Abstract
A specific signaling role for H2O2 in Chlamydomonas reinhardtii was demonstrated by the definition of a promoter that specifically responded to this ROS. Expression of a nuclear-encoded reporter gene driven by this promoter was shown to depend not only on the level of exogenously added H2O2 but also on light. In the dark, the induction of the reporter gene by H2O2 was much lower than in the light. This lower induction was correlated with an accelerated disappearance of H2O2 from the culture medium in the dark. Due to a light-induced reduction in catalase activity, H2O2 levels in the light remained higher. Photosynthetic electron transport mediated the light-controlled down-regulation of the catalase activity since it was prevented by 3-(3′4′-dichlorophenyl)-1,1-dimethylurea (DCMU), an inhibitor of photosystem II. In the presence of light and DCMU, expression of the reporter gene was low while the addition of aminotriazole, a catalase inhibitor, led to a higher induction of the reporter gene by H2O2 in the dark. The role of photosynthetic electron transport and thioredoxin in this regulation was investigated by using mutants deficient in photosynthetic electron flow and by studying the correlation between NADP-malate dehydrogenase and catalase activities. It is proposed that, contrary to expectations, a controlled down-regulation of catalase activity occurs upon a shift of cells from dark to light. This down-regulation apparently is necessary to maintain a certain level of H2O2 required to activate H2O2-dependent signaling pathways.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Life in an oxygen-rich atmosphere has to cope with the danger of oxidative stress. Plants are particularly exposed to oxidative stress caused by photosynthetic processes. Reactive oxygen species (ROS) are produced during normal cell metabolism but their production is drastically enhanced when plants are exposed to stresses such as high light, low temperatures, or drought, or combinations thereof. Due to their high reactivity, ROS can damage macromolecules essential for the integrity of the cell such as lipids, proteins, and nucleic acids. Thus, the concentration and accumulation of ROS needs to be strictly controlled. For this purpose, cells have evolved a variety of very efficient non-enzymatic (e.g., ascorbate, glutathione, tocopherols) and enzymatic (e.g., catalase, ascorbate peroxidase, superoxide dismutase, peroxiredoxins, thioredoxins, glutaredoxins) antioxidant systems (Apel and Hirt 2004; Foyer and Noctor 2005). The requirement for a control of ROS accumulation implicates that these reactive intermediates are sensed by cells in order to activate stress responses allowing adaptation to a new set of environmental conditions. It is now widely accepted that ROS, and especially H2O2, play a major role in cellular signaling pathways and regulation of gene expression in a wide variety of organisms including plants (Apel and Hirt 2004; Laloi et al. 2004). Several studies have used microarray analyses to study alterations in gene expression in response to ROS (Gadjev et al. 2006 and references therein). These studies have demonstrated that increased ROS levels alter the expression of a rather large set of genes (up to one-third of the genome). They also have shown that different ROS have specific signaling properties; a result supported by the recent discovery that H2O2 and singlet oxygen employ specific targets for the activation of an ROS-responsive promoter (Shao et al. 2007). The mechanisms that allow transduction of ROS signals to the nucleus remain unknown. In plants, the signaling pathways mediated by ROS involve heterotrimeric G-proteins (Joo et al. 2005) and protein phosphorylation mediated by MAP kinases and protein phosphatases (Kovtun et al. 2000; Gupta and Luan 2003; Rentel et al. 2004).
Under conditions of intracellular H2O2 production or treatment with exogenous H2O2, several plant H2O2 detoxifying enzymes, such as catalases and ascorbate peroxidases, are activated at the transcript, protein, and/or activity levels (Gechev et al. 2002; Vandenabeele et al. 2004; Yabuta et al. 2004; Davletova et al. 2005). Several studies have also shown that some of these enzymes may be inactivated by H2O2 (Miyake and Asada 1996; Shikanai et al. 1998; Hiner et al. 2000; Miyake et al. 2006).
Ascorbate peroxidases (APX) are the main H2O2 detoxifying enzymes in plants. Several isoforms of APX exist in plants and are located in the chloroplasts, the cytosol and peroxisomes (Shigeoka et al. 2002; Panchuk et al. 2005; Teixeira et al. 2006). Incubation of tobacco cell cultures in the presence of an H2O2 generating system (glucose + glucose oxidase) has been shown to result in inhibition of cytosolic APX activity (De Pinto et al. 2006). Also the endogenous generation of H2O2 via photosynthetic electron transfer resulted in inhibition of chloroplastic APX activity in tobacco plants (Miyake et al. 2006). APX were recently found among new putative targets of thioredoxins (TRX) identified by proteomic studies (Marchand et al. 2004; Wong et al. 2004; Yamazaki et al. 2004). Moreover, purified recombinant cytosolic APX from poplar was shown to be inhibited by reduced cytosolic thioredoxin h, reduced glutathione, and dithiothreitol. The molecular mechanism of this inhibition though remains to be determined (Gelhaye et al. 2006).
In plants, also catalases and peroxiredoxins play an important role in detoxifying H2O2 (for review see Feierabend 2005; Dietz et al. 2006). Light inactivation of catalase has been observed for enzymes of both plant and animal origin. Different isoforms of plant catalases show different sensitivities towards light (Grotjohann et al. 1997; Engel et al. 2006). The action spectrum of photoinactivation is identical to the absorption spectrum of catalase, indicating that the light inactivation of catalases is caused by direct light absorption by the heme moieties. However, when irradiation was performed in the presence of chloroplasts the in vitro inactivation of catalase was enhanced. Under these conditions red light which is not absorbed by catalase but by photosynthetic pigments is sufficient to inhibit catalase (Feierabend and Engel 1986).
Catalases from Chlamydomonas, wheat, spinach, pea and potato were found among the potential targets of thioredoxin (TRX) identified by proteomic studies (Balmer et al. 2004; Lemaire et al. 2004; Maeda et al. 2004; Wong et al. 2004; Michelet et al. 2006). TRX are small disulfide oxidoreductases which play a major role in light signaling and in the oxidative stress responses (Vieira Dos Santos and Rey 2006; Lemaire et al. 2007). We have previously shown that in total soluble extracts of Chlamydomonas, the activity of catalase could be modulated by cytosolic thioredoxin h: a 50% inhibition in catalase activity was observed after treatment with the cytosolic TRX system (NADPH, NADPH-thioredoxin reductase, TRX h) which could be mimicked by treatment with the strong reductant dithiothreitol (DTT) (Lemaire et al. 2004).
