Abstract
Main conclusion
Australian native species of sorghum contain negligible amounts of dhurrin in their leaves and the cyanogenesis process is regulated differently under water-stress in comparison to domesticated sorghum species.
Abstract
Cyanogenesis in forage sorghum is a major concern in agriculture as the leaves of domesticated sorghum are potentially toxic to livestock, especially at times of drought which induces increased production of the cyanogenic glucoside dhurrin. The wild sorghum species endemic to Australia have a negligible content of dhurrin in the above ground tissues and thus represent a potential resource for key agricultural traits like low toxicity. In this study we investigated the differential expression of cyanogenesis related genes in the leaf tissue of the domesticated species Sorghum bicolor and the Australian native wild species Sorghum macrospermum grown in glasshouse-controlled water-stress conditions using RNA-Seq analysis to analyse gene expression. The study identified genes, including those in the cyanogenesis pathway, that were differentially regulated in response to water-stress in domesticated and wild sorghum. In the domesticated sorghum, dhurrin content was significantly higher compared to that in the wild sorghum and increased with stress and decreased with age whereas in wild sorghum the dhurrin content remained negligible. The key genes in dhurrin biosynthesis, CYP79A1, CYP71E1 and UGT85B1, were shown to be highly expressed in S. bicolor. DHR and HNL encoding the dhurrinase and α-hydroxynitrilase catalysing bio-activation of dhurrin were also highly expressed in S. bicolor. Analysis of the differences in expression of cyanogenesis related genes between domesticated and wild sorghum species may allow the use of these genetic resources to produce more acyanogenic varieties in the future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Water-stress is a key abiotic stress factor in crop production causing reduced yields (Farooq et al. 2009) and inducing vastly altered expression profiles for genes related to plant growth and development (Takahashi et al. 2018; Luo et al. 2014; Shankar et al. 2013; Ahuja et al. 2010). Water-stress tolerance in sorghum is regulated by the combined effects of genotype and environment (Borrell et al. 2006). Domesticated sorghum, Sorghum bicolor (L.) Moench is tolerant to water-stress but is susceptible to severe droughts (Borrell et al. 2014). RNA sequencing technologies have been used in sorghum to analyse drought tolerance (Dugas et al. 2011), fungal resistance (Mizuno et al. 2012) and the role of miRNAs (Calviño et al. 2011). The differential gene expression of S. bicolor in response to water-stress has been investigated in several studies (Abdel-Ghany et al. 2020; Johnson et al. 2014; Dugas et al. 2011) and has identified many up-regulated and down-regulated genes encoding formation of protective molecules and regulatory factors such as dehydrins (DHN), aquaporins (AQP), abscisic acid (ABA) responsive proteins, drought-responsive element binding proteins (DREB), cryoprotectants, detoxification related genes, osmoprotectants as well as membrane composition and signalling molecules (Abdel-Ghany et al. 2020; Azzouz-Olden et al. 2020; Gosal et al. 2009; Fetter et al. 2004).
In sorghum, water-stress induces the accumulation of the cyanogenic glucoside, dhurrin ((S)-4-hydroxymandelonitrile-β-d-glucopyranoside) (Gleadow et al. 2016; Neilson et al. 2015; Gleadow and Moller 2014; O’Donnell et al. 2013). Cyanogenic glycosides are amino acid derived bioactive compounds identified in more than 3000 plant species (Gleadow and Moller 2014). In addition to sorghum (Kojima et al. 1979), species such as almonds (Sánchez-Pérez et al. 2019), cassava (Jorgensen et al. 2005), macadamia nuts (Dahler et al. 1995) and barley (Knoch et al. 2016) produce cyanogenic glycosides.
Dhurrin is present in all major tissues of domesticated sorghum species except the mature grain (Nielsen et al. 2016a; Kahn et al. 1997). Dhurrin is part of a two-component defence system activated upon cell destruction e.g., as caused by a chewing herbivore. The bio-activation process is termed cyanogenesis and results in detonation of a hydrogen cyanide (HCN) bomb and concomitant production of stoichiometric amounts of p-hydroxybenzaldehyde (Gleadow and Moller 2014; Møller 2010) and is catalysed by specific β-glucosidases (dhurrinases, DHR1 and DHR2) and an α-hydroxynitrilase (HNL). HCN inhibits metalloenzymes in cytochrome c oxidase and may disrupt the mitochondrial respiratory electron transport chain (Nielsen et al. 2016a) and when ingested at sufficiently high concentrations be lethal in humans (Loyd and Gray 1970) and animals (Finnie et al. 2011). Cyanogenic glucosides may therefore serve as effective agents against generalist herbivores (Ballhorn et al. 2008; Zagrobelny et al. 2004; Gleadow and Woodrow 2002b). The dhurrin content in forage sorghum may be high and upon bio-activation generate a HCN content exceeding the 600-ppm maximum content for safe grazing by cattle (Gleadow et al. 2016). Humans may inadvertently have selected for cyanogenic plants over non-cyanogenic plants during evolution (Cowan et al. 2020; Jones 1998) either because their improved resistance to herbivores (McKey et al. 2010; Jones 1998) or better nitorgen use efficiency (Myrans et al. 2020; Rosati et al. 2019b). Dhurrin levels decrease with tissue age in sorghum and the highest biosynthesis and accumulation rate is observed in young sorghum seedlings (Busk and Møller 2002; Gleadow and Woodrow 2002b).
The biosynthetic pathway of CNglcs has been extensively studied in S. bicolor (Møller and Conn 1979) and the pathway intermediates and enzymes have been identified (Jones et al. 1999; Bak et al. 1998; Kahn et al. 1997; Sibbesen et al. 1994). In the first step, tyrosine is converted to (E)-p-hydroxyphenyl acetaldoxime by CYP79A1 (Sibbesen et al. 1995). Then CYP71E1 catalyses the conversion of the oxime into the cyanohydrin p-hydroxymandelonitrile (Bak et al. 1998) which upon glycosylation, catalysed by UGT85B1, is converted into dhurrin (Jones et al. 1999). In these processes, the NADPH-dependent cytochrome P450 oxidoreductase (POR) serves as an obligatory electron donor to the two P450s (Jensen et al. 2021; Halkier and Møller 1991) and is localized in the endoplasmic reticulum forming an enzyme complex (metabolon) with CYP79A1 and CYP71E1 that recruits the UGT85B1 (Laursen et al. 2015; Jensen et al. 2011; Nielsen et al. 2008). The formation of a metabolon prevents the escape of toxic intermediates. In sorghum, uptake of excess nitrogen is primarily stored as nitrate but a trade-off mechanism in partitioning nitrogen between dhurrin and nitrate may operate (Gleadow et al. 2016).
In addition to its role in defence against herbivores, dhurrin also serves as a storage form of reduced nitrogen (Bjarnholt et al. 2018; Nielsen et al. 2016b; Pičmanová et al. 2015; Jenrich et al. 2007). The recycle pathway proceeds without the release of HCN and is catalysed by a glutathione transferase (GSTL1 or GSTL2) and a heteromer of the nitrilases, NIT4A and NIT4B2 (Bjarnholt et al. 2018). In a detoxification reaction catalysed by β-cyanoalanine synthase (CAS) and using cysteine as substrate, HCN released by the bio-activation reaction may be incorporated into β-cyanoalanine (Piotrowski and Volmer 2006) and further converted to asparagine, aspartate and ammonia by heteromers of nitrilases (NIT4) of the A and B types (Jenrich et al. 2007).
Seventeen wild sorghum species endemic to Australia (Ananda et al. 2020) are known and vary in their resistance to water-stress (Myrans et al. 2020). The effects of severe water-stress conditions on the growth, morphology, physiological and biological characteristics of wild sorghum species from different subgenera have recently been reported (Cowan et al. 2020). Whereas water-stress significantly increases the dhurrin levels in the above ground tissue (leaves and sheath) of S. bicolor, dhurrin levels remained unchanged in the wild relatives, except for a significant decrease in the sheath tissue in some wild species. Water-stress caused a significant growth reduction in S. bicolor whereas the wild species were more tolerant. The wild sorghum species, S. macrospermum E. D. Garber maintained a high relative growth rate and an insignificant aboveground dhurrin content under water-stress. The dhurrin levels in the leaves of S. macrospermum was 1000-fold lower than in the leaves of S. bicolor, while water-stress did not significantly increase the leaf dhurrin levels of S. macrospermum. Another recent study (Cowan et al. 2022) supports these findings showing that the leaf dhurrin content of wild sorghum species are significantly lower than those in S. bicolor. However, the dhurrin levels in the roots of wild sorghum species are similar to the levels in the domesticated S. bicolor (Cowan et al. 2021). The observed differences in dhurrin regulation between domesticated and wild species may mirror the differences in selective pressures encountered in natural and cultivated habitats (Bredeson et al. 2016). The maintained dhurrin content in the roots of wild sorghum may represent a recyclable reduced nitrogen store facilitating growth in the nutrient poor environments, characteristic of the native ranges of the wild species (Cowan et al. 2020; Myrans et al. 2020; Pičmanová et al. 2015; Dillon et al. 2007).
