Abstract
DNA damage is one of the most important effects induced by chemical agents. We report a comparative analysis of DNA fragmentation on three different cell lines using terminal deoxynucleotidyl transferase dUTP nick end labeling (TUNEL) assay, generally applied to detect apoptosis. Our approach combines cytogenetic techniques and investigation in detached cellular structures, recovered from the culture medium with the aim to compare the DNA fragmentation of three different cell line even beyond the cells adherent to substrate. Consequently, we detect any fragmentation points on single chromosomes, whole nuclei and other cellular structures. Cells were exposed to resveratrol (RSV) and doxorubicin (Doxo), in single and combined treatments. Control and treated astrocytes showed DNA damage in condensed nuclei and detached structures. Caco-2 cells showed fragmented DNA only after Doxo-treatment, while controls showed fragmented chromosomes, indicating DNA damage in replicating cells. MDA-MB-231 cells showed nuclear condensation and DNA fragmentation above all after RSV-treatment and related to detached structures. This model proved to perform a grading of genomic instability (GI). Astrocytes show a hybrid level of GI. Caco-2 cells showed fragmented metaphase chromosomes, proving that the DNA damage was transmitted to the daughter cells probably due to an absence of DNA repair mechanisms. Instead, MDA–MB-231 cells showed few or no fragmented metaphase, suggesting a probable activation of DNA repair mechanisms. By applying this alternative approach of TUNEL test, we obtained data that can more specifically characterize DNA fragmentation for a suitable application in various fields.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
DNA fragmentation refers to the process of DNA strand breakage, resulting in the generation of smaller DNA fragments (Yoshida et al. 2006). This phenomenon has garnered significant attention in scientific research due to its relevance in various biological contexts (González-Marín et al. 2012). The distinctive nature of DNA fragmentation serves as a crucial cellular signature, providing valuable insights into fundamental cellular processes and pathological conditions (Kroemer et al. 2009; Kitazumi and Tsukahara 2011).
Understanding the mechanisms and consequences of DNA fragmentation is of paramount importance as it has implications in various biological phenomena, such as aging, cancer, response to therapeutic agents (Bouwman and Jonkers 2012; Chatterjee and Walker 2017; Ou and Schumacher 2018; Razak et al. 2019; Carusillo and Mussolino 2020), and embryonic development (Chiarelli et al. 2014, 2019, 2021, 2022c; Martino et al. 2021).
Interestingly, DNA fragmentation represents a relevant marker of the damage induced by physical or chemical stresses (Chiarelli and Roccheri 2012), especially when the cytoprotection mechanisms are not enough to deal with the stress (Agnello et al. 2016; Chiarelli et al. 2016, 2022a, b; Martino et al. 2021).
DNA fragmentation can also occur in response to various stressors, including genotoxic insults, oxidative damage, or DNA replication errors. Unrepaired or improperly repaired DNA breaks may result in aberrant DNA fragmentation patterns, leading to genomic instability and the potential for the development of genetic disorders or cancer (Jackson and Bartek 2009; Kroemer et al. 2009; Bouwman and Jonkers 2012; Caradonna 2015; Chatterjee and Walker 2017; Luparello et al. 2021).
In physiological conditions, DNA fragmentation occurs as a controlled and regulated process during specific stages of the cell cycle, or programmed cell death (Xu et al. 2019). During apoptosis, cells undergo a series of well-defined molecular events, culminating in the fragmentation of genomic DNA into smaller fragments of distinct sizes (Zhang and Xu 2000). For these reasons, DNA fragmentation is now considered a distinctive trait and a marker of irreversible cell death (Moore et al. 2021).
This characteristic DNA fragmentation pattern can be detected and analyzed using techniques such as DNA laddering gel electrophoresis, comet assay, terminal deoxynucleotidyl transferase dUTP nick end labeling (TUNEL) assay (Majtnerová and Roušar 2018), or next-generation sequencing (NGS) approaches (van Dijk et al. 2018). DNA laddering assay is not quantitative (Tsukada et al. 2001). Comet assay is limited to in vitro (cultured cells) only, and its quantification is labor-intensive (Kim et al. 2002). TUNEL is the most sensitive, accurate, and quantitative test (Moore et al. 2021), applicable to different biological samples.
