Abstract
The amazing array of diversity among insect wings offers a powerful opportunity to study the mechanisms guiding morphological evolution. Studies in Drosophila (the fruit fly) have identified dozens of genes important for wing development. These genes are often called candidate genes, serving as an ideal starting point to study wing development in other insects. However, we also need to explore beyond the candidate genes to gain a more comprehensive view of insect wing evolution. As a first step away from the traditional candidate genes, we utilized Tribolium (the red flour beetle) as a model and assessed the potential involvement of a group of developmental toolkit genes (embryonic patterning genes) in beetle wing development. We hypothesized that the highly pleiotropic nature of these developmental genes would increase the likelihood of finding novel wing genes in Tribolium. Through the RNA interference screening, we found that Tc-cactus has a less characterized (but potentially evolutionarily conserved) role in wing development. We also found that the odd-skipped family genes are essential for the formation of the thoracic pleural plates, including the recently discovered wing serial homologs in Tribolium. In addition, we obtained several novel insights into the function of these developmental genes, such as the involvement of mille-pattes and Tc-odd-paired in metamorphosis. Despite these findings, no gene we examined was found to have novel wing-related roles unique in Tribolium. These results suggest a relatively conserved nature of developmental toolkit genes and highlight the limited degree to which these genes are co-opted during insect wing evolution.




Similar content being viewed by others
References
Abouheif E, Wray GA (2002) Evolution of the gene network underlying wing polyphenism in ants. Science 297(5579):249–252. doi:10.1126/science.1071468
Angelini DR, Kikuchi M, Jockusch EL (2009) Genetic patterning in the adult capitate antenna of the beetle Tribolium castaneum. Dev Biol 327(1):240–251. doi:10.1016/j.ydbio.2008.10.047
Angelini DR, Smith FW, Jockusch EL (2012) Extent with modification: leg patterning in the beetle Tribolium castaneum and the evolution of serial homologs. G3 (Bethesda) 2(2):235–248. doi:10.1534/g3.111.001537
Beermann A, Schroder R (2004) Functional stability of the aristaless gene in appendage tip formation during evolution. Dev Genes Evol 214(6):303–308. doi:10.1007/s00427-004-0411-7
Beermann A, Aranda M, Schroder R (2004) The Sp8 zinc-finger transcription factor is involved in allometric growth of the limbs in the beetle Tribolium castaneum. Development 131(4):733–742. doi:10.1242/dev.00974
Benedyk MJ, Mullen JR, DiNardo S (1994) odd-paired: a zinc finger pair-rule protein required for the timely activation of engrailed and wingless in Drosophila embryos. Genes Dev 8(1):105–117
Bowsher JH, Wray GA, Abouheif E (2007) Growth and patterning are evolutionarily dissociated in the vestigial wing discs of workers of the red imported fire ant, Solenopsis invicta. J Exp Zool B Mol Dev Evol 308(6):769–776. doi:10.1002/jez.b.21200
Brook WJ, Diaz-Benjumea FJ, Cohen SM (1996) Organizing spatial pattern in limb development. Annu Rev Cell Dev Biol 12:161–180. doi:10.1146/annurev.cellbio.12.1.161
Brown SJ, Mahaffey JP, Lorenzen MD, Denell RE, Mahaffey JW (1999) Using RNAi to investigate orthologous homeotic gene function during development of distantly related insects. Evol Dev 1(1):11–15
Brown SJ, Denell RE, Beeman RW (2003) Beetling around the genome. Genet Res 82(3):155–161
Bucher G, Klingler M (2014) iBeetle: functional genomics of insect development and metamorphosis. http://ibeetle.uni-goettingen.de
Bucher G, Scholten J, Klingler M (2002) Parental RNAi in Tribolium (Coleoptera). Curr Biol 12(3):R85–R86
Carroll SB, Grenier JK, Weatherbee SD (2001) From DNA to diversity. Science, Blackwell
Choe CP, Miller SC, Brown SJ (2006) A pair-rule gene circuit defines segments sequentially in the short-germ insect Tribolium castaneum. Proc Natl Acad Sci U S A 103(17):6560–6564. doi:10.1073/pnas.0510440103