Crop wild relatives are expected to harbour valuable genetic traits that can be used in crop improvement (Ananda et al. 2020). A major constraint in utilising the genetic resources in wild crop relatives is hindrance in gene transfer between domesticated crops and their wild relatives (Bevan et al. 2017). However, the Australian endemic wild sorghum species, S. macrospermum is closely related to domesticated S. bicolor (Ananda et al. 2021), with successful introgression of the two species reported by Kuhlman et al. (2010). The regulation of the synthesis of dhurrin in wild sorghum species in comparison to domesticated S. bicolor has not been studied. This study provides new knowledge on the differences in gene expression profiles in wild and domesticated sorghum including dhurrin biosynthesis, bio-activation and recycling.
Materials and methods
Plant material, sample collection and processing
The domesticated species of sorghum S. bicolor (L.) Moench (AGG 314746) and one wild species S. macrospermum E. D. Garber (AGG 302367) were selected based on the results of Cowan et al. (2020). Seeds of each accession were obtained from the Australian Grains Genebank (AGG), Horsham, Victoria and germinated according to the reported optimised protocol (Cowan et al. 2020). The two genotypes were grown at the glasshouse complex at the University of Queensland (UQ), QLD (27.4975° S, 153.0137° E) using a complete randomised design. The experiment was conducted in July–October 2020. Seeds were planted in 4 L ANOVA pots in a UQ23 soil mix (250 L) (70% Composite Pine Bark 0–5 mm, 30% Coco Peat and Fertilizers and other augments/M3 (1 kg Yates Flowtrace, 1 kg iron sulphate heptahydrate, 0.4 kg superphosphate [Ca (H2PO4)2], 0.03 kg copper sulphate, 1 kg gypsum, 1 kg dolomite, 6 g Osmocte L−1)), with five replicates for each well-watered condition (control) and water-stressed (stress) conditions. At the latter conditions, only four replicates of S. macrospermum survived to the very end of the experiment. The glasshouse was maintained at 28 °C and 18 °C ± 1.5 °C day/night with an average photoperiod of 14 h and with an average light intensity of 1100 µmol quanta m−2 s−1. The growth room was equipped with four Heliospectra LX602-G growth lights (blue 450 nm, white 5700 K, red 660 nm). For S. bicolor, all plants were watered daily to 100% soil water capacity for the first 10 d. After 10 d, the well-watered (control) plants were maintained at 100% soil water capacity, whilst the water-stressed (stress) plants were maintained at a soil water capacity of 15% obtained after decreasing the water content gradually over 10 days period following Cowan et al. (2020). For S. macrospermum, the same method was followed except that the water-stressed period was initiated using 47 days old plants and continued only for 18 days due to the slower growth of the plants. Plants were watered daily according to the requirements. For the water-stressed S. macrospermum plants, the weight of each individual pot was measured daily as the weights varied quite a lot.
The soil water capacity was assessed using the following equation (Hasanuzzaman et al. 2017). Target soil water content WT:
Wp—weight of an empty pot, WD—dry soil weight, %RSWC relative soil water content (15% in this study), Ws—soil water content (wet soil pot weight-dry soil pot weight).
The weight of the plants/pots was measured daily, and plants were watered according to the following equation
Leaf samples were collected from each individual plant after 10 days of either well-watered or water-stressed growth (20 days old plants) and 47 days of water-stressed growth (57 days old plants) for S. bicolor and 10 days of either well-watered or water-stressed growth (57 days old plants) and 18 days water-stressed growth (65 days old plants) for S. macrospermum using destructive harvesting (Fig. S1b). The use of different harvest times for the two species was necessary due to their different growth rates. From each plant, the three youngest leaves which were fully expanded were collected. Each leaf was divided vertically along the mid rib into two equal parts and the left side was used for chemical analysis while the right side was used for RNA extraction. The material collected for chemical analysis was oven dried at 65 °C, then the three samples from each replicate were mixed and finely ground using a Qiagen Tissue Lyser II. The leaves collected for RNA extraction were immediately frozen in liquid nitrogen, briefly stored on dry ice, and then transferred to a − 80 °C freezer for long term storage. The leaves were pulverised using a Qiagen Tissue Lyser II under cryogenic conditions and stored at − 80 °C until RNA extraction.
RNA extraction and sequencing
Total RNA was extracted from the pulverised leaf tissues of 36 sorghum samples (two time points and two treatments, five biological replicates for S. bicolor and four biological replicates for S. macrospermum) using an optimised methodology based on the Trizol—Qiagen RNAeasy mini kit (Furtado 2014). RNA was extracted separately from each of the three leaves harvested from each plant. The quality and quantity of the RNA were assessed using spectrometry with A260/280 and A260/230 absorbance ratios (NanoDrop, Thermofisher Scientific, USA) as well as using a 2100 Agilent Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA) to monitor and secure 260/280 nm absorption ratios between 1.9 and 2.0 and RIN number above 6.5 for all RNA samples used for sequencing. All three RNA samples extracted from one replicate were combined affording a final RNA sample of 150–200 ng µL−1 that was sequenced at the Ramaciotti Centre, University of NSW, Australia using an Illumina NovaSeq 6000 platform.
Transcriptome analysis and determination of gene expression levels
RNA-Seq analysis was performed using CLC Genomic Workbench (CLC-GWB) software (CLC Genomics Workbench 11.0, http://www.clcbio.com). Raw reads were imported to CLC-GWB, and the sequence quality trimmed at 0.01 quality limits (Phred score equivalent to > 20, 95% of the reads had a Phred score greater than 35, averaged across all bases) at a sequence length of 1000 bp. The trimmed reads were mapped to the reference mRNA transcriptome of S. bicolor (GCF_000003195.3). Then a count data table was generated in CLC, and differential gene expression was analysed for different comparison groups (Table S1). Results were filtered based on the FDR p value (≤ 0.01) and Principal Component Analysis (PCA) was conducted in CLC. An expression browser was created in CLC and exported to Omics Box (https://www.biobam.com/omicsbox/) for functional annotation and finally the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways were generated for the up- and down-regulated genes. A Venn diagram was drawn for the differentially expressed genes using the online tool in Bioinformatics and Evolutionary Genomics (http://bioinformatics.psb.ugent.be/webtools/Venn/). The top 10 up- and down-regulated genes identified in each comparison group were filtered based on the fold change values and KEGG pathway analysis was undertaken in Omics Box. Using the MapMan software (v. 3.0.0) (Thimm et al. 2004) metabolism overview and regulation overview of different comparison groups were designed. Furthermore, the differential expression of genes in the cyanogenesis pathway in sorghum (Table 1) was studied.
To identify the source of the unmapped reads in RNA-Seq analysis, the unmapped reads were mapped against the ribosomal database SILVA (Quast et al. 2013) followed by the remaining unmapped reads mapping against the long noncoding RNA database (PNRD: Plant Non-coding RNAs database; cau.edu.cn). Then the remaining unmapped reads were subjected to de-novo assembly in CLC GWB and BLAST analysis and coding potential analysis was conducted with the resulting contigs in Omics Box.
Chemical analysis
Hydrogen cyanide potential (HCNp), nitrate and phenylpropanoids were determined using finely ground oven-dried leaf tissue. HCNp is the total amount of HCN produced per unit plant material following addition of excess amounts of β-glucosidase (β-d-glucoside glucohydrolase, G4511, Sigma-Aldrich, Sydney, Australia) to ensure complete hydrolysis of the total amount of dhurrin present. Evolved HCN was determined colorimetrically (O’Donnell et al. 2013) as modified by Cowan et al. (2020). Each milligram of HCN is equivalent to 11.5 mg of dhurrin in the plant tissue. The nitrate concentration was determined colorimetrically (O’Donnell et al. 2013). Phenylpropanoids were measured using 10–20 mg dried tissue following Myrans et al. (2021).
