Abstract
In yeast and mammals, ATP-dependent chromatin remodelling complexes of the SWI/SNF family play critical roles in the regulation of transcription, cell proliferation, differentiation and development. Homologues of conserved subunits of SWI/SNF-type complexes, including Snf2-type ATPases and SWI3-type proteins, participate in analogous processes in Arabidopsis. Recent studies indicate a remarkable similarity between phenotypic effects of mutations in the SWI3 homologue ATSWI3C and bromodomain-ATPase BRM genes. To verify the extent of functional similarity between BRM and ATSWI3C, we have constructed atswi3c brm double mutants and compared their phenotypic traits to those of simultaneously grown single atswi3c and brm mutants. In addition to inheritance of characteristic developmental abnormalities shared by atswi3c and brm mutants, some additive brm-specific traits were also observed in the atswi3c brm double mutants. Unlike atswi3c, the brm mutation results in the enhancement of abnormal carpel development and pollen abortion leading to complete male sterility. Despite the overall similarity of brm and atswi3c phenotypes, a critical requirement for BRM in the differentiation of reproductive organs suggests that its regulatory functions do not entirely overlap those of ATSWI3C. The detection of two different transcript isoforms indicates that BRM is regulated by alternative splicing that creates an in-frame premature translation stop codon in its SNF2-like ATPase coding domain. The analysis of Arabidopsis mutants in nonsense-mediated decay suggests an involvement of this pathway in the control of alternative BRM transcript level.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Based on the recent advances in chromatin research, it is now firmly established that mechanisms affecting the accessibility of chromatin DNA play key roles in the regulation of transcription, replication, repair and recombination of nuclear genomes in eukaryotes (Martens and Winston 2003; Roberts and Orkin 2004; Smith and Peterson 2005). These mechanisms involve ATP-dependent chromatin remodelling that acts in conjunction with DNA and histone modifications. ATP-dependent chromatin remodelling is mediated by multi-subunit complexes that use energy from ATP hydrolysis to destabilise and translocate DNA on nucleosomes (Saha et al. 2006). Four major classes of chromatin remodelling complexes (CRCs) (SWI/SNF, ISWI, Mi-2 and Ino80) characterised thus far are distinguished by their central catalytic ATPase and unique composition of auxiliary subunits. Complexes belonging to the SWI/SNF class were first discovered in S. cerevisiae and carry a Snf2-type (Sth1, BRAHMA) ATPase with a C-terminal signature called the bromodomain. In addition to Snf2-type ATPases, the SWI/SNF complexes purified from yeast, Drosophila and mammals share at least two other evolutionarily conserved subunits representing homologues of yeast SNF5 and SWI3 proteins. Both SNF5 and SWI3 (the latter occurring as a dimer) are critical for the assembly, stability and proper targeting of SWI/SNF complexes (Mohrmann and Verrijzer 2005).
To date, no SWI/SNF-type CRC has been purified from higher plants. However, sequencing of the Arabidopsis thaliana and rice genomes has identified genes encoding homologues of Snf2 ATPase, SNF5 and SWI3 subunits of SWI/SNF complexes (Brzeski et al. 1999; Verbsky and Richards 2001; Sarnowski et al. 2002; Wagner and Meyerowitz 2002; Farrona et al. 2004; Jerzmanowski 2007; Kwon and Wagner 2007; see also the Plant Chromatin Database at http://chromdb.org). Of the four potential Arabidopsis orthologues of the Snf2-type ATPase, only BRAHMA (BRM) carries a bromodomain. In SPLAYED (SYD), the closest homologue of BRM, the bromodomain is replaced by a divergent C-terminal domain of unknown function (Su et al. 2006; Jerzmanowski 2007; Kwon and Wagner 2007). The Arabidopsis genome encodes a single SNF5 orthologue, BUSHY (BSH, Brzeski et al. 1999) and four different (ATSWI3A, ATSWI3B, ATSWI3C and ATSWI3D) SWI3-type SWI/SNF core subunits (Sarnowski et al. 2002).
Combinatorial assembly of SWI/SNF subunits into structurally and functionally diverse CRCs is connected to the control of key pathways in mammalian development (Lessard et al. 2007). Analogously, CRCs carrying different combinations of Snf2-type ATPases with SNF5/BSH and homo- or heterodimeric forms of different ATSWI3 subunits have been implicated in various developmental processes in Arabidopsis (Jerzmanowski 2007; Kwon and Wagner 2007). Recent genetic analysis of the ATSWI3 gene family demonstrated that both ATSWI3A and ATSWI3B are essential for early embryonic development, whereas ATSWI3C and ATSWI3D affect different phases of vegetative and reproductive development (Sarnowski et al. 2005). Functional studies of Snf2-type ATPases showed that the Arabidopsis syd mutant is viable and its phenotypic characteristics are clearly different from those of atswi3c and atswi3d mutants (Wagner and Meyerowitz 2002; Su et al. 2006). In contrast, phenotypic traits of brm and atswi3c insertion mutants are intriguingly similar (Sarnowski et al. 2005; Hurtado et al. 2006; Tang et al. 2008). The interaction of ATSWI3C with BRM in the yeast two-hybrid system (Farrona et al. 2004) suggests that ATSWI3C is a core subunit of a BRM ATPase-associated SWI/SNF complex. To verify the functional similarity of BRM and ATSWI3C, we have performed a comparative study of developmental defects of atswi3c brm double mutants and previously characterised atswi3c and brm null mutants. Our data show that most developmental defects caused by single brm and atswi3c mutations are indistinguishable from those observed in the atswi3c brm double mutants. This indicates that BRM and ATSWI3C perform largely complementary functions and most likely represent interacting subunits of a CRC. Nonetheless, inactivation of BRM results in more severe disturbance of differentiation of female and male reproductive organs than do atswi3c mutations, suggesting that BRM has some unique regulatory functions. The detection of two different isoforms of BRM mRNA indicates that the transcription of this SNF2-like ATPase is regulated by alternative splicing. The occurrence of a premature termination codon (PTC) in the coding region of one of the isoforms implicates a role for nonsense-mediated decay (NMD) in the modulation of the level of this mRNA isofom in plants. Accumulation of aberrant BRM mRNA isoform in the upf1-5 and upf3-1 mutants indicates that the stability of this isoform is indeed controlled by NMD.