Hence, the question arises whether a down-regulation of the activity of ROS detoxifying enzymes, like APX and catalases, might be necessary to allow diffusion and perception of H2O2 under stress conditions that would result in the activation of genes that allow acclimation to changes in the environment. In order answer this question, we investigated whether photosynthetic electron transfer may be involved in the regulation of H2O2 detoxifying enzymes and thus the regulation of gene expression in response to H2O2. As a model system, we have chosen Chlamydomonas reinhardtii which was transformed with a construct containing a Renilla reniformis luciferase reporter gene under the control of an HSP70A promoter fragment that specifically responds to H2O2 (Shao et al. 2007).
Materials and methods
Algal strains and culture conditions
Chlamydomonas reinhardtii strains 325 (CW15, arg7-8) and D66 (CW15) were kindly provided by R. Matagne (University of Liège, Belgium) and Arthur Grossman (Carnegie Institute, Stanford, CA, USA), respectively. The wild-type strains CC-124 (mt−) and WT34 (mt+) (a derivative of strain 137c) were from the Chlamydomonas Genetics Center (Duke University, Durham, NC, USA) and the Institut de Biologie Physico-Chimique in Paris, respectively. Mutants ΔpetD (Kuras and Wollman 1994) and rbcL lacking both, the small and large subunit of Rubisco (rbcL-18-5B, Spreitzer et al. 1985) were kindly provided by Olivier Vallon (IBPC, Paris). Cultures were grown photomixotrophically in Tris, acetate, phosphate (TAP) medium (Harris 1989) on a rotary shaker at 23–25°C under continuous irradiation with white light (70 μmol photons m−2 s−1). TAP medium was supplemented with 100 mg l−1 of arginine when required.
Nuclear transformation of Chlamydomonas
Plasmids used for Chlamydomonas transformation were purified by PEG precipitation. Prior to transformation, the plasmid containing the ARG7 gene (pCB412) was linearized by EcoRI digestion. Plasmids containing the reporter construct, P HSP70A-ΔHSE -LUC (Shao et al. 2007) were linearized by ScaI digestion. Chlamydomonas reinhardtii strain 325 was cotransformed with pCB412 (selection plasmid) and the plasmid containing the P HSP70A-ΔHSE -LUC reporter construct using the glass beads method (Kindle 1990). For transformation, cells were grown to 3–5 × 106 cells ml−1 and concentrated to 3 × 108 cells ml−1. 1 × 108 cells were mixed with 500 ng of linearized LUC reporter plasmid, 100 ng of linearized pCB412, 0.3 g of acid-washed glass beads, and 100 μl of 10% PEG 6000. Immediately after vortexing for 20 s, cells were spread onto TAP-agar (1% agar) plates for the selection of arginine autotrophic clones. Transformants that harbored the LUC constructs were detected by luciferase assay.
Bioluminescence assay
Bioluminescence assays were performed essentially as described by Minko et al. (1999) at room temperature with a luminescence counter (MicroBeta TriLux) set in “flash” mode with single auto-injection. After sampling, cells were spun down, resuspended in the same volume of sample buffer [1.5 mM Tris-HCI (pH 7.8), 1 mM EDTA], and frozen at −80°C for at least 20 min. After thawing, 20 μl were transferred to 96-well microtiter plates and 125 μl assay buffer [0.1 M K2HPO4 (pH 7.6); 0.5 M NaCl; 1 mM EDTA] were added to each well. After incubation at room temperature for 15 min in the dark, bioluminescence was assayed using the MicroBeta TriLux. The substrate (0.01 mM Coelenterazine, Biosynth AG, Staad, Switzerland) was auto-injected into the wells, and luminescence was recorded over a 20 s period, following a 1 s delay window. The background was normalized using wells containing only buffer or buffer with cells lacking the LUC gene. Expression of the LUC reporter gene in transgenic C. reinhardtii cells was normalized for cell numbers. The induction factor was calculated by comparison with untreated cells.
RNA isolation and RNA-blot analyses
RNA extraction, electrophoretic separation of RNA and hybridizations were performed as described previously (von Gromoff et al. 1989). The probe for LUC (accession number: AY004213) was a 0.9-kb BamHI/XhoI fragment from pCrLUC (Fuhrmann et al. 2004). The CBLP gene encoding a Chlamydomonas Gβ-like polypeptide was employed as loading control gene (von Kampen et al. 1993).
Measurement of hydrogen peroxide
At each time point, 0.5-ml aliquots of the cultures were centrifuged at 13,000g in a microcentrifuge. The supernatants were mixed with an equal volume of 1 M KI. After 15 min at room temperature, the mixture was assayed for iodine formation by determining the OD at 390 nm. The absorbance at 390 nm was stable for at least 3 h (Waffenschmidt et al. 1993). Concentrations were determined using a standard calibration curve with known amounts of H2O2 (Sigma).
Measurement of enzyme activities
Cultures of C. reinhardtii strain D66 were grown to 1–2 × 106 cells ml−1, transferred in the dark for 16 h, and subsequently exposed to light (70 μmol photons m−2 s−1) or maintained in the dark. When indicated, DCMU (final concentration 6 μM) was added 45 min prior to illumination. Some cultures were supplemented with H2O2 (2 mM final concentration). Cells to be collected (1 × 108 cells) were pelleted by centrifugation (microcentrifuge, maximal speed) and resuspended in 200 μl 50 mM Hepes buffer (pH 8). After two consecutive freeze/thawing cycles in liquid nitrogen, the enzyme activities in crude extracts of Chlamydomonas cultures were measured.
Catalase activity was measured polarographically at 20°C with a Clark-type electrode in 50 mM Hepes buffer (pH 8) in the presence of 1 mM H2O2 as substrate using a final protein concentration of 5 μg ml−1. The protein concentration of the crude extracts was determined using the amido black assay.
Ascorbate peroxidase activity was measured photometrically at 290 nm in 20 mM phosphate buffer, pH 7.0, containing 0.5 mM ascorbate and 1 mM H2O2 as substrates using crude extract with a final protein concentration of 5 μg ml−1.
Initial NADP-malate dehydrogenase activity was measured photometrically at 340 nm in 100 mM Tris/HCl, pH 7.9, containing 0.75 mM oxalacetate and 0.27 mM NADPH as substrates using crude extract with a final protein concentration of 10 μg ml−1. To activate MDH to its maximal activity, samples were incubated with 10 mM DTT, 20-μM thioredoxin (cytosolic thioredoxin h1 from Chlamydomonas) in 100 mM Tris/HCl, pH 7.9 for 10 min prior to start of the assay by addition of substrates. The ratio between initial activity and total activity gives the percentage of activated MDH at the time point of harvest.
Statistics
Data represent means or representative examples from measurements repeated 3–6 times. Typical standard errors are shown, in Figs. 1b, 6 and 8; they are omitted for some data for clarity.