In this study, we aim to conduct a comparative analysis of DNA fragmentation in different cell lines using the TUNEL assay on metaphase chromosomes samples, nuclei and cellular structures recovered from cultures media.
The TUNEL assay, initially designed as a standard histochemical method to detect DNA fragmentation resulting from apoptosis (Gavrieli et al. 1992), is the conventional approach followed in literature for analyzing cultured cells, cells in suspension and tissues.
Here, we propose an alternative approach that introduces two new and important variants: (1) cytogenetic techniques for selecting only those cells undergoing mitosis and (2) extension of the analysis of DNA fragmentation signals beyond the cells adherent to substrate. In fact, it has been recently described that the conditioned medium is a reservoir of useful information which until now was excluded from the various analyses because it was judged to be waste (Rosochowicz et al. 2023).
To achieve our objectives, clone-7 astrocytes and Caco-2 and MDA-MB-231 cells were subjected to single and combined treatments with resveratrol (RSV) and doxorubicin (Doxo) to modulate the level of DNA damage (Lin et al. 2014; Liu et al. 2014; Bong et al. 2020; Gomes et al. 2020; Jadid et al. 2021; Maszczyk et al. 2022; Volpes et al. 2023).
Investigating DNA fragmentation patterns at different levels and their underlying mechanisms contributes to the understanding of fundamental cellular processes, disease pathogenesis, and the development of potential diagnostic and therapeutic strategies. In particular, this study, by using an alternative approach of TUNEL assay, seeks to shed some light on the contribution of DNA fragmentation in different cell lines, elucidating the global profile of DNA damage.
Materials and methods
Cell lines and treatments
Clone-7 astrocytes cells were obtained by limited dilution technique starting from primary astrocytes, isolated as previously described (Caradonna et al. 2020).
Cells were cultured with Dulbecco’s modified Eagle’s medium (DMEM) high glucose (Sigma-Aldrich, D6429, USA) and F-12 K (Corning, 10–025-CV, USA) in a 2:1 ratio, enriched with 10% fetal bovine serum, 2 mM l-glutamine (Sigma-Aldrich, G7513, USA), and antibiotics penicillin/streptomycin 1% (Sigma-Aldrich, P4333, USA), until subconfluent, changing the culture medium approximately every 2 days and keeping cells at 37 °C in a controlled atmosphere (5% CO2).
Caco-2 and MDA-MB-231 cell lines were obtained from the American Type Culture Collection (Rockville, Md., USA) and cultured in DMEM (Sigma-Aldrich, D6429, USA) supplemented with 10% fetal bovine serum, 2 mM l-glutamine (Sigma, Life Science, G7513, U.K), and antibiotics penicillin/streptomycin 1% (Sigma-Aldrich, P4333, USA), and were maintained at 37 °C in 5% CO2 and 95% humidity. Caco-2 and MDA-MB-231 cell lines were cultured as previously described (Naselli et al. 2015; Luparello et al. 2019).
To identify potential modulatory effects on DNA fragmentation, clone-7 astrocytes and Caco-2 and MDA-MB-231 cells were treated with doxorubicin and resveratrol, solubilized in DMSO verifying that the vehicle volume was always less than 0.01% of the cell medium to avoid self-effect on the biological and epigenetic/molecular endpoints (Galvao et al. 2014).
The selected concentration of resveratrol for each cell line was 50 μM, chosen as it is under the IC50 reported in the literature (Lin et al. 2014; Liu et al. 2014; Gomes et al. 2019, 2020; Jadid et al. 2021; Volpes et al. 2023). The treatment extended for 48 h.
Clone-7 astrocytes and Caco-2 cells were also treated with 1 μM doxorubicin for 48 h (Jadid et al. 2021; Maszczyk et al. 2022), unlike MDA-MB-231 cells that were treated for 48 h with 0.6 μM doxorubicin chosen as it is under the IC50 reported in the literature (Sadeghi-Aliabadi et al. 2012).
Combined treatments with both selected molecules were carried out at the same concentrations and times used in the single treatments.