Statistical analysis
Results of the chemical analysis (HCNp, NO3− and phenolics) were analysed using SigmaPlot v.14.5 (Systat Software) by two-way ANOVA, with the species and each stress treatment as the variables. Average values ± the standard error was plotted for S. bicolor (n = 5 for each treatment) and S. macrospermum (n = 4 for each treatment). Data sets were tested for normality (Shapiro–Wilk) and equal variance (Brown–Forsythe). All data sets passed these tests, except the analysis of nitrate in S. macrospermum, where reciprocal transformation was carried out. For all tests, a p value of < 0.05 was considered significant. When an interaction was detected, post-hoc comparisons were undertaken using Tukey’s test.
Results
Imposition and assessment of water-stress
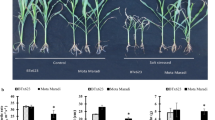
To monitor and adjust the level of imposed water-stress, the weight of randomly selected pots was measured daily, and the required amount of water added as described in the methods. According to the weight measurements, the water-stressed S. bicolor plants had a lower actual weight than the target weight for the day. The daily water usage of the S. macrospermum plants was higher than that of the S. bicolor plants (Fig. S1a). This is likely due to the higher number of tillers formed by the S. macrospermum plants. Compared to the S. bicolor plants, the growth of the S. macrospermum plants was slower. Accordingly, identical water-stress periods could not be maintained for the two species. In both species, the water-stress treatment was effective with the water-stressed plants being smaller than the well-watered plants and carrying severely wilted leaves (Fig. 1).
Transcriptome analysis and determination of gene expression levels
RNA-Seq followed by differential gene expression analysis was performed for S. bicolor and S. macrospermum to compare gene expression profiles at different time points and growth conditions within and between the two species (Table S1). In RNA-Seq, the average mapping percentage of the paired-end trimmed reads was 37% (Table S2). The unmapped reads were mapped against the ribosomal database SILVA (Quast et al. 2013) affording a mapping percentage of 15% of the total reads and against the long noncoding RNA database (PNRD: plant non-coding RNAs database; cau.edu.cn) affording an additional mapping percentage of 15% of the total reads. The remaining 33% of the total reads were mapped against the whole genome of S. bicolor. The total mapping percentage was therefore 99% (Table S3).
In the BLAST analysis of the contigs, 71% of the contigs had no BLAST hits whereas in the coding potential analysis only 50% of the contigs had complete Open Reading Frames (ORF) while the rest had 3′ or 5′ partial ORFs (Table S4).
Age and water-stress result in differential expression of distinct genes
Differential gene expression analysis was performed at the FDR p value ≤ 0.01 for each comparison group. Comparison of the transcript profiles of S. bicolor control at day 47 (47 days) and water-stressed at 47 days showed 4069 differentially expressed genes (DEGs). This is the highest number of DEGs observed between the analysed groups and demonstrate a highly complex response to long term water-stressed growth. The lowest number of DEGs was observed between the S. macrospermum control at day 10 and day 18 (18 days). The 76 DEGs identified indicate that an 8 day additional growth under well-watered conditions results in minute changes in the gene expression profile. The number of up- and down-regulated DEGs within and between S. bicolor and S. macrospermum independent of plant age and imposed water-stressed growth is shown on Fig. 2a.
Analysis of the Venn diagrams identifies 10 DEGs in S. bicolor that were up-regulated in all treatments and 2 DEGs that were down-regulated (Fig. 2b and Table S7). In S. macrospermum, a single DEG was down-regulated in all treatments whereas none of the DEGs were up-regulated in all treatments (Fig. 2b, Table S7). In the water-stressed S. bicolor plants at 10 days and 47 days (Fig. 2b), 8 and 1628 DEGs were up-regulated exclusively whereas 50 and 1892 DEGs were down-regulated (Fig. 2b), respectively. Similarly, 1316 and 434 DEGs were up-regulated in the water-stressed S. macrospermum plants at 10 days and 18 days (Fig. 2b), while 1,182 and 585 genes were exclusively down-regulated (Fig. 2b), respectively.
Principal component scatter plots of DEGs in S. bicolor and S. macrospermum show that the total variance present in the data set was 40.4% and 44.1%, respectively. In S. bicolor, PC1 contributed to 17.1%, PC2 contributed to 15.9% and the PC3 contributed to 7.4% of the total variance (Fig. 3a). The corresponding values in S. macrospermum were 22.1%, 12.6% and 9.4%. PCA was able to differentiate the samples based on plant age and water-stressed growth. PCA was not able to differentiate the 10 days control and water-stressed growth samples in S. bicolor and the 10 days and 18 days control samples in S. macrospermum (Fig. 3b).
KEGG pathway analysis of top 10 DEGs
The top 10 up- and down-regulated DEGs from each of the comparison groups were selected based on the fold change values of all the comparison groups and KEGG pathway analysis was used to identify the associated pathways. Up-regulated DEGs were associated with common pathways such as purine and thiamine metabolism pathways in the comparison groups of S. bicolor control 10 days vs 47 days, S. bicolor water-stressed growth 10 days vs 47 days, S. bicolor 47 days control vs water-stressed growth, and S. macrospermum control 10 days vs 18 days. Down-regulated DEGs were linked to purine and thiamine metabolism in the comparison groups of S. macrospermum control 10 days vs 18 days and S. macrospermum water-stressed growth 10 days vs 18 days. No KEGG defined pathways could be associated with the top 10 up- or down-regulated genes in S. bicolor control 47 days vs water-stressed growth 47 days (Table 2). The details of the top 10 up- and down-regulated DEGs are given in the supplementary information (Tables S5 and S6).
Among the top 10 up- and down-regulated genes from all eight comparison groups, 50% and 37.5% of the up- and down-regulated genes in S. bicolor, respectively, were functionally uncharacterized. In S. macrospermum, the corresponding numbers of genes with unknown functions were 27.5% and 30%. Among these top 10 DEGs, the only commonly expressed gene in both the species was XM_021447547.1, which encodes a late embryogenesis abundant (LEA) protein. Some of the individual comparison groups include DEGs encoding LEA proteins highly up-regulated upon water-stressed growth in S. bicolor (gene ID: XM_021447547.1 and XM_002444176.2) as well as in S. macrospermum (gene ID: XM_021448835.1), which have been linked with water-stress in other studies (Johnson et al. 2014). In addition, genes which encode lipid transfer proteins, ATP binding proteins, chromatin regulators, electron transfer proteins, cysteine type peptidases, ion binding proteins, transcription regulators and proteins involved in protein metabolism pathway are highly differentially expressed in all individual comparison groups (Tables S5 and S6). Supplementary Figs. S2–S9 provide a metabolism-based overview of the genes which are differentially expressed in S. bicolor and S. macrospermum independent of plant age and imposed water-stress. These results match the results of the PCA (Fig. 3) by showing few differences in gene expression between S. bicolor control 10 days and water-stressed growth 10 days (Fig. S4) and S. macrospermum control 10 days and 18 days (Fig. S6). Marked differences in the expression of genes related to photorespiration, C1-metabolism, lipid metabolism and mitochondrial electron transport were observed dependent on plant tissue age and imposed water-stressed growth (Figs. S2–S9).
Role of hormones in water-stress response
An overview of the effect of plant tissue age and water-stressed growth on the transcript levels of genes involved in plant hormone metabolism is presented in Fig. 4. Genes which encode enzymes catalysing biosynthesis of abscisic acid (ABA), benzyl adenine (BA), cytokinin, indole acetic acid (IAA), jasmonate, ethylene and gibberellic acid were highly expressed in water-stressed S. bicolor plants. In older plant tissues, the genes which produce IAA, jasmonate and ABA were down-regulated. In S. macrospermum, genes which regulate ABA, BA, cytokinin, jasmonate, and ethylene production were up-regulated in water-stressed plants in young as well as older tissues (Fig. 4), except for the jasmonate biosynthetic transcripts in the 10 days experiment which were down-regulated.