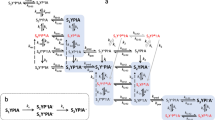
a–c Response of a P HSP70A -Luc reporter gene in Chlamydomonas transformants to the addition of H2O2 in the light and in the dark. a A truncated version (ΔHSE) of the HSP70A promoter construct known to be inducible by H2O2 (Shao et al. 2007) was fused to a luciferase reporter gene that at its upstream end harbored sequences of HSP70B. Dark shaded bars represent exons, light shaded bars indicate introns. The third exon of HSP70B was fused to a Renilla luciferase reporter gene with C. reinhardtii adapted codon usage (Fuhrmann et al. 2004) (LUC), indicated by a dark grey bar. The 3′ UTR of the fusion gene is encoded by the corresponding region of RBCS2. b Induction of P HSP70A -LUC in Chlamydomonas transformants by exogenously applied hydrogen peroxide. Light grown C. reinhardtii cells were incubated in the presence of 2 mM H2O2 with white light of an intensity of 70 μmol photons m−2 s−1 (open circles) or in the dark (closed circles). H2O was added to light-incubated control cultures (open squares) and to control cultures transferred to the dark (closed squares). Samples for luciferase assays were taken at the time points indicated and processed as described in Sect. “Materials and methods”. The activity was normalized with respect to cell numbers and the induction factors were calculated by comparison with untreated cells at time point 0 h. c Dependence of the degree of induction of the P HSP70A -LUC reporter construct in Chlamydomonas transformants in the light on the concentration of exogenous H2O2. Experiments were performed as described in b. Samples for luciferase assays were taken after incubation for 3 h (diamonds) and after 5 h (triangles)
Results
Induction of an H2O2-responsive reporter gene by H2O2 is strongly stimulated by light
In the present work, we investigated the regulation of an H2O2-responsive reporter gene by exogenously added H2O2. For these assays we employed transformants harboring an H2O2-responsive fragment of the HSP70A promoter fused to a Renilla-derived luciferase gene (LUC) (Shao et al. 2007). A schematic representation of the LUC reporter construct is presented in Fig. 1a. Transformants with this construct did not respond to heat shock but expression of the reporter gene is induced by hydrogen peroxide, either produced inside the chloroplast or added exogenously to the culture medium (Shao et al. 2007).
A strong increase in luciferase activity was observed in transformants with this construct after treatment by H2O2, but only when the cultures were incubated in the light (Fig. 1b). In the light, the activity increased about 200-fold within 5 h of incubation in the presence of 2 mM H2O2. In the dark, a distinctly weaker induction was observed. The luc activity was 11-fold higher after 3 h, and 19-fold higher after 5 h incubation. Light itself (70 μmol photons m−2 s−1) had no effect since no induction of the reporter gene could be observed in the light-grown control cultures supplemented with H2O. In the absence of H2O2, no difference was observed between light-grown cultures and cultures transferred from the light to the dark. When cultures were grown in the dark for 20 h and then shifted to light in the absence of H2O2, a weak increase in luciferase activity was observed (Shao et al. 2007). This increase was 2.5- to 3-fold smaller than the one observed in the presence of H2O2 in the dark.
We next tested the level of expression in dependence on the concentration of H2O2 added to the culture medium. Luciferase activity was shown to increase strongly with the concentration of H2O2 as can be seen by plotting luciferase activity 3 and 5 h after H2O2 addition as a function of the concentration of externally added H2O2 (Fig. 1c). We have shown previously that much lower H2O2 concentrations are sufficient to induce the reporter gene when H2O2 is produced inside the chloroplast by the photosynthetic electron transfer chain in the presence of metronidazole as electron acceptor. In this case, a high level of induction of the reporter gene was observed already after 2 h, when the detectable H2O2 concentration in the medium was 3 μM (Shao et al. 2007).
Catalase activity contributes to a faster degradation of H2O2 in the dark
The strong induction of transformants with the P HSP70A-ΔHSE -LUC reporter construct by externally added H2O2 observed only in the light may possibly be linked to lower rate of H2O2 degradation by Chlamydomonas cells incubated in the light than in the dark. This was tested by measuring the rate of H2O2 disappearance from the culture medium kept in light and or darkness. Indeed, H2O2 was more rapidly degraded by cultures in the dark than in the light (Fig. 2a). In the dark, H2O2 almost completely disappeared from the culture medium within 4 h and more than 50% of the initial H2O2 had disappeared already within 1 h. By contrast, in the light, about 75% of the H2O2 was still present after 1 h and some H2O2 could still be detected after 4 h. The slower disappearance of the H2O2 from the culture medium in the light may possibly be explained by the continuous production of H2O2 by the photosynthetic electron transport chain. However, the Mehler reaction as potential internal source of H2O2 cannot explain the difference between H2O2 degradation in the light and in the dark, because little H2O2 was produced by cultures, even in the presence of metronidazole, an artificial electron acceptor of PSI (less than 2 μM H2O2 per hour of irradiation was detectable in the medium; Shao et al. 2007).
a, b Degradation of exogenously added H2O2 by Chlamydomonas cultures. a Time course of degradation of H2O2 added to the medium of Chlamydomonas cultures in the light (open circles) or in the dark (closed circles). The change in H2O2 content in cell-free medium served as a control (open squares). 2 mM H2O2 was added at time point zero. Samples were taken at the times points indicated and the H2O2 concentration was determined as described in Sect. “Materials and methods”. Light-grown C. reinhardtii cells were irradiated with 70 μmol photons m−2 s−1 of white light. b Time course of H2O2 degradation in the medium of Chlamydomonas cultures in the dark in the absence (closed circles) or in the presence of 2 mM aminotriazole, a catalase inhibitor (closed triangles). The inhibitor was added 45 min prior to the addition of 2 mM H2O2. All other conditions were as described in a
This faster degradation of H2O2 in the dark may be caused by a light-dependent inhibition of H2O2 detoxifying enzymes. In plants, catalases and ascorbate peroxidases constitute the two major classes of enzymes involved in H2O2 detoxification. To test whether catalase is involved in the observed differences in H2O2 degradation between light and dark conditions, we measured the rates of H2O2 degradation in the dark in the presence or absence of the catalase inhibitor aminotriazole (AT; Fig. 2b). The concentration of H2O2 in the culture medium decreased considerably slower in the presence of AT than in its absence. This indicates that, under the conditions tested, catalase appears to be one of the major enzymes involved in H2O2 detoxification in Chlamydomonas cells. This result also suggested that a light-dependent regulation of catalase activity may account for the differences in H2O2 degradation observed between light and dark conditions (Fig. 2) and consequently for the light-dependence of the H2O2 inducibility of the reporter gene (Fig. 1b).
