 µl of ligase-65 mixture (dilution buffer containing 2.6 mM MgCl2, 5 mM Tris-HCl, pH 8.5, 0.013% non-ionic detergents, 0.2 mM of nicotinamide adenine dinucleotide (NAD), and 1 U of ligase-65 enzyme) was added to each sample for ligation of hybridized hemiprobes during a 10–15 min of incubation at 54°C, followed by a 5 min of incubation at 98°C to inactivate the ligase.
Polymerase chain reaction (PCR) was performed with 10 µl of polymerase mixture containing the PCR primers (10 pmol), dNTPs (2.5 nmol), and 2.5 U Taq polymerase (promega), 4 µl of PCR buffer (2.6 mM MgCl2, 5 mM Tris-HCl, pH 8.5, 0.013% non-ionic detergents, 0.2 mM NAD), 26 µl of water and 10 µl of MLPA ligation reaction.
Multiplex ligation-dependent probe amplification data analysis
Analysis of the MLPA PCR products for each gene was performed on an ABI 3100 capillary sequencer (Applied Biosystems, Warrington, UK) in a mixture of 8.5 µl of deionized formamide (Applied Biosystems, Warrington, UK), 1 µl of PCR product and 0.5 µl marker including a ROX-labeled internal size standard (ROX-500 Genescan; Applied Biosystems, Warrington, UK). Data analysis was performed using the MLPAnalyzer version 8.0 (http://www.mlpa.com/coffalyser/) [30]. For each tumor, peak heights for every probe were derived from the ABI output and median peak heights of at least two different ligation reactions and three different PCR reactions were calculated. For each sample, tumor to normal DNA copy-number ratios was calculated per probe by dividing the median peak heights in the tumor tissue by the median peak heights in the reference DNA. All ratios were normalized by setting the median of the tumor to reference DNA copy-number ratios of the control genes in the probe mixture to 1.0. When multiple probes were present for one gene, the mean value of the probes was calculated and used for further analysis. Tumor to normal ratios below 0.7 and above 1.3 was considered as a loss or gain, respectively. TMEV software 3.1 (http://www.tigr.org/) was used to present descriptive data.
Statistical analysis
Box and scatter plots were used to present descriptive statistics. Chi-square test was used to evaluate associations of DNA copy-number gain of chromosomes 8q, 13q, and 20q with clinicopathological variables. t Test was used to evaluate differences in DNA copy-number aberrations and age of the patients. Mann-Whitney U test and Kruskal-Wallis H test were used to evaluate differences in DNA copy-number changes of each gene between lymph node status and histological tumor type according to the Laurén classification [31], respectively. Correlation coefficients between the log2 ratios for the array CGH and MLPA analysis were obtained by Spearman correlation (SPSS 12.0.1 for Windows; SPSS Inc. Chicago, IL, USA). A threshold of 0.05 for significance was used.
Results
Array CGH analysis and correlation with lymph node status and histological type
Of the 63 gastric adenocarcinomas analyzed by array CGH, 49 (77.8%) showed gains on chromosome 8, 25 (39.7%) showed gains on chromosome 13, and 49 (77.8%) showed gains on chromosome 20. Concurrent gains of chromosomes 8 and 13, 8 and 20, and 13 and 20 were observed in four (6.3%), 24 (38.1%), and three (4.8%) of the gastric adenocarinomas, respectively. Concurrent gains of chromosomes 8, 13, and 20 were observed in 17 (27%) gastric adenocarcinomas. Gain of chromosome 20q was significantly correlated with lymph node status (p = 0.05) and histological type of the tumor (p = 0.02), being more common in lymph node-positive gastric cancers (85%) than in lymph node-negative gastric cancers (64%) and more common in intestinal-type (87%) and mixed-type (88%) gastric cancers than in diffuse-type (53%) gastric cancers. No significant (NS) correlation was found between 20q gain and age and gender of the patients and tumor size. No correlation of 8q gain or 13q gain with clinicopathological variables was found. An overview of the frequencies of copy-number gains on chromosomes 8q, 13q, and 20q detected by array CGH analysis is given in Table 1. An overview of patient and tumor characteristics is given in Table 2.