Preparation of metaphase chromosomes
To perform the cytogenetic analysis, the cells were cultured in T75 cm2 flasks until reaching 70–80% confluence, thus ensuring the maintenance of the exponential phase of growth without reaching the stationary phase. Metaphases were obtained by adding colcemid (KaryoMAX™ Colcemid™ solution in HBSS, Gibco, 15,210–040, USA) at a final concentration of 0.1 μg/ml, 3 h before the trypsinization. The cultures were then processed according to the conventional air-drying protocol, as previously described (Sirchia and Luparello 2007; Mannino et al. 2019; Caradonna et al. 2020).
Preparation of detached cell population
After cell culture, the medium was recovered and the pellet was washed two times in phosphate-buffered saline (PBS). The pellet was resuspended and fixed in 3.7% paraformaldehyde for 60 min, harvested by centrifugation at 200g for 7 min and resuspended in PBS. The slides were prepared by cytospin centrifugation at 60 g for 5 min on polylysine-coated glass slides. The glass permeabilization was performed at 4 °C in 0.1% Triton X-100 plus 0.1% sodium citrate in PBS. Therefore, after these steps we proceeded with the application of the TUNEL assay.
TUNEL assay
The TUNEL assay (Promega, G3250, USA) was used to identify DNA fragmentation as described with some modifications for metaphase chromosomes and the detached cell population. The slide areas containing the biological material were delimited using a PAP Pen Liquid Blocker to create a visible hydrophobic barrier.
The slides were placed in a humid chamber and 50 μl of equilibration buffer (200 mM potassium cacodylate, pH 6.6 at 25 °C; 25 mM Tris–HCl, pH 6.6 at 25 °C; 0.2 mM DTT 0.25 mg/ml BSA; 2.5 mM cobalt chloride) was added in each area for 10 min. After, a mix containing: 1 μl of recombinant terminal deoxynucleotidyl transferase ( (rTdT) independent template DNA polymerase; 5 μl of nucleotide mix (fluorescein-12-dUTP) and 45 μl of equilibration buffer was added.
The reaction was carried on for 75 min at 37 °C and then stopped by immersing the slides in 2× salinesodium citrate (SSC) for 15 min. After two washes in PBS, propidium iodide (1:1000 dilution) was added for 10 min. After two washes in PBS, 10 μl of 1,4-diazabicyclo (2-2-2) octane, Tris–HCl 1 M pH 8, glycerol (DABCO) was added on each area. Finally, a coverslip was positioned and the fragmented labeled DNA in nuclei or chromosomes or in apoptotic body-like structures was observed by fluorescence microscopy (Olympus BX50, Japan) under a 60× objective.
Several microscope fields were acquired to obtain a significant number of observed cells. In addition, negative and positive controls were analyzed: negative controls were incubated with a buffer containing the fluorescent nucleotide mix (fluorescein 12-dUTP) without the TdT enzyme; positive controls were pretreated with DNAse (10 mg/ ml) for 10 min before the TdT assay, to fragment the total DNA.
Image acquisition was performed by measuring the intensity of autofluorescence in the negative control; this value was considered as the threshold level for the analysis of other samples. Any fluorescence intensity above the negative control’s threshold was therefore considered a positive TdT signal.
The amount of fragmented DNA and nuclei area were analyzed and quantified by ImageJ 1.46r image analyzing software after acquisition of different images using a digital camera (Nikon Sight DS-U1, Japan). This analysis was not applied to the images of the detached cells as there were no defined nuclei present but cellular structures in suspension containing DNA, fragmented or not.
The relative nuclei area was reported by setting the control equal to 1 and comparing the treated samples to this value. In this way, the values relating to the variation of the area of the nuclei in the treated cells indicate a variation of this parameter compared with the nuclear area of the control cells.
Statistical analysis
Data of nuclei fragmentation and DNA fragmentation from detached cellular structures were presented as the mean of three independent experiments ± standard deviation (SD). The data were analyzed by one-way analysis of variance (ANOVA), using the Levene’s test to check homogeneity of variance and the Tukey’s HSD test as post hoc test analysis of significant differences. The area of fragmented and unfragmented nuclei in clone-7 astrocytes, and Caco-2 and MDA-MB-231 cells was presented as the mean of three independent experiments ± SD and comparisons between mean values for each group were carried using a t-test. The analyses were performed using the Statistica 13.2 software (StatSoft, Tulsa, OK, USA), and the level of significance was set to p < 0.05.