Light has previously been shown to inactivate plant catalases (Grotjohann et al. 1997; for review see Feierabend 2005). Generally, this inactivation was observed under high light conditions (800 μmol photons m−2 s−1). To test whether catalase activity was already affected by the low light intensities used in the experiments shown in Figs. 1 and 2, cell extracts of Chlamydomonas were illuminated with white light of an intensity of 70 μmol photons m-2 s−1 for up to 5 h. Under these conditions, no significant photoinactivation of the enzyme could be observed within 5 h of illumination (Fig. 6). We therefore concluded that a direct photoinactivation of catalase is unlikely to play a role under the low light conditions that were employed for studying the induction of the reporter gene by H2O2.
The redox state of the photosynthetic electron transfer chain affects H2O2 detoxification and catalase activity
Recently, it was shown that the activity of Chlamydomonas catalase is inhibited by reduced TRX in vitro (Lemaire et al. 2004). In chloroplasts, TRXs are reduced by the photosynthetic electron transfer chain in the light. The reduction of TRXs may thus possibly be responsible for the inactivation of catalases; and therefore, the slower H2O2 degradation observed in the light. Such a mechanism may account for the light dependence of the induction of the reporter gene by H2O2. In order to test whether photosynthetic electron transfer plays a role in this light dependence, we investigated the effect of DCMU, an inhibitor of photosystem II, on the expression of the reporter gene, the degradation of H2O2, and the activity of H2O2 detoxifying enzymes. In the presence of DCMU, linear photosynthetic electron transfer is blocked and, among other components which may play a role in redox regulation, chloroplastic TRXs remain oxidized. In the presence of DCMU, only a small increase in luciferase activity could be detected after treatment with H2O2 in the light (Fig. 3), indicating only a weak induction of the reporter gene. The level of luciferase activity observed was similar to that seen after addition of H2O2 in the dark. This suggests that linear photosynthetic electron flow is required for an efficient induction of the reporter gene by H2O2 in the light.
Effect of DCMU on the induction of the P HSP70A -LUC reporter construct by H2O2 in Chlamydomonas transformants. Cells were incubated in the presence of 6 μM DCMU for 45 min prior to the addition of 2 mM H2O2. After H2O2 addition, cells were incubated for 5 h in the light (70 μmol photons m−2 s−1) or in the dark. Luciferase activity was normalized with respect to cell numbers. The luciferase activity of light-grown cultures in the absence of DCMU was set as one
If this requirement is linked to a light-dependent inactivation of catalase, implicating TRXs or other mediators, then DCMU was predicted to affect the rate of H2O2 degradation in the light and the activity of catalases. Therefore, the time course of H2O2 consumption was measured in the light in the presence and absence of DCMU (Fig. 4). In the light, the H2O2 concentration decreased significantly faster in the presence of DCMU than in its absence. This suggests that photosynthetic electron flow accounts for the effect of light on the rate of H2O2 degradation in the culture medium.
Effect of DCMU on the degradation of H2O2 in the medium by Chlamydomonas cells. Cells were incubated with irradiation (white light, 70 μmol photons m−2 s−1) in the presence of 6 μM DCMU for 45 min (open triangles) or its absence (open circles) prior to addition of 2 mM H2O2. Results from cells incubated in the absence of DCMU in the dark (closed circles) are also shown. Samples were taken at the times points indicated and the H2O2 concentration in the media was determined as described in Sect. “Materials and methods”
To investigate the effect of the redox state of photosynthetic electron flow further, the degradation of H2O2 was tested using mutants lacking either the cytochrome b6f complex (ΔpetD) or deficient in ribulose-1,5-bisphosphate carboxylase (Rubisco) (rbcL). As shown in Fig. 5, only a small difference in the kinetics of H2O2 degradation was observed between dark-treated and light-treated cultures of the petD mutant lacking the cytochrome b6f complex. The differences were statistically not significant according to the Student’s t test. In this mutant, forward electron transport to PSI is completely abolished, the plastoquinone pool stays reduced and components downstream from PSI remain oxidized. In the mutant lacking the large subunit of Rubisco, the difference in H2O2 degradation between light and dark is comparable to that observed in the wild-type strain. In this mutant, upon irradiation, the electron transport chain, including electron acceptors like NADP+ and thioredoxins stays highly reduced.
Degradation of exogenously added H2O2 in the medium by photosynthesis mutants. Time course of degradation of H2O2 added to the medium of Chlamydomonas cultures in the light (white light, 55 μmol photons m−2 s−1) (open symbol) or in the dark (closed symbol). The rbcL and ΔpetD mutants were grown in very dim light (<10 μmol photons m−2 s−1) for 5 days and then diluted to 1 × 106 cell ml−1. The parental strain of the mutants (WTS34), named WT, was incubated with irradiation (white light, 55 μmol photons m−2 s−1) for 3 days and then diluted to 1 × 106cell ml−1. Right after dilution, H2O2 (final concentration 2 mM) was added. Samples were taken at the time points indicated and the H2O2 concentration in the media was determined as described in Sect. “Materials and methods”
These data suggest that photosynthetic electron flow regulates the activity of H2O2 detoxifying enzymes, possibly involving TRXs. In order to test this possibility, we measured catalase activity in cells opened up by freeze/thawing. The activity could be inhibited by 75% by addition of 5-mM DTT (Fig. 6). This indicates that the catalase can undergo redox dependent inactivation. Figure 6 shows that incubation of the cells in the light clearly inhibited the activity of catalase. Indeed, about 40% of the activity was lost within 2 h and only 25% remained after 5 h of incubation in the light. The remaining activity after 5 h illumination corresponded to the activity of the catalase in the presence of DTT. By contrast, in cells maintained in the dark, the activity remained above 85% of the initial activity throughout the experiment. Presence of DCMU during the light incubation protected against the loss of activity (Fig. 6), supporting the idea that the activity of H2O2-consuming enzymes is regulated via the redox state of the electron transfer chain. The kinetics of inactivation were not dependent on the presence of H2O2 in the culture medium, since the same degree of inhibition was observed when no H2O2 was added prior to the shift of the cultures from dark to light (data not shown).