Clark-Hachtel CM, Linz DM, Tomoyasu Y (2013) Insights into insect wing origin provided by functional analysis of vestigial in the red flour beetle, Tribolium castaneum. Proc Natl Acad Sci U S A 110(42):16951–16956. doi:10.1073/pnas.1304332110
Crowson RA (1981) The biology of the Coleoptera. Academic, London
de Celis Ibeas JM, Bray SJ (2003) Bowl is required downstream of Notch for elaboration of distal limb patterning. Development 130(24):5943–5952. doi:10.1242/dev.00833
Denell R (2008) Establishment of tribolium as a genetic model system and its early contributions to evo-devo. Genetics 180(4):1779–1786. doi:10.1534/genetics.104.98673
Finn RD, Bateman A, Clements J, Coggill P, Eberhardt RY, Eddy SR, Heger A, Hetherington K, Holm L, Mistry J, Sonnhammer EL, Tate J, Punta M (2013) Pfam: the protein families database. Nucleic Acids Res. doi:10.1093/nar/gkt1223
Friedman AA, Tucker G, Singh R, Yan D, Vinayagam A, Hu Y, Binari R, Hong P, Sun X, Porto M, Pacifico S, Murali T, Finley RL Jr, Asara JM, Berger B, Perrimon N (2011) Proteomic and functional genomic landscape of receptor tyrosine kinase and ras to extracellular signal-regulated kinase signaling. Sci Signal 4(196):rs10. doi:10.1126/scisignal.2002029
Galindo MI, Pueyo JI, Fouix S, Bishop SA, Couso JP (2007) Peptides encoded by short ORFs control development and define a new eukaryotic gene family. PLoS Biol 5(5):e106. doi:10.1371/journal.pbio.0050106
Gilbert SF (2014) Developmental biology. Tenth Edition edn. Sinauer Associates, Sunderland
Grimaldi D, Engel MS (2005) Evolution of the insects. Cambridge University Press, Cambridge
Hall BG (2011) Phylogenetic trees made easy: a how-to manual, 4th edn. Sinauer Associates, Sunderland
Hao I, Green RB, Dunaevsky O, Lengyel JA, Rauskolb C (2003) The odd-skipped family of zinc finger genes promotes Drosophila leg segmentation. Dev Biol 263(2):282–295
Hart MC, Wang L, Coulter DE (1996) Comparison of the structure and expression of odd-skipped and two related genes that encode a new family of zinc finger proteins in Drosophila. Genetics 144(1):171–182
Ip YT, Reach M, Engstrom Y, Kadalayil L, Cai H, Gonzalez-Crespo S, Tatei K, Levine M (1993) Dif, a dorsal-related gene that mediates an immune response in Drosophila. Cell 75(4):753–763
Kim HS, Murphy T, Xia J, Caragea D, Park Y, Beeman RW, Lorenzen MD, Butcher S, Manak JR, Brown SJ (2010) BeetleBase in 2010: revisions to provide comprehensive genomic information for Tribolium castaneum. Nucleic Acids Res 38(Database issue):D437–D442. doi:10.1093/nar/gkp807
Klingler M (2004) Tribolium. Curr Biol 14(16):R639–R640. doi:10.1016/j.cub.2004.08.004
Knorr E, Bingsohn L, Kanost MR, Vilcinskas A (2013) Tribolium castaneum as a model for high-throughput RNAi screening. Adv Biochem Eng Biotechnol 136:163–178. doi:10.1007/10_2013_208
Kondo T, Hashimoto Y, Kato K, Inagaki S, Hayashi S, Kageyama Y (2007) Small peptide regulators of actin-based cell morphogenesis encoded by a polycistronic mRNA. Nat Cell Biol 9(6):660–665. doi:10.1038/ncb1595
Kondo T, Plaza S, Zanet J, Benrabah E, Valenti P, Hashimoto Y, Kobayashi S, Payre F, Kageyama Y (2010) Small peptides switch the transcriptional activity of Shavenbaby during Drosophila embryogenesis. Science 329(5989):336–339. doi:10.1126/science.1188158
Kukalova-Peck J (1983) Origin of the insect wing and wing articulation from the arthropodan leg. Can J Zool 61(7):1618–1669. doi:10.1139/z83-217
Kukalova-Peck J (2008) Phylogeny of higher taxa in Insecta: finding synapomorphies in the extant fauna and separating them from homoplasies. Evol Biol 35(1):4–51. doi:10.1007/s11692-007-9013-4
Lawrence JF, Britton EB (1991) Coleoptera (beetles). In: The insects of Australia: a textbook for students and research workers, vol 2. Second Edition edn. CSIRO Division of Entomology, Carlton, Australia: Melbourne University Press, pp pp. 543–564
Linz DM, Clark-Hachtel CM, Borras-Castells F, Tomoyasu Y (2014) Larval RNA interference in the red flour beetle, Tribolium castaneum. J Vis Exp (92). doi:10.3791/52059
Lorenzen MD, Berghammer AJ, Brown SJ, Denell RE, Klingler M, Beeman RW (2003) piggyBac-mediated germline transformation in the beetle Tribolium castaneum. Insect Mol Biol 12(5):433–440