Results
To evaluate a possible DNA damage in control and treated cells, analyses on several levels of DNA organization were carried out in all the considered cell lines: nuclei, metaphases and cellular structures recovered from the culture medium.
Analysis of DNA fragmentation in clone-7 astrocytes
Chromosomal suspensions and nuclei of clone-7 astrocytes, isolated in our laboratory, were subjected to the TUNEL assay (Fig. 1a–g). In this cell line, DNA fragmentation was only found at the nuclear level, while no metaphase chromosomes with fragmentation points were observed.
Fluorescent TUNEL assay in astrocytes. Images of representative fields showing nuclei from control and treated cells. DNA fragmentation is shown (a1–f1). Nuclei marked with propidium iodide (a2–f2). Merging of both signals (a3–f3 and g). Control cells are shown(a1–a3). RSV-treated cells are shown (b1–b3). Doxo-treated cells are shown (c1–c3). Doxo + RSV-treated cells (d1–d3). The following controls are shown: positive control (e1–e3), negative control (f1–f3). Enlargement (g) of a particular of control preparation (a3). White arrows indicate two fragmented pyknotic nuclei. Scale bar, 20 μm. Histogram showing TUNEL-positive nuclei, percentage of the total cells (h). Histograms showing the relative area in unfragmented or fragmented nuclei of control (i) and RSV-treated cells (j). Doxo-treated cells (k). Doxo + RSV-treated cells (l). Experiments were performed in triplicate and data are expressed as means ± standard deviation (n = 3 ± SD); there were 100 analyzed cells in each experimental group. Asterisks indicate responses significantly different from those of the controls (h) and significantly different in fragmented versus unfragmented samples (i–l) (*p < 0.05)
An aliquot of 36% of fragmented nuclei was observed in control cells (Fig. 1h).
This percentage decreased to 16% in RSV-treated cells, and it was established to 36% after Doxo treatment. RSV + Doxo cotreatments slightly increased the value to 39%.
In this cell line, a peculiarity about the area of fragmented nuclei was observed. In both the controls and all the treated cells, DNA fragmentation concerned only small-sized nuclei (pyknosis-like nuclei), while all other normal sized nuclei showed unfragmented DNA (Fig. 1g).
The differences between an unfragmented nuclei area and fragmented nuclei area were found significant in all the samples.
The above qualitative and quantitative analyses suggest that in these cells, DNA fragmentation could be related to apoptotic phenomena since the DNA fragmentation signal did not concern the replicating cells (no metaphase chromosomes showed fragmentation points) and all nuclei with characteristics of pyknosis showed fragmented DNA.
Qualitatively, we observed that the culture medium was rich in suspended cellular structures containing DNA. These showed partially fragmented nuclei, as in control cells (Fig. 2a1–a3) and in RSV + Doxo-treated cells (Fig. 2d1–d3). DNA fragmentation and partitioning into small spherical structures were found in RSV-treated cells (Fig. 2b1–b3) and Doxo-treated cells (Fig. 2c1–c3).
Fluorescent TUNEL assay of detached cellular structures, recovered from astrocytes culture medium. Images of representative fields showing fragmented DNA in control and treated cells. DNA fragmentation (a1–d1). DNA marked with propidium iodide (a2–d2). Merging of both signals (a3–d3). Control samples are shown(a1–a3). RSV-treated samples (b1–b3). Doxo-treated samples (c1–c3). Doxo + RSV-treated samples (d1–d3). White arrows indicate spherical corpuscles containing fragmented DNA. Scale bar, 20 μm. Histogram showing TUNEL-positive nuclei, percentage of total cells. Experiments were performed in triplicate and data are expressed as mean ± standard deviation (n = 3 ± SD); there are 100 analyzed cells in each experimental group . Asterisks indicate responses that are significantly different from those of the controls (*p < 0.05)
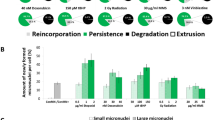
For this detached population, we detected high fragmentation rates, about 33% in controls, 90% in RSV-treated cells, 100% in Doxo-treated cells and 89% in RSV + Doxo-treated cells. However, in all treatments, the amount of detached cellular structures with fragmented DNA was always greater and with a statistically significant difference compared with control cells.