Materials and methods
Plant lines and growth conditions
Arabidopsis thaliana L. Heynh., Columbia-0 (Col-0) (Lehle seeds, Round Rock, TX, USA) was used in all experiments. The atswi3c-1 (Koncz_27320) and brm-1 (SALK_030046) mutant alleles were previously characterised by Sarnowski et al. (2005) and Hurtado et al. (2006), respectively. The brm-6 (Koncz_77269) mutant allele was identified by PCR screening of our T-DNA mutant collection (Ríos et al. 2002). The NMD pathway mutants upf1-5 (SALK_112922) and upf3-1 (SALK_025175) were also described previously (Hori and Watanabe 2005; Arciga-Reyes et al. 2006). The atswi3c-1 brm-1 and atswi3c-1 brm-6 double mutants were generated either by crossing heterozygous mutant lines or by pollination of a brm-6 null homozygote with atswi3c-1 pollen. Genotypes of all single and double mutants were confirmed by PCR analysis using allele-specific primers (Table S1).
Owing to sterility of homozygous brm and atswi3c brm lines and low yield of atswi3c mutant seeds, F2 segregating progeny used for genotyping the [atswi3c-1/+ brm-1/brm-1] [atswi3c-1/+ brm-6/brm-6] [atswi3c-1/atswi3c-1 brm-1/+] and [atswi3c-1/atswi3c-1 brm-6/+] lines were planted in soil and grown under long day (LD) conditions (16 h light/8 h dark) at 18–23°C, with 70% humidity and 200 μM m−2 s−1 light intensity. Seedlings were cultivated in 1/2 Murashige and Skoog (MS) seed germination medium with 0.5% (w/v) sucrose and 0.8% agar (Koncz et al. 1994).
Characterisation of the brm-6 T-DNA insertion mutant
The brm-6 (Koncz_77269) mutant allele was identified using combinations of gene-specific primers with the T-DNA left border primer as described in Sarnowski et al. (2005) (Table S1). The brm-6 mutant line contains an inverted T-DNA repeat flanked by two left borders in exon 2. The T-DNA insertion event resulted in a target site deletion of 40 bp (nucleotides 1,364–1,404 of the BRM coding sequence). The junctions between BRM exon 2 and the ends of the inverted T-DNA were LB1 (5′-GGACAGGGGAatctacatggat-3′) and LB2 (5′-gcacccgcgacCTCCATGTTCT-3′, Fig. 1a). Upper and lower case letters denote plant DNA and T-DNA sequences, respectively.
Characteristics of brm insertion and point mutations. a Positions of T-DNA insertions and point mutations in the BRM gene. Exons and introns are represented by grey boxes and lines, respectively. Boundaries of the T-DNA inserts are marked LB and RB, whereas the positions of point mutations are indicated by arrows. The brm-6 mutant is described in this work. Insertion mutations brm-1, -2, -3, and -4 and point mutations brm-101, -102, -103, and brm-5 were characterised previously (Hurtado et al. 2006; Kwon et al. 2006; Farrona et al. 2007; Tang et al. 2008). The brm-1 and brm-6 mutations were used in the genetic analyses described in this work. The positions and orientation of primers are marked with small arrows: pair 1–2 was used to identify the brm-6 mutation, whereas pair 3–4 was employed in RT-PCR assays. b RT-PCR assay with cDNA templates prepared from wild-type and homozygous brm-1 and brm-6 mutant plants with gene-specific primers (3–4) and control ACTIN2 primers. The full-length BRM transcript is not detectable in the brm-1 and brm-6 mutants. c Western analysis of nuclear extracts of 30-day-old wild-type (Col-0), brm-1 and brm-6 plants with an anti-BRM antibody (α-BRM, top) and with a control anti-histone H3 antiserum (α-H3, bottom)
RNA isolation, northern hybridisation, cDNA synthesis and RT-PCR
Isolation of total RNA from plant tissues was performed as described previously (Sarnowski et al. 2002). For northern-blot analysis 10 μg of total RNA isolated from whole plants were loaded per lane, run on 1.2% formaldehyde-agarose gels and blotted on Hybond N-plus membrane (Amersham), which was then hybridised with a probe labelled with [α –32P]dCTP using a random priming DecaLabel DNA Labeling Kit (Fermentas). The analysis of hybridisations was carried out using a PhosphorImager (Molecular Dynamics). cDNA synthesis was performed as described by Sarnowski et al. (2005). Amplification of a BRM cDNA fragment of 2,206 bp shown in Fig. 1b was performed with primers 3 and 4 (Table S1). For amplification of control ACTIN2 cDNA, primers ACT1 and ACT2 were used (Sarnowski et al. 2005; Table S1). Amplification of a cDNA corresponding to alternative splice variant of BRM transcript (BRM Δ ) was performed with primers PL and PRΔ (Table S1). UBQ11 cDNA was used as internal PCR control (Tyler et al. 2004; Table S1).
BRM antibody and Western blotting
To produce a BRM antigen, a 300 bp fragment of BRM cDNA encoding amino acids 131–230 was inserted into plasmid pQE60 (Qiagen) to create pQE-N. This plasmid encodes a 13-kDa N-terminal fragment of the BRM protein fused to 6xHis tag at its N terminus. The fusion protein was expressed in Escherichia coli and purified on Ni-NTA agarose affinity matrix as recommended by Qiagen. Against the 13-kDa His-tagged BRM fusion protein polyclonal antibody was raised by immunisation of a rabbit (Eurogentec). A portion of anti-BRM antibody was subsequently purified by affinity chromatography (Eurogentec). For Western-blot analysis of the BRM protein, 30-day-old Col-0, brm-1, and brm-5 plants grown in soil were harvested. Purification and extraction of nuclei were according to Gendrel et al. (2005) with modifications. A 20 μg aliquots of nuclear extracts were separated by 6% SDS-PAGE and the BRM protein was detected by immunoblotting using a 2,000-fold dilution of affinity-purified anti-BRM polyclonal antibody. The blots were then incubated with SuperSignal West Femto substrate (Pierce) following the detection of signals by chemiluminescence. Histone H3 immunoblotting was performed after electrophoresis using 13% SDS-PAGE gels. Membranes were probed with anti-histone H3 antibody (1:5,000; 07-690, Upstate).
