Abstract
Main conclusion
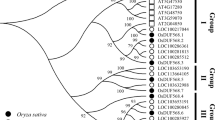
Exhaustive searches of the rice genome have revealed 30 different potential OsDof (Oryza sativa DNA binding with One Finger) genes. Their subcellular localization, phylogenetic relationship, conserved motifs identification, chromosomal allocation, expression patterns, and interaction networks were analyzed.
Abstract
The Dof (DNA binding with One Finger) family of transcription factors represents a particular class of plant-specific transcriptional regulators, contain a highly conserved region of 50–52 amino acids (Dof domain) and involved in various plant developmental processes and response to various environmental stresses. Few (Oryza sativa) OsDof genes have been demonstrated previously for their biological functions but there is no comprehensive study on most of the Dof genes of rice. In the current study, exhaustive searches of the rice genome revealed 30 different potential OsDof genes, and then their subcellular localization, phylogenetic relationship, conserved motifs identification, chromosomal allocation, expression patterns, and interaction networks were analyzed. Phylogenetic analysis of Dof proteins in rice showed that they are distributed in 4 groups. By genome-wide observation of gene expression profiles, we found that OsDof genes showed significant variances in expression levels in different tissues across multiple developmental stages. Protein–protein correlation network analysis, shows a statically significant overlap of some OsDofs, suggesting their similar functions and a high degree of co-expression. The Dof family transcription factors have been reported for their involvement in the regulation of various gene expression processes in rice but still, most of the Dof genes are not characterized for their specific physiological functions. This study revealed useful information and clues about predicting the potential roles of OsDofs in rice by combining their genome-wide characterization, expression profiling, protein–protein interactions, and for further studies to develop high-quality rice varieties.







Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410
Azam SM, Liu Y, Rahman ZU, Ali H, Yan C, Wang L, Priyadarshani S, Hu B, Huang X, Xiong J (2018) Identification, characterization and expression profiles of Dof transcription factors in pineapple (Ananas comosus L). Trop Plant Biol 11(1–2):49–64
Cai X, Zhang Y, Zhang C, Zhang T, Hu T, Ye J, Zhang J, Wang T, Li H, Ye Z (2013) Genome-wide analysis of plant-specific Dof transcription factor family in tomato. J Integr Plant Biol 55(6):552–566
Chen Y, Cao J (2015) Comparative analysis of Dof transcription factor family in maize. Plant Mol Biol Rep 33(5):1245–1258
Chen F, Chen Y, Dong Y, Li X, Xu M, Zhang C, Yan Y, Zhang G (2005) OsDof28: A new member of the DOF transcription factor family from rice. Tsinghua Sci Technol 10(4):454–460
Ferreira LM, Rocha JGd, Santos LA, Souza SRd, Fernandes MS (2015) Absorption kinetics and nitrogen fractions in rice as an expression of the OsDof26 transcription factor. Rev Cienc Agron 46(2):379–387
Gupta S, Kushwaha H, Singh VK, Bisht NC, Sarangi BK, Yadav D (2014) Genome wide in silico characterization of Dof transcription factor gene family of sugarcane and its comparative phylogenetic analysis with Arabidopsis, rice and sorghum. Sugar Tech 16(4):372–384
Kang W-H, Kim S, Lee H-A, Choi D, Yeom S-I (2016) Genome-wide analysis of Dof transcription factors reveals functional characteristics during development and response to biotic stresses in pepper. Sci Rep 6:33332
Krebs J, Mueller-Roeber B, Ruzicic S (2010) A novel bipartite nuclear localization signal with an atypically long linker in DOF transcription factors. J Plant Physiol 167(7):583–586
Lijavetzky D, Carbonero P, Vicente-Carbajosa J (2003) Genome-wide comparative phylogenetic analysis of the rice and Arabidopsis Dof gene families. BMC Evol Biol 3(1):17
Liu M, Ma Z, Sun W, Huang L, Wu Q, Tang Z, Bu T, Li C, Chen H (2019) Genome-wide analysis of the NAC transcription factor family in Tartary buckwheat (Fagopyrum tataricum). BMC Genomics 20(1):113
Ma J, Li M-Y, Wang F, Tang J, Xiong A-S (2015) Genome-wide analysis of Dof family transcription factors and their responses to abiotic stresses in Chinese cabbage. BMC Genomics 16(1):1–15
Noguero M, Atif RM, Ochatt S, Thompson RD (2013) The role of the DNA-binding One Zinc Finger (DOF) transcription factor family in plants. Plant Sci 209:32–45
Park SH, Lim H, Hyun SJ, Yun D-W, Yoon U-H, Ji H, Kim T-H, Eun M-Y, Kim Y-H, Lee G-S (2014) Wound-inducible expression of the OsDof1 gene promoter in a Ds insertion mutant and transgenic plants. Plant Biotechnol Rep 8(4):305–313
Pruitt KD, Tatusova T, Maglott DR (2007) NCBI reference sequences (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res. 35(suppl_1):61-65.