Multiplex ligation-dependent probe amplification analysis of DNA copy-number aberrations and correlation with lymph node status and histological type
All 63 gastric adenocarcinomas analyzed by array CGH were analyzed by MLPA. An overview of the DNA copy-number ratios of all individual genes on chromosomes 8, 13, and 20 in all tumor samples is presented as a heatmap in Fig. 1.
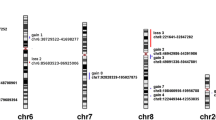
Gains of genes on chromosome 8q were observed in 9.5%–73.0% of the gastric adenocarcinomas, with the highest frequencies of gains observed in c-myc (73.0%). Gains of genes on chromosome 13q were detected in 11.1%–28.6%, with the highest frequencies of gains observed in FOXO1A and ATP7B (both 28.6%). Frequent DNA copy-number gains of multiple genes on chromosome 20q were observed. Gain of TOP1, PTPN1, CYP24A1, and GATA5 was observed in more than 60% of the gastric adenocarcinomas, and gain of BCL2L1, MYBL2, STX16, and GNAS was observed in more than 70% of the gastric adenocarcinomas. When correlating gene-specific copy-number status to lymph node status, EIF3S6 (p = 0.04), located on chromosome 8q22 and TNFRSF6B (p = 0.02), located on chromosome 20q13.3, were significantly different between lymph node-positive and lymph node-negative gastric adenocarcinomas. Although Mann-Whitney U test yielded a significant difference in copy-number of the gene EIF3S6 between lymph node-positive and lymph node-negative gastric adenocarcinomas, mean copy-number ratio of both carcinoma groups were within the normal copy-number range (DNA copy-number ratio between 0.7 and 1.3). Lymph node-positive gastric adenocarcinomas showed gain (DNA copy-number ratio >1.3) of TNFRSF6B while lymph node negative gastric adenocarcinomas showed normal copy-number ratios of this gene (Fig. 2). When correlating gene-specific copy-number ratio to histological tumor type, Kruskall-Wallis H test yielded five significant genes. Mean copy-number ratios of FBXO25 (p = 0.01), located on 8p23.3, RASL11A (p = 0.03), located on 13q12.2 and CDX2 (p = 0.03), located on 13q12, were within the normal copy-number range for all three tumor types. Mean copy-number ratio of ZNF217 (p = 0.02), located on 20q13.2, showed gain in intestinal and mixed histological gastric cancer types versus normal in the diffuse histological types (Fig. 3). STX16 (p = 0.03), located on 20q13.32, showed mean copy-number gain in all three tumor types, but higher mean copy-number ratios in the intestinal and mixed types of gastric carcinomas compared with diffuse-type gastric carcinomas (1.90, 2.03, and 1.47, respectively). A detailed overview of the frequencies of gains per gene and mean copy-number ratio per tumor group, i.e., lymph node-positive versus lymph node-negative gastric adenocarcinomas and intestinal- versus diffuse- versus mixed-type gastric carcinomas are given in Table 3.
Box plot of DNA copy number ratios of the gene TNFRFS6B between lymph node-negative and lymph node-positive gastric adenocarcinomas. Lymph node-positive gastric adenocarcinomas have significantly higher DNA copy number ratios of TNFRFS6B compared with lymph node-negative gastric adenocarcinomas (p = 0.02). The central box covers the middle 50% of the data values between the upper and lower quartiles. The line across the box indicates the median. The whiskers extend from the box to the minimum and maximum values with the exception of outliers, which are marked by circles
Correlation of copy-number status of MLPA and array CGH
The BAC clones RP4-583P15 and RP4-724E16 comprised the location of the TNFRSF6B and ZNF217 genes, respectively. To evaluate the correlation between the array CGH and MLPA data, the MLPA DNA copy-number ratios were transformed to a log2 scale. Spearman correlation yielded a significant correlation between array CGH and MLPA data for both TNFRSF6B (r = 0.57, p < 0.001; Fig. 4a) and ZNF217 (r = 0.51, p < 0.001; Fig. 4b).
Discussion
Gastric cancer is a common disease with a poor prognosis [1]. Chromosomal instability is a major mechanism of genetic instability in gastric cancers which has been widely studied by array CGH analysis [2, 32]. Frequent copy-number gains of chromosomes 8q, 13q, and 20q have been detected in gastric adenocarcinomas by this technique [3–10]. In this study, we report a detailed analysis of DNA copy-number changes of genes located on chromosomes 8, 13, and 20 in gastric adenocarcinomas aiming to correlate gene-specific alterations to clinicopathological data.