Quantitative real-time PCR
Total RNA was extracted from approximately 50 mg samples of different wild-type Arabidopsis tissues: leaf rosettes and roots of 15-day-old seedlings, as well as rosette leaves, cauline leaves, stems, flowers, and siliques of 45-day-old plants. Flowers and siliques were harvested at various stages of development. RNA was extracted using the RNeasy Plant Mini kit (Qiagen), and DNA was removed by DNase treatment with a TURBO DNA-free kit (Ambion). A first-strand cDNA synthesis kit (Roche) was used to prepare cDNA from 1 μg of RNA. Aliquots (1 μl) of cDNA samples were used as templates in 20 μl reactions containing LightCycler 480 SYBR Green I Master mix (Roche) and specific primers (see below) for PCR amplification in a LightCycler 480 System (Roche) as recommended by the manufacturer (Roche). The qRT-PCR data were analysed with LightCycler 480 Software version 1.3. The number of gene-specific cDNA copies was determined for each sample, averaged over three replicates and normalised to PP2A as described previously (Czechowski et al. 2005). Linearised plasmids encoding BRM, BRM Δ , ATSWI3C and a PP2A PCR product were used to generate standard curves. The gene-specific primers used for BRM splice variant profiling and measurements of BRM and ATSWI3C expression levels, as well as for PP2A (as described previously by Oh et al. 2007) are listed in Table S1. Final concentrations of qRT-PCR primers were 0.5 μM and the annealing temperature was set at 60°C in all assays. Each experiment was performed using at least two independent biological replicates, and the specificity of real-time PCR products was confirmed by melting curve analysis and electrophoresis on agarose gels.
Microscopic analyses
To characterise defects in stamen development, separated stamens were stained with 1% (w/v) acetoorcein for 30 min, washed in 80% (v/v) glycerol and examined under a light microscope (Leica Aristoplan). For anther sectioning and light microscopy, flowers were treated as described in Sarnowski et al. (2005). For scanning electron microscopy (SEM), flowers were fixed with 3% glutaraldehyde in 25 mM sodium phosphate buffer (pH 7.0) at 4°C overnight. The samples were rinsed with the same buffer and then fixed with 1% osmium tetroxide in phosphate buffer at 4°C overnight. After rinsing with phosphate buffer, the samples were dehydrated in graded ethanol and subsequently in acetone series. The material was then critical point dried in liquid carbon dioxide. Individual flowers were mounted on to SEM stubs. Pollen grains were dissected from anthers and mounted on SEM stubs without the above fixation steps. The mounted specimens were coated with gold and examined using a LEO 1430VP SEM at an accelerating voltage of 20 kV.
Accession numbers
Sequence data described in this article have been deposited at the GenBank/EMBL database under the accession number FJ168468 (BRM alternative splice variant).
Results
Characteristics of allelic series of BRM insertion and point mutations
In a previously described mutant screen (Ríos et al. 2002), we identified a new BRM T-DNA insertion mutant allele designated brm-6 (Fig. 1a). Characterisation of this new mutant allele revealed that brm-6 carries an inverted T-DNA repeat in exon 2, which replaces a target site deletion of 40 bp and is flanked by two T-DNA left border junctions. The T-DNA insertion in brm-6 has fused exon 2 to a short open reading frame carrying a stop codon 52 bp downstream of the 3′ T-DNA insert junction. RT-PCR analysis of the homozygous brm-6 line using primers 3 and 4 (Fig. 1a) revealed the absence of wild-type BRM transcript indicating that, like the previously characterised brm-1 allele (Fig. 1b; see below), brm-6 also corresponds to a null mutation. This is consistent with the results of Western-blot analysis showing that brm-6 plants, similarly to the brm-1 mutant, do not contain BRM protein (Fig. 1c).
The phenotype of brm-6 mutant was indistinguishable from that of brm-1 throughout all stages of plant development (Fig. S2). Both brm null mutants displayed a number of characteristic traits: complete male sterility, delayed seedling development, semi-dwarf growth habit and notably shorter root system showing increased growth of lateral roots when grown in sucrose containing MS medium. The rosette and cauline leaves of brm mutants were similarly distorted as a result of twisting along the proximodistal axis and downward curvature of leaf edges. In addition to the retardation of vegetative development, the flowers of brm-1 and brm-6 plants showed highly characteristic abnormalities, including the occurrence of fused sepals and stamen filaments, deformed and nonfused gynoecia, and retardation of anther development. Both brm mutants displayed strongly inhibited elongation of fruits resulting in very short siliques.
The positions of insertion and point mutations identified so far in the Arabidopsis BRM gene are depicted schematically in Fig. 1a. The brm-1 (SALK-030046) and brm-2 (GABI-kat-854D01) insertion mutations were originally identified by Hurtado et al. (2006). Similar to brm-4 (WiscDs/Lox436E9), which carries a transposon insertion (Tang et al. 2008), both brm-1 and brm-2 were shown to represent null mutations. In contrast, the brm-3 (SALK-088462) insertion mutant expresses a truncated BRM protein that lacks a C-terminal segment of 454 amino acids encompassing the bromodomain motif (Farrona et al. 2007). Point mutations brm-101, brm-102 and brm-103 were originally identified and designated atbrm-1, atbrm-2 and atbrm-3 by Kwon et al. (2006). These alleles correspond to nonsense mutations that presumably permit the synthesis of truncated protein products lacking various domains important for the biological function of BRM (Kwon et al. 2006). The brm-5 point mutation results in the exchange of Gly to Arg in the catalytic domain of BRM protein (Tang et al. 2008). We have compared brm-6 (identified in this study) and the published phenotypic analyses of all aforementioned mutant lines and found that the brm-1, -2, -4 and -6 insertion mutations and brm-101, -102 and -103 point mutations resulted in the manifestation of all highly reproducible brm phenotypic traits described above. The brm-3 and brm-5 mutants displayed less severe developmental alterations showing intermediate elongation and growth defects compared with wild-type and brm null mutant plants (Hurtado et al. 2006; Tang et al. 2008).
