Abstract
Main conclusion
The Arabidopsis DORNRÖSCHEN-LIKE enhancer 2 comprises a high-occupancy target region in the IM periphery that integrates signals for the spiral phyllotactic pattern and cruciferous arrangement of sepals.
Abstract
Transcription of the DORNRÖSCHEN-LIKE (DRNL) gene marks lateral organ founder cells (LOFCs) in the peripheral zone of the inflorescence meristem (IM) and enhancer 2 (En2) in the DRNL promoter upstream region essentially contributes to this phyllotactic transcription pattern. Further analysis focused on the phylogenetically highly conserved 100-bp En2core element, which was sufficient to promote the phyllotactic pattern, but was recalcitrant to further shortening. Here, we show that En2core functions independent of orientation and create a series of mutations to study consequences on the transcription pattern. Their analysis shows that, first, in addition to in the inflorescence apex, En2core acts in the embryo; second, cis-regulatory target sequences are distributed throughout the 100-bp element, although substantial differences exist in their function between embryo and IM. Third, putative core auxin response elements (AuxREs) spatially activate or restrict DRNL expression, and fourth, according to chromatin configuration data, En2core enhancer activity in LOFCs correlates with an open chromatin structure at the DRNL transcription start. In combination, mutational and chromatin analyses imply that En2core comprises a high-occupancy target (HOT) region for transcription factors, which implements phyllotactic information for the spiral LOFC pattern in the IM periphery and coordinates the cruciferous array of floral sepals. Our data disfavor a contribution of activating auxin response factors (ARFs) but do not exclude auxin as a morphogenetic signal.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The DORNRÖSCHEN-LIKE (DRNL) gene is one of the earliest available markers for cellular determination in the peripheral zone of the Arabidopsis thaliana shoot apical meristem (SAM) (Chandler et al. 2011b). DRNL encodes an AP2-type transcription factor (Kirch et al. 2003) and acts redundantly with its close relatives DORNRÖSCHEN (DRN) (Chandler et al. 2007) and PUCHI (Chandler and Werr 2017) to control meristem identity and organ initiation. DRNL transcription has been analysed by imaging DRNL::erGFP transgenic plants and starts in the apical domain of the early embryo proper (Chandler et al. 2011a). Later in development, DRNL::GFP expression is restricted to founder cells of lateral organ primordia and starts with cotyledons in the embryo or leaves in the vegetative phase and prepatterns floral primordia in the peripheral zone of the inflorescence meristem (IM) or organs in all four floral whorls during the reproductive phase (Chandler et al. 2011b). The DRNL transcription pattern in lateral organ founder cells (LOFCs) is highly dynamic and is controlled by three enhancer elements designated En1, En2 and En3 (see top of Fig. 1a), which are distributed within 5.6 kb of upstream DRNL promoter sequences (Comelli et al. 2016). Enhancer En2 is located approximately 2.5 kb upstream from the DRNL transcription start site (TSS) and in the IM controls the spiral LOFC arrangement that prepatterns floral primordia redundantly with the more distal element En1 (Comelli et al. 2016). According to progressively increasing external and extended internal deletions of the DRNL promoter, enhancer En2 in the absence of En1 is necessary to activate transcription in the spiral LOFC pattern. The central 100-bp En2core sequence that is highly conserved and shares 87 invariant nucleotide positions in six distant Brassicaceae species is sufficient for En2core expression (Comelli et al. 2016).
DRNL promoter structure and mutations in the En2core enhancer element. a A schematic representation of the microsyntenic DRNL promoter upstream region up to the flanking At1g24600 gene (DRNLLONG) and previously established enhancer elements En1, En2 and En3 (Comelli et al. 2016). Depicted below are the external DRNLSHORT deletion that identified the redundancy between enhancer elements En1 and En2 and the 400-bp deletion SΔEn2 within the DRNLSHORT promoter construct that functionally identified the role of En2 in the IM. Reinsertion of the 100-bp En2core fragment into the 400-bp SΔEn2 deletion restored enhancer activity, whereas smaller overlapping 50-bp-bp fragments of En2core in En2coreD, En2coreE and Encore2F lines did not. Only the central En2coreD fragment that spans AuxRE1 and AuxRE2 showed occasional weak activity in sepals of a few transgenic lines (Comelli et al. 2016). b Molecular structure of new En2core mutations created within the DRNLSHORT promoter construct, which from top to bottom depict inversion of the 100-bp En2core fragment in SEn2core−inv, the position of nine scanning mutations (Scan1-9) and the 5-bp deletions Δ 5.1, Δ5.2 and Δ5.3 within the spacer region between AuxRE1 and AuxRE2 or the point mutations created within the core AuxRE tetranucleotides. The brown marked regions represent putative PIF protein binding sites (Mironova et al. 2014)
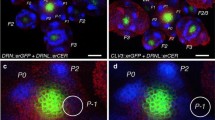
Imaging of the inflorescence apex has revealed that DRNL::GFP expression initiates in close proximity to the central stem-cell zone (Seeliger et al. 2016) and ongoing cell divisions displace DRNL::GFP-expressing LOFCs to the IM periphery, where the GFP signal bifurcates into two domains of which the outer or basal signal transiently marks the position of the bract; whereas, the inner or apical domain prepatterns the abaxial sepal (Chandler et al. 2011b; Chandler and Werr 2014). Expression in the abaxial sepal is followed by two expression foci that simultaneously prepattern the positions of both lateral sepals before DRNL::GFP expression is activated in LOFCs for the adaxial sepal. At the end of floral stage 1 (Smyth et al. 1990), the specification of LOFCs for the adaxial sepal coincides with the histological appearance of a furrow that separates the floral meristem (FM) from the IM (Chandler et al. 2011b). The four groups of sepal LOFCs are, thus, all specified before activation of the WUSCHEL and CLAVATA3 stem-cell markers indicates FM autonomy from the IM (Goldshmidt et al. 2008; Seeliger et al. 2016). Accordingly, DRNL transcription accompanies cellular fate decisions that begin close to the stem-cell zone (Seeliger et al. 2016), are perpetuated to the outer IM periphery and extend into the stereotypic cruciferous arrangement of sepals in the outer floral whorl before the floral meristem gains autonomy (Chandler and Werr 2014). En2 and particularly its central, most conserved En2core part in the absence of distal DRNL promoter elements are sufficient to report phyllotactic information (Comelli et al. 2016).
Plant phyllotaxis exemplifies the highly reproducible patterns within biological systems (Wolpert 1969) and the most widely accepted theory assumes the presence of inhibitory fields that originate from developing lateral primordia at the SAM periphery (Reinhardt et al. 2003; reviewed by Traas 2013). In concert, overlapping inhibitory fields that originate from incipient primordia might generate a dynamic spatiotemporal pattern that allows local organ initiation below a threshold inhibitor level. Auxin has been implicated in this network (Reinhardt et al. 2000) and the polarly localised PIN-FORMED 1 (PIN1) efflux carrier directs auxin to the sites of organ initiation and, thus, creates a concentration gradient around newly initiated organs (Reinhardt et al. 2003). Although auxin functioning as a morphogen was initially considered to be sufficient to generate appropriate phyllotaxy (Reinhardt et al. 2000, 2003), an inhibitory field involving non-cell-autonomous cytokinin signalling by the ARABIDOPSIS HISTIDINE PHOSPHOTRANSFER PROTEIN 6 (AHP6) inhibitor was identified to regulate the position of lateral organ initiation or the timing of successive primordia initiation in crosstalk with auxin (Besnard et al. 2014). The AHP6 promoter is a potential target of the DRNL transcription factor (Ikeda et al. 2006) and the transcription patterns of both genes in the IM peripheral zone are largely congruent (Chandler and Werr 2015). In contrast, DR5-reported auxin response maxima reside at the outer margin of the DRNL and AHP6 expression domains (Besnard et al. 2014; Chandler and Werr 2014), although point mutations in two core auxin response elements (AuxREs) have provided evidence that the DRNL En2 enhancer integrates an auxin response (Comelli et al. 2016). Two core GACA AuxRE motifs (Yamaguchi et al. 2013) in inverted orientation reside centrally within the 100-bp En2core region at a distance of 19 bp from each other, which approximates to two helical turns, suggesting that both face to one side of the DNA double helix. The entire 27-bp region is invariant in six Brassicaceae species and inactivating GCCA point mutations (Yamaguchi et al. 2013) in both core AuxREs abolishes En2 enhancer function (Comelli et al. 2016).