Differential expression of genes involved in cyanogenesis
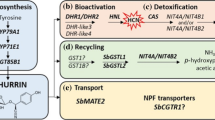
The expression values and the transcripts per million of the selected cyanogenesis related genes for the two species under all conditions are given in the supplementary table S8. The expression values of the cyanogenesis genes were comparatively much higher in the S. bicolor samples relative to the S. macrospermum samples (Table S8). The genes which are differentially expressed in the cyanogenesis pathway were determined by KEGG pathway analysis for each comparison group in both S. bicolor and S. macrospermum. In S. bicolor, between the control plants at 10 days and 47 days, the genes CYP79A1 (1.14.14.36), CYP71E1 (1.14.14.37), UGT85B1 (2.4.1.85), HNL (4.1.2.11), and cysteine synthase 2 (CAS C1) (4.4.1.9), were down-regulated whereas isoaspartyl peptidase/l-asparaginase 2 (3.5.1.1) was up-regulated. Compared to the water-stressed plants at 10 days, the genes CYP79A1 (1.14.14.36), CYP71E1 (1.14.14.37), and beta-glucosidase (3.2.1.21) were down-regulated while HNL (4.1.2.11) was up-regulated in the water-stressed grown plants at 47 days. In the comparison between the 10 days control and water-stressed plants, none of the genes related to dhurrin synthesis, recycling or bio-activation were differentially expressed. In water-stressed compared to control plants at 47 days, isoaspartyl peptidase/l-asparaginase 1/2 (3.5.1.1) and dhurrinase-1 (3.2.1.21) were down-regulated whereas UGT85B1 (2.4.1.85), HNL (4.1.2.11) were up-regulated. Furthermore, the dhurrin transporter SbMATE2 was down-regulated in the 47 days control plants compared to the 10 days control plants (Table S9, Fig. 5). In S. macrospermum, none of the genes in the dhurrin biosynthetic pathway were differentially expressed in the control plants at 18 days compared to the control plants at day 10. However, the gene NIT4A (3.5.5.4/1) was down-regulated whereas CYP71E1 (1.14.14.37) was up-regulated in the water-stressed 18 days plants relative to the water-stressed 10 days plants. Compared to the control 10 days plants, the genes cysteine synthase 2 (CAS C1) (4.4.1.9), and isoaspartyl peptidase/l-asparaginase 1/2 (3.5.1.1) were down-regulated while dhurrinase-1, dhurrinase-2 and dhurrinase-like 3 (3.2.1.21), and NIT4A (3.5.5.4/1) were up-regulated in the water-stressed plants at 10 days. In the final combination of the control 18 days plants relative to water-stressed 18 days plants, the isoaspartyl peptidase/l-asparaginase 1/2 (3.5.1.1) gene was down-regulated whereas dhurrinase-1, dhurrinase-2 and dhurrinase-like 3 (3.2.1.21) were up-regulated Moreover, the putative dhurrin transporter SbCGTR1 was down-regulated in the 10 days water-stressed growth compared to the 10 days control plants (Table S9, Fig. 5).
Chemical analysis
The average HCNp of leaf tissues of sorghum plants was measured under control and water-stressed growth conditions as a means to assess the total content of dhurrin. At all full-watered and water-stressed conditions tested, the dhurrin content of the S. bicolor leaves was around 1000-fold higher than that of the corresponding S. macrospermum leaf samples. In S. bicolor, the dhurrin level decreased with tissue age but showed significant increases under water-stressed growth conditions. The highest dhurrin levels were found in leaves of S. bicolor harvested from 47 days water-stressed plants. In S. macrospermum leaves, the dhurrin levels were minute and showed no significant changes with plant age and following water-stressed growth conditions (Fig. 6).
The average NO3 concentration was comparatively higher in S. bicolor tissues, with a significant difference between control and water-stressed plants at 47 days with lower NO3 concentrations in the well-watered plants (Fig. 6). There were no significant differences between the NO3 concentrations in the S. macrospermum samples.
The total phenylpropanoid content in S. bicolor leaves did not show significant changes with tissue age and water-stressed growth (Fig. 6). In S. macrospermum, the total content of phenylpropanoids showed a significant increase in the 10 days water-stressed leaf tissues but did not increase with extended growth under water-stressed conditions (Fig. 6).
Discussion
Plants respond to different biotic and abiotic stresses by altering their gene expression profiles as monitored by transcriptome analysis (Takahashi et al. 2018; Shankar et al. 2013). Water-stress increases the dhurrin content in leaves of domesticated sorghum (O'Donnell et al. 2013) but not in the leaves of wild sorghum (Cowan et al. 2020). Several studies of the effects of biotic and abiotic stresses have also been carried out on other cyanogenic species, including cassava (Brown et al. 2016), lima bean (Ballhorn et al. 2011) and eucalypt (Gleadow and Woodrow 2002a). However, there have been no studies investigating the differences in gene expression in wild sorghum species in response to water-stress. In the current study, growth of the domesticated sorghum species S. bicolor was shown to differ from that of the wild sorghum species S. macrospermum plants. Under the glasshouse conditions used, the S. macrospermum plants grew more slowly than the S. bicolor plants. As a result of producing more tillers, the transpiration rate in S. macrospermum plants was higher than that of S. bicolor, which led to a higher daily water usage. S. macrospermum plants were severely affected by the water-stressed growth conditions and showed more severe symptoms compared to the S. bicolor plants.
Differential gene expression analysis following imposed water-stressed growth conditions showed down-regulation of the expression of many genes suggesting that many of metabolic processes were slowed down or stopped due to the stress. Comparison of the differential gene expression between the control treatments of both species at the two time points show approximately the same number of up and down regulated genes within each species. These genes would be involved in general plant metabolism related to growth and development and the total number is much higher in S. bicolor than in S. macrospermum. The Venn diagrams illustrate the number of unique genes expressed in water-stressed plants of both species. Many of the genes were involved in functions such as protein binding and ion binding but further analysis is required to identify their specific roles related to water-stress. In the PCA, the variance between the control 10 days and water-stressed 10 days plants of S. bicolor and between control 10 days and 18 days plants of S. macrospermum were not able to differentiate them into separate groups. As reflected by the metabolism overview also, there is not much difference in gene expression levels in response to short-term water-stressed growth in younger S. bicolor plants. The high number of differentially expressed genes in the S. bicolor water-stressed plants at 47 days compared to the control plants indicate a complex response to long term water-limitation. In S. macrospermum control plants, the difference in gene expression profiles with age were minor.
Plant hormones control the growth of the plant under stress conditions (Miransari 2016; Hasanuzzaman et al. 2013). This study highlights the differential expression of genes which regulate stress related hormones. ABA is a plant hormone that improves resistance to drought stress (Ullah et al. 2018) by regulating stomatal activities, protein and lipid synthesis, root development and leaf senescence (Tuteja 2007). Drought and salinity stress are reported to increase ABA levels in plants (Hasanuzzaman et al. 2013) and thereby regulate the function of aquaporins (Li et al. 2014) and enhance root growth (Lynch and Brown 2008). Jasmonic acid, ethylene, IAA, gibberellins, and cytokinins play other important roles in drought stress tolerance (Ullah et al. 2018). Thus, IAA enhances drought tolerance by increasing ABA and jasmonic content and activating auxin responsive genes in white clover (Zhang et al. 2020). Further studies are required to understand the relationship of these genes with the age of sorghum plants as different responses were observed between the young and old tissues of the two species.
KEGG pathway analysis was performed on the top 10 up- and down-regulated genes in each comparison group in an attempt to identify metabolic pathways in which these highly differentially expressed genes were involved. Among these genes, only the gene XM_021447547.1, which encodes a LEA protein was commonly expressed in both species while the others were uniquely expressed. In addition, and in agreement with previous studies (Johnson et al. 2014), we report that genes encoding LEA proteins were among the top 10 up-regulated genes in stressed plants of both the species with very high fold change values. LEA proteins are a group of hydrophilic proteins which accumulate under water deficit conditions (Battaglia and Covarrubias 2013) and act e.g. by preventing inactivation of lactate dehydrogenase enzymes (Reyes et al. 2008), stabilizing membranes (Stacy and Aalen 1998), and supporting ion sequestration (Hara et al. 2005). Furthermore, lipid transfer proteins were up-regulated in the S. bicolor water-stressed plants (Johnson et al. 2014).
Cyanogenesis in forage sorghum can be problematic, especially in countries like Australia where sorghum is an important source of forage (Cowan et al. 2020). The HCNp is high in young plants, in heavily fertilised plants and in sorghum exposed to drought (O’Donnell et al. 2013). Efforts have been made to develop low-cyanide sorghum using mutagenesis (Blomstedt et al. 2012). Another option is to exploit the naturally low dhurrin content of other species of sorghum in the tertiary gene pool (Cowan et al. 2020). In the present study we demonstrate that, even under severe stress, S. bicolor leaves have around a 1000-fold higher HCNp than the wild sorghums. In agreement with previous studies, a high HCNp was found in young S. bicolor plants and that the HCNp increased with stress in older S. bicolor plants (Cowan et al. 2020; Blomstedt et al. 2018; Gleadow et al. 2016). The high concentration of dhurrin in stressed S. bicolor plants may be a direct stress response. S. macrospermum plants showed an opposite response to water-stressed growth by maintaining very low concentrations of dhurrin in their leaf tissue independent of tissue age or imposed water-stress, although the low dhurrin concentrations made exact comparisons difficult.