Effect of light and DCMU on the activity of catalase measured in vitro. Cells were incubated in the presence of 2 mM H2O2 in the light (white light, 70 μmol photons m−2 s−1) in the absence (circles) or presence of 6 μM DCMU (open triangles) and in the dark without DCMU (closed circles) for the time indicated. H2O2 and DCMU were added at time point zero. Cells were harvested at the time indicated and opened up by freeze/thawing. H2O2 consumption by the crude extracts was measured polarographically. 100% activity corresponds to an activity of 10 μmol O2 mg protein−1 min−1 produced when 1 mM H2O2 was added as substrate. The catalase activity in the crude extract of cells incubated in the light was strongly inhibited when 5 mM DTT was added (stars). Also shown is the catalase activity of crude extract of dark-grown cells (diamonds) that were illuminated (white light, 70 μmol photons m−2 s−1) for the time indicated
The activity of ascorbate peroxidases did not show any inhibition during the time course of illumination of the cultures in the presence of 2 mM H2O2 (Fig. 7), indicating that indeed the redox-dependent inhibition of catalase is the important factor leading to a loss of H2O2 degradation and subsequent expression of the reporter gene. To show a correlation between the state of reduction of thioredoxins and the activity of catalase, we measured the activity of the NADP-dependent malate dehydrogenase (MDH), one of the classical enzymes activated by reduction of its disulfide bonds by reduced thioredoxin (e.g., Lemaire et al. 2005, 2007). MDH activity increased upon incubation in the light while catalase activity decreased (Fig. 7). These data support the hypothesis that the activity of catalase is linked to the redox state of thioredoxin in the chloroplast. DCMU inhibited the activation of MDH as expected (Fig. 7).
Effect of light on the activity of ascorbate peroxidase and on the activity of NADP-malate dehydrogenase (MDH) in vitro. Cells were incubated in the presence of 2 mM H2O2 in the light (white light, 70 μmol photons m−2 s−1) for the time indicated. H2O2 and DCMU were added at time point zero. Cells were harvested at the time indicated and opened up by freeze/thawing. Ascorbate peroxidase (squares) activity was measured photometrically at 290 nm in the crude extract in the presence of ascorbate and H2O2 as outlined in Sect. “Materials and methods”. 100% activity corresponds to an activity of 18 μmol dehydroascorbate mg protein−1 min−1 produced when 1 mM H2O2 was added as substrate. The activity of MDH was measured photometrically at 340 nm in the presence of NADPH and oxalacetate in the absence (inverted triangles) and in the presence of 6 μM DCMU (diamonds). Maximal activity (100%) was achieved by incubation of samples with DTT and thioredoxin h (see Sect. “Materials and methods” for details”). For comparison the catalase activity (circles) is shown (data taken from Fig. 6)
It has to be mentioned that H2O2 inhibits CO2 fixation, since the transketolase reactions of the Calvin cycle are very sensitive to inhibition by H2O2 (Kaiser 1976). Under our conditions, photosynthetic oxygen evolution measured in the presence of 1 mM NaHCO3 was inhibited by 50% upon addition of 2 mM H2O2 (data not shown) in accordance with data reported for a different green algal species (Drabokova et al. 2007). This effect, however, seems not to interfere with our observations, neither with the induction of the reporter gene nor with the light-dependent inactivation of catalase.
Aminotriazole and DCMU affect the expression of the reporter gene at the mRNA level
We have shown above that a down-regulation of catalase activity mediated either by photosynthetic electron flow or the addition of AT appears to be responsible for a slower degradation of H2O2 in the light. This suggested that a down-regulation of catalase activity in the light also accounts for the light dependence of the LUC reporter gene induction by H2O2. In order to confirm that the activity of H2O2 degrading enzymes and photosynthetic electron flow indeed are involved in the regulation of the reporter gene and to rule out the possibility that DCMU and AT interfere with luciferase activity directly, we measured the effect of AT and DCMU on H2O2 induction of the reporter at the mRNA level (Fig. 8). In the light, an increase of LUC mRNA level was observed after H2O2 treatment. This induction was abolished by the presence of DCMU, confirming that photosynthetic electron flow is required for induction of the reporter gene, as already observed at the activity level (Fig. 3). In the dark, a small increase in LUC mRNA levels was observed upon H2O2 addition, consistent with the low level increase of LUC activity observed under these conditions (Fig. 1b). Addition of AT in the dark allowed a stronger increase in LUC mRNA levels after H2O2 treatment, consistent with a lower degradation of this ROS (Fig. 2b). These results are in agreement with those observed at the LUC activity level and indicate that down-regulation of the activity of H2O2 degrading enzymes is a prerequisite for an efficient induction of the reporter gene by H2O2.
Effect of aminotriazole and DCMU on the H2O2 induction of the P HSP70A -LUC reporter gene at the RNA level in Chlamydomonas transformants. Cultures incubated in the dark (D) were treated with H2O2 (2 mM final concentration) in the presence (lane 4) or absence (lane 3) of the catalase inhibitor aminotriazole (2 mM AT). AT was added 45 min prior to the addition of H2O2. Cells incubated in the light (L) were treated with H2O2 (2 mM final concentration) in the presence (lane 5) or absence (lane 2) of the PSII inhibitor DCMU (6 μM). DCMU was added 45 min prior to the addition of H2O2. A control culture that did not receive H2O2 was incubated in the light (lane 1). Samples for RNA isolation were taken 2 h after H2O2 addition. RNA blots (10 μg total RNA per lane) were hybridized with a probe specific for the LUC transgene and CBLP, the latter serving as a loading control as described in Sect. “Materials and methods”
Discussion
The data presented here show that the expression of the H2O2-responsive reporter construct not only depended on the H2O2 concentration in the media (Fig. 1c) but also on the activity state of H2O2 degrading enzymes (Figs. 2, 3, 8). The decrease in the activity of H2O2 degrading enzymes observed upon illumination was prevented by DCMU (Fig. 4) and by mutations perturbing photosynthetic electron flow (Fig. 5), indicating that catalase activity was correlated with the redox state of the photosynthetic electron transport chain (Fig. 6). These data illustrate that Chlamydomonas cells have evolved mechanisms that ensure maintenance of a certain level of H2O2 in order to allow activation of H2O2-responsive genes. In photosynthetic organisms, ROS are mainly formed in the light as a consequence of electron transport reactions under conditions where the electron flow system is saturated and the electron acceptors are reduced. In the so-called Mehler reaction, the electron acceptors of photosystem I react with oxygen when the ferredoxin-NADP system and alternative electron sinks are already reduced. Superoxide anions and H2O2 are formed. These ROS are released into the stroma where they encounter an efficient antioxidant system and detoxifying enzymes (for review see Asada 1999; Foyer and Noctor 2005). In general, it is assumed that detoxifying enzymes have to be present in an active state during oxidative stress conditions.