Auxin is essential for lateral organ initiation (Przemeck et al. 1996; Galweiler et al. 1998; Reinhardt et al. 2003), but auxin responses are complex. At the level of transcriptional control, the auxin response factor (ARF) family functions by binding to AuxREs in promoters of auxin-response genes. However, the ARF family contains activating or repressing members, which can recruit members of a second family of transcription factors, the Aux/IAA repressors, which are degraded depending on the intracellular auxin concentration. In addition, ARFs also homo- or heterodimerise (Boer et al. 2014), which involves combinatorial cooperativity and spacing constraints between AuxREs, such as the 19 bp between the putative core AuxREs in En2core (Comelli et al. 2016). Furthermore, canonical AuxRE motifs co-exist with non-canonical variants (Boer et al. 2014; Mironova et al. 2014) and are possibly recognised with different affinities. Additionally, ARFs functionally interact with other transcription factors (Guilfoyle and Hagen 2012) and well-studied examples are BRASSINAZOLE-RESISTANT1 (BZR) and PHYTOCHROME INTERACTING FACTOR 4 (PIF4) (Oh et al. 2014). G-boxes, the cis-regulatory target sites of these transcription factors, are enriched genome wide in the vicinity of AuxREs in Arabidopsis (Mironova et al. 2014) and both are present within En2core (Comelli et al. 2016). Together, the conservation of sequence and attributes suggests that En2core potentially functions as a unit and serves to recruit multiple transcription factors into a functional enhanceosome (Panne 2008). This interpretation is supported by an open chromatin configuration in IM cells and LOFCs (Frerichs et al. 2019).
Here, we describe a detailed mutational analysis of the DRNL En2core element (Comelli et al. 2016) and compare the effects of mutations in the IM and the embryo. The data reveal differences in the cis-regulatory sequences that are active in the embryo or IM and that include the core AuxREs, which spatially restrict DRNL transcription in the embryo. In the IM peripheral zone, expression in LOFCs appears to be closely coupled to DRNL expression in the sepals. Apparently, En2core combines cis-regulatory elements used to integrate phyllotactic information for LOFC specification in the IM periphery with elements that contribute to the determination of sepal anlagen in the outer floral whorl.
Materials and methods
DRNL promotor constructs
The DRNLLONG::GFP and DRNLSHORT::GFP promoter constructs have been described previously (Comelli et al. 2016) and consist of 5634 bp or 3176 bp upstream of the DRNL translation start codon, respectively. A unique XmaI restriction site in front of the translation start allowed isolation of the DRNL promoter as an XmaI-AscI fragment, which for mutagenesis was subcloned into pBluescript II KS ( +). The Phusion® site-directed mutagenesis kit (NEB) was used to delete the 100-bp En2core element and to create a unique FseI site with which to insert the modified En2core fragments. The double AuxRE1 + 2mut mutation has already been described in Comelli et al. (2016), the single AuxRE1mut and AuxRE2mut (TGTC to TGGC) mutations were created by primers in Table 1. The 5-bp deletion mutations Δ5.1, Δ5.2 andΔ5.3 were also created by the Phusion® site-directed mutagenesis kit (NEB) with primers listed in Table 1.
The Scan1–9 mutations were generated by the In-Fusion HD Cloning Kit (Clontech/Takara); the SΔEn2core promoter construct in pBluescript II KS ( +) served as a recipient, which was linearised at the FseI site and the overhanging 3′ termini were trimmed with T4 polymerase. All nine mutated 100-bp En2core fragments were chemically synthesised as 130-bp single-strand oligonucleotides that contained 13-bp extra DRNL promoter sequences at either end plus a residual 2 bp from the FseI site, which served as targets for PCR amplification and recombination. Homologous recombination followed the manufacturer’s protocol; the resulting Scan1–9 mutation constructs were confirmed by DNA sequencing. Promoter fragments carrying En2core mutations were generally converted to a GFP expression cassette with flanking AscI sites for cloning into the binary pGPTV Asc-BAR vector (Überlacker and Werr 1996). Transformation of Agrobacterium tumefaciens (GV3101) and Arabidopsis thaliana L. Col-0 plants was performed as described in Bechtold and Pelletier (1998). Transgenic progeny were selected via BASTA resistance and were grown on soil in the greenhouse in long-day conditions (16 h light: 8 h dark).
Confocal imaging
Expression of GFP was monitored using a Zeiss LSM 700 confocal laser scanning microscope. GFP was excited at 488 nm and emission was analysed between 502 and 525 nm. Image Z-stacks were converted into 3D images using Imaris software (Bitplane, Zürich, Switzerland) to visualise DRNL expression at all floral developmental stages. Floral staging was assigned according to Smyth et al. (1990) and images were processed using Photoshop CS2 software (Adobe). At least six independent transgenic lines for each En2 core mutation were compared for expression in the embryo, IM and stage1 FMs.
Results
En2 mutagenesis
The DRNL promoter constructs used in previous analyses are schematically summarised in Fig. 1a and their characterisation identified three enhancer elements designated En1, En2 and En3 in the DRNL promoter upstream region extending to the upstream At1G24600 gene (Comelli et al. 2016). Both distal enhancers En1 and En2 synergise in the DRNLlong promoter construct to enhance transcription in the IM or early FM. In contrast, when En1 is absent in the DRNLSHORT promoter version, En2 is necessary and sufficient to promote DRNL transcription in the spiral LOFC pattern at the IM periphery and in the stage 1 floral buttress (Comelli et al. 2016). This En2 function correlated phylogenetically with a high degree of sequence conservation in the 100-bp En2core element, which is sufficient and fully functional when inserted into a larger 400 bp deletion SΔEn2 (Fig. 1a) that removes En2 in addition to flanking sequences (Comelli et al. 2016). Unfortunately, the 100-bp En2core element was recalcitrant to further dissection and three overlapping 50-bp sub-fragments (En2coreD–F in Fig. 1a) tested previously essentially lacked enhancer function.
We, therefore, adopted a less disruptive strategy and firstly created a set of linker-scanning mutations that shifted a 10-bp stretch of alternating AT nucleotides throughout the En2core sequences (Scan1–9). Secondly, we generated 5-bp deletions (Δ5.1, Δ5.2 or Δ5.3) in the evolutionarily invariant 19-bp spacer that separates the putative core AuxRE1 and AuxRE2 motifs. Although these 5-bp deletions overlapped with scanning mutations Scan3, 4 or 5, the deletions differed substantially; whereas the scanning mutations preserved distances, each 5-bp deletion reduced the distance between AuxRE1 and AuxRE2. More importantly, however, each 5-bp deletion reduced half a helical turn, which might affect the relative orientation of AuxREs from both being oriented to one face of the DNA double helix to being on opposite faces, which potentially also affects sequences at the outer flank of AuxRE1 and 2. Lastly, as in the previous study, we had only simultaneously mutated both AuxREs (AuxRE1 + 2mut in Fig. 1b); we now additionally mutated each putative core AuxRE element by converting GACA to GCCA in each (AuxRE1mut, AuxRE2mut) (Ulmasov et al. 1999; Yamaguchi et al. 2013). All the internal En2core mutations are depicted in Fig. 1b, which also includes SEn2−inv, in which the En2core sequences have been inverted.