Nitrate concentrations in plants are dependent on the genotype, tissue type, and water availability (Rosati et al. 2019a). Our results show that the nitrate concentration in leaves was higher in domesticated sorghum than the wild sorghum plants. Nitrate concentration increased at water-stressed growth in both species but only significantly in S. bicolor. The rate of nitrate uptake from the soil and nitrate reductase enzyme activity are key determinants of the nitrate concentration in plants (Rosati et al. 2019a). Studies have shown that drought stress reduces nitrate reductase activity (Fresneau et al. 2007; Kaiser and Huber 2001).
Phenylpropanoids are a large class of natural products. They are known to provide protection against biotic and abiotic challenges (Myrans et al. 2021; Varela et al. 2016; Hura et al. 2008). Drought-stress increases the content of phenylpropanoids in Populus euphratica (Guignard et al. 2005), Populus nigra (Hale et al. 2005), and Amaranthus tricolor (Sarker and Oba 2018). In the current study, water-stressed S. macrospermum plants had a significantly higher content of phenylpropanoids indicating that they also play a role in stress tolerance in wild sorghums. The metabolic relationship between the formation of dhurrin and other phenylpropanoids requires further investigation.
In the comparative study of transcript levels of genes responsible for the biosynthesis of dhurrin and its recycling and bio-activation in domesticated and wild sorghum subjected to water-stressed growth (Fig. 5), most of the gene expression profiles obtained mirrored the analyses of dhurrin content. None of the genes involved in these processes were registered among the top 10 DEGs in the eight comparison groups (Table 1). CYP79A1 (1.14.14.36) is the first committed enzyme in dhurrin biosynthesis, and its transcript shows a 19.4- and 4.9-fold higher expression in the leaves of 10 days S. bicolor seedlings in comparison with leaves from 47 days old sorghum plants grown under well-watered and water-stressed conditions, respectively (Table S9). CYP71E1 (1.14.14.37) is the second enzyme in the dhurrin pathway and the corresponding fold reduction in its transcript is 3.9 and 1.4-fold, implying that dhurrin formation shows an age dependent decrease and is induced as a result of water-stress (Fig. 5) in agreement with previous data (Cowan et al. 2020; Busk and Møller 2002; Gleadow and Woodrow 2002b). Dhurinase2, the key gene in the bio-activation pathway was down-regulated in the older leaves as was CAS1 which is active in preventing auto-toxicity from released HCN (Table S9).
The dhurrin content of the wild sorghum S. macrospermum is minute and no significant increase was observed water-stressed growth conditions (Fig. 6). No significant expression was observed for the gene CYP79A1. Nevertheless, significant increased expression of the CYP71E1 gene involved in the biosynthesis of dhurrin and of several dhurrinase genes was observed under water-stressed conditions. The CAS C1 and NIT4A genes are involved in detoxification of HCN resulting from bio-activation of dhurrin as well as HCN release concomitant with the synthesis of ethylene (Jenrich et al. 2007). This may explain the observed shifting expression patterns. The increased expression of the transcripts encoding dhurrinase-1 and 2 and of dhurrinase-like 3 under water-stressed growth implies that these β-glycosidases may have other in vivo substrates than dhurrin.
Although, some studies have indicated that the wild sorghum species are more tolerant to drought (Cowan et al. 2020), we observed that the wild sorghum S. macrospermum is more susceptible to water-stressed growth and grows slower than domesticated S. bicolor. It might be that wild sorghum species were more adapted to natural harsh environments and show a different pattern of growth under artificial conditions.
Cyanogenesis is believed to come at a metabolic cost to the plant by utilizing the resources which could be used in plant growth and development (Herms and Mattson 1992). Although, cultivated sorghum plants use cyanogenesis as an herbivore defence mechanism, wild sorghum leaves might have roles other than defence. S. macrospermum is endemic to northern Australia, and the low nitrogen availability of the soil (Dillon et al. 2007) in their habitat, might result in the available nitrogen being utilized in general metabolic and growth process rather than cyanogenesis also, dhurrin may be instantly converted into free nitrogen (Cowan et al. 2022). Therefore, wild sorghum species have low dhurrin content in the leaves and the plants might rely on other defence mechanisms such as trichomes on the leaf surface (Johnson 1975).
Identification of potential candidate genes (transcription factors) that control the differential regulation of dhurrin biosynthesis in domesticated and wild sorghum is a vital step in developing more drought tolerant sorghum species. On the other hand, wild sorghum species such as S. macrospermum with a very low dhurrin content in their leaves can be “redomesticated” by introducing genes related to higher yield and biomass from the already domesticated sorghum species e.g., as shown in wild tomato (Lemmon et al. 2018; Zsögön et al. 2018). The close phylogenetic relationship of domesticated S. bicolor and the wild species S. macrospermum (Ananda et al. 2021) provides a great advantage to the introgression of beneficial traits of sorghum species.
Conclusions
In this study, we identified genes which are differentially expressed in response to water-stress in domesticated and wild sorghum species and specifically highlighted the different expression patterns of the genes in the dhurrin synthesis and bio-activation pathways in response to water-stressed growth conditions. Under all conditions tested, the dhurrin content was around 1000-fold higher in S. bicolor leaves as compared to S. macrospermum leaves. In S. bicolor, dhurrin concentration decreased with age and increased with stress while in S. macrospermum the levels remained minute. Beneficial traits of S. macrospermum such as its acyanogenic leaves may be introgressed into domesticated sorghum to provide safe fodder for livestock especially under conditions of drought.
Author contribution statement
RJH: conceptualization, methodology, supervision, project administration, funding acquisition, resources and writing—review and editing. RG: conceptualization, supervision, funding acquisition, writing—review and editing. BLM: conceptualization, funding acquisition, writing—review and editing. AF: methodology, formal analysis, supervision, data curation, writing—review and editing. SLN: methodology, resources and writing—review and editing. CB: methodology, data collection, formal analysis, resources and writing—review and editing. GKSA: formal analysis, data curation, investigation, writing—original draft.
Data availability statement
All data and materials used and described in this study are made available for non-commercial research purposes. The data that support the findings of this study are openly available in Sequence Read Archive (SRA) under the BioProject number PRJNA736757 (http://www.ncbi.nlm.nih.gov/bioproject/736757).
Abbreviations
- ABA:
-
Abscisic acid
- AGG:
-
Australian Grains Genebank
- AQP:
-
Aquaporins
- BA:
-
Benzyl adenine
- BGD:
-
β-Glucosidase
- CAS:
-
β-Cyanoalanine synthase
- CLC-GWB:
-
CLC Genomic Workbench
- CNglcs:
-
Cyanogenic glycosides
- d:
-
Day
- DEGs:
-
Differentially expressed genes
- DHN:
-
Dehydrin proteins
- DREB:
-
Drought-responsive element binding proteins
- HCN:
-
Hydrogen cyanide
- HCNp:
-
Hydrogen cyanide potential
- HNL:
-
α-Hydroxynitrilase
- IAA:
-
Indole acetic acid