In the present study of 63 gastric adenocarcinomas studied by array CGH, nearly 80% of the cancers showed gain of chromosomes 8q and 20q. These frequencies of gains are consistent with previous studies where gains of chromosomes 8q and 20q were reported in 40–80% and 50–80% of gastric cancers, respectively [4, 10, 13, 14, 17, 33]. Both gains and losses of chromosome 13q have been detected in gastric cancers. Losses of chromosome 13q have been described in up to 30% of gastric cancers and gains of 13q have been described in up to 40% of gastric cancers [4, 10, 13–15, 17, 33], which is consistent with the findings of the present study.
When analyzing DNA copy-number gain at the gene level, mean copy-number ratio of two genes, EIF3S6E (8q22) and TNFRSF6B (20q12.3), correlated significantly with lymph node status. Mean copy-number ratios of EIF3S6E were within the normal range for both lymph node-positive and lymph node-negative gastric cancers, corresponding to the array CGH results in this study. Mean copy-number ratio of TNFRSF6B was significantly higher in lymph node-positive compared with lymph node-negative gastric cancers and was within the range of DNA copy-number gain and normal, respectively. Although 20q gain CGH data correlated with lymph node metastasis, of the 20q genes tested, only DNA copy-number gain of TNFRSF6B was significantly correlated with lymph node metastasis. TNFRSF6B is a member of the tumor necrosis factor receptor superfamily and is also known as decoy receptor 3 (DrR3). TNFRSF6B binds to FasL and inhibits FasL-induced apoptosis [34]. Overexpression of this gene has been observed in multiple cancer types, including cancers of the gastrointestinal tract. Interestingly, gastric cancers with pN2 and pN3 diseases have been shown to have significantly higher TNFRSF6B expression compared with gastric cancers with pN0 and pN1 diseases. Serum levels of TNFRSF6B were correlated with TNM stage in gastric cancers [35, 36]. We found TNFRSF6B DNA copy-number ratios on average to be gained in gastric cancers metastasized to the lymph nodes (pN1–3) compared with on average normal DNA copy-number ratios of TNFRSF6B in non-metastasized gastric cancers (pN0). Although previous studies have reported overexpression of TNFRSF6B without gene amplification [37], results of the present study indicate that DNA copy-number status of TNFRSF6B could be a valuable marker for identifying lymph node-positive gastric cancers.
DNA copy-number status of five genes, FBXO25 (8p23.3), RASL11A (13q12.2), CDX2 (13q12), ZNF217 (20q13.2), and STX16 (20q13.32), correlated significantly with histological tumor type, i.e., intestinal-, diffuse-, or mixed-type gastric cancer. Mean DNA copy-number ratios of FBXO25, RASL11A, and CDX2, however, were within the normal copy-number range for all three histological types, in consistence with the array CGH data of this study. The biological meaning of this difference, therefore, is not immediately clear, but could, e.g., relate to only a subpopulation of tumor cells being affected.
STX16 showed significantly higher mean copy-number ratios in intestinal- and mixed-type compared with diffuse-type gastric cancers; however, DNA copy-number gain was observed in all three tumor groups. STX16 encodes a syntaxin protein which plays a role in intracellular trafficking [38], but its role in cancer still has to be elucidated.
ZNF217 encodes a transcription factor which was shown to be involved in immortalization of breast cancer cells when overexpressed [39]. High-level amplifications of ZNF217 have been previously described in approximately 10% of gastric cancers [9, 10, 40]. In the study of Weiss et al. on three tumors, all of the intestinal-type, out of 27 gastric adenocarcinomas, showed an amplification on 20q13. Fluorescence in situ hybridization (FISH) analysis was performed on these three tumors, confirming the copy-number increase at this locus [9]. Results from the present study suggest that DNA copy-number gain of ZNF217 plays a more important role in intestinal- and mixed-type compared with diffuse-type gastric cancers, since gains of mean copy-number ratios are observed in intestinal- and mixed-type gastric cancers and normal mean copy-number ratio in diffuse-type gastric cancers.