Analysis of DNA fragmentation in Caco-2 cells
A completely different scenario was observed in Caco-2 cells. A peculiar feature was that DNA fragmentation affected chromosomes, uncondensed chromatin nuclei, and, only in treated cells, detached cellular structures.
In control cells, we observed several metaphases rich in fragmented chromosomes and nuclei with uncondensed chromatin but reporting defined points of fragmentation (Fig. 3a1–a3 and g).
Fluorescent TUNEL assay in Caco-2 cells. Images of representative fields showing nuclei and metaphases from control and treated cells. DNA fragmentation is shown (a1–f1). DNA marked with propidium iodide (a2–f2). Merging of both signals (a3–f3 and g). Control cells are shown (a1–a3). RSV-treated cells (b1–b3). Doxo-treated cells (c1–c3). Doxo + RSV-treated cells (d1–d3). Positive control (e1–e3) and negative control (f1–f3) are shown. Enlargement of a representative fragmented metaphase of control cells (g). Scale bar, 20 μm. Histogram showing TUNEL-positive nuclei, percentage of total cells (h). Histograms showing the relative area in unfragmented or fragmented nuclei of control (i); RSV-treated cells (j). Doxo-treated cells (k). Doxo + RSV-treated cells (l). Experiments were performed in triplicate and data are expressed as mean ± standard deviation (n = 3 ± SD). There are 100 analyzed cells in each experimental group. Asterisks indicate responses significantly different from those of the controls (h), and which are significantly different in fragmented versus fragmented samples (i–l) (*p < 0.05)
In RSV-treated cells, on the other hand, no DNA fragmentation signals were found in cells adherent to the substrate (Figs. 3b1–b3). In Doxo-treated cells, many nuclei containing fragmented DNA were found with no distinction between normal or pyknotic nuclei. Specifically, the nuclear fragmentation did not concern the whole area but only some points that appeared as fragmented spots (Figs. 3c1–c3). Cells exposed to the combined treatment (RSV + Doxo), however, showed nuclei with more diffuse fragmented DNA (Figs. 3d1–d3).
As reported in the histogram (Fig. 3h), a high percentage of nuclei containing fragmented DNA (66%) was found only in Doxo-treated cells, for which, compared to control, a statistically significant difference was found.
Analyzing the area of fragmented and unfragmented nuclei, no statistically significant difference was found between the two parameters (Figs. 3i–l), except for Doxo-treated cells and excluding RSV-treated cells, which had no nuclei containing fragmented DNA. Consequently, since metaphase chromosomes represent a clear sign of cell replication, the presence of DNA fragmentation, also in chromosomes, suggested an unlikely relationship with the ongoing death phenomena.
In parallel, an analysis on detached cellular structures was performed. No fragmented or unfragmented nuclear structures were detected in the medium from control cells (Figs. 4a1–a3). Large clusters of nuclei with a high number of fragmented DNA spots were found in the medium of RSV-treated cells (Figs. 4b1–b3). Groups of nuclei with a lower fragmentation rate than the RSV-treated ones were identified in the culture medium recovered from Doxo-treated cells (Figs. 4c1–c3). A high number of nuclei was found in the culture medium of cells treated with RSV + Doxo. All these nuclei exhibited diffuse fragmentation, albeit with a very slight intensity (Figs. 4d1–d3).
Fluorescent TUNEL assay in detached cellular structures, recovered from Caco-2 cells’ culture medium. Images of representative fields showing fragmented DNA in control and treated cells. DNA fragmentation is shown (a1–d1). DNA stained with propidium iodide (a2–d2). Merging of both signals (a3–d3). Control samples are shown (a1–a3). RSV-treated samples (b1–b3). Doxo-treated samples (c1–c3). Doxo + RSV-treated samples (d1–d3). Scale bar, 20 μm. Histogram showing TUNEL-positive nuclei, percentage of total cells. Experiments were performed in triplicate and data are expressed as mean ± standard deviation (n = 3 ± SD). There are 100 analyzed cells in each experimental group. Asterisks indicate responses that are significantly different from those of the controls (*p < 0.05)
Analysis of DNA Fragmentation in MDA–MB–231 cells
Considering the DNA damage, MDA–MB–231 cells showed a different behavior compared with the other two cell lines. The control cells did not show signals of DNA fragmentation, neither at the nuclei level nor at the chromosomes level (Fig. 5a1–a3). Following treatment with RSV, many nuclei containing fragmented DNA were observed (Fig. 5b1–b3). The nuclei fragmentation rate was increased following treatment with Doxo (Fig. 5c1–c3) and greatly reduced in RSV + Doxo cotreated cells. Surprisingly, only 1% of the analyzed metaphases showed some fragmented chromosomes (Fig. 5g). The increase in nuclear fragmentation reached statistical significance, especially for cells exposed to RSV or Doxo (Fig. 5h).