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- LEA:
-
Late embryogenesis abundant protein
- NIT4:
-
Nitrilase 4
- ORF:
-
Open reading frames
- PCA:
-
Principal component analysis
- POR:
-
P450 oxidoreductase
- UGT:
-
UDP-glucosyltransferase
- UQ:
-
University of Queensland
References
Abdel-Ghany SE, Ullah F, Ben-Hur A, Reddy ASN (2020) Transcriptome analysis of drought-resistant and drought-sensitive sorghum (Sorghum bicolor) genotypes in response to PEG-induced drought stress. Int J Mol Sci. https://doi.org/10.3390/ijms21030772
Ahuja I, de Vos RCH, Bones AM, Hall RD (2010) Plant molecular stress responses face climate change. Trends Plant Sci 15(12):664–674. https://doi.org/10.1016/j.tplants.2010.08.002
Ananda GK, Myrans H, Norton SL, Gleadow R, Furtado A, Henry RJ (2020) Wild sorghum as a promising resource for crop improvement. Front Plant Sci 11:1108. https://doi.org/10.3389/fpls.2020.01108
Ananda GKS, Norton SL, Blomstedt C, Furtado A, Møller BL, Gleadow R, Henry RJ (2021) Phylogenetic relationships in the Sorghum genus based on sequencing of the chloroplast and nuclear genes. Plant Genome. https://doi.org/10.1002/tpg2.20123
Azzouz-Olden F, Hunt AG, Dinkins R (2020) Transcriptome analysis of drought-tolerant sorghum genotype SC56 in response to water-stress reveals an oxidative stress defense strategy. Mol Biol Rep 47(5):3291–3303. https://doi.org/10.1007/s11033-020-05396-5
Bak S, Kahn RA, Nielsen HL, Møller BL, Halkier BA (1998) Cloning of three A-type cytochromes P450, CYP71E1, CYP98, and CYP99 from Sorghum bicolor (L.) Moench by a PCR approach and identification by expression in Escherichia coli of CYP71E1 as a multifunctional cytochrome P450 in the biosynthesis of the cyanogenic glucoside dhurrin. Plant Mol Biol 36(3):393–405. https://doi.org/10.1023/A:1005915507497
Ballhorn DJ, Schiwy S, Jensen M, Heil M (2008) Quantitative variability of direct chemical defense in primary and secondary leaves of lima bean (Phaseolus lunatus) and consequences for a natural herbivore. J Chem Ecol 34(10):1298–1301. https://doi.org/10.1007/s10886-008-9540-1
Ballhorn DJ, Kautz S, Jensen M, Schmitt I, Heil M, Hegeman AD (2011) Genetic and environmental interactions determine plant defences against herbivores. J Ecol 99(1):313–326. https://doi.org/10.1111/j.1365-2745.2010.01747.x
Battaglia M, Covarrubias A (2013) Late embryogenesis abundant (LEA) proteins in legumes. Front Plant Sci. https://doi.org/10.3389/fpls.2013.00190
Bevan MW, Uauy C, Wulff BBH, Zhou J, Krasileva K, Clark MD (2017) Genomic innovation for crop improvement. Nature 543(7645):346–354. https://doi.org/10.1038/nature22011
Bjarnholt N, Neilson EHJ, Crocoll C, Jørgensen K, Motawia MS, Olsen CE, Dixon DP, Edwards R, Møller BL (2018) Glutathione transferases catalyze recycling of auto-toxic cyanogenic glucosides in sorghum. Plant J 94(6):1109–1125. https://doi.org/10.1111/tpj.13923
Blomstedt CK, Gleadow RM, O’Donnell N, Naur P, Jensen K, Laursen T, Olsen CE, Stuart P, Hamill JD, Møller BL, Neale AD (2012) A combined biochemical screen and TILLING approach identifies mutations in Sorghum bicolor L. Moench resulting in acyanogenic forage production. Plant Biotechnol J 10(1):54–66. https://doi.org/10.1111/j.1467-7652.2011.00646.x
Blomstedt CK, Rosati VC, Møller BL, Gleadow R (2018) Counting the costs: nitrogen partitioning in Sorghum mutants. Funct Plant Biol 45(7):705–718. https://doi.org/10.1071/FP17227
Borrell A, Jordan D, Mullet H, Hammer G (2006) Drought adaptation in Sorghum. In: Ribaut J-M (ed) Drought adaptation in cereals. The Haworth Press, New York, pp 335–399
Borrell AK, Mullet JE, George-Jaeggli B, van Oosterom EJ, Hammer GL, Klein PE, Jordan DR (2014) Drought adaptation of stay-green sorghum is associated with canopy development, leaf anatomy, root growth, and water uptake. J Exp Bot 65(21):6251–6263. https://doi.org/10.1093/jxb/eru232
Bredeson JV, Lyons JB, Prochnik SE, Wu GA, Ha CM, Edsinger-Gonzales E, Grimwood J, Schmutz J, Rabbi IY, Egesi C (2016) Sequencing wild and cultivated cassava and related species reveals extensive interspecific hybridization and genetic diversity. Nat Biotechnol 34(5):562. https://doi.org/10.1038/nbt.3535
Brown AL, Cavagnaro TR, Gleadow R, Miller RE (2016) Interactive effects of temperature and drought on cassava growth and toxicity: implications for food security? Glob Change Biol 22(10):3461–3473. https://doi.org/10.1111/gcb.13380
Busk PK, Møller BL (2002) Dhurrin synthesis in sorghum is regulated at the transcriptional level and induced by nitrogen fertilization in older plants. Plant Physiol 129(3):1222–1231. https://doi.org/10.1104/pp.000687
Calviño M, Bruggmann R, Messing J (2011) Characterization of the small RNA component of the transcriptome from grain and sweet sorghum stems. BMC Genom. https://doi.org/10.3929/ethz-b-000046889
Cowan MF, Blomstedt CK, Norton SL, Henry RJ, Møller BL, Gleadow R (2020) Crop wild relatives as a genetic resource for generating low-cyanide, drought-tolerant Sorghum. Environ Exp Bot 169:103884. https://doi.org/10.1016/j.envexpbot.2019.103884
Cowan M, Blomstedt CK, Møller BL, Henry R, Gleadow RM (2021) Variation in production of cyanogenic glucosides during early plant development: a comparison of wild and domesticated Sorghum. Phytochemistry 184:112645. https://doi.org/10.1016/j.phytochem.2020.112645
Cowan M, Møller BL, Norton S, Knudsen C, Crocoll C, Furtado A, Henry R, Blomstedt C, Gleadow RM (2022) Cyanogenesis in the Sorghum genus: from genotype to phenotype. Genes 13(1):140
Dahler JM, Mcconchie C, Turnbull CGN (1995) Quantification of cyanogenic glycosides in seedlings of three Macadamia (Proteaceae) species. Aust J Bot 43(6):619–628. https://doi.org/10.1071/BT9950619
Dillon SL, Shapter FM, Henry RJ, Cordeiro G, Izquierdo L, Lee LS (2007) Domestication to crop improvement: genetic resources for Sorghum and Saccharum (Andropogoneae). Ann Bot 100(5):975–989. https://doi.org/10.1093/aob/mcm192
Dugas DV, Monaco MK, Olsen A, Klein RR, Kumari S, Ware D, Klein PE (2011) Functional annotation of the transcriptome of Sorghum bicolor in response to osmotic stress and abscisic acid. BMC Genom 12:514. https://doi.org/10.1186/1471-2164-12-514
Farooq M, Wahid A, Kobayashi N, Fujita D, Basra SMA (2009) Plant drought stress: effects, mechanisms and management. In: Lichtfouse E, Navarrete M, Debaeke P, Véronique S, Alberola C (eds) Sustainable agriculture. Springer, Dordrecht. https://doi.org/10.1007/978-90-481-2666-8_12
Fetter K, Van Wilder V, Moshelion M, Chaumont F (2004) Interactions between plasma membrane aquaporins modulate their water channel activity. Plant Cell 16(1):215. https://doi.org/10.1105/tpc.017194
Finnie JW, Windsor PA, Kessell AE (2011) Neurological diseases of ruminant livestock in Australia. II: toxic disorders and nutritional deficiencies. Aust Vet J 89(7):247–253. https://doi.org/10.1111/j.1751-0813.2011.00793.x
Fresneau C, Ghashghaie J, Cornic G (2007) Drought effect on nitrate reductase and sucrose-phosphate synthase activities in wheat (Triticum durum L.): role of leaf internal CO2. J Exp Bot 58(11):2983–2992. https://doi.org/10.1093/jxb/erm150
Furtado A (2014) DNA extraction from vegetative tissue for next-generation sequencing. In: Henry RJ, Furtado A (eds) Cereal genomics: methods and protocols, methods in molecular biology. Springer, New York, pp 1–5. https://doi.org/10.1007/978-1-62703-715-0_1
Gleadow RM, Moller BL (2014) Cyanogenic glycosides: synthesis, physiology, and phenotypic plasticity. Annu Rev Plant Biol 65(1):155–185. https://doi.org/10.1146/annurev-arplant-050213-040027
Gleadow RM, Woodrow IE (2002a) Defense chemistry of cyanogenic Eucalyptus cladocalyx seedlings is affected by water supply. Tree Physiol 22(13):939–945. https://doi.org/10.1093/treephys/22.13.939