Genetic interactions between atswi3c and brm mutations
Phenotypic traits of brm null mutants showed striking similarity to developmental alterations observed previously in atswi3c insertion mutants (Sarnowski et al. 2005). As detection of interaction between BRM and ATSWI3C also suggested that ATSWI3C could be a core subunit of a BRM ATPase-associated SWI/SNF complex (Hurtado et al. 2006; Jerzmanowski 2007), we have systematically examined the features of the brm (At2g46020) and atswi3c (At1g21700) mutations by performing crosses of homo- or heterozygous brm-1 and brm-6 mutants with an atswi3c-1/+ and atswi3c-1 lines. Although homozygous brm-1 and brm-6 mutants failed to produce viable hybrid progeny when used as pollen donors in reciprocal crosses, they yielded some hybrid seed upon pollination when plants were grown under optimal long day (16 h light/8 h dark) conditions (i.e., 21–22°C with 60–75% humidity during the day and 18–20°C with 50–65% humidity during the night). However, when the average humidity level was below 60% and the day temperature exceeded 22°C, none of the examined homozygous brm mutants set seed. This indicated that female fertility of the brm mutants is highly sensitive to environmental conditions, such as mild drought stress. In contrast, male fertility of the brm mutants was extremely reduced independently of the growth conditions. In comparison, the atswi3c mutants displayed a lower frequency of aberrant stamen differentiation and pollen abortion (see below) and their female fertility was reduced but not fully abolished at lower humidity levels.
Analysis of F2 progeny of self-pollinated atswi3c-1/+ brm/+ F1 hybrids revealed that the phenotypes of the brm, atswi3c and double atswi3c brm lines were indistinguishable during early development. The observed segregation for wild-type and mutant traits significantly differed from the expected ratio of 9 wild-type to 7 mutant in each F2 family. For example, upon self-fertilisation an atswi3c-1/+ brm-6/+ F1 line segregated wild-type and mutant offspring at a ratio of approximately 5:2, for which the high χ2 value clearly indicated a significant deviation from the expected 9:7 complementary ratio (Table 1). Subsequent PCR genotyping of the mutant class showed 25 and 50% reduction in the number of expected homozygous brm-6 and atswi3c offspring, respectively, whereas the frequency of double mutant atswi3c-1 brm-6 class was about 30% lower than the expected value (4.3 instead of 6.25, Table 2). Similar results were obtained for F2 segregation of other atswi3c-1/+ brm-1/+ F1 hybrids (data not shown) indicating that both reduced female and male transmission of brm and atswi3c mutant alleles contributed to the lower than expected appearance of single and double mutant classes.
The analysis of simultaneously grown F2 populations provided suitable material for comparative characterisation of developmental defects observed in the single brm and atswi3c, and double atswi3c brm mutants (Table 3; Fig. 2). As all traits of brm-1 and brm-6 lines were identical, we shall not refer to specific brm alleles from here onwards. As mentioned above, the appearance of leaf rosette of young (26-day-old) atswi3c, brm and atswi3c brm plants was very similar (Table 3; Fig. 2a) and characterised by twisting and downward curvature of leaves (Fig. 2d). The appearance of cauline leaves was also indistinguishable in all mutants (Fig. 2e). Under long day conditions, the number of rosette leaves was very similar at the onset of flowering (Table 3). Both single and double mutant classes flowered slightly earlier than wild-type based on their rosette leaf number, the difference being statistically significant (P < 0.01). However, due to their overall slower development, the mutant lines flowered on average 5 days later than control wild-type plants (Table 3).
Comparative analysis of atswi3c, brm, and atswi3c brm double mutants grown under identical long day conditions. a Rosette phenotype of wt, atswi3c-1, brm-1, brm-6 and atswi3c-1 brm-6. b–c Comparison of adult wild-type and mutant plants grown under suboptimal (average humidity level below 60% and the day temperature exceeding 22°C, b or optimal (21–22°C with 60–75% humidity during the day and 18–20°C with 50–65% humidity during the night, c, conditions. d Rosette leaves of wild-type and mutant plants. e Cauline leaves of wild-type and mutant plants. f Siliques of wild-type and mutant plants at 12 days after pollination. g Flowers of wild-type and mutant plants. Flowers of atswi3c-1, brm, and atswi3c-1 brm mutants display similar developmental defects, including fused stamen filaments (h), staminoid filaments (i), stamens with petaloid tissues in place of anthers (petaloid stamens, j), and fused sepals (k). Some sepals and petals were removed to show the abnormalities. l Scanning electron micrographs showing open gynoecia in brm-1 and atswi3c-1 brm-1 flowers. Bars 1 cm (d, e), 5 mm (f), 1 mm (g–k)
When compared with optimal growth conditions (Fig. 2c, see above), the brm and atswi3c brm mutants showed retarded development in response to lower humidity (45–55%) and changes in temperature range between 21 and 25°C (Fig. 2b). Under optimal conditions, flowers of single atswi3c and brm and double atswi3c brm mutants displayed similar defects, including occasional (1–5%) occurrence of fused stamens and staminoid petal filaments, fused sepals and petaloid tissues in place of anthers (Fig. 2g–k). Interestingly, under suboptimal growth conditions the frequency of aberrantly differentiating flower organs was significantly increased (30% or higher) in the brm and atswi3c brm mutants compared with atswi3c. Also, the frequency of flowers carrying open gynoecia (Fig. 2l) was as high as 15% in brm and brm/atswi3c but was <1% in atswi3c mutant flowers under suboptimal conditions. A high frequency (15%) of opened gynoecia was also observed previously by Hurtado et al. (2006) in the brm-1 and brm-2 mutants.
The most notable and growth condition-independent difference between the single and double mutant lines was that all lines homozygous for either brm-1 or brm-6 mutation displayed male sterility. Compared with wild-type Arabidopsis, the atswi3c mutant developed shorter siliques, which contained viable seeds. Remarkably, both brm and atswi3c brm lines (Fig. 2f) also showed the initiation of dwarfed siliques, but produced no seed (Table 3). Thus, the unique sterility trait of brm mutants appeared to be an additive character in the atswi3c brm double mutants. Here, we note that although Hurtado et al. (2006) reported both female and male sterility of brm-1 and brm-2 mutants, we observed that pollination of homozygous brm-6 null mutant with either atswi3c or wild-type pollen yielded viable progeny (see above). Given the aforementioned high stress sensitivity of brm flower phenotype, this contradiction is probably due to differences in growth conditions used here and by Hurtado et al. (2006).