In summary, we present a detailed DNA copy-number analysis of a panel of genes located on chromosomes 8, 13, and 20 in gastric adenocarcinomas. We found that DNA copy-number gain of ZNF217 plays a role mainly in intestinal- and mixed-type but not in diffuse-type gastric adenocarcinomas. In addition, we showed that DNA copy-number gain of TNFRSF6B is significantly correlated to gastric cancers with lymph node metastasis, indicating a potential role for this gene as a biomarker for identifying patients at high risk of lymph node metastasis, who might, therefore, benefit from extended lymph node resection or neoadjuvant chemotherapy to improve gastric cancer outcome.
References
Parkin DM, Bray F, Ferlay J et al (2005) Global cancer statistics, 2002. CA Cancer J Clin 55:74–108
Lengauer C, Kinzler KW, Vogelstein B (1998) Genetic instabilities in human cancers. Nature 396:643–649
Hidaka S, Yasutake T, Kondo M et al (2003) Frequent gains of 20q and losses of 18q are associated with lymph node metastasis in intestinal-type gastric cancer. Anticancer Res 23:3353–3357
Kimura Y, Noguchi T, Kawahara K et al (2004) Genetic alterations in 102 primary gastric cancers by comparative genomic hybridization: gain of 20q and loss of 18q are associated with tumor progression. Mod Pathol 17:1328–1337
Koo SH, Kwon KC, Shin SY et al (2000) Genetic alterations of gastric cancer: comparative genomic hybridization and fluorescence In situ hybridization studies. Cancer Genet Cytogenet 117:97–103
Sakakura C, Mori T, Sakabe T et al (1999) Gains, losses, and amplifications of genomic materials in primary gastric cancers analyzed by comparative genomic hybridization. Genes Chromosomes Cancer 24:299–305
Takada H, Imoto I, Tsuda H et al (2005) Screening of DNA copy-number aberrations in gastric cancer cell lines by array-based comparative genomic hybridization. Cancer Sci 96:100–110
van Grieken NC, Weiss MM, Meijer GA et al (2000) Helicobacter pylori-related and -non-related gastric cancers do not differ with respect to chromosomal aberrations. J Pathol 192:301–306
Weiss MM, Snijders AM, Kuipers EJ et al (2003) Determination of amplicon boundaries at 20q13.2 in tissue samples of human gastric adenocarcinomas by high-resolution microarray comparative genomic hybridization. J Pathol 200:320–326
Weiss MM, Kuipers EJ, Postma C et al (2004) Genomic alterations in primary gastric adenocarcinomas correlate with clinicopathological characteristics and survival. Cell Oncol 26:307–317
Wu MS, Shun CT, Wang HP et al (1997) Genetic alterations in gastric cancer: relation to histological subtypes, tumor stage, and Helicobacter pylori infection. Gastroenterology 112:1457–1465
Buffart TE, Carvalho B, Mons T et al (2007) DNA copy-number profiles of gastric cancer precursor lesions. BMC Genomics 8:345
Kim YH, Kim NG, Lim JG et al (2001) Chromosomal alterations in paired gastric adenomas and carcinomas. Am J Pathol 158:655–662
Kokkola A, Monni O, Puolakkainen P et al (1998) Presence of high-level DNA copy-number gains in gastric carcinoma and severely dysplastic adenomas but not in moderately dysplastic adenomas. Cancer Genet Cytogenet 107:32–36
van Dekken H, Alers JC, Riegman PH et al (2001) Molecular cytogenetic evaluation of gastric cardia adenocarcinoma and precursor lesions. Am J Pathol 158:1961–1967
Weiss MM, Kuipers EJ, Postma C et al (2003) Genome wide array comparative genomic hybridisation analysis of premalignant lesions of the stomach. Mol Pathol 56:293–298
Buffart T, Carvalho B, Hopmans E et al (2007) Gastric cancers in young and elderly patients show different genomic profiles. J Pathol 211:45–51
Weiss MM, Kuipers EJ, Postma C et al (2003) Genomic profiling of gastric cancer predicts lymph node status and survival. Oncogene 22:1872–1879
Carvalho B, Postma C, Mongera S et al (2008) Integration of DNA and expression microarray data unravels seven putative oncogenes on 20Q amplicon involved in colorectal adenoma to carcinoma progression. Cell Oncol 30:145–146