Gleadow RM, Woodrow IE (2002b) Mini-review: constraints on effectiveness of cyanogenic glycosides in herbivore defense. J Chem Ecol 28(7):1301–1313. https://doi.org/10.1023/A:1016298100201
Gleadow RM, Ottman MJ, Kimball BA (2016) Drought–induced changes in nitrogen partitioning in sorghum are not moderated by elevated CO2 in FACE studies. Field Crops Res 185:97–102
Gosal SS, Wani SH, Kang MS (2009) Biotechnology and drought tolerance. J Crop Improv 23(1):19–54. https://doi.org/10.1080/15427520802418251
Guignard C, Jouve L, Bogéat-Triboulot MB, Dreyer E, Hausman JF, Hoffmann L (2005) Analysis of carbohydrates in plants by high-performance anion-exchange chromatography coupled with electrospray mass spectrometry. J Chromatogr A 1085(1):137–142. https://doi.org/10.1016/j.chroma.2005.05.068
Hale BK, Herms DA, Hansen RC, Clausen TP, Arnold D (2005) Effects of drought stress and nutrient availability on dry matter allocation, phenolic glycosides, and rapid induced resistance of poplar to two lymantriid defoliators. J Chem Ecol 31:2601–2620. https://doi.org/10.1007/s10886-005-7616-8
Halkier BA, Møller BL (1991) Involvement of cytochrome p-450 in the biosynthesis of dhurrin in Sorghum bicolor (L.) Moench. Plant Physiol 96(1):10. https://doi.org/10.1104/pp.96.1.10
Hara M, Fujinaga M, Kuboi T (2005) Metal binding by citrus dehydrin with histidine-rich domains. J Exp Bot 56(420):2695–2703. https://doi.org/10.1093/jxb/eri262
Hasanuzzaman MD, Shabala L, Brodribb TJ, Zhou M, Shabala S (2017) Assessing the suitability of various screening methods as a proxy for drought tolerance in barley. Funct Plant Biol 44(2):253–266. https://doi.org/10.1071/FP16263
Hasanuzzaman M, Nahar K, Fujita M (2013) Plant response to salt stress and role of exogenous protectants to mitigate salt-induced damages. In: Ahmad P, Azooz M, Prasad M (eds) Ecophysiology and responses of plants under salt stress. Springer, New York, NY, pp 25–87. https://doi.org/10.1007/978-1-4614-4747-4_2
Herms DA, Mattson WJ (1992) The dilemma of plants—to grow or defend. Q Rev Biol 67:283–335. https://doi.org/10.1086/417659
Hura T, Hura K, Grzesiak S (2008) Contents of total phenolics and ferulic acid, and PAL activity during water potential changes in leaves of maize single-cross hybrids of different drought tolerance. J Agron Crop Sci 194(2):104–112. https://doi.org/10.1111/j.1439-037X.2008.00297.x
Jenrich R, Trompetter I, Bak S, Olsen CE, Møller BL, Piotrowski M (2007) Evolution of heteromeric nitrilase complexes in Poaceae with new functions in nitrile metabolism. Proc Natl Acad Sci USA 104(47):18848–18853. https://doi.org/10.1073/pnas.0709315104
Jensen K, Osmani SA, Hamann T, Naur P, Møller BL (2011) Homology modeling of the three membrane proteins of the dhurrin metabolon: catalytic sites, membrane surface association and protein–protein interactions. Phytochemistry 72(17):2113–2123. https://doi.org/10.1016/j.phytochem.2011.05.001
Jensen SB, Thodberg S, Parween S, Moses ME, Hansen CC, Thomsen J, Sletfjerding MB, Knudsen C, Del Giudice R, Lund PM, Castaño PR, Bustamante YG, Velazquez MNR, Jørgensen FS, Pandey AV, Laursen T, Møller BL, Hatzakis NS (2021) Biased cytochrome P450-mediated metabolism via small-molecule ligands binding P450 oxidoreductase. Nat Commun 12(1):2260. https://doi.org/10.1038/s41467-021-22562-w
Johnson HB (1975) Plant pubescence: an ecological perspective. Bot Rev 41:233–258. https://doi.org/10.1007/BF02860838
Johnson SM, Lim F-L, Finkler A, Fromm H, Slabas AR, Knight MR (2014) Transcriptomic analysis of Sorghum bicolor responding to combined heat and drought stress. BMC Genom 15(1):456. https://doi.org/10.1186/1471-2164-15-456
Jones DA (1998) Why are so many food plants cyanogenic? Phytochemistry 47(2):155–162. https://doi.org/10.1016/S0031-9422(97)00425-1
Jones PR, Møller BL, Høj PB (1999) The UDP-glucose: p-hydroxymandelonitrile-O-glucosyltransferase that catalyzes the last step in synthesis of the cyanogenic glucoside dhurrin in Sorghum bicolor isolation, cloning, heterologous expression, and substrate specificity. J Biol Chem 274(50):35483–35491. https://doi.org/10.1074/jbc.274.50.35483
Jorgensen K, Bak S, Busk PK, Sorensen C, Olsen CE, Puonti-Kaerlas J, Moller BL (2005) Cassava plants with a depleted cyanogenic glucoside content in leaves and tubers. Distribution of cyanogenic glucosides, their site of synthesis and transport, and blockage of the biosynthesis by RNA interference technology. Plant Physiol 139(1):363. https://doi.org/10.1104/pp.105.065904
Kahn RA, Bak S, Svendsen I, Halkier BA, Moller BL (1997) Isolation and reconstitution of cytochrome P450ox and in vitro reconstitution of the entire biosynthetic pathway of the cyanogenic glucoside dhurrin from sorghum. Plant Physiol 115(4):1661–1670. https://doi.org/10.1104/pp.115.4.1661
Kaiser WM, Huber SC (2001) Post-translational regulation of nitrate reductase: mechanism, physiological relevance and environmental triggers. J Exp Bot 52(363):1981–1989. https://doi.org/10.1093/jexbot/52.363.1981
Knoch E, Motawia MS, Olsen CE, Møller BL, Lyngkjær MF (2016) Biosynthesis of the leucine derived alpha-, beta- and gamma-hydroxynitrile glucosides in barley (Hordeum vulgare L.). Plant J 88:247–256. https://doi.org/10.1111/tpj.13247
Kojima M, Poulton JE, Thayer SS, Conn EE (1979) Tissue distributions of dhurrin and of enzymes involved in its metabolism in leaves of Sorghum bicolor. Plant Physiol 63(6):1022. https://doi.org/10.1104/pp.63.6.1022
Kuhlman LC, Burson BL, Stelly DM, Klein PE, Klein RR, Price H, Rooney WL (2010) Early-generation germplasm introgression from Sorghum macrospermum into sorghum (S. bicolor). Genome 53(6):419–429. https://doi.org/10.1139/g10-027
Laursen T, Møller BL, Bassard J (2015) Plasticity of specialized metabolism as mediated by dynamic metabolons. Trends Plant Sci 20(1):20–32. https://doi.org/10.1016/j.tplants.2014.11.002
Lemmon ZH, Reem NT, Dalrymple J, Soyk S, Swartwood KE, Rodriguez-Leal D, Van Eck J, Lippman ZB (2018) Rapid improvement of domestication traits in an orphan crop by genome editing. Nat Plants 4(10):766–770. https://doi.org/10.1038/s41477-018-0259-x
Li G, Santoni V, Maurel C (2014) Plant aquaporins: roles in plant physiology. Biochim Biophys Acta (BBA) Gen Subj 1840(5):1574–1582. https://doi.org/10.1016/j.bbagen.2013.11.004
Loyd RC, Gray E (1970) Amount and distribution of hydrocyanic acid potential during the life cycle of plants of three sorghum cultivars. Agron J 62(3):394–397. https://doi.org/10.2134/agronj1970.00021962006200030025x
Luo H, Mocoeur A, Jing HC (2014) Next-generation sequencing technology for genetics and genomics of Sorghum. In: Wang Y, Upadhyaya HD, Kole C (eds) Genetics, genomics and breeding of Sorghum. CRC Press, London, pp 226–250. https://doi.org/10.1201/b17153
Lynch JP, Brown KM (2008) Root strategies for phosphorus acquisition. In: White PJ, Hammond JP (eds) The ecophysiology of plant–phosphorus interactions. Springer, pp 83–116. https://doi.org/10.1007/978-1-4020-8435-5_5
McKey D, Cavagnaro TR, Cliff J, Gleadow R (2010) Chemical ecology in coupled human and natural systems: people, manioc, multitrophic interactions and global change. Chemoecology 20(2):109–133. https://doi.org/10.1007/s00049-010-0047-1
Miransari M (2016) Soybeans and plant hormones. In: Miransari M (ed) Environmental stresses in soybean production. Academic Press, San Diego, pp 131–156. https://doi.org/10.1016/B978-0-12-801535-3.00006-1
Mizuno H, Kawahigashi H, Kawahara Y, Kanamori H, Ogata J, Minami H, Itoh T, Matsumoto T (2012) Global transcriptome analysis reveals distinct expression among duplicated genes during sorghum-interaction. BMC Plant Biol. https://doi.org/10.1186/1471-2229-12-121
Møller BL (2010) Functional diversifications of cyanogenic glucosides. Curr Opin Plant Biol 13(3):337–346. https://doi.org/10.1016/j.pbi.2010.01.009