Qin H, Wang J, Chen X, Wang F, Peng P, Zhou Y, Miao Y, Zhang Y, Gao Y, Qi Y (2019) Rice Os DOF 15 contributes to ethylene-inhibited primary root elongation under salt stress. New Phytol 223(2):798–813
Riechmann JL, Heard J, Martin G, Reuber L, Jiang C-Z, Keddie J, Adam L, Pineda O, Ratcliffe O, Samaha R (2000) Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 290(5499):2105–2110
Rojas-Gracia P, Roque E, Medina M, López-Martín MJ, Cañas LA, Beltrán JP, Gómez-Mena C (2019) The DOF transcription factor SlDOF10 regulates vascular tissue formation during ovary development in tomato. Front Plant Sci 10:216
Sani HM, Hamzeh-Mivehroud M, Silva AP, Walshe JL, Mohammadi SA, Rahbar-Shahrouziasl M, Abbasi M, Jamshidi O, Low JK, Dastmalchi S (2018) Expression, purification and DNA-binding properties of zinc finger domains of DOF proteins from Arabidopsis thaliana. BioImpacts 8(3):167
Seifert E (2014) OriginPro 9.1: Scientific Data Analysis and Graphing Software Review.
Shangguan L, Chen M, Fang X, Xie Z, Zhang K, Zheng T, Pu Y, Fang J (2020) Comparative study of DAM, Dof, and WRKY gene families in fourteen species and their expression in Vitis vinifera. 3 Biotech 10 (2):72.
Shim Y, Kang K, An G, Paek N-C (2019) Rice DNA-binding one zinc finger 24 (OsDOF24) delays leaf senescence in a jasmonate-mediated pathway. Plant Cell Physiol 60(9):2065–2076
Stecher G, Tamura K, Kumar S (2020) Molecular evolutionary genetics analysis (MEGA) for macOS. Mol Biol Evol 37(4):1237–1239
Wang L, Guo K, Li Y, Tu Y, Hu H, Wang B, Cui X, Peng L (2010) Expression profiling and integrative analysis of the CESA/CSL superfamily in rice. BMC Plant Biol 10(1):282
Wang W, Liu Z, Guo Z, Song G, Cheng Q, Jiang D, Yang D (2011) Comparative transcriptomes profiling of photoperiod-sensitive male sterile rice Nongken 58S during the male sterility transition between short-day and long-day. BMC Genomics 12(1):1–10
Wang H, Zhao S, Gao Y, Yang J (2017) Characterization of Dof transcription factors and their responses to osmotic stress in poplar (Populus trichocarpa). PLoS ONE 12(1):e0170210
Wei F, Coe E, Nelson W, Bharti AK, Engler F, Butler E, Kim H, Goicoechea JL, Chen M, Lee S (2007) Physical and genetic structure of the maize genome reflects its complex evolutionary history. PLoS Genet 3(7):e123
Wu Q, Li D, Li D, Liu X, Zhao X, Li X, Li S, Zhu L (2015) Overexpression of OsDof12 affects plant architecture in rice (Oryza sativa L.). Front Plant Sci 6:833
Wu Z, Cheng J, Cui J, Xu X, Liang G, Luo X, Chen X, Tang X, Hu K, Qin C (2016) Genome-wide identification and expression profile of Dof transcription factor gene family in pepper (Capsicum annuum L.). Front Plant Sci 7:574
Wu Q, Liu X, Yin D, Yuan H, Xie Q, Zhao X, Li X, Zhu L, Li S, Li D (2017a) Constitutive expression of OsDof4, encoding a C 2-C 2 zinc finger transcription factor, confesses its distinct flowering effects under long-and short-day photoperiods in rice (Oryza sativa L.). BMC Plant Biol 17(1):166
Wu Y, Yang W, Wei J, Yoon H, An G (2017b) Transcription factor OsDOF18 controls ammonium uptake by inducing ammonium transporters in rice roots. Mol Cells 40(3):178
Wu Y, Lee S-K, Yoo Y, Wei J, Kwon S-Y, Lee S-W, Jeon J-S, An G (2018) Rice transcription factor OsDOF11 modulates sugar transport by promoting expression of sucrose transporter and SWEET genes. Mol Plant 11(6):833–845
Zhang H-M, Chen H, Liu W, Liu H, Gong J, Wang H, Guo A-Y (2012) AnimalTFDB: a comprehensive animal transcription factor database. Nucleic Acids Res 40(D1):D144–D149
Zhang Y, Verhoeff N, Chen Z, Chen S, Wang M, Zhu Z, Ouwerkerk P (2015) Functions of OsDof25 in regulation of OsC4PPDK. Plant Mol Biol 89(3):229–242
Zhang Y, Giuliani R, Zhang Y, Zhang Y, Araujo WL, Wang B, Liu P, Sun Q, Cousins A, Edwards G (2018) Characterization of maize leaf pyruvate orthophosphate dikinase using high throughput sequencing. J Integr Plant Biol 60(8):670–690
Zhou Y, Cheng Y, Wan C, Li J, Yang Y, Chen J (2020) Genome-wide characterization and expression analysis of the Dof gene family related to abiotic stress in watermelon. PeerJ 8:e8358
Zou Z, Zhu J, Zhang X (2019) Genome-wide identification and characterization of the Dof gene family in cassava (Manihot esculenta). Gene 687:298–307
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflicts of interests
All the authors declare that they have no financial or competing interests.
Additional information
Communicated by Anastasios Melis.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Khan, I., Khan, S., Zhang, Y. et al. Genome-wide analysis and functional characterization of the Dof transcription factor family in rice (Oryza sativa L.). Planta 253, 101 (2021). https://doi.org/10.1007/s00425-021-03627-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-021-03627-y