Lynch JA, Roth S (2011) The evolution of dorsal-ventral patterning mechanisms in insects. Genes Dev 25(2):107–118. doi:10.1101/gad.2010711
Macdonald WP, Martin A, Reed RD (2010) Butterfly wings shaped by a molecular cookie cutter: evolutionary radiation of lepidopteran wing shapes associated with a derived Cut/wingless wing margin boundary system. Evol Dev 12(3):296–304. doi:10.1111/j.1525-142X.2010.00415.x
Martin A, Papa R, Nadeau NJ, Hill RI, Counterman BA, Halder G, Jiggins CD, Kronforst MR, Long AD, McMillan WO, Reed RD (2012) Diversification of complex butterfly wing patterns by repeated regulatory evolution of a Wnt ligand. Proc Natl Acad Sci U S A 109(31):12632–12637. doi:10.1073/pnas.1204800109
Miller SC, Miyata K, Brown SJ, Tomoyasu Y (2012) Dissecting systemic RNA interference in the red flour beetle Tribolium castaneum: parameters affecting the efficiency of RNAi. PLoS One 7(10):e47431. doi:10.1371/journal.pone.0047431
Moussian B, Roth S (2005) Dorsoventral axis formation in the Drosophila embryo—shaping and transducing a morphogen gradient. Curr Biol 15(21):R887–R899. doi:10.1016/j.cub.2005.10.026
Niwa N, Akimoto-Kato A, Niimi T, Tojo K, Machida R, Hayashi S (2010) Evolutionary origin of the insect wing via integration of two developmental modules. Evol Dev 12(2):168–176. doi:10.1111/j.1525-142X.2010.00402.x
Nunes da Fonseca R, von Levetzow C, Kalscheuer P, Basal A, van der Zee M, Roth S (2008) Self-regulatory circuits in dorsoventral axis formation of the short-germ beetle Tribolium castaneum. Dev Cell 14(4):605–615. doi:10.1016/j.devcel.2008.02.011
Nusslein-Volhard C, Wieschaus E (1980) Mutations affecting segment number and polarity in Drosophila. Nature 287(5785):795–801
Ohde T, Yaginuma T, Niimi T (2013) Insect morphological diversification through the modification of wing serial homologs. Science 340(6131):495–498. doi:10.1126/science.1234219
Philip BN, Tomoyasu Y (2011) Gene knockdown analysis by double-stranded RNA injection molecular methods for evolutionary genetics. In: Orgogozo V, Rockman MV (eds), vol 772. Methods in molecular biology. Humana Press, pp 471–497. doi:10.1007/978-1-61779-228-1_28
Pueyo JI, Couso JP (2008) The 11-aminoacid long Tarsal-less peptides trigger a cell signal in Drosophila leg development. Dev Biol 324(2):192–201. doi:10.1016/j.ydbio.2008.08.025
Rambaut A (2014) FigTree. http://tree.bio.ed.ac.uk/software/figtree/
Rasnitsyn AP (1981) A modified paranotal theory of insect wing origin. J Morphol 168(3):331–338. doi:10.1002/jmor.1051680309
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Hohna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61(3):539–542. doi:10.1093/sysbio/sys029
Roth S, Stein D, Nusslein-Volhard C (1989) A gradient of nuclear localization of the dorsal protein determines dorsoventral pattern in the Drosophila embryo. Cell 59(6):1189–1202
Rushlow CA, Han K, Manley JL, Levine M (1989) The graded distribution of the dorsal morphogen is initiated by selective nuclear transport in Drosophila. Cell 59(6):1165–1177
Sanson B (2001) Generating patterns from fields of cells. Examples from Drosophila segmentation. EMBO Rep 2(12):1083–1088. doi:10.1093/embo-reports/kve255
Savard J, Marques-Souza H, Aranda M, Tautz D (2006) A segmentation gene in tribolium produces a polycistronic mRNA that codes for multiple conserved peptides. Cell 126(3):559–569. doi:10.1016/j.cell.2006.05.053
Shirai LT, Saenko SV, Keller RA, Jeronimo MA, Brakefield PM, Descimon H, Wahlberg N, Beldade P (2012) Evolutionary history of the recruitment of conserved developmental genes in association to the formation and diversification of a novel trait. BMC Evol Biol 12:21. doi:10.1186/1471-2148-12-21
St Pierre SE, Ponting L, Stefancsik R, McQuilton P (2014) FlyBase 102—advanced approaches to interrogating FlyBase. Nucleic Acids Res 42(Database issue):D780–D788. doi:10.1093/nar/gkt1092
Steward R (1989) Relocalization of the dorsal protein from the cytoplasm to the nucleus correlates with its function. Cell 59(6):1179–1188