However, it was recently observed that higher plant ascorbate peroxidase is inhibited in the light under conditions where both, the Calvin cycle and photorespiration, are inhibited (Miyake et al. 2006). This paradoxical situation makes sense when the function of ROS in gene regulation is taken into account. Maintenance of a certain H2O2 concentration inside the chloroplast may be achieved in higher plants via control of the activity of ascorbate peroxidases, the main H2O2-detoxifying enzyme in this organelle. The H2O2 concentration in the cell will depend on the activity of the whole set of detoxifying enzymes; also on those located in different organelles because H2O2 can easily diffuse across biological membranes (Bienert et al. 2007). As shown for different organisms, the expression of a number of genes, encoding, e.g., heat shock proteins, catalases, and ascorbate peroxidases involved in the abrogation of oxidative stress, is regulated via H2O2 signaling. Thus, it can be postulated that a down-regulation of the activity of detoxifying enzymes under stress conditions is a necessary prerequisite for the activation of those genes that allow acclimation of the organism to oxidative stress conditions. In Chlamydomonas, this may be achieved via inactivation of catalase; the activity of ascorbate peroxidase under these conditions being constant (Fig. 7). Inactivation of plant catalases by high intensity irradiation (Feierabend and Engel 1986; Grotjohann et al. 1997) needs to be distinguished from low light induced inactivation as studied here by exposing Chlamydomonas cells to light of an intensity of 70 μmol photons m−2 s−1 or less. It has also been reported previously that the catalase activity in Chlamydomonas is higher in cells grown in the dark than in the light (Kato et al. 1997).
The inactivation of H2O2 degrading enzymes by light of moderate intensity as observed in Chlamydomonas could be linked to photosynthetic electron transfer since this inactivation was prevented by DCMU known to block the photosynthetic apparatus at PSII (Fig. 4). In vitro measurements revealed that catalase is the target for this inactivation (Fig. 6). The redox sensor which mediates the inactivation of H2O2 degrading enzymes appears to be located downstream from PSI, since a mutation leading to an inactivation of the cytochrome b6f complex (ΔpetD) resulted in a protection against inhibition of the activity of H2O2 degrading enzymes by light (Fig. 5). The reduction state of the plastoquinone pool (Pfannschmidt and Liere 2005) can thus be ruled out to be of importance in the regulation of catalase activities, because mutations leading to an inactivation of the cytochrome b6f complex are characterized by a high reduction state of the plastoquinone pool while in the presence of DCMU, the plastoquinone pool stays oxidized. Since a reduction of TRX by the photosynthetic electron transfer chain is prevented by DCMU and in the petD mutant, these molecules are candidates for regulators of H2O2 degrading enzymes. Indeed, several H2O2 detoxifying enzymes have been shown to be inactivated by TRXs in vitro (Lemaire et al. 2004; Gelhaye et al. 2006). Proteomic studies revealed that also several ROS scavenging enzymes which do not use TRXs as substrates such as catalase, superoxide dismutase, or APX, are putative TRX targets since they are bound to TRX affinity columns (Balmer et al. 2004; Lemaire et al. 2004; Maeda et al. 2004; Wong et al. 2004; Michelet et al. 2006).
In the genomic sequence of Chlamydomonas (http://genome.jgi-psf.org/Chlre3/Chlre3.home.html), only a single gene encoding a typical catalase (gene model Chlre3/scaffold_30:1312932-1317626) was observed. A gene (gene model Chlre3/scaffold_1:4550894-4558523) encoding a putative catalase–peroxidase homologous to those found in prokaryotes (Jakopitsch et al. 2005) was also found. However, as judged on the basis of the number of EST sequences, expression of the latter gene appears to be very weak as compared to the expression of the typical catalase gene, even under oxidative stress conditions. Consequently, the catalase activities reported here are likely to correspond to the activity of the typical catalase, although it cannot be excluded that the putative catalase–peroxidase also contributes to H2O2 degrading activity measured in extracts.
In higher plants, catalases were localized to peroxisomes (Heazlewood et al. 2005), while in Chlamydomonas the enzyme is most likely located in mitochondria (Kato et al. 1997; Lemaire et al. 2004). This implies that a signal about the redox state of the photosynthetic electron transfer chain has to be transmitted to targets outside of the chloroplast. This signal could be initiated by a component whose redox state depends on the redox state of a functional photosynthetic electron transfer chain such as TRXs. The signal is possibly transmitted from the chloroplast by redox mediators such as glutathione or NADPH. Indeed, reducing equivalents of chloroplastic NADPH have been reported to be transferred outside of the chloroplast in the form of malate (Scheibe 2004). This malate is formed by reduction of oxalacetate using NADPH as electron donor. Interestingly, the NADP-malate dehydrogenase of the chloroplast is activated by light by a mechanism dependent on the redox state of plastidic TRXs (Lemaire et al. 2005, 2007). Indeed, a reverse correlation between the activities of catalase and NADP-malate dehydrogenase was observed (Fig. 7). Once outside of the chloroplast, the signal may be transduced to a redox regulator of the catalase via different redox mediators. Since the activity of Chlamydomonas catalase has been shown to be regulated by TRXs in vitro (Lemaire et al. 2004), the redox regulator of catalase is likely to be a TRX or involves TRX-related proteins like glutaredoxins. There is some evidence that thioredoxins belonging to the TRX o and TRX h subtypes are present in the mitochondria of Chlamydomonas (Lemaire et al. 2003).
In conclusion, the data presented provide support for the idea that Chlamydomonas has evolved mechanisms that allow maintenance of a certain level of H2O2. We assume that these mechanisms ensure a regulation of gene expression in response to ROS, allowing for an adaptation of the organisms to stress conditions. A fast inactivation of catalase activity is seen when cells are exposed to a 10-fold higher light intensity (data not shown). Regulation of the activity of ROS detoxifying enzymes appears to play an important role. A regulation via TRX or other components of the regulatory redox network like glutaredoxins, peroxiredoxins or glutathione provides an attractive strategy for the regulation of such enzyme activities. These compounds may ensure a fast and reversible inactivation of the enzymes, thereby allowing for a rapid sensing of changes in environmental conditions. Most importantly, through thioredoxins or other redox compounds, the regulation of the activity of H2O2-consuming enzymes may directly be coupled to photosynthetic electron flow, and thus to the main source for ROS production in photosynthetic organisms.