En2 mutation affects DRNL expression in the embryo
To directly compare the expression pattern observed with individual En2core mutants to the wild-type pattern, Fig. 2a–f combines stage-specific expression patterns of the DRNLSHORT::GFP construct, which provided the basis for the mutations. Although comparison of DRNLLONG::GFP and DRNLSHORT::GFP expression patterns in developing wild-type embryos revealed no obvious differences, we have combined the DRNLLONG::GFP embryo patterns in Supplementary Fig. S1a, as they refine a previous analysis described in Chandler et al. (2011a). Following expression in the apical domain of the 8- to 16-cell proembryo (Fig. 2a), expression becomes focused to incipient cotyledon primordia at the late globular stage and is repressed in the prospective SAM (Fig. 2b). From the heart to early torpedo stage, DRNLSHORT::GFP expression was present throughout the emerging cotyledons (Fig. 2c, d), but became restricted to a small subset of cells in the cotyledon tips by the late torpedo stage (Fig. 2e). Expression in the cotyledons initially included the L1 layer (Fig. 2g), but was restricted to cells in the sub-epidermal layer during the torpedo stage (Fig. 2e), a layering that was less pronounced in later walking stick embryos (Fig. 2f). This expression pattern strictly depended on En2 activity and was completely lost when En2 was absent, in external deletions starting from DRNLLONG (ΔEX5) or internal deletions of the DRNLSHORT promoter (SΔEn2), both described in Comelli et al. (2016). For direct comparison with the expression of DRNLSHORT::GFP pattern on top, the second row (Fig. 2g-l) depicts embryos of SΔEn2core::GFP lines, where the 100-bp En2core element was deleted. Supplementary Fig. S1b shows similar results at two representative stages resulting from the larger 400-bp SΔEn2 deletion, where GFP expression is absent but restored in SΔEn2 + En2core::GFP lines, i.e. when the 100-bp En2core element replaced the deleted native 400-bp sequences (compare Fig. S1b and c). Embryonic GFP signals independent of En2 enhancer activity were only observed late in the walking-stick embryo at the flanks of the embryonic SAM in incipient vegetative leaf primordia. This leaf aspect of the expression pattern was robust and was only absent in external deletions when most of the DRNL promoter upstream sequences were removed, i.e. when enhancers En1, En2 and En3 were absent (Comelli et al. 2016).
DRNL::GFP expression during embryogenesis and the contribution of the En2core enhancer. a–f GFP expression pattern observed with the DRNLSHORT promoter constructs at successive embryonic stages as indicated above each CLSM picture. The pattern is identical to that of the DRNLLONG promoter version included in Suppl. Fig. S1a. g–l Results obtained following deletion of the 100-bp En2core element in SΔEn2core lines and representative examples with the longer 400-bp deletion SΔEn2 (Comelli et al. 2016) are shown in Suppl. Fig. S1b. The only remaining expression was observed in incipient leaf primordia initiated after the late torpedo stage, robust transcriptional activity that is redundantly controlled in the DRNL promoter (Comelli et al. 2016). m Tabular summary of changes in expression patterns observed with individual En2core mutations; columns correspond to the series of embryonic stages in a–f. Dark green indicates no difference from the wild-type pattern; no colour means the absence of DRNL expression at this embryo stage, whereas light green indicates quantitative changes in expression, which either consisted of reduced or increased (enhanced) GFP signal intensity or a spatially aberrant (ectopic) GFP pattern. The lower panel (n–t) shows altered expression patterns specific for individual En2core mutations. In AuxRE1,2mut lines (n, t). Activity was normal in the emerging cotyledons during the late globular/early heart stage (n), but ectopic activity in the L1 layer was detected at the torpedo stage (o); a similar result was observed for AuxRE1mut (p). Scan4 lines exhibited reduced GFP signal intensity during the early torpedo stage (q), whereas Scan6 or Scan9 (data not shown) lines showed an increased GFP signal, which was also spatially expanded at this stage (r). Subtle dynamic changes were observed later in Scan9 lines: the GFP signal was still absent in the L1 layer at the tip of the cotyledons at the torpedo stage (s) but was observed here later (t), together with epidermis-specific expression in the hypocotyl
Changes observed in the embryonic GFP expression pattern with individual En2core mutations are tabulated in Fig. 2m, where the horizontal axis corresponds to the embryo stages shown in Fig. 2a–f. At least 10 and on average 15 embryos per stage and line were analysed for six independent homozygous transgenic lines for each embryonic stage. Accordingly, the upper vertical row (SΔEn2/En2core) in Fig. 2m depicts deletion of En2 and corresponds to Fig. 2g–l, where GFP signals are absent, except in incipient leaf primordia. The non-stained tabular fields in row 1 mean that no signal was observed in at least 60 embryos from six independent lines per stage, although a few (< 0.5%) exceptions were occasionally observed in single plants, probably due to cross-fertilisation. In contrast, inversion of En2core in SEn2−inv::GFP lines generated patterns that were indistinguishable from those of DRNLshort::GFP lines and are indicated by green tabular fields in row 2. The overlapping 50-bp sub-fragments En2coreD and En2coreF (Comelli et al. 2016) were insufficient to restore expression of SΔEn2 (Fig. 2m); whereas, we observed expression with the En2coreE insert in the SΔEn2 400-bp deletion in two out of six lines in several late torpedo or walking-stick stage embryos with signals at the cotyledon tips (Suppl. Fig. S1e). This partial rescue is indicated by the light green-filled fields in row four. The late cotyledon activity depends on the integrity of both core AuxREs, because GFP expression in the AuxRE1 + 2mut construct was absent in the cotyledon tips from the torpedo stage onwards (Suppl. Fig. S1d). By contrast, both AuxREs were not required for expression at the transition from the globular to the heart stage, when DRNL promoter activity was restricted to cotyledon founder cells and was lost in the SAM anlage (Fig. 2n, Suppl. Fig. S1d). However, single or double AuxRE mutations led to reduced expression in the emerging cotyledons at the heart stage (Fig. 2m) and remaining expression was often asymmetric and occasionally confined to a single cotyledon or ectopically extended into the L1 layer (Fig. 2o, p). According to the 5-bp deletions Δ5.1, Δ5.2 or Δ5.3, neither the distance nor the relative orientation of AuxRE1 and AuxRE2 affected DRNL expression at these stages.
The early globular embryo stage was most sensitive to En2core mutations, in which expression was abolished (i) by the AuxRE1 + 2mut double point mutation but not by single AuxRE mutations; (ii) by each of the three central core 5-bp deletions (Δ5.1, Δ5.2, Δ5.3) and (iii) by six scanning mutations, Scan1, 3, 4, 5, 7 and 8. Strikingly, although individual point mutations in the AuxRE1mut or AuxRE2mut constructs did not affect early GFP expression, the overlapping Scan3 or Scan5 mutations and Scan4, which affected sequences in the invariant 19-bp spacer that separates AuxRE1 and AuxRE2, abolished En2core activity. This central region of En2core is not only particularly important for enhancer activity in the early Arabidopsis embryo, but also consistent with the unique partial rescue of the late embryonic pattern observed with SΔEn2 + En2coreE and a similar partial rescue previously detected in the IM (Comelli et al. 2016).
The central Scan3, 4 and 5 mutants also concertedly and quantitatively reduced En2core activity at the late heart and early torpedo stage (Fig. 2q), when other mutations that affected the earliest embryonic DRNLSHORT::GFP expression, i.e. Scan1, 7 and 8, Δ5.1, Δ5.2 and Δ5.3 did not affect DRNL expression. Notably, DRNL promoter activity in the incipient cotyledons at the late globular stage was not affected by AuxRE1mut, AuxRE2mut and AuxRE1 + 2mut; whereas, these quantitatively reduced or caused ectopic expression in the L1 layer at subsequent stages (compare Fig. 2p–e). After the heart stage, several En2core mutations outside of AuxRE1 or 2 quantitatively affected GFP activity, for example: Scan4 (Fig. 2q) reduced and Scan6 (Fig. 2r) enhanced expression in early torpedo stage embryos. These quantitative effects normalised towards the late torpedo stage, when GFP expression in the cotyledon tips of Scan1, Scan3–8 and Scan9 lines again was similar to that of wild type (compare Fig. 2s, e). Scan9, however, was unique in causing ectopic DRNL expression at the walking stick stage; i.e. in the epidermis at the base of the cotyledons or the hypocotyl (compare Fig. 2s, t). The only En2core mutation without any phenotypic consequences in the early or late embryo was Scan2.