Møller BL, Conn EE (1979) The biosynthesis of cyanogenic glucosides in higher plants. N-Hydroxytyrosine as an intermediate in the biosynthesis of dhurrin by Sorghum bicolor (Linn) Moench. J Biol Chem 254(17):8575–8583. https://doi.org/10.1016/S0021-9258(19)86931-4
Myrans H, Diaz MV, Khoury CK, Carver D, Henry RJ, Gleadow R (2020) Modelled distributions and conservation priorities of wild sorghums (Sorghum Moench). Divers Distrib 26:1727–1740. https://doi.org/10.1111/ddi.13166
Myrans H, Vandegeer R, Henry R, Gleadow RM (2021) Nitrogen availability and allocation in sorghum and its wild relatives: divergent roles for cyanogenic glucosides. J Plant Physiol 258–259:e153393. https://doi.org/10.1016/j.jplph.2021.153393
Nielsen KA, Tattersall DB, Jones PR, Møller BL (2008) Metabolon formation in dhurrin biosynthesis. Phytochemistry 69(1):88–98. https://doi.org/10.1016/j.phytochem.2007.06.033
Neilson EH, Edwards AM, Blomstedt CK, Berger B, Møller BL, Gleadow RM (2015) Utilization of a high-throughput shoot imaging system to examine the dynamic phenotypic responses of a C 4 cereal crop plant to nitrogen and water deficiency over time. J Exp Bot 66(7):1817–1832. https://doi.org/10.1093/jxb/eru526
Nielsen LJ, Bjarnholt N, Blomstedt C, Gleadow RM, Møller BL (2016a) Biosynthesis, regulation, and significance of cyanogenic glucosides. In: Arimura G, Maffei M (eds) Plant specialized metabolism. CRC Press, pp 143–168. https://doi.org/10.1201/9781315370453-8
Nielsen LJ, Stuart P, Pičmanová M, Rasmussen S, Olsen CE, Harholt J, Møller BL, Bjarnholt N (2016b) Dhurrin metabolism in the developing grain of Sorghum bicolor (L.) Moench investigated by metabolite profiling and novel clustering analyses of time-resolved transcriptomic data. BMC Genom 17(1):1021. https://doi.org/10.1186/s12864-016-3360-4
O’Donnell NH, Møller BL, Neale AD, Hamill JD, Blomstedt CK, Gleadow RM (2013) Effects of PEG-induced osmotic stress on growth and dhurrin levels of forage sorghum. Plant Physiol Biochem 73:83–92. https://doi.org/10.1016/j.plaphy.2013.09.001
Pičmanová M, Neilson EH, Motawia MS, Olsen CE, Agerbirk N, Gray CJ, Flitsch S, Meier S, Silvestro D, Jørgensen K, Sanchez-Perez R, Møller BL, Bjarnholt N (2015) A recycling pathway for cyanogenic glycosides evidenced by the comparative metabolic profiling in three cyanogenic plant species. Biochem J 469(3):375–389. https://doi.org/10.1042/BJ20150390
Piotrowski M, Volmer JJ (2006) Cyanide metabolism in higher plants: cyanoalanine hydratase is a NIT4 homolog. Plant Mol Biol 61(1–2):111–122. https://doi.org/10.1007/s11103-005-6217-9
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41(Database issue):D590–D596. https://doi.org/10.1093/nar/gks1219
Reyes JL, Campos F, Wei H, Arora R, Yang Y, Karlson DT, Covarrubias AA (2008) Functional dissection of hydrophilins during in vitro freeze protection. Plant Cell Environ 31(12):1781–1790. https://doi.org/10.1111/j.1365-3040.2008.01879.x
Rosati V, Quinn A, Fromhold S, Gleadow R, Blomstedt C (2019a) Investigation into the role of DNA methylation in cyanogenesis in sorghum (Sorghum bicolor L. Moench). Int J Plant Growth Dev 88(1):73–85. https://doi.org/10.1007/s10725-019-00489-z
Rosati VC, Blomstedt CK, Møller BL, Garnett T, Gleadow R (2019b) The interplay between drought stress, dhurrin and nitrate in the low-cyanogenic sorghum mutant adult cyanide deficient class 1. Front Plant Sci 10:1458. https://doi.org/10.3389/fpls.2019.01458
Sánchez-Pérez R, Pavan S, Mazzeo R, Moldovan C, Aiese Cigliano R, Del Cueto J, Ricciardi F, Lotti C, Ricciardi L, Dicenta F, López-Marqués RL, Møller BL (2019) Mutation of a bHLH transcription factor allowed almond domestication. Science (new York, NY) 364(6445):1095–1098. https://doi.org/10.1126/science.aav8197
Sarker U, Oba S (2018) Drought stress enhances nutritional and bioactive compounds, phenolic acids and antioxidant capacity of Amaranthus leafy vegetable. BMC Plant Biol 18(1):258. https://doi.org/10.1186/s12870-018-1484-1
Shankar A, Singh A, Kanwar P, Srivastava AK, Pandey A, Suprasanna P, Kapoor S, Pandey GK (2013) Gene expression analysis of rice seedling under potassium deprivation reveals major changes in metabolism and signaling components. PLoS One 8(7):e70321. https://doi.org/10.1371/journal.pone.0070321
Sibbesen O, Koch B, Halkier BA, Møller BL (1994) Isolation of the heme-thiolate enzyme cytochrome P-450TYR, which catalyzes the committed step in the biosynthesis of the cyanogenic glucoside dhurrin in Sorghum bicolor (L.) Moench. Proc Natl Acad Sci 91(21):9740–9744. https://doi.org/10.1073/pnas.91.21.9740
Sibbesen O, Koch B, Halkier BA, Møller BL (1995) Cytochrome P-450TYR is a multifunctional heme-thiolate enzyme catalyzing the conversion of l-tyrosine to p-hydroxyphenylacetaldehyde oxime in the biosynthesis of the cyanogenic glucoside dhurrin in Sorghum bicolor (L.) Moench. J Biol Chem 270(8):3506–3511. https://doi.org/10.1074/jbc.270.8.3506
Stacy RAP, Aalen RB (1998) Identification of sequence homology between the internal hydrophilic repeated motifs of Group 1 late-embryogenesis-abundant proteins in plants and hydrophilic repeats of the general stress protein GsiB of Bacillus subtilis. Planta 206(3):476–478. https://doi.org/10.1007/s004250050424
Takahashi F, Kuromori T, Sato H, Shinozaki K (2018) Regulatory gene networks in drought stress responses and resistance in plants. Adv Exp Med Biol 1081:189–214. https://doi.org/10.1007/978-981-13-1244-1_11
Thimm O, Blaesing O, Gibon Y, Nagel A, Meyer S, Krüger P, Selbig J, Müller LA, Rhee SY, Stitt M (2004) MAPMAN: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J 37(6):914–939
Tuteja N (2007) Abscisic acid and abiotic stress signaling. Plant Signal Behav 2(3):135–138. https://doi.org/10.4161/psb.2.3.4156
Ullah A, Manghwar H, Shaban M, Khan AH, Akbar A, Ali U, Ali E, Fahad S (2018) Phytohormones enhanced drought tolerance in plants: a coping strategy. Environ Sci Pollut Res Int 25(33):33103–33118. https://doi.org/10.1007/s11356-018-3364-5
Varela MC, Arslan I, Reginato MA, Cenzano AM, Luna MV (2016) Phenolic compounds as indicators of drought resistance in shrubs from Patagonian shrublands (Argentina). Plant Physiol Biochem 104:81–91. https://doi.org/10.1016/j.plaphy.2016.03.014
Zagrobelny M, Bak S, Rasmussen AV, Jørgensen B, Naumann CM, Lindberg Møller B (2004) Cyanogenic glucosides and plant–insect interactions. Phytochemistry 65(3):293–306. https://doi.org/10.1016/j.phytochem.2003.10.016
Zhang Y, Li Y, Hassan MJ, Li Z, Peng Y (2020) Indole-3-acetic acid improves drought tolerance of white clover via activating auxin, abscisic acid and jasmonic acid related genes and inhibiting senescence genes. BMC Plant Biol 20(1):150. https://doi.org/10.1186/s12870-020-02354-y
Zsögön A, Čermák T, Naves ER, Notini MM, Edel KH, Weinl S, Freschi L, Voytas DF, Kudla J, Peres LEP (2018) De novo domestication of wild tomato using genome editing. Nat Biotechnol. https://doi.org/10.1038/nbt.4272
Acknowledgements
The authors acknowledge the University of Queensland Research Computing Centre (UQ-RCC) for providing all the computing resources.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions. This research is funded by a Grant of Australian research Council Discovery Project. Grant ID DP180101011.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by Anastasios Melis.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ananda, G.K.S., Norton, S.L., Blomstedt, C. et al. Transcript profiles of wild and domesticated sorghum under water-stressed conditions and the differential impact on dhurrin metabolism. Planta 255, 51 (2022). https://doi.org/10.1007/s00425-022-03831-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-022-03831-4