Stoehr AM, Walker JF, Monteiro A (2013) Spalt expression and the development of melanic color patterns in pierid butterflies. Evodevo 4(1):6. doi:10.1186/2041-9139-4-6
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739. doi:10.1093/molbev/msr121
Tomoyasu Y, Denell RE (2004) Larval RNAi in Tribolium (Coleoptera) for analyzing adult development. Dev Genes Evol 214(11):575–578. doi:10.1007/s00427-004-0434-0
Tomoyasu Y, Wheeler SR, Denell RE (2005) Ultrabithorax is required for membranous wing identity in the beetle Tribolium castaneum. Nature 433 (7026):643–647. doi:http://www.nature.com/nature/journal/v433/n7026/suppinfo/nature03272_S1.html
Tomoyasu Y, Miller SC, Tomita S, Schoppmeier M, Grossmann D, Bucher G (2008) Exploring systemic RNA interference in insects: a genome-wide survey for RNAi genes in Tribolium. Genome Biol 9(1):R10. doi:10.1186/gb-2008-9-1-r10
Tomoyasu Y, Arakane Y, Kramer KJ, Denell RE (2009) Repeated co-options of exoskeleton formation during wing-to-elytron evolution in beetles. Curr Biol 19(24):2057–2065. doi:10.1016/j.cub.2009.11.014
Trauner J, Schinko J, Lorenzen MD, Shippy TD, Wimmer EA, Beeman RW, Klingler M, Bucher G, Brown SJ (2009) Large-scale insertional mutagenesis of a coleopteran stored grain pest, the red flour beetle Tribolium castaneum, identifies embryonic lethal mutations and enhancer traps. BMC Biol 7:73. doi:10.1186/1741-7007-7-73
Tribolium Genome Sequencing Consortium (2008) The genome of the model beetle and pest Tribolium castaneum. Nature 452(7190):949–955. doi:10.1038/nature06784
Wang L, Coulter DE (1996) bowel, an odd-skipped homolog, functions in the terminal pathway during Drosophila embryogenesis. EMBO J 15(12):3182–3196
Weatherbee SD, Nijhout HF, Grunert LW, Halder G, Galant R, Selegue J, Carroll S (1999) Ultrabithorax function in butterfly wings and the evolution of insect wing patterns. Curr Biol 9(3):109–115
Acknowledgments
We thank Susan Brown and the Brown lab at Kansas State University for the provided clones, the Center for Bioinformatics and Functional Genomics at Miami University for the technical support, and the members of the Tomoyasu lab for the discussion. This work was supported by a Miami University start-up grant (to Y.T.) and National Science Foundation Grant IOS 0950964 (to Y.T.).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by: Siegfried Roth
Electronic supplementary material
Below is the link to the electronic supplementary material.
Document S1
Sequence alignments of odd-skipped family. a Amino acid alignment. Orange box highlights the region used for tree building (pfam defined Zn fingers). b Nucleotide alignment. Green box highlights the region used for tree building. These sequences correspond to the amino acid regions highlighted in orange in a. Alignments created and curated by the ClustalW module with the default setting in MEGA 5.2.1. Dm: Drosophila melanogaster, Tc: Tribolium castaneum, and Am: Apis mellifera (PDF 70 kb)
Document S2
Nucleotide alignments of the two Zn finger coding regions that are conserved among all odd-skipped family members (odd, sob, bowl, drm), and alignments of the Zn finger coding regions that are conserved among three members (odd, sob, and bowl). Identity matrices showing percent similarity among the family members are included. ClustalW2 with the default setting for nucleotide sequences was used for each alignment (PDF 71 kb)
Table S1
dsRNA fragments used in this study. *Templates for dsRNA synthesis were made with a primer designed for the pCR4-TOPO vector: TOPO RNAi (taatacgactcactatagggcgaatt). This primer works both the forward and the reverse direction. **Templates for dsRNA synthesis were made with primers designed for pGEM-T Easy vector: pGEMTE_RNAi F1 (taatacgactcactatagggcggccg) and pGEMTE_RNAi R1 (taatacgactcactatagggccgcga). Underline indicates the T7 promoter sequence. ***cDNA clones obtained from the Brown laboratory at Kansas State University (PDF 40 kb)
Fig. S1