Effects of En2 mutations on DRNL expression in the inflorescence apex
Within the reference construct DRNLSHORT::GFP (Comelli et al. 2016), En2core activity is essential to promote expression in the phyllotactic LOFC pattern in the IM and accordingly, the SΔEn2core deletion construct was not active in the IM peripheral zone (compare Fig. 3a, b); whereas, inversion of the En2core sequences in SEn2core−inv did not affect expression (compare Fig. 3c to a). However, En2core deletion affected not only the LOFC-specific pattern in the IM peripheral zone, but also expression in early sepal primordia (compare Fig. 3a, b). This is similar to previous results with the larger SΔEn2 and also, due to established redundancy between En1 and En2, is only observed in the context of the DRNLSHORT promoter (Comelli et al. 2016). Expression patterns in the IM and sepals were extremely dynamic and therefore, we have grouped aspects of the pattern that were affected by En2core mutation into four categories that are schematically depicted in the top row of Fig. 3g. The diagrams are simplifications deduced from the scheme in Fig. 3d that illustrates the IM, associated developing flowers and signals therein that prepattern individual organs in the top view on the IM of the DRNLSHORT::GFP reference pattern in Fig. 3a. The most frequent phenotype was (i) the absence of GFP signals in the IM and sepal primordia, similar to SΔEn2core (Fig. 3e) or (ii) the selective absence of GFP signals in the IM but their presence in early sepals (Fig. 3e); permutations thereof were (iii) reduced GFP signal intensity in the IM peripheral zone and the absence of promoter activity in the abaxial and lateral sepal primordia, but remaining activity at the position of the adaxial sepal until floral stage 2 or (iv) no En2 activity in the IM and in lateral sepals but activity in the abaxial and adaxial sepals (Fig. 3f).
DRNLSHORT:: GFP expression in the inflorescence apex and consequences of En2core mutations. a–f Merged CLSM Z-stacks of characteristic GFP expression patterns observed with different promoter versions. aDRNLSHORT:: GFP promoter construct, b Expression pattern observed with 100-bp SΔEn2core deletion lines. Similar pictures with the 400-bp (SΔEn2) are published in Comelli et al. (2016). c Restored expression pattern when En2core is inserted in inverse orientation in SEn2core−inv lines. d Histology of the DRNLSHORT:: GFP pattern shown in in a and a schematic differentiation between the IM and associated flowers (F1–F10) or incipient primordia (P0, I1 and I2) based on DRNLSHORT::GFP expression. GFP expression up to F7 is sepal-specific; in later F8–10 flowers, GFP expression ceases in the sepal tips. The remaining expression is associated with stamen founder cells and is not dependent on En2 activity. Some signals, as in F3, are out of the range of the CLSM Z-stacks. The abaxial (ab), adaxial (ad) or lateral (l) sepals are marked in 6, 7 and 8. The arrow towards Table g indicates a further reduction in this DRNLSHORT::GFP reference pattern, which lacks older F5–10 flowers and ignores the sequential activity of the DRNL promoter in abaxial, medial and adaxial flowers. e The absence of signals in LOFCs within the IM peripheral zone is exemplified by mutation Δ5.3, and f the concerted absence of the En2 enhancer activity in the IM periphery and in lateral sepals by mutation Scan4. g Tabular summary of changes in the expression patterns observed with individual En2core mutations, indices (i)–(iv) in the top row relate to the phenotypic categories described in the main text. The symbol + indicates a reproducible phenotype within a single category; + ± , ± or ± – indicate increasingly quantitative reductions in expression, which for Scan5 prevent an unambiguous classification; for example, whether only the adaxial or both medial, i.e. abaxial and adaxial sepal signals remain. In analogy to the embryo, these classifications are based on six independent lines per En2core mutation and at least 10 IMs per line. A + symbol means without exception, and + ± , ± and ± represent 80%, 50% and below 30% of analysed apices, respectively. The latter category relates to SΔEn2 + En2F only and to the same transgenic lines that partially rescued the embryonic expression pattern. The asterisk (*) associated with Scan6 and Scan9 indicates that both mutations selectively affect the morphogenetic phase of sepal development, i.e. late DRNLSHORT::GFP activity in the tips of the growing sepals after floral stage 3 is absent
We previously showed that when both core AuxREs are mutated, En2 activity in the IM is lost (Comelli et al. 2016). Here, however, only AuxRE1mut abolished enhancer activity in the IM and in sepals; whereas, AuxRE2mut led to only mild reductions in DRNL promoter expression. This selectivity contrasts with the phenotype of scanning mutations Scan3 and Scan5 that covered AuxRE1 or AuxRE2, respectively, and that both eliminated En2core activity in the IM peripheral zone. Clear differences in the effect of both scanning mutations were observed in sepal primordia: for Scan5, DRNL expression was observed in the adaxial sepal and occasionally somewhat weaker in the abaxial sepal; whereas similar to AuxRE1 + 2mut and AuxRE1mut, expression was absent in the sepals of Scan3 lines. The third-strongest phenotype after AuxRE1mut and Scan3 was obtained with Scan7, which eliminated DRNL expression in the IM but retained weak signal in the adaxial sepal. More subtle effects on expression were caused by Scan1, 4, 5 and 8, which led to weaker DRNL promoter activity in the spiral LOFC pattern in the IM but differed with respect to the degree of relative expression in the abaxial and adaxial or lateral sepal primordia. Scan4 is located between core AuxRE1 and AuxRE2 and partly or fully overlaps the three 5-bp deletions (Δ5.1, Δ5.2 and Δ5.3), which all reduced DRNL expression in the IM periphery, but not in sepals, a phenotype that was strongest with Δ5.3 (Fig. 3e). Only Scan2 and Scan9 did not affect DRNLSHORT::GFP expression in the IM or early FM; however, Scan9 and Scan6 (see asterisks in Fig. 3g) both affected late DRNL promoter activity in the tips of growing sepals after floral stage 3.
In vivo footprinting of the active En2 enhancer
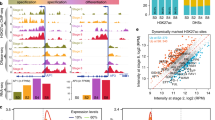
We previously applied fluorescence-activated cell sorting (FACS) with assay for transposase-accessible chromatin with high-throughput sequencing (ATAC-seq) and compared the chromatin state of DRNL::erGFP-expressing LOFC protoplasts in the ap1 cal IM (GFP +) with that of non-fluorescing (GFP-) cells (Frerichs et al. 2019). Peak calling revealed multiple transposase hypersensitive sites (THSs) throughout the microsyntenic region between DRNL and the upstream flanking AtG24600 gene (Fig. 4a). The qualitative difference between DRNL-expressing GFP + LOFCs and GFP- IM cells was greatest upstream of the DRNL TSS, where the chromatin in the proximal promoter region was only accessible in GFP + protoplasts (Fig. 4a). By contrast, peak calling of ATAC-seq reads across the En2 region revealed an open chromatin configuration in GFP- cells but further increased openness in LOFCs, with highest read frequencies within the 100-bp En2core element (Fig. 4b).
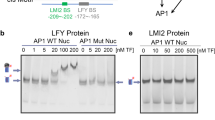
Chromatin configuration of the DRNL promoter upstream region within the ap1 cal inflorescence meristem with special emphasis on the En2core element. a Distribution of read termini obtained via ATAC-seq in DRNL::GFP-expressing LOFCs (green graph) and non-expressing IM cells (blue graph), depicted as normalised read counts (nrc) on the y-axis. Note the higher accessibility in En2, En3 and at the transcription start site in LOFCs. b Close-up of the En2 open chromatin region and position of the En2core element. c Distribution of ATAC-seq read termini within the nucleotide sequence; green bars depict numbers in LOFCs; blue bars those for DRNL::GFP-negative IM cells, below. Note the mostly significantly higher read numbers in LOFCs, the difference in transposase accessibility between the AuxRE1 and AuxRE2 regions and the LOFC-specific extra terminus (arrow) at position 68, which resides in the Scan7 mutant and a putative G-box motif
Calculation of mean normalised read frequencies and read termini in three biological replicates in the En2core region on the nucleotide level (Fig. 4c) revealed minimal variability in the frequency and position of read termini between the GFP + or GFP- replicates, but consistently showed substantially higher frequencies of read termini in DRNL-expressing LOFCs. Accordingly, in the ap1 cal IM, the En2core element locates within an open chromatin region that becomes significantly more accessible (P < 0.001) in LOFCs, when DRNL is actively transcribed. Novel cleavage sites unique to the transcriptionally active enhancer element were rare; a single potential candidate site is marked by an arrow at position 68 (Fig. 4c) and resides within Scan7, which eliminated En2core activity. However, En2core activity was also abolished by Scan3 or Scan5, which span AuxRE1 and AuxRE2, respectively. According to the AuxRE1mut phenotype, the distal element is functionally important, which correlates with few read termini within the AuxRE1 region in GFP + and GFP- cells. Although ATAC-seq read termini exhibit a sequence bias (Calviello et al. 2019), this contrasts to read termini within AuxRE2 in GFP- chromatin and even higher numbers in transcriptionally active chromatin from GFP + LOFCs. Differential in vivo transposase accessibility, thus, supports the phenotypic differences between AuxRE1mut and AuxRE2mut, i.e. accessibility in the latter is consistent with En2core activity that is unaffected in AuxRE2mut lines. By contrast, an adjacent 4-bp sequence towards the DRNL TSS that is devoid of read termini suggests it is protected from transcription and might explain the phenotypic discrepancy between the effects of AuxRE2mut and the overlapping Scan5 mutation.