Scanning electron microscopy examinations revealed that even the highly deformed open gynoecia of brm and atswi3c brm plants contained 14% of normally developing ovules (Fig. 3a). Nonetheless, consistent with a possible role of BRM in ovule development, most ovules in open gynoecia of brm and atswi3c brm plants showed aberrant differentiation mostly affecting the initiation and growth of integument. This resulted in the lack of the characteristic bent (Fig. 3b, c), outgrowth of the outer integument (resembling defects observed in sup mutants; Meister et al. 2002; Fig. 3d), and inner integument protrusion (resembling defects observed in the tsl mutant; Roe et al. 1997; Fig. 3e, f). Some of the ovules did not form any integument but developed instead into finger-like protrusions (Fig. 3b, e). In addition, leaf-like structures that arose from placental tissue, which resembled those seen in ap2-6/bel1-3 double mutants, were also found (Fig. 3f; Western and Haughn 1999). Although we did not analyse in similar detail the defects in ovule development observable in a small fraction of atswi3c plants, the notably higher frequency of these defects in the brm and atsw3c brm mutants suggests that BRM is more critical than ATSWI3C for carpel and ovule development.
Scanning electron micrographs of ovule types found in open gynoecia of brm and atswi3c-1 brm mutants. a Normal wild-type-like ovule. b Ovule lacking integuments (arrow) and an ovule with altered integument growth lacking the characteristic bent (arrowhead). c Ovule with irregular integument growth resembling ovules of ap2-6/bel1-3 double mutant. The two integuments (arrowheads) are visible only on one side of the nucellus. d A sup-like ovule showing outer integument outgrowth. e tsl-1-like ovules showing inner integument protrusion (arrow) and ovules developed into finger-like protrusions (arrowhead). f Fused tsl-1-like ovules converted into a leaf-like structure
Comparison of pollen maturation in the atswi3c and brm mutants
In order to understand the cause of male sterility conferred by the brm mutations, we compared the development of anthers and pollen in the brm and atswi3c mutants. Both atswi3c and brm plants were reported to have less stamens than wild-type plants (Sarnowski et al. 2005; Hurtado et al. 2006). The typical appearance of anthers in an atswi3c-1 flower is shown in Fig. 4b. Although atswi3c anthers contained less pollen compared with wild-type anthers (Fig. 4a, b), the shape and size of atswi3c pollen grains was similar to normal. However, a proportion of atswi3c pollen grains appeared to be glued together and showed partial deformation of their walls (Fig. 4g, j). In contrast, brm plants carried only about 20% of atswi3c-1- like anthers (Fig. 4c). The remaining 80% of brm anthers displayed severe deformations roughly half of them carrying coalesced pollen material (Fig. 4d) and half showing no intact pollen grains (Fig. 4e). Compared with atswi3c, the rare pollen grains that could be observed in the brm anthers were highly deformed (compare Fig. 4g, j with Fig. 4 h, k). In general, mature pollen sacs in brm anthers either had no pollen or contained abnormal pollen grains that could not be easily released. In conclusion, compared with atswi3c, the brm mutations resulted in more severe anther development and pollen maturation defects, which are fully consistent with complete male sterility of brm and atswi3c brm mutants. Despite aforementioned specific effects of the brm mutation on differentiation of reproductive organs, all other phenotypic traits of brm, atswi3c and atswi3c brm mutants proved to be indistinguishable. This, together with the genetic data, fully supports the model that BRM and ATSWI3C act in the same regulatory complex. On the other hand, differences between the severity of differentiation defects in reproductive organs in the brm and atswi3c mutants suggests that BRM may also have some unique functions.
Comparison of defects of anther and pollen development in the atswi3c and brm mutants. Acetoorcein staining of stamens of wild-type (a), atswi3c-1 (b) and brm (c–e) flowers. The brm mutants display defects, which are either identical to those of atswi3c-1 (c) or much stronger, such as coalescence of pollen (d) or lack of pollen grains (e). The picture to the right in (d) is a light micrograph of a cross-section through a brm mutant anther. f–h Scanning electron micrographs showing deformed pollen grains in the anthers of atswi3c-1 and brm mutants. Note that brm pollen grains are more abnormal and display a more circular shape. i–k Scanning electron micrographs of pollen grains dissected from anthers at higher magnification. Pollen grains of brm and atswi3c-1 appear glued together
Comparison of transcription profiles of ATSWI3C and BRM
If ATSWI3C and BRM function in the same putative CRC, one would also expect that the transcription of their genes is co-regulated in all organs and stages of plant development that are similarly affected by the brm and atswi3c mutations. To confirm this assumption, we inspected the transcript profiling data publicly available in the Genevestigator (Zimmermann et al. 2005) and AtGenExpress (Schmid et al. 2005) databases. This indicated that the patterns of ATSWI3C and BRM transcription are indeed very similar in most organs and developmental stages examined, but the levels of BRM transcript are consistently slightly higher (Fig. S3). To verify the transcript profiling data, the levels of ATSWI3C and BRM transcripts in different organs were compared using quantitative real-time PCR (qRT-PCR) with gene-specific primers (Fig. 5). In full agreement with the Genevestigator and AtGenExpress databases, our data indicated that ATSWI3C and BRM are transcribed ubiquitously in all organs tested. The BRM transcript level was 1.5-fold higher than that of ATSWI3C in all organs. These results were also consistent with a previous RT-PCR study of ATSWI3C transcript levels by Bezhani et al. (2007).
Alternative splicing of BRM transcript
In our RT-PCR studies using different BRM primers, we observed the existence of two splicing isoforms of BRM mRNA (Fig. 6a). Compared with the major BRM mRNA, carrying all 14 exons and encoding the full-size BRM protein, the alternatively spliced transcript contained truncated exon 9 sequences due to the use of a different 3′ splice site within this exon. This alternatively spliced BRM transcript is only 109 nucleotides shorter but contains a premature translation stop codon (PTC) in the coding sequence of N-terminal segment of SNF2 ATPase domain (Fig. 6c, d). The alternatively spliced BRM transcript could be detected by RT-PCR in all tissues examined (Fig. 6b). However, qRT-PCR analysis showed that in different tissues of wild-type plants the BRM Δ transcript was maintained at very low level, not exceeding 1–1.5% of that of main BRM splice variant. To examine whether any other splicing isoforms of BRM transcript were present in Arabidopsis, we performed northern RNA hybridisations using total RNA from wild-type Col-0 plants and two different probes complementary to 5′ and 3′ sequences of the BRM gene. This analysis did not resolve the PTC-containing alternatively spliced mRNA isoform, which has a size comparable to that of the major BRM transcript, and failed to reveal any other shorter splice isoforms (results not shown).