Fluorescent TUNEL assay in MDA-MB-231 cells. Images of representative fields showing nuclei and metaphases from control and treated cells. DNA fragmentation is shown (a1–f1). DNA stained with propidium iodide (a2–f2). Merging of both signals (a3–f3 and g). Control cells are shown(a1–a3). RSV-treated cells (b1–b3). Doxo-treated cells (c1–c3). Doxo + RSV-treated cells (d1–d3). Positive controls (e1–e3) and negative controls (f1–f3) are shown. Enlargements of a representative fragmented metaphase of control cells (g). White arrows indicate chromosomes with fragmented points. Scale bar, 20 μm. Histogram showing the percentage of fragmented nuclei (h). Histogram showing TUNEL-positive nuclei, percentage of total cells (i); RSV-treated cells (j). Doxo-treated cells (k). Doxo + RSV-treated cells (l). Experiments were performed in triplicate and data are expressed as mean ± standard deviation (n = 3 ± SD). There are 100 analyzed cells in each experimental group. Asterisks indicate responses significantly different from those of the controls (h) and that are significantly different in fragmented versus unfragmented samples (i–l) (*p < 0.05)
A relationship was observed between the reduction of the nuclei area and the DNA fragmentation. As reported in the histograms (Fig. 5i–l), the differences between unfragmented nuclei area and fragmented nuclei areas in all the samples were found to be significant.
These data suggest that in MDA–MB–231 cells, DNA fragmentation could be related to apoptotic phenomena, although some cells showed fragmented chromosomes, indicative of DNA damage in replicating cells.
Except for control cells, numerous cellular structures (apoptotic body-like corpuscles) were observed in all treatments in the suspended cell population belonging to the MDA–MB–231 cell line (Fig. 6).
Fluorescent TUNEL assay of detached cellular structures, recovered from MDA-MB-231 medium culture. Images of representative fields showing fragmented DNA from control and treated cells. DNA fragmentation is shown (a1–d1). DNA stained with propidium iodide (a2–d2). Merging of both signals (a3–d3). Control samples are shown (a1–a3). RSV-treated samples (b1–b3). Doxo-treated samples (c1–c3). Doxo + RSV-treated samples (d1–d3). Scale bar, 20 μm. Histogram showing TUNEL-positive nuclei, percentage of total cells. Experiments were performed in triplicate and data are expressed as mean ± standard deviation (n = 3 ± SD). There are 100 analyzed cells in each experimental group. Asterisks indicate responses that are significantly different from those of the controls (*p < 0.05)
As shown in the histogram, 100% of fragmented spots for RSV-treated cells, 58% for Doxo-treated cells, and 88% for combined RSV + Doxo-treated cells was observed.
Since the culture medium recovered from RSV-treated cells showed a greater variety of detached cellular structures, a more detailed analysis was performed.
In all cases, a breakdown of fragmented DNA within the nuclear structure was observed, highlighting the simultaneous presence of intact DNA and fragmented DNA. The latter appeared in spherical body structures clearly detached from the portions of unfragmented DNA (Figs. 7a1–d3).