Hermsen M, Postma C, Baak J et al (2002) Colorectal adenoma to carcinoma progression follows multiple pathways of chromosomal instability. Gastroenterology 123:1109–1119
Aust DE, Muders M, Kohler A et al (2004) Prognostic relevance of 20q13 gains in sporadic colorectal cancers: a FISH analysis. Scand J Gastroenterol 39:766–772
Postma C, Terwischa S, Hermsen MA et al (2007) Gain of chromosome 20q is an indicator of poor prognosis in colorectal cancer. Cell Oncol 29:73–75
Schouten JP, McElgunn CJ, Waaijer R et al (2002) Relative quantification of 40 nucleic acid sequences by multiplex ligation-dependent probe amplification. Nucleic Acids Res 30:e57
Buffart TE, Coffa J, Hermsen MA et al (2005) DNA copy-number changes at 8q11–24 in metastasized colorectal cancer. Cell Oncol 27:57–65
Postma C, Hermsen MA, Coffa J et al (2005) Chromosomal instability in flat adenomas and carcinomas of the colon. J Pathol 205:514–521
Bonenkamp JJ, Hermans J, Sasako M et al (1999) Extended lymph-node dissection for gastric cancer. Dutch Gastric Cancer Group. N Engl J Med 340:908–914
Buffart TE, Tijssen M, Krugers T et al (2007) DNA quality assessment for array CGH by isothermal whole genome amplification. Cell Oncol 29:351–359
Snijders AM, Nowak N, Segraves R et al (2001) Assembly of microarrays for genome-wide measurement of DNA copy-number. Nat Genet 29:263–264
van de Wiel MA, Kim KI, Vosse SJ et al (2007) CGHcall: calling aberrations for array CGH tumor profiles. Bioinformatics 23:892–894
Coffa J, van de Wiel MA, Diosdado B et al (2008) MLPAnalyzer: data analysis tool for reliable automated normalization of MLPA fragment data. Cell Oncol 30:323–335
Laurén P (1965) The two histological main types of gastric carcinoma: diffuse and so-called intestinal-type carcinoma. An attemt at a histo-clinical classification. Acta Pathol Microbiol Scand 64:31–49
Pinkel D, Albertson DG (2005) Array comparative genomic hybridization and its applications in cancer. Nat Genet 37(Suppl):S11–S17
Carvalho B, Buffart TE, Reis RM et al (2006) Mixed gastric carcinomas show similar chromosomal aberrations in both their diffuse and glandular components. Cell Oncol 28:283–294
Pitti RM, Marsters SA, Lawrence DA et al (1998) Genomic amplification of a decoy receptor for Fas ligand in lung and colon cancer. Nature 396:699–703
Wu Y, Han B, Sheng H et al (2003) Clinical significance of detecting elevated serum DcR3/TR6/M68 in malignant tumor patients. Int J Cancer 105:724–732
Wu Y, Guo E, Yu J et al (2008) High DcR3 expression predicts stage pN2–3 in gastric cancer. Am J Clin Oncol 31:79–83
Bai C, Connolly B, Metzker ML et al (2000) Overexpression of M68/DcR3 in human gastrointestinal tract tumors independent of gene amplification and its location in a four-gene cluster. Proc Natl Acad Sci U S A 97:1230–1235
Simonsen A, Bremnes B, Ronning E et al (1998) Syntaxin-16, a putative Golgi t-SNARE. Eur J Cell Biol 75:223–231
Nonet GH, Stampfer MR, Chin K et al (2001) The ZNF217 gene amplified in breast cancers promotes immortalization of human mammary epithelial cells. Cancer Res 61:1250–1254
Yang SH, Seo MY, Jeong HJ et al (2005) Gene copy-number change events at chromosome 20 and their association with recurrence in gastric cancer patients. Clin Cancer Res 11:612–620
Acknowledgements
This study was financially supported by the Dutch Cancer Society, grant KWF2004-3051.
Conflict of interest statement
We declare that we have no conflict of interest.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Buffart, T.E., van Grieken, N.C.T., Tijssen, M. et al. High resolution analysis of DNA copy-number aberrations of chromosomes 8, 13, and 20 in gastric cancers. Virchows Arch 455, 213–223 (2009). https://doi.org/10.1007/s00428-009-0814-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-009-0814-y