RNAi for a subset of genes revealed evolutionarily conserved roles. a-c wild-type. The ventral view of the entire adult (a), antenna (b), and the tarsal segments of a metathoracic leg (c). Normal segmented structures in the leg and antenna are indicated by an arrow (b, c). d, e Sp8 RNAi. The ventral view (d) and a magnified image of the white box in d (e), showing the fusion of leg and antennal segments (indicated by an arrow in d and e). f, g Tc-bab RNAi. The ventral view (f) and a magnified image of the white box in f (g), showing the fusion of tarsal segments (indicated by arrows in f and g). h, i Tc-al RNAi. The ventral view (h) and a magnified image of the white box in h (i), showing the fusion of distal club segments in the antennae (arrow in i) (GIF 144 kb)
Fig. S2
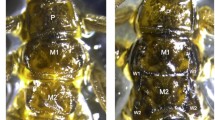
RNAi phenotypes of odd-skipped family genes. a wild-type showing proper leg joint (arrow) and trochantin formation (arrowhead). b Tc-drm cross RNAi. c Tc-sob cross RNAi. d Tc-bowl cross RNAi. e Tc-odd cross RNAi. RNAi with Tc-drm cross, Tc-sob cross, and Tc-bowl cross dsRNA fragments showed strong defects in leg segmentation (arrows in b–d) and trochantin formation (arrowhead in b-d), most likely due to the cross-reacting dsRNA molecules. Although Tc-odd cross dsRNA also contains the highly conserved region, Tc-odd cross RNAi induced no detectable abnormalities (e). f Tc-sob unique RNAi. g Tc-bowl unique RNAi. Tc-sob single knock down (f) was sufficient to produce defects in leg segmentation (arrow in f) and trochantin formation (arrowhead in f), while Tc-bowl single RNAi did not produce any abnormalities (g). h Tc-sob unique+Tc-odd cross+Tc-bowl unique triple RNAi. The simultaneous knock down of Tc-odd and Tc-bowl with Tc-sob RNAi did not enhance the Tc-sob unique RNAi phenotype (h) (GIF 256 kb)
Fig. S3
Domain architecture of the odd-skipped family members. The green regions represent the Zn finger domains. The light green represent extended conserved regions associated with the Zn finger domains seen in Bowl, Odd, and Sob. Other colors represent motifs outside of the Zn finger domains that are conserved in each class (GIF 24 kb)
Fig. S4
Phylogenetic analysis of odd-skipped family members from three holometabolous insect orders by maximum likelihood. The maximum likelihood (ML) tree is based on the alignment of the conserved Zn finger domains (See Document S1 for the alignment). Topology of the ML tree is similar to the NJ tree (Fig. 4), further supporting an ancient origin of these paralogs. Dm: Drosophila melanogaster, Tc: Tribolium castaneum, and Am: Apis mellifera (GIF 183 kb)
Fig. S5
Phylogenetic analysis of odd-skipped family members from three holometabolous insect orders by Bayesian method. The Bayesian tree is based on the alignment of the conserved Zn finger domains (see Document S1 for the alignment). a Bayesian tree constructed for Odd, Sob, and Bowl using amino acid sequences corresponding to the conserved Zn finger regions (the same region used in the NJ and ML analyses. See Document S1 for the alignment). b Bayesian tree constructed for odd, sob, and bowl using nucleotide sequences corresponding to Zn finger regions (Document S1). The amino acid-based Bayesian tree (a) is consistent with the idea of an ancient origin of the paralogs, however, the nucleotide-based tree suggests lineage specific duplications (b). Dm: Drosophila melanogaster, Tc: Tribolium castaneum, and Am: Apis mellifera (GIF 20 kb)
Fig. S6
odd-skipped family paralog orientation in the Drosophila (top) and Tribolium (bottom) genomes. The conserved microsynteny among odd-skipped family paralogs of the two evolutionarily distant insects supports the idea that the odd-skipped paralog duplications preceded the split of beetles and flies. Genome structure annotation is based on Gbrowse on BeetleBase and FlyBase (Kim et al. 2010; St Pierre et al. 2014) (PDF 57 kb)
Rights and permissions
About this article
Cite this article
Linz, D.M., Tomoyasu, Y. RNAi screening of developmental toolkit genes: a search for novel wing genes in the red flour beetle, Tribolium castaneum . Dev Genes Evol 225, 11–22 (2015). https://doi.org/10.1007/s00427-015-0488-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-015-0488-1