Activating ARFs and En2core function
The phenotype resulting from the point mutation AuxRE1 + 2mut and AuxRE1mut suggests that AuxREs are functionally relevant for DRNL promoter activity, which overlaps with auxin response maxima in the IM. We, therefore, introduced DRNL::GFP into loss-of-function backgrounds of activating ARFs (Guilfoyle and Hagen 2007), i.e. arf5, arf6, arf7, arf8 or arf19 single mutants and due to known redundancy, into arf6 arf8 (Nagpal et al. 2005) and arf7 arf19 (Okushima et al. 2005) double mutants. No difference in the wild-type DRNL::GFP expression pattern was observed in inflorescence apices or embryos in any of these mutant backgrounds. Among arf mutants, arf5/monopteros is unique in exhibiting a lowly penetrant basal embryo domain phenotype followed by early seedling arrest (Hardtke and Berleth 1998). We, therefore, analysed DRNL::GFP expression in three arf5 alleles: the strong arf5-1 or mp-U55 alleles (Mayer et al. 1991) and the hypomorphic mp-S319/SALK_021319/arf5-2 allele (Alonso et al. 2003; Cole et al. 2009; Donner et al. 2009). Among a population segregating for the strong arf5-1 and mp-U55 alleles, wild-type DRNL::GFP expression was observed in embryos until the early globular stage. Homozygous mutant embryos only become morphologically distinguishable later and DRNL::GFP expression was detectable in the apical domain and in discrete foci in the presumptive cotyledons during the heart stage. Expression remained in the sub-epidermal layer of the cotyledon tips, although often in a broader expression domain than in wild-type embryos (Fig. 5a–d). By contrast, DRNL::GFP expression in the weak mp-S319 background differed spatially from that in both strong alleles and wild type; from the late heart stage, ectopic DRNL::GFP expression was observed in a central embryo domain and subsequently in the apical hypocotyl, where it weakly extended into the SAM and the cotyledon tips (Fig. 5e–h). By contrast to in strong mp alleles, DRNLSHORT::GFP expression in mp-S319 cotyledon tips remained weak.
DRNL::GFP expression pattern in mp mutant backgrounds. a–hDRNLLONG::GFP expression in homozygous mp-U55(a–d) or mp-S319 (e–h) embryos at successive developmental stages, which are difficult to stage due to aberrant and delayed development. Note the focus of GFP expression to the prospective cotyledons in the absence of MP in the mp-U55 loss-of-function allele (a) relative to the unstructured apical domain in the hypomorphic mp-S319 background (f), which possibly relates to subsequent ectopic activity in the hypocotyl domain in g or h. i Phenotype of the mp-S319 inflorescence with occasional flowers. j and k Top view on the mp-S319 IM and expression of the DRNLLONG::GFP construct, with the IM and two adjacent FMs in j or an isolated IM in k. The circular DRNLLONG expression domains in the IM or FMs overlap but are not identical to the MP::MP-GFP pattern depicted for comparison in n. l, m Expression of the DRNLSHORT::GFP construct. Compare the similar patterns and signal intensities of both promoter versions in the FMs between l or j and note the selectively reduced intensity of DRNLSHORT::GFP activity in the IM relative to the FM
Because arf5-1 and mpU55 alleles show the developmental arrest of early seedlings, post-embryonic DRNL promoter activity can only be studied in the weak mp-S319 allele (Fig. 5i), for which we compared expression of DRNLLONG::GFP and DRNLSHORT::GFP. Within DRNLLONG, En2 synergises with the distal enhancer element En1 that is absent in DRNLSHORT (Comelli et al. 2016). In the occasional flowers that develop in the mp-S319 inflorescence, both promoter versions showed a patchy, discontinuous ring of DRNL::GFP expression (Fig. 5j, l). This contrasts with that in the IM, where DRNLLONG::GFP showed a similar ring-shaped expression (Fig. 5j, k), whereas expression of DRNLSHORT::GFP was substantially weaker and was frequently restricted to isolated cells (Fig. 5l, m). This reduction in signal intensity with DRNLSHORT in the IM relates to the synergy between En1 and En2 (Comelli et al. 2016), but in the mp-S319 background, neither the DRNLLONG::GFP nor the DRNLSHORT::GFP transgenes showed a similar phyllotactic expression pattern to wild type (compare Fig. 5 j–m with Fig. 3a).
To date, the mp-S319 allele has been described to contain a left-border T-DNA insertion at position 6,890,755 of chromosome 1, which has been estimated from the MP 3′ terminus and is located within the penultimate exon 12 of the MP transcription unit (Alonso et al. 2003). Given the ectopic embryonic DRNL::GFP expression in the weak mp-S319 allele compared to the strong arf-1 and mp-U55 alleles, we queried whether a truncated transcript was present. Analysis of the mp-S319 right border revealed a large truncated T-DNA insertion exceeding the potential of long-range PCR reactions and joining two left T-DNA borders. This truncation removes 31 bp (6,890,746–6,890,774) of the MP transcription unit that include the 5′ splice-acceptor site of exon 12. RT-PCR amplifications with several MP-forward primers located in different exons and left-border T-DNA primers confirmed the presence of a spliced mp-S319 transcript that extends into the T-DNA sequence. This truncated mp-S319 mRNA is translatable into a shorter 815 amino-acid (aa) protein that deviates from the native MP gene product after aa 788, which is encoded close to the 3′ splice-donor site of exon 11. Read-through into non-spliced intron 11 sequences adds a 17-aa peptide (KRLSLIHSTNFLCHNLT), which terminates at a stop codon prior to the T-DNA insertion site.
When translated, this truncation causes deletion of the conserved ARF domains III and IV (Ulmasov et al. 1999) and results in a mp-S319 protein similar to the semi-dominant MP Δ transgene (Krogan et al. 2012), especially MPΔ-2 (1–794 aa). Therefore, we analysed expression of DRNL::GFP in MP Δ transgenic plants, but we observed no difference from the wild-type DRNL expression pattern in embryos or inflorescences either in heterozygous or in homozygous MP Δ progeny, which can be discriminated according to leaf shape, vascular density or floral phenotypes (Krogan et al. 2012). Even when we reduced competing endogenous MP activity by crossing the MP Δ transgene into the arf5-1 mutant background we observed no alteration in the spiral DRNL::GFP expression pattern in LOFCs at the IM periphery. However, the MP Δ transgene failed to rescue homozygous arf5-1 mutants and whereas genotyping revealed Mendelian inheritance of the MP Δ transgene, we could not confirm a single homozygous arf5-1 plant among 50 progeny that according to strength of leaf and floral phenotypes, were homozygous for MP Δ. The changes in the DRNL::GFP expression pattern observed in the mp-S319 IM (Fig. 5j–m), thus, can hardly be explained merely by the absence of domains III and IV in the C-terminally truncated MP polypeptide.
Discussion
Deletion analysis of the DRNL promoter with emphasis on the IM peripheral zone previously identified En2 as an enhancer element that synergistically activates transcription in LOFCs (Comelli et al. 2016), together with a more distal element, enhancer En1. The phylogenetically highly conserved central 100-bp En2core sequences were essential for this function, and these sequences were subjected to a detailed molecular analysis here and in addition to acting in the IM, also function in the embryo, where similar to its paralogue DRN, DRNL is initially expressed (Chandler et al. 2011a). The comparison of individual En2core mutations in the IM and the embryo showed: first, potential cis-regulatory target sequences are clustered throughout the 100-bp element; second, cis-elements function synergistically in the early embryo and the IM peripheral zone; third, putative core AuxRE sequences can spatially activate or restrict DRNL expression.