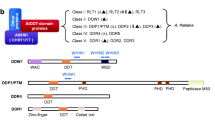
Alternative splicing of the BRM transcript. a Exon–intron structure of the BRM gene: exons are indicated by grey rectangles and introns by black lines. Enlarged section shows the alternatively spliced region with the 5′ (GU) and additional 3′ (AG) splice site located in exon 9. b RT-PCR analysis of alternatively spliced BRMΔ transcript in different organs of wild-type plants. The UBQ11 (At4g05050) transcript was used as an internal standard. The positions of primers used for RT-PCR analysis are shown on the diagram to the right. c Schematic representation of the BRM Δ splice isoform. The alternatively spliced mRNA variant (upper line) is only 109 bases shorter than the major mRNA isoform and carries a premature translation stop codon (UGA) in the region encoding the ATPase domain. d Positions of functional domains in the BRM protein indicated as described by Knizewski et al. (2008). The upper line illustrates the hypothetical truncated protein product of the alternatively spliced BRM transcript
Premature termination codon-containing transcripts are known to be substrates for degradation via the nonsense-mediated decay (NMD) pathway (Stalder and Mühlemann 2008). To determine whether the alternatively spliced BRM Δ mRNA was a target for NMD, we have analysed transcript levels in the Arabidopsis upf1-5 and upf 3-1 mutants that are defective in key factors of the NMD pathway and therefore accumulate PTC-containing transcripts (Hori and Watanabe 2005; Arciga-Reyes et al. 2006). Consistently with some previous data (Arciga-Reyes et al. 2006), we found by RT-PCR analysis that the level of PTC-containing BRM Δ transcript was 2- to 3-fold higher in the NMD-deficient mutants (Fig. 7) indicating that the BRM Δ splice variant was indeed an NMD substrate.
Possible role of NMD pathway in the regulation of BRM Δ levels. Transcript levels of full-length BRM and truncated BRM Δ splice variant were assayed by qRT-PCR in different organs of upf1-5 and upf3-1 mutants and wild-type plants. Because of low amounts of BRM Δ compared with the major BRM mRNA isoform, the expression levels are shown as logarithm of number of cDNA copies per 200 copies of PP2A. The figure shows the mean expression values from six replicates ±SD
Discussion
Recently, several Arabidopsis loci encoding putative homologues of conserved subunits of yeast and human SWI/SNF-type CRCs have been identified and characterised by the help of T-DNA insertion and EMS-induced point mutations (Jerzmanowski 2007; Kwon and Wagner 2007). During our work on functional characterisation of different Arabidopsis SWI3 homologues, we observed that the phenotypic traits of atswi3c insertion mutants are very similar to those of brm mutations that inactivate BRAHMA, the only Arabidopsis SNF2-like ATPase that carries a C-terminal bromodomain for interaction with acetylated histones (Farrona et al. 2004). Both atswi3c and brm mutants display unique and characteristic developmental alterations including semi-dwarf appearance, shortened and branched root system, twisted rosette and cauline leaves, downward curling of leaf edges, defects in proper differentiation of several flower organs, and shortened curved siliques (Sarnowski et al. 2005; Hurtado et al. 2006; Kwon et al. 2006). Furthermore, transcript levels of flower homeotic genes are changed similarly in both brm and atswi3c mutants (Sarnowski et al. 2005; Hurtado et al. 2006). These phenotypic similarities were particularly striking because mutations in none of the three other Arabidopsis ATSWI3 genes appeared to produce developmental changes that resembled the phenotypes of brm (Sarnowski et al. 2005) or syd mutations affecting SNF2-type ATPase homologues (Wagner and Meyerowitz 2002).
To genetically challenge the hypothesis that BRM and ATSWI3C act in a single complex, we constructed and characterised Arabidopsis lines with double atswi3c brm mutations. The double mutants did not reveal any neomorph phenotype but displayed all characteristic traits of the two single mutants. This genetic evidence suggests that ATSWI3C and BRM are functionally interdependent and act in a single functional unit, in which elimination of either of the two partners renders the unit inactive. This conclusion is also supported by our qRT-PCR data indicating that BRM and ATSWI3C are co-expressed. Moreover, we found that the ratio of BRM and ATSWI3C transcripts is similar in various organs suggesting transcriptional co-regulation of these genes.
Similarities between the phenotypes conferred by null mutations of SWI3 and bromodomain-containing ATPases have also been reported in yeast (Peterson and Herskowitz 1992) and mammals (Kim et al. 2001), and functional interaction of these proteins was demonstrated by isolation and characterisation of SWI2/SNF2 CRCs (Mohrmann and Verrijzer 2005). In yeast two-hybrid assays, Farrona et al. (2004) found that the N-terminal region of Arabidopsis BRM interacts with ATSWI3C. Previously, it has been reported that BRM interacts with AtSWI3B but not with ATSWI3A, and unlike ATSWI3B, neither ATSWI3C nor BRM can interact with SNF5/BSH in yeast two-hybrid assays (Farrona et al. 2004; Sarnowski et al. 2005; Hurtado et al. 2006; our unpublished results). As Arabidopsis SWI/SNF complexes harbour two SWI3-type subunits, these observations suggest that ATSWI3B is probably the second SWI3-type subunit of BRM and ATSWI3C-containing Arabidopsis CRCs.
The genetic data described above indicate that compared with atswi3c, the brm mutation results in some unique flower developmental defects that appear as additive traits in the atswi3c brm double mutants. One of the traits conferred by the brm mutation is complete male sterility, which is fully manifested independently of environmental stress in the double mutants. Male sterility of the brm mutants correlates with more severe defects of pollen development in comparison with the atswi3c mutants. Other brm-specific traits, such as slower development, higher degree of dwarfism, and increased frequency of open gynoecia appear only under suboptimal growth conditions and are likely to represent stress-related regulatory functions of BRM. The ovule defects occurring at high frequency in the brm mutants resemble the effects of mutations of BEL1, SUP, TSL and other key genes controlling ovule development. This suggests possible regulatory interactions involving these genes that require BRM-mediated chromatin remodelling. TSL was recently shown to be involved in chromatin modifications (Wang et al. 2007). Interestingly, the BRAHMA SWI/SNF CRC in Drosophila melanogaster was found to act together with a histone chaperone ASF1, which is a target for a TSL-like kinase (Moshkin et al. 2002).