Fluorescent TUNEL assay of detached cellular structures, recovered from MDA-MB-231 cells’ culture medium. Images of representative fields showing fragmented DNA from RSV-treated cells. DNA fragmentation is shown (a1–d1). DNA stained with propidium iodide (a2–d2). Merging of both signals (a3–d3). Nuclear structure with unfragmented central body and fragmented peripheral DNA islands (a1–a3). Nuclear structure with fragmented central body and two agglomerates of unfragmented peripheral DNA (b1–b3). Nuclear structure divided into two compartments of fragmented and unfragmented DNA (c1–c3). Nuclear structure was divided into three compartments, one with unfragmented DNA and two with fragmented DNA (d1–d3). Scale bar, 5 μm
By considering these morphological aspects, it seems that this cell line is very active in eliminating cells with fragmented DNA (which we found in detached cellular structures), but at the same time, although in a small percentage of the replicating cells showed fragmented DNA as chromosomal evidence.
Discussion
The characterization of cell line markers is of considerable importance in biomedical research (Geraghty et al. 2014). This aspect arouses considerable interest when the toxic or beneficial effects induced by new chemical agents need to be tested. Moreover, it is a common notion that different cell lines respond differently to cytotoxic stimuli.
Numerous experimental approaches show that the genetic characterization of cells, as well as the karyotype analysis, needs to be coupled to other markers (Gäberlein et al. 2023).
DNA is the only molecule in the cell that can be repaired but cannot be completely resynthesized after damage (Moore et al. 2021).
Most of the experimental investigations in the literature aim to observe variations in gene expression or in the modulation of proteins marker levels. The presence of genome fragmentation points represents a difficult component to study since there are no unique and indicative markers of this damage.
DNA damage is one of the main markers that can be used to study multiple effects induced by chemical or physical agents in a multifactorial and extremely complex process. The multifactoriality derives from the fact that DNA fragmentation can be triggered following an exogenous factor, but also in the context of programmed cell death phenomena. In fact, final stages of apoptosis involve the DNA fragmentation and the structuring of chromatin inside apoptotic bodies, particularly extracellular vesicles that could be involved in both programmed cell death and also intercellular communication (Battistelli and Falcieri 2020).
Apoptosis is an active process that occurs in normal cell turnover as well as in stress-induced cell death, representing a homeostatic mechanism for maintaining regular cell populations in tissues (Atkin-Smith and Poon 2017).
Apoptosis progresses through several stages, the first being nuclear chromatin condensation, then nuclear splitting and the frequent appearance of micronuclei and corpuscles containing chromatin remnants, cytosol portions, degraded proteins and DNA fragments.
DNA fragmentation can affect replicating cells and, in this case, the damage can be repaired or transmitted to daughter cells (Huang and Zhou 2021).
Another important point is the research of fragmented DNA in detached cells, since this data contribute to the definition of global toxicity induced by chemicals. In fact, the conditioned medium is a reservoir of useful information that until now was excluded from the various analyses, as reported by Rosochowicz et al. (2023).
In the past, we determined genetic details about three rat astrocyte cell lines with increasingly transformed properties and analyzed them at both cytogenetic, epigenetic (Caradonna et al. 2020), and proteomic levels (Schiera et al. 2023). We found that the most modified cell line (A-FC6) showed epigenetic and chromosomal alterations typical of tumor cells, such as the cytogenetic marker i(8q) in 100% of metaphases. Subsequently, we isolated the clone-7 astrocytes selected for a shortening of cell doubling time. This clone further showed the same epigenetics features of A-FC6 and more,interestingly, the cytogenetic marker i(8q) was present in double copy in about 15% of metaphases (data not shown). In a schematic line of increasing genomic instability, we put the clone-7 astrocytes at the second place, after the normal cells. Therefore, regarding the here described in vitro model system, starting from the normality to the genetically more unstable transformed cells, we assign the point “0” to normal cells and the point “1” to the clone-7 astrocytes.
It is known the peculiarity of the widely used Caco-2 cell line. These cells, although tumoral, assume a different behavior in long-term cultures, as they show the ability to differentiate into “normal” cells. It is evident that their genomic instability is still at reversible levels, far from that of an aggressive tumor cell that does not possess this possibility. For this peculiarity we assign to these cells, here cultured for the short-term, the point “2” of the described in vitro model system.
The MDA-MB-231 triple negative breast cancer cell line is composed of cells with a high degree of genomic instability and also a high metastatic potency (Huang et al. 2020). For these features, we assign to this cell line the point “3” of the described in vitro model system.