The greatest sensitivity to En2core mutations in the embryo was revealed in an early developmental window, when apical–basal polarity is established and 10 mutations (Fig. 2m) interfered with enhancer activity. This high sensitivity to sequence changes within En2core during early embryonic development suggests that the concerted activity of multiple cis-regulatory elements is required to establish a functional enhanceosome (Panne 2008). Subsequent maintenance of expression might involve fewer cis-regulatory sequences, because 7 out of 10 En2core mutations (Scan1, 2, 7 and 8 or Δ5-1, 2 and 3) had few transcriptional consequences at later embryonic stages. Their distribution within En2core agrees with the result of substitution experiments, in which only the central fragment (En2coreE) out of three overlapping 50-bp fragments (En2coreD,E or F in Fig. 1) weakly restored late DRNL promoter activity in the cotyledon tips, a rescue that according to AuxRE1 + 2mut lines, depends on the integrity of both AuxREs. A feature during late development is that many mutations, including point mutations in the putative core AuxREs, cause ectopic DRNL transcription, suggesting that some cis-elements potentially restrict DRNL promoter activity.
This loss of cell-type specificity in the embryo differs from that in the IM, where the most prominent phenotype is the absence of DRNL transcriptional activity, either in the spiral LOFC pattern within the peripheral zone or in early sepal primordia (Fig. 3b, g). The 5-bp deletions Δ5.1, Δ5.2 and Δ5.3 specifically affect DRNL transcription in LOFCs at the IM periphery (Fig. 3e, g) and suggest that the En2core centre is apparently critical to activate transcription in the phyllotactic LOFC pattern.
The LOFC-specific En2core activity in the IM and in sepal anlagen that encloses the prospective FM is observed with several mutations and relates to the development of the floral phytomer, which consists of a node, an axillary meristem, an internode and a bract. Bract outgrowth is suppressed in Arabidopsis, but occurs in puchi and leafy mutants, where DRNL::GFP expression persists from LOFCs into the incipient bracts (Chandler and Werr 2014). Bract outgrowth is a modulation of proximo–distal development and does not affect synchronised DRNL activity in the lateral sepals, but alters the sequence in which sepals are initiated, from abaxial, lateral then adaxial in wild type, to bract, lateral, abaxial and then adaxial sepal in puchi and leafy mutants (Chandler and Werr 2014). Within En2core, several mutations (Scan1, 4, 5 and 8) specifically affect DRNL expression in the two lateral sepals (Fig. 3f) and the IM and, thus, affect cis-regulatory sequences common to both patterning aspects. The most robust DRNL expression is observed in the adaxial sepal, which is specified last and reflects the bifurcation of the floral primordium from the inflorescence apex, the initiation of which at the end of floral stage 1 correlates with the de novo initiation of a stem-cell population in the FM (Goldshmidt et al. 2008). Dissection of En2core, thus, reveals: (i) cis-regulatory elements that monitor the phyllotactic position of new floral primordia within the IM periphery are either identical or in direct proximity to cis-motifs that provide coordinates for the robust crucifer arrangement of sepals; and (ii) such cis-regulatory sequences distinguish between lateral (Scan1,4) and abaxial sepals (Scan7) before the FM acquires autonomy.
Available ATAC-seq data depicted the chromatin configuration of DRNL::GFP-expressing LOFCs and non-expressing neighbours in the ap1 cal IM (Frerichs et al. 2019) and in seedling stem cells (Sijacic et al. 2018). The comparison of available chromatin configurations allows three conclusions: (i) En2 and especially En2core acquire an open chromatin configuration in the ap1 cal IM, which is not detectable in seedling stem cells (Frerichs et al. 2019), (ii) chromatin is more open in DRNL::GFP-positive LOFCs than in DRNL::GFP-negative cells. However, increased openness in LOFCs is not due to largely altered transposase cleavage sites, which would indicate changes in transcription factor binding, but reflects local hypersensitivity in LOFCs at several discrete En2core positions. (iii) This chromatin hypersensitivity in En2core contrasts with that in the DRNL TSS, which acquires an open configuration only in LOFCs, a long-range effect that is often associated with active transcription due to enhancer activity.
Although ATAC-seq data provide little evidence that transcription factors are recruited de novo to En2core for its expression within LOFCs, they support mutational analyses and indicate a compact genomic region targeted by multiple transcription factors. Such elements either guide the assembly of a multifactorial enhanceosome (Panne 2008), as is suggested by the orientation-independent function of the En2core element, or comprise a high-occupancy target (HOT) region (Foley and Sidow 2013). HOT regions in animals typically integrate signals from diverse regulatory pathways to quantitatively fine-tune immediate upstream promoter regions for RNA polymerase II recruitment. The cell-type-specific opening of chromatin in LOFCs at the DRNL TSS is compatible with RNA polymerase II entry (Frerichs et al. 2019); whereas, mutational analysis of En2core shows that phyllotactic signals in the IM peripheral zone integrate with those that pattern sepal anlagen in the flower.
The integration of auxin signalling into En2core function
Current models of phyllotactic signalling mainly focus on auxin as a morphogen (Reinhardt et al. 2003), although cytokinin signalling has also been acknowledged as an additional component (Besnard et al. 2014). The presence of two conserved putative core AuxREs in En2core and the essential integrity of AuxRE1, which also resides within a poorly accessible chromatin region, suggest that auxin contributes to En2core expression. By contrast, the AuxRE2mut mutation only subtly quantitatively affected DRNL expression and ATAC-seq reads terminate within the TGTC motif; which differs from directly flanking sequences that are inaccessible and are mutated in the phenotypic Scan5 mutant that overlaps AuxRE2. Although the detrimental effect of AuxRE1mut and its inaccessible chromatin configuration suggest that auxin contributes to En2core activity in LOFCs, the spiral phyllotactic pattern of DRNL expression in the IM is selectively affected only by the deletions Δ5.1, Δ5.2 and Δ5.3. Each deletion removes 5 bp from the 19-bp spacer region, which either fully or partially overlaps with scanning mutation Scan4 (Fig. 1b), which in addition to the spiral LOFCs in the IM, also affects En2core activity in lateral sepal anlagen. It is, therefore, unclear whether En2core transcription in LOFCs is directly due to sequence alterations in the AuxRE spacer region or to the 180° twist that results from deletion of half a helical turn from the centre of the En2core element. Concerted ATAC-seq hypersensitivity at the outer flanks of both AuxRE motifs suggests synergetic interactions between transcription factors bound to motifs at the outer flanks.
The effects of AuxRE point mutations differed in the early embryo, where En2core activity in the apical domain is abolished by the double mutation AuxRE1 + 2mut, but single mutations in either AuxRE1mut or AuxRE2mut had no detectable consequences. Based on the hypothesis that changes in DRNL::GFP expression are dependent on activating ARFs, we analysed DRNL::GFP expression in loss-of-function alleles of MP (ARF5), ARF6, ARF7, ARF8 and ARF19. However, the DRNL expression pattern and level was similar to that of wild type in the embryos or IMs of single mutant arf6 arf8 arf7 arf19 alleles or arf6 arf8 and arf7 arf19 double mutants. MONOPTEROS is particularly relevant, as it is essential for transcription of the DRNL paralogue DRN in the cotyledon tips and physically binds to canonical AuxREs in the DRN promoter upstream region (Cole et al. 2009). By contrast to DRN, we observed DRNL::GFP expression in homozygous arf5-1 and mp-U55 phenotypic embryos (Fig. 5a–d); whereas, DRN is expressed in the L1 layer of the cotyledon tips, DRNL is present in subtending provascular cells (Chandler et al. 2011a). The differential dependence of DRN and DRNL transcription on MP, thus, relates to discrete cell-type specificity and is also consistent with distinct genetic interactions between drn or drnl loss-of-function alleles and pinformed1 or pinoid mutants, which are hampered in auxin efflux or signal transduction, respectively (Chandler et al. 2011a).