There are two possible explanations for additional effects of the brm mutations. The first is that BRM may perform ATSWI3C-independent regulatory functions, acting either alone or as a subunit of an alternative complex. In this respect, it is relevant that small differences were also observed in the phenotypes conferred by knockout mutations in Brg1 ATPase (a homologue of BRM) and Srg3 (a homologue of SWI3) in mice. In addition, Srg3 shows differential expression in various organs suggesting extra regulatory functions besides those it fulfils as a subunit of the SWI/SNF complex (Kim et al. 2001). Alternatively, it is also plausible that the inactivation of ATSWI3C is compensated by one of the other AtSWI3 subunits expressed during certain developmental stages, such as floral organ differentiation, whereas the lack of BRM cannot be compensated by any other Snf2-type ATPases in Arabidopsis.
In this work, we also show that the BRM pre-mRNA undergoes alternative splicing which results in the production of a PTC-containing splice isoform. A 2–3-fold upregulation of the level of this BRM Δ splice variant in upf1-5 and upf3-1 mutants suggests that the level of this transcript is controlled by the NMD pathway. NMD has been shown not only to degrade aberrant transcripts but also to regulate the steady-state level of many mRNAs involved in numerous cellular processes, such as DNA repair, cell cycle and metabolism (Rehwinkel et al. 2005; Stalder and Mühlemann 2008). The answer to the question whether the detected alternative splicing event may be used in the regulation of BRM, for example in response to stress or other signals (Palusa et al. 2007) requires further studies.
Abbreviations
- ATSWI3C:
-
Arabidopsis homologue of SWI3, a subunit of the yeast SWI/SNF complex
- BRM:
-
BRAHMA, Arabidopsis Snf2 family protein
- SYD:
-
SPLAYED, Arabidopsis Snf2 family protein
- SWI/SNF:
-
SWItch/Sucrose NonFermentable, a yeast nucleosome remodelling complex
- SNF5:
-
Subunit of SWI/SNF chromatin remodelling complex
- NMD:
-
Nonsense-Mediated Decay
References
Arciga-Reyes L, Wootton L, Kieffer M, Davies B (2006) UPF1 is required for nonsense-mediated mRNA decay (NMD) and RNAi in Arabidopsis. Plant J 47:480–489
Bezhani S, Winter C, Hershman S, Wagner JD, Kennedy JF, Kwon CS, Pfluger J, Su Y, Wagner D (2007) Unique, shared, and redundant roles for the Arabidopsis SWI/SNF chromatin remodeling ATPases BRAHMA and SPLAYED. Plant Cell 19:403–416
Brzeski J, Podstolski W, Olczak K, Jerzmanowski A (1999) Identification and analysis of the Arabidopsis thaliana BSH gene, a member of the SNF5 gene family. Nucleic Acids Res 27:2393–2399
Czechowski T, Stitt M, Altmann T, Udvardi MK, Scheible WR (2005) Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol 139:5–17
Farrona S, Hurtado L, Bowman JL, Reyes JC (2004) The Arabidopsis thaliana SNF2 homolog AtBRM controls shoot development and flowering. Development 131:4965–4975
Farrona S, Hurtado L, Reyes JC (2007) A nucleosome interaction module is required for normal function of Arabidopsis thaliana BRAHMA. J Mol Biol 373:240–250
Gendrel AV, Lippman Z, Martienssen R, Colot V (2005) Profiling histone modification patterns in plants using genomic tiling microarrays. Nat Methods 3:213–218
Hori K, Watanabe Y (2005) UPF3 suppresses aberrant spliced mRNA in Arabidopsis. Plant J 43:530–540
Hurtado L, Farrona S, Reyes JC (2006) The putative SWI/SNF complex subunit BRAHMA activates flower homeotic genes in Arabidopsis thaliana. Plant Mol Biol 62:291–304
Jerzmanowski A (2007) SWI/SNF remodeling and linker histones in plants. Biochim Biophys Acta 1769:330–345
Kim JK, Huh SO, Choi H, Lee KS, Shin D, Lee C, Nam JS, Kim H, Chung H, Lee HW, Park SD, Seong RH (2001) Srg3, a mouse homolog of yeast SWI3, is essential for early embryogenesis and involved in brain development. Mol Cell Biol 21:7787–7795
Knizewski L, Ginalski K, Jerzmanowski A (2008) Snf2 proteins in plants: gene silencing and beyond. Trends Plant Sci 13:557–565
Koncz C, Martini N, Szabados L, Hrouda M, Bachmair A, Schell J (1994) Specialized vectors for gene tagging and expression studies. In: Gelvin S, Schilperoort B (eds) Plant molecular biology manual, vol.B2. Kluwer, Dordrecht, pp 1–22
Kwon SB, Wagner D (2007) Unwinding chromatin for development and growth: a few genes at a time. Trends Genet 23:403–412
Kwon CS, Hibara K, Pfluger J, Bezhani S, Matha H, Aida M, Tasaka M, Wagner D (2006) A role for chromatin remodeling in regulation of CUC gene expression in the Arabidopsis cotyledon boundary. Development 133:3223–3230
Lessard J, Wu JI, Ranish JA, Wan M, Winslow MM, Staahl BT, Wu H, Aebersold R, Graef IA, Crabtree GR (2007) An essential switch in subunit composition of a chromatin remodeling complex during neural development. Neuron 55:201–215
Martens JA, Winston F (2003) Recent advances in understanding chromatin remodeling by Swi/Snf complexes. Curr Opin Genet Dev 13:136–142
Meister RJ, Kotow LM, Gasser CS (2002) SUPERMAN attenuates positive INNER NO OUTER autoregulation to maintain polar development of Arabidopsis ovule outer integuments. Development 129:4281–4289
Mohrmann L, Verrijzer CP (2005) Composition and functional specificity of SWI2/SNF2 class chromatin remodeling complexes. Biochim Biophys Acta 1681:59–73