Abbreviations
- APX:
-
Ascorbate peroxidase
- AT:
-
Aminotriazole
- DCMU:
-
3-(3′4′-Dichlorophenyl)-1, 1-dimethylurea
- DTT:
-
Dithiothreitol
- MDH:
-
Malate dehydrogenase
- LUC:
-
Luciferase
- ROS:
-
Reactive oxygen species
- TRX:
-
Thioredoxin
References
Apel K, Hirt H (2004) Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu Rev Plant Biol 55:373–399
Asada K (1999) The water–water cycle in chloroplasts: scavenging of active oxygens and dissipation of excess photons. Annu Rev Plant Physiol Plant Mol Biol 50:601–639
Balmer Y, Vensel WH, Tanaka CK, Hurkman WJ, Gelhaye E, Rouhier N, Jacquot JP, Manieri W, Schürmann P, Droux M, Buchanan BB (2004) Thioredoxin links redox to the regulation of fundamental processes of plant mitochondria. Proc Natl Acad Sci USA 101:2642–2647
Bienert GP, Møller AL, Kristiansen KA, Schulz A, Møller IM, Schjoerring JK, Jahn TP (2007) Specific aquaporins facilitate the diffusion of hydrogen peroxide across membranes. J Biol Chem 282:1183–1192
Davletova S, Rizhsky L, Liang H, Shengqiang Z, Oliver DJ, Coutu J, Shulaev V, Schlauch K, Mittler R (2005) Cytosolic ascorbate peroxidase 1 is a central component of the reactive oxygen gene network of Arabidopsis. Plant Cell 17:268–281
Dietz KJ, Jacob S, Oelze ML, Laxa M, Tognetti V, de Miranda SM, Baier M, Finkemeier I (2006) The function of peroxiredoxins in plant organelle redox metabolism. J Exp Bot 57:1697–1709
Drabokova M, Admiraal W, Marsalek B (2007) Combined exposure to hydrogen peroxide and light-selective effects on cyanobacteria, green algae, and diatoms. Environ Sci Technol 41:309–314
de Pinto MC, Paradiso A, Leonetti P, De Gara L (2006) Hydrogen peroxide, nitric oxide and cytosolic ascorbate peroxidase at the crossroad between defence and cell death. Plant J 48:784–795
Engel N, Schmidt M, Lutz C, Feierabend J (2006) Molecular identification, heterologous expression and properties of light-insensitive plant catalases. Plant Cell Environ 29:593–607
Feierabend J (2005) Catalases in plants: molecular and functional properties and role in stress defence. In: Smirnoff N (ed) Antioxidants and reactive oxygen species in plants. Blackwell, Oxford, pp 101–140
Feierabend J, Engel S (1986) Photoinactivation of catalase in vitro and in leaves. Arch Biochem Biophys 251:567–576
Foyer CH, Noctor G (2005) Redox homeostasis and antioxidant signaling: a metabolic interface between stress perception and physiological responses. Plant Cell 17:1866–1875
Fuhrmann M, Hausherr A, Ferbitz L, Schodl T, Heitzer M, Hegemann P (2004) Monitoring dynamic expression of nuclear genes in Chlamydomonas reinhardtii by using a synthetic luciferase reporter gene. Plant Mol Biol 55:869–881
Gadjev I, Vanderauwera S, Gechev TS, Laloi C, Minkov IN, Shulaev V, Apel K, Inze D, Mittler R, van Breusegem F (2006) Transcriptomic footprints disclose specificity of reactive oxygen species signaling in Arabidopsis. Plant Physiol 141:436–445
Gechev T, Gadjev I, van Breusegem F, Inze D, Dukiandjiev S, Toneva V, Minkov I (2002) Hydrogen peroxide protects tobacco from oxidative stress by inducing a set of antioxidant enzymes. Cell Mol Life Sci 59:708–714
Gelhaye E, Navrot N, Macdonald IK, Rouhier N, Raven EL, Jacquot JP (2006) Ascorbate peroxidase–thioredoxin interaction. Photosynth Res 89:193–200
Grotjohann N, Janning A, Eising R (1997) In vitro photoinactivation of catalase isoforms from cotyledons of sunflower (Helianthus annuus L.). Arch Biochem Biophys 346:208–218
Gupta R, Luan S (2003) Redox control of protein tyrosine phosphatases and mitogen-activated protein kinases in plants. Plant Physiol 132:1149–1152
Harris E (1989) The Chlamydomonas sourcebook: a comprehensive guide to biology and laboratory use. Academic Press, San Diego
Heazlewood JL, Tonti-Filippini J, Verboom RE, Millar AH (2005) Combining experimental and predicted datasets for determination of the subcellular location of proteins in Arabidopsis. Plant Physiol 139:598–609
Hiner AN, Rodriguez-Lopez JN, Arnao MB, Lloyd Raven E, Garcia-Canovas F, Acosta M (2000) Kinetic study of the inactivation of ascorbate peroxidase by hydrogen peroxide. Biochem J 348:321–328
Jakopitsch C, Wanasinghe A, Jantschko W, Furtmuller PG, Obinger C (2005) Kinetics of interconversion of ferrous enzymes, compound II and compound III, of wild-type synechocystis catalase–peroxidase and Y249F: proposal for the catalytic mechanism. J Biol Chem 280:9037–9042
Joo JH, Wang S, Chen JG, Jones AM, Fedoroff NV (2005) Different signaling and cell death roles of heterotrimeric G protein alpha and beta subunits in the Arabidopsis oxidative stress response to ozone. Plant Cell 17:957–970
Kaiser W (1976) The effect of hydrogen peroxide on CO2 fixation of isolated intact chloroplasts. Biochim Biophys Acta 440:476–482
Kato J, Yamahara T, Tanaka K, Takio S, Satoh T (1997) Characterization of catalase from green alga Chlamydomonas reinhardtii. J Plant Physiol 151:262–268
Kindle KL (1990) High-frequency nuclear transformation of Chlamydomonas reinhardtii. Proc Natl Acad Sci USA 87:1228–1232
Kovtun Y, Chiu WL, Tena G, Sheen J (2000) Functional analysis of oxidative stress-activated mitogen-activated protein kinase cascade in plants. Proc Natl Acad Sci USA 97:2940–2945
Kuras R, Wollman FA (1994) The assembly of cytochrome b6/f complexes: an approach using genetic transformation of the green alga Chlamydomonas reinhardtii. EMBO J 13:1019–1027
Laloi C, Apel K, Danon A (2004) Reactive oxygen signalling: the latest news. Curr Opin Plant Biol 7:323–328
Lemaire SD, Collin V, Keryer E, Issakidis-Bourguet E, Lavergne D, Miginiac-Maslow M (2003) Chlamydomonas reinhardtii: a model organism for the study of the thioredoxin family. Plant Physiol Biochem 41:513–521
Lemaire SD, Guillon B, Le Maréchal P, Keryer E, Miginiac-Maslow M, Decottignies P (2004) New thioredoxin targets in the unicellular photosynthetic eukaryote Chlamydomonas reinhardtii. Proc Natl Acad Sci USA 101:7475–7480