Although AuxRE1mut links an AuxRE core motif with enhancer activity in LOFCs at the IM periphery, except for the pattern obtained in the mp-S319 background, we have obtained little evidence for a role of activating ARFs within the En2core element. Our data do not exclude that En2core responds to auxin, but mutational analyses and the chromatin configuration suggest that such a response would potentially have to integrate into a multifactorial protein–DNA complex. Two imperfect G-boxes (Figs. 1, 4) near to the putative core AuxREs within En2core are potential binding sites for PIF transcription factors, which interact with ARFs (Oh et al. 2014), and binding motifs for both transcription factor families co-occur genome wide (Mironova et al. 2014). In this context, the availability or local degradation of Aux/IAA proteins might be functionally relevant and be indirectly affected by the absence of domains III and IV in the mp-S319 background. The MP gene is transcriptionally up-regulated in LOFCs and in the ap1cal IM and belongs to the class of highly expressed ARFs (Frerichs et al. 2016); thus, the absence of domains III and IV might affect the interaction with cognate AUX/IAA inhibitory proteins (Guilfoyle and Hagen 2007). AUX/IAA proteins are locally degraded when auxin concentrations are high, which has been visualised in the IM via the DII-VENUS degron reporter (Brunoud et al. 2012). The DII-VENUS expression pattern spatially coincides with DR5::GFP-marked auxin response maxima in the IM, and both expression domains partially overlap with DRNL::GFP expression in LOFCs. The increased accessibility of the En2 chromatin to transposase cleavage in LOFCs could indicate the degradation of an inhibitory AUX/IAA interaction and gain of enhancer activity without recruitment of an extra transcription factor.
In conclusion, a detailed analysis of the En2core enhancer element reveals that its evolutionary sequence conservation relates to two major functions: embryonic patterning and LOFC specification in the peripheral zone of the IM. The effect of En2core mutations on DRNL expression shows that: first, potential cis-regulatory target sequences are distributed throughout the evolutionarily conserved 100-bp element; second, major differences exist in En2core function between the embryo and IM; third, cis-elements function synergistically in the early embryo and the IM peripheral zone and fourth, putative core AuxRE sequences can spatially activate or restrict DRNL expression. The En2core mutational data are consistent with available chromatin configuration data and suggest that En2core comprises a HOT region that is potentially occupied by multiple transcription factors. The essential integrity of a single putative core AuxRE element within En2core for IM activity suggests that a potential regulation of En2core by auxin integrates into a pre-assembled multi-factorial enhanceosome, although chromatin data for the active enhancer element in LOFCs provide little evidence for massive chromatin remodelling due to the entry of novel transcription factors.
Author contribution statement
DG, AF, JWC and WW conceived and designed the research. PC and DG conducted the experiments. AF and JE contributed new methods and analysed data. JWC and WW wrote the manuscript. All authors read and approved the manuscript.
Abbreviations
- ARF:
-
Auxin response factor
- AuxRE:
-
Auxin response element
- DRNL:
-
DORNRÖSCHEN-LIKE
- FM:
-
Floral meristem
- IM:
-
Inflorescence meristem
- LOFC:
-
Lateral organ founder cell
- SAM:
-
Shoot apical meristem
- TSS:
-
Transcription start site
References
Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H, Shinn P, Stevenson DK, Zimmerman J, Barajas P, Cheuk R, Gadrinab C, Heller C, Jeske A, Koesema E, Meyers CC, Parker H, Prednis L, Ansari Y, Choy N, Deen H, Geralt M, Hazari N, Hom E, Karnes M, Mulholland C, Ndubaku R, Schmidt I, Guzman P, Aguilar-Henonin L, Schmid M, Weigel D, Carter DE, Marchand T, Risseeuw E, Brogden D, Zeko A, Crosby WL, Berry CC, Ecker JR (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301(5633):653–657. https://doi.org/10.1126/science.1086391
Bechtold N, Pelletier G (1998) In planta Agrobacterium-mediated transformation of adult Arabidopsis thaliana plants by vacuum infiltration. Methods Mol Biol 82:259–266
Besnard F, Refahi Y, Morin V, Marteaux B, Brunoud G, Chambrier P, Rozier F, Mirabet V, Legrand J, Laine S, Thevenon E, Farcot E, Cellier C, Das P, Bishopp A, Dumas R, Parcy F, Helariutta Y, Boudaoud A, Godin C, Traas J, Guedon Y, Vernoux T (2014) Cytokinin signalling inhibitory fields provide robustness to phyllotaxis. Nature 505(7483):417–421. https://doi.org/10.1038/nature12791
Boer DR, Freire-Rios A, van den Berg WA, Saaki T, Manfield IW, Kepinski S, Lopez-Vidrieo I, Franco-Zorrilla JM, de Vries SC, Solano R, Weijers D, Coll M (2014) Structural basis for DNA binding specificity by the auxin-dependent ARF transcription factors. Cell 156(3):577–589. https://doi.org/10.1016/j.cell.2013.12.027
Brunoud G, Wells DM, Oliva M, Larrieu A, Mirabet V, Burrow AH, Beeckman T, Kepinski S, Traas J, Bennett MJ, Vernoux T (2012) A novel sensor to map auxin response and distribution at high spatio-temporal resolution. Nature 482(7383):103–106. https://doi.org/10.1038/nature10791
Chandler JW, Werr W (2014) Arabidopsis floral phytomer development: auxin response relative to biphasic modes of organ initiation. J Exp Bot 65(12):3097–3110. https://doi.org/10.1093/jxb/eru153
Chandler JW, Werr W (2015) Cytokinin-auxin crosstalk in cell type specification. Trends Plant Sci 20(5):291–300. https://doi.org/10.1016/j.tplants.2015.02.003
Chandler JW, Werr W (2017) DORNRÖSCHEN, DORNRÖSCHEN-LIKE, and PUCHI redundantly control floral meristem identity and organ initiation in Arabidopsis. J Exp Bot 68(13):3457–3472. https://doi.org/10.1093/jxb/erx208
Chandler JW, Cole M, Flier A, Grewe B, Werr W (2007) The AP2 transcription factors DORNRÖSCHEN and DORNRÖSCHEN-LIKE redundantly control Arabidopsis embryo patterning via interaction with PHAVOLUTA. Development 134(9):1653–1662. https://doi.org/10.1242/dev.001016
Chandler JW, Cole M, Jacobs B, Comelli P, Werr W (2011a) Genetic integration of DORNRÖSCHEN and DORNRÖSCHEN-LIKE reveals hierarchical interactions in auxin signalling and patterning of the Arabidopsis apical embryo. Plant Mol Biol 75(3):223–236. https://doi.org/10.1007/s11103-010-9721-5
Chandler JW, Jacobs B, Cole M, Comelli P, Werr W (2011b) DORNRÖSCHEN-LIKE expression marks Arabidopsis floral organ founder cells and precedes auxin response maxima. Plant Mol Biol 76(1–2):171–185. https://doi.org/10.1007/s11103-011-9779-8
Cole M, Chandler J, Weijers D, Jacobs B, Comelli P, Werr W (2009) DORNRÖSCHEN is a direct target of the auxin response factor MONOPTEROS in the Arabidopsis embryo. Development 136(10):1643–1651. https://doi.org/10.1242/dev.032177
Comelli P, Glowa D, Chandler JW, Werr W (2016) Founder-cell-specific transcription of the DORNRÖSCHEN-LIKE promoter and integration of the auxin response. J Exp Bot 67(1):143–155. https://doi.org/10.1093/jxb/erv442
Donner TJ, Sherr I, Scarpella E (2009) Regulation of preprocambial cell state acquisition by auxin signaling in Arabidopsis leaves. Development 136(19):3235–3246. https://doi.org/10.1242/dev.037028
Foley JW, Sidow A (2013) Transcription-factor occupancy at HOT regions quantitatively predicts RNA polymerase recruitment in five human cell lines. BMC Genomics 14:720. https://doi.org/10.1186/1471-2164-14-720