Moshkin YM, Armstrong JA, Maeda RK, Tamkun JW, Verrijzer P, Kennison JA, Karch F (2002) Histone chaperone ASF1 cooperates with the Brahma chromatin-remodelling machinery. Gen Dev 16:2621–2626
Oh E, Yamaguchi S, Hu J, Yusuke J, Jung B, Paik I, Lee HS, Sun TP, Kamiya Y, Choi G (2007) PIL5, a phytochrome-interacting bHLH protein, regulates gibberellin responsiveness by binding directly to the GAI and RGA promoters in Arabidopsis seeds. Plant Cell 19:1192–1208
Palusa SG, Ali GS, Reddy AS (2007) Alternative splicing of pre-mRNAs of Arabidopsis serine/arginine-rich proteins: regulation by hormones and stresses. Plant J 49:1091–1107
Peterson CL, Herskowitz I (1992) Characterization of the yeast SWI1, SWI2 and SWI3 genes, which encode a global activator of transcription. Cell 68:573–583
Rehwinkel J, Letunic I, Raes J, Bork P, Izaurralde E (2005) Nonsense-mediated mRNA decay factors act in concert to regulate common mRNA targets. RNA 11:1530–1544
Ríos G, Lossow A, Hertel B, Breuer F, Schaefer S, Broich M, Kleinow T, Jásik J, Winter J, Ferrando A, Farrás R, Panicot M, Henriques R, Mariaux J-B, Oberschall A, Molnár G, Berendzen K, Shukla V, Lafos M, Koncz Z, Rédei GP, Schell J, Koncz C (2002) Rapid identification of Arabidopsis insertion mutants by nonradioactive detection of T-DNA tagged genes. Plant J32:243–253
Roberts CWM, Orkin SH (2004) The SWI/SNF complex: chromatin and cancer. Nat Rev Cancer 4:133–142
Roe JL, Nemhauser JL, Zambryski PC (1997) TOUSLED participates in apical tissue formation during gynoecium development in Arabidopsis. Plant Cell 9:335–353
Saha A, Wittmeyer J, Cairns BR (2006) Chromatin remodelling: the industrial revolution of DNA around histones. Mol Cell Biol 7:437–447
Sarnowski T, Świeżewski S, Pawlikowska K, Kaczanowski S, Jerzmanowski A (2002) AtSWI3B, an Arabidopsis homolog of SWI3, a core subunit of yeast Swi/Snf chromatin remodeling complex, interacts with FCA, a regulator of flowering time. Nucleic Acids Res 30:3412–3421
Sarnowski T, Ríos G, Jasik J, Świeżewski S, Kaczanowski S, Kwiatkowska A, Pawlikowska K, Koźbiał M, Koźbiał P, Koncz C, Jerzmanowski A (2005) SWI3 subunits of putative SWI/SNF chromatin remodeling complex play distinct roles during Arabidopsis development. Plant Cell 17:2454–2472
Schmid M, Davison TS, Henz SR, Pape UJ, Demar M, Vingron M, Schölkopf B, Weigel D, Lohmann JU (2005) A gene expression map of Arabidopsis thaliana development. Nat Genet 37:501–506
Smith CL, Peterson CL (2005) ATP-dependent chromatin remodeling. Curr Top Dev Biol 65:115–148
Stalder L, Mühlemann O (2008) The meaning of nonsense. Trends Cell Biol 18:315–321
Su Y, Kwon CS, Bezhani S, Huvermann B, Chen C, Peragine A, Kennedy JF, Wagner D (2006) The N-terminal ATPase AT-hook-containing region of the Arabidopsis chromatin-remodeling protein SPLAYED is sufficient for biological activity. Plant J 46:685–699
Tang X, Hou A, Babu M, Nguyen V, Hurtado L, Lu Q, Reyes JC, Wang A, Keller WA, Harada JJ, Tsang EWT, Cui Y (2008) The Arabidopsis BRAHMA chromatin remodeling ATPase is involved in repression of seed maturation genes in leaves. Plant Physiol 147:1143–1157
Tyler L, Thomas SG, Hu J, Dill A, Alonso JM, Ecker JR, Sun TP (2004) Della proteins and gibberellin-regulated seed germination and floral development in Arabidopsis. Plant Physiol 135:1008–1019
Verbsky ML, Richards EJ (2001) Chromatin remodeling in plants. Curr Opin Plant Biol 4:494–500
Wagner D, Meyerowitz EM (2002) SPLAYED, a novel SWI/SNF ATPase homolog, controls reproductive development in Arabidopsis. Curr Biol 12:85–94
Wang Y, Liu J, Xia R, Wang J, Shen J, Cao R, Hong X, Zhu JK, Gong Z (2007) The protein kinase TOUSLED is required for maintenance of transcriptional gene silencing in Arabidopsis. EMBO Rep 8:77–83
Western TL, Haughn GW (1999) BELL1 and AGAMOUS genes promote ovule identity in Arabidopsis thaliana. Plant J 18:329–336
Zimmermann P, Hennig L, Gruissem W (2005) Gene expression analysis and network discovery using Genevestigator. Trends Plant Sci 10:407–409
Acknowledgments
We thank M. Kuras and M. Sobolewska (University of Warsaw, Poland) for assistance with microscopy and Ingrid Reintsch and Sabine Schäfer (Max-Planck Institut für Züchtungsforschung, Germany) for excellent technical assistance, and A. Jarmolowski (University of Poznan, Poland) for upf3-1 homozygous seeds. This work was supported by the Deutsche Forschungsgemeischaft (DFG) SFB635 and AFGN grants for C.K., a Marie-Curie Intra-European Fellowship grant (PIEF-GA-2008-220291) for T.J.S. and by Ministerstwo Nauki i Szkolnictwa Wyzszego (MNiSW) grants: N302 060434 for R.A., N301 034 31/1151 for T.J.S., PBZ-MNiI-2/1/2005 for M.P-B. and T.J.S., and PO4A03928 for A.J.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
R. Archacki, T. J. Sarnowski, C. Koncz, and A. Jerzmanowski contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Archacki, R., Sarnowski, T.J., Halibart-Puzio, J. et al. Genetic analysis of functional redundancy of BRM ATPase and ATSWI3C subunits of Arabidopsis SWI/SNF chromatin remodelling complexes. Planta 229, 1281–1292 (2009). https://doi.org/10.1007/s00425-009-0915-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-009-0915-5