Lemaire SD, Quesada A, Merchan F, Corral JM, Igeno MI, Keryer E, Issakidis-Bourguet E, Hirasawa M, Knaff DB, Miginiac-Maslow M (2005) NADP-malate dehydrogenase from unicellular green alga Chlamydomonas reinhardtii. A first step toward redox regulation? Plant Physiol 137:514–521
Lemaire SD, Michelet L, Zaffagnini M, Massot V, Issakidis-Bourguet E (2007) Thioredoxins in chloroplasts. Curr Genet 51:343–355
Maeda K, Finnie C, Svensson B (2004) Cy5 maleimide labelling for sensitive detection of free thiols in native protein extracts: identification of seed proteins targeted by barley thioredoxin h isoforms. Biochem J 378:497–507
Marchand C, Le Maréchal P, Meyer Y, Miginiac-Maslow M, Issakidis-Bourguet E, Decottignies P (2004) New targets of Arabidopsis thioredoxins revealed by proteomic analysis. Proteomics 4:2696–2706
Michelet L, Zaffagnini M, Massot V, Keryer E, Vanacker H, Miginiac-Maslow M, Issakidis-Bourguet E, Lemaire SD (2006) Thioredoxins, glutaredoxins, and glutathionylation: new crosstalks to explore. Photosynth Res 89:225–245
Minko I, Holloway SP, Nikaido S, Carter M, Odom OW, Johnson CH, Herrin DL (1999) Renilla luciferase as a vital reporter for chloroplast gene expression in Chlamydomonas. Mol Gen Genet 262:421–425
Miyake C, Asada K (1996) Inactivation mechanism of ascorbate peroxidase at low concentrations of ascorbate; hydrogen peroxide decomposes compound I of ascorbate peroxidase. Plant Cell Physiol 37:423–430
Miyake C, Shinzaki Y, Nishioka M, Horiguchi S, Tomizawa K (2006) Photoinactivation of ascorbate peroxidase in isolated tobacco chloroplasts: Galdieria partita APX maintains the electron flux through the water–water cycle in transplastomic tobacco plants. Plant Cell Physiol 47:200–210
Panchuk II, Zentgraf U, Volkov RA (2005) Expression of the Apx gene family during leaf senescence of Arabidopsis thaliana. Planta 222:926–932
Pfannschmidt T, Liere K (2005) Redox regulation and modification of proteins controlling chloroplast gene expression. Antioxid Redox Signal 7:607–618
Rentel MC, Lecourieux D, Ouaked F, Usher SL, Petersen L, Okamoto H, Knight H, Peck SC, Grierson CS, Hirt H, Knight MR (2004) OXI1 kinase is necessary for oxidative burst-mediated signalling in Arabidopsis. Nature 427:858–861
Scheibe R (2004) Malate valves to balance cellular energy supply. Physiol Plant 120:21–26
Shao N, Krieger-Liszkay A, Schroda M, Beck CF (2007) A reporter system for the individual detection of hydrogen peroxide and singlet oxygen: its use for the assay of reactive oxygen species produced in vivo. Plant J 50:475–487
Shigeoka S, Ishikawa T, Tamoi M, Miyagawa Y, Takeda T, Yabuta Y, Yoshimura K (2002) Regulation and function of ascorbate peroxidase isoenzymes. J Exp Bot 53:1305–1319
Shikanai T, Takeda T, Yamauchi H, Sano S, Tomizawa KI, Yokota A, Shigeoka S (1998) Inhibition of ascorbate peroxidase under oxidative stress in tobacco having bacterial catalase in chloroplasts. FEBS Lett 428:47–51
Spreitzer RJ, Goldschmidt-Clermont M, Rahire M, Rochaix JD (1985) Nonsense mutations in the Chlamydomonas chloroplast gene that codes for the large subunit of ribulosebisphosphate carboxylase/oxygenase. Proc Natl Acad Sci USA 82:5460–5464
Teixeira FK, Menezes-Benavente L, Galvao VC, Margis R, Margis-Pinheiro M (2006) Rice ascorbate peroxidase gene family encodes functionally diverse isoforms localized in different subcellular compartments. Planta 224:300–314
Vandenabeele S, Vanderauwera S, Vuylsteke M, Rombauts S, Langebartels C, Seidlitz HK, Zabeau M, Van Montagu M, Inze D, van Breusegem F (2004) Catalase deficiency drastically affects gene expression induced by high light in Arabidopsis thaliana. Plant J 39:45–58
Vieira Dos Santos C, Rey P (2006) Plant thioredoxins are key actors in the oxidative stress response. Trends Plant Sci 11:329–334
von Gromoff ED, Treier U, Beck CF (1989) Three light-inducible heat shock genes of Chlamydomonas reinhardtii. Mol Cell Biol 9:3911–3918
von Kampen J, Nieländer U, Wettern M (1993) Stress-dependent transcription of a gene encoding a Gβ-like polypeptide from Chlamydomonas reinhardtii. J Plant Physiol 143:756–758
Waffenschmidt S, Woessner JP, Beer K, Goodenough UW (1993) Isodityrosine cross-linking mediates insolubilization of cell walls in Chlamydomonas. Plant Cell 5:809–820
Wong JH, Cai N, Balmer Y, Tanaka CK, Vensel WH, Hurkman WJ, Buchanan BB (2004) Thioredoxin targets of developing wheat seeds identified by complementary proteomic approaches. Phytochemistry 65:1629–1640
Yabuta Y, Maruta T, Yoshimura K, Ishikawa T, Shigeoka S (2004) Two distinct redox signaling pathways for cytosolic APX induction under photooxidative stress. Plant Cell Physiol 45:1586–1594
Yamazaki D, Motohashi K, Kasama T, Hara Y, Hisabori T (2004) Target proteins of the cytosolic thioredoxins in Arabidopsis thaliana. Plant Cell Physiol 45:18–27
Acknowledgments
The authors are grateful to Olivier Vallon (IBPC, Paris) for many suggestions and the providing of strains, Myroslawa Miginiac-Maslow (IBP, Orsay) for valuable suggestions and to Bill Rutherford, and Diana Kirilovsky (both at CEA, Saclay) for critical reading of the manuscript. This work was supported in part by grants of the Deutsche Forschungsgemeinschaft to C.F.B. (Be 903/13-2) and A.K.L. (Li883/10-1) and the Agence Nationale de la Recherche Grant JC05-45751 to S.D.L.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License ( https://creativecommons.org/licenses/by-nc/2.0 ), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Shao, N., Beck, C.F., Lemaire, S.D. et al. Photosynthetic electron flow affects H2O2 signaling by inactivation of catalase in Chlamydomonas reinhardtii . Planta 228, 1055–1066 (2008). https://doi.org/10.1007/s00425-008-0807-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-008-0807-0