Frerichs A, Thoma R, Abdallah AT, Frommolt P, Werr W, Chandler JW (2016) The founder-cell transcriptome in the Arabidopsis apetala1 cauliflower inflorescence meristem. BMC Genomics 17:855. https://doi.org/10.1186/s12864-016-3189-x
Frerichs A, Engelhorn J, Altmuller J, Gutierrez-Marcos J, Werr W (2019) Specific chromatin changes mark lateral organ founder cells in the Arabidopsis inflorescence meristem. J Exp Bot 70(15):3867–3879. https://doi.org/10.1093/jxb/erz181
Galweiler L, Guan C, Muller A, Wisman E, Mendgen K, Yephremov A, Palme K (1998) Regulation of polar auxin transport by AtPIN1 in Arabidopsis vascular tissue. Science 282(5397):2226–2230
Goldshmidt A, Alvarez JP, Bowman JL, Eshed Y (2008) Signals derived from YABBY gene activities in organ primordia regulate growth and partitioning of Arabidopsis shoot apical meristems. Plant Cell 20(5):1217–1230. https://doi.org/10.1105/tpc.107.057877
Guilfoyle TJ, Hagen G (2007) Auxin response factors. Curr Opin Plant Biol 10(5):453–460. https://doi.org/10.1016/j.pbi.2007.08.014
Guilfoyle TJ, Hagen G (2012) Getting a grasp on domain III/IV responsible for auxin response factor-IAA protein interactions. Plant Sci 190:82–88. https://doi.org/10.1016/j.plantsci.2012.04.003
Hardtke CS, Berleth T (1998) The Arabidopsis gene MONOPTEROS encodes a transcription factor mediating embryo axis formation and vascular development. EMBO J 17(5):1405–1411. https://doi.org/10.1093/emboj/17.5.1405
Ikeda Y, Banno H, Niu QW, Howell SH, Chua NH (2006) The ENHANCER OF SHOOT REGENERATION 2 gene in Arabidopsis regulates CUP-SHAPED COTYLEDON 1 at the transcriptional level and controls cotyledon development. Plant Cell Physiol 47(11):1443–1456. https://doi.org/10.1093/pcp/pcl023
Karabacak Calviello A, Hirsekorn A, Wurmus R, Yusuf D, Ohler U (2019) Reproducible inference of transcription factor footprints in ATAC-seq and DNase-seq datasets using protocol-specific bias modeling. Genome Biol 20(1):42. https://doi.org/10.1186/s13059-019-1654-y
Kirch T, Simon R, Grunewald M, Werr W (2003) The DORNROSCHEN/ENHANCER OF SHOOT REGENERATION1 gene of Arabidopsis acts in the control of meristem ccll fate and lateral organ development. Plant Cell 15(3):694–705
Krogan NT, Ckurshumova W, Marcos D, Caragea AE, Berleth T (2012) Deletion of MP/ARF5 domains III and IV reveals a requirement for Aux/IAA regulation in Arabidopsis leaf vascular patterning. New Phytol 194(2):391–401. https://doi.org/10.1111/j.1469-8137.2012.04064.x
Mayer U, Ruiz RAT, Berleth T, Miséra S, Jürgens G (1991) Mutations affecting body organization in the Arabidopsis embryo. Nature 353(6343):402–407. https://doi.org/10.1038/353402a0
Mironova VV, Omelyanchuk NA, Wiebe DS, Levitsky VG (2014) Computational analysis of auxin responsive elements in the Arabidopsis thaliana L. genome. BMC Genomics 15(Suppl 12):S4. https://doi.org/10.1186/1471-2164-15-S12-S4
Nagpal P, Ellis CM, Weber H, Ploense SE, Barkawi LS, Guilfoyle TJ, Hagen G, Alonso JM, Cohen JD, Farmer EE, Ecker JR, Reed JW (2005) Auxin response factors ARF6 and ARF8 promote jasmonic acid production and flower maturation. Development 132(18):4107–4118. https://doi.org/10.1242/dev.01955
Oh E, Zhu JY, Bai MY, Arenhart RA, Sun Y, Wang ZY (2014) Cell elongation is regulated through a central circuit of interacting transcription factors in the Arabidopsis hypocotyl. Elife 3:e03031. https://doi.org/10.7554/eLife.03031
Okushima Y, Overvoorde PJ, Arima K, Alonso JM, Chan A, Chang C, Ecker JR, Hughes B, Lui A, Nguyen D, Onodera C, Quach H, Smith A, Yu G, Theologis A (2005) Functional genomic analysis of the AUXIN RESPONSE FACTOR gene family members in Arabidopsis thaliana: unique and overlapping functions of ARF7 and ARF19. Plant Cell 17(2):444–463. https://doi.org/10.1105/tpc.104.028316
Panne D (2008) The enhanceosome. Curr Opin Struct Biol 18(2):236–242. https://doi.org/10.1016/j.sbi.2007.12.002
Przemeck GK, Mattsson J, Hardtke CS, Sung ZR, Berleth T (1996) Studies on the role of the Arabidopsis gene MONOPTEROS in vascular development and plant cell axialization. Planta 200(2):229–237
Reinhardt D, Mandel T, Kuhlemeier C (2000) Auxin regulates the initiation and radial position of plant lateral organs. Plant Cell 12(4):507–518
Reinhardt D, Pesce ER, Stieger P, Mandel T, Baltensperger K, Bennett M, Traas J, Friml J, Kuhlemeier C (2003) Regulation of phyllotaxis by polar auxin transport. Nature 426(6964):255–260. https://doi.org/10.1038/nature02081
Seeliger I, Frerichs A, Glowa D, Velo L, Comelli P, Chandler JW, Werr W (2016) The AP2-type transcription factors DORNROSCHEN and DORNROSCHEN-LIKE promote G1/S transition. Mol Genet Genomics 291(5):1835–1849. https://doi.org/10.1007/s00438-016-1224-x
Sijacic P, Bajic M, McKinney EC, Meagher RB, Deal RB (2018) Changes in chromatin accessibility between Arabidopsis stem cells and mesophyll cells illuminate cell type-specific transcription factor networks. Plant J 94(2):215–231. https://doi.org/10.1111/tpj.13882
Smyth DR, Bowman JL, Meyerowitz EM (1990) Early flower development in Arabidopsis. Plant Cell 2(8):755–767. https://doi.org/10.1105/tpc.2.8.755
Traas J (2013) Phyllotaxis. Development 140(2):249–253
Überlacker B, Werr W (1996) Vectors with rare-cutter restriction enzyme sites for expression of open reading frames in transgenic plants. Mol Breeding 2(3):293–295
Ulmasov T, Hagen G, Guilfoyle TJ (1999) Activation and repression of transcription by auxin-response factors. Proc Natl Acad Sci USA 96(10):5844–5849. https://doi.org/10.1073/pnas.96.10.5844
Wolpert L (1969) Positional information and the spatial pattern of cellular differentiation. J Theor Biol 25(1):1–47
Yamaguchi N, Wu MF, Winter CM, Berns MC, Nole-Wilson S, Yamaguchi A, Coupland G, Krizek BA, Wagner D (2013) A molecular framework for auxin-mediated initiation of flower primordia. Dev Cell 24(3):271–282. https://doi.org/10.1016/j.devcel.2012.12.017
Acknowledgements
Open Access funding provided by Projekt DEAL. This work was supported by DFG grant WE 1262/11-2; W.W. is a member of the Cluster of Excellence on Plant Science (CEPLAS; EXC 1028), which provided infrastructural support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material Supplementary Fig. S1 Additional embryonic expression patterns. a Transgenic DRNLLONG::GFP expression in the developing Arabidopsis embryo; successive stages are indicated above each CLSM picture. b Early heart and late torpedo stage embryos of SΔEn2::GFP plants devoid of GFP signals. c Replacement of the 400-bp SΔEn2 deletion by the 100-bp En2core element and signals in SΔEn2+En2core::GFP transgenic lines. d DRNLLONG promoter carrying the AuxRE1 + 2mut mutation fused to the GFP marker showing expression in the cotyledon founder cells in the late globular stage embryo but the absence of GFP expression in the cotyledon tips after the torpedo stage. e Partial rescue observed with the central 50-bp En2coreE fragment in the context of the 400-bp SΔEn2 deletion in two out of six lines analysed. Scale bars correspond to 20 μm
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Comelli, P., Glowa, D., Frerichs, A. et al. Functional dissection of the DORNRÖSCHEN-LIKE enhancer 2 during embryonic and phyllotactic patterning. Planta 251, 90 (2020). https://doi.org/10.1007/s00425-020-03381-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-020-03381-7