In the present paper, the comparative analysis carried out with three different cell lines (clone-7 astrocytes, Caco-2 cells, and MDA-MB-231 cells) proposes a model of increasing genomic instability. We report that cells respond in a different way to the effects induced by toxic agents, since there were different levels of fragmentation in nuclear DNA (whole nuclei), single chromosomes, and DNA associated with extracellular structures (recovered from the culture medium). Although our aim was not to study apoptosis but the presence of DNA damage at different levels, in some cases we noticed that such apoptotic body-like structures were abundantly present in the culture medium and were of considerable importance for the overall study of DNA fragmentation.
Our integrated and comparative approach allows us to highlight some key points. In many cases, when apoptotic phenomena or other DNA fragmentation processes are investigated, it is not enough to study the phenomena at the cellular level, but it is necessary to integrate the data with the phenomena that are occurring at the extracellular level.
As an example, clone-7 astrocytes showed a nuclear fragmentation rate in adherent cells, both in controls and in Doxo-treated and RSV + Doxo cotreated cells ranging from 16% to 40%. Integrating the data obtained from the culture medium, compared with the control, we note that all the treated cells showed a nuclear fragmentation rate equal to about 1.5 times higher than those treated with RSV and about 2 times higher than those treated with Doxo and cotreated with RSV + Doxo. These cells, overall, were more sensitive to the exposure of Doxo.
On the other hand, Caco-2 cells showed a low rate of nuclear fragmentation in substrate-adherent cells, except for Doxo-treated cells, for which a value equal to about 65% was reached. Integrating the data detected by the analysis of the culture medium, we noted that in the controls we did not detect any signal of DNA fragmentation (this is in accordance with the type of fragmentation, which involved replicating cells and not dying cells, given that the points of fragmentation were evident in metaphase chromosomes). Although RSV− and RSV+ Doxo-treated cells did not show a high level of nuclear fragmentation in substrate-attached cells, with our integrated analysis we noted that in the culture medium the rate of nuclei containing fragmented DNA reached values ranging from 20 to 25-fold higher than those of the control, proving that the nuclear fragmentation phenomena were in an advanced stage, such as to promote a loss of adhesion of the cells. The cells exhibited sensitivity to both RSV and Doxo, to such an extent that cotreatment of RSV + Doxo resulted in increased nuclear fragmentation.
The MDA–MB-231 control cells did not exhibit nuclear fragmentation in the substrate-adherent cells. These cells showed a high rate of nuclear fragmentation after exposure to Doxo (about three times higher compared with those treated with RSV alone). In cotreatment, at least for substrate-adherent cells, the nuclear fragmentation rate was basal. By integrating the data obtained from the culture medium, we noted that there was a compensation as the levels of nuclear fragmentation were low in the cells that showed high values of nuclear fragmentation at the level of substrate-adherent cells (Doxo-treated cells) and high in cells showing low levels of nuclear fragmentation related to substrate-adherent cells (RSV-treated and RSV + Doxo cotreated cells). This integrated view allows us to understand that these cells are sensitive to treatment with both molecules, with a slightly higher entity for treatments with RSV.
Conclusions
The data, obtained with a modern revisitation of a TUNEL assay, showed a comparative vision of three genomically different cell lines and allowed us to gain data for the study of DNA fragmentation at different levels. Through our analysis, we were able to observe that the clone-7 astrocytes were the only cells demonstrating the absence of DNA fragmentation at the chromosomal level. These cells presented chromatin condensation and pyknotic nuclei corresponding to the fragmented DNA signals. This aspect appeared to be present in control cells and was modulated by exposure to single or combined treatments with RSV/Doxo. No chromosomal fragmentation was found in any case. Evidently, the cells choose to either repair the damage or simply activate apoptotic phenomena. Caco-2 cells showed fragmented metaphase chromosomes, proving that the DNA damage was transmitted to the daughter cells. The treatments seemed to reduce this effect and, in particular, RSV promoted a major induction of fragmentation at the nuclear level, most noticeable in the suspended than in the substrate-adherent cells. Thus, we can hypothesize that Caco-2 cells have no DNA repair mechanism activated. However, MDA–MB-231 cells, analyzed by this alternative TUNEL assay approach, showed a very low or no fragmented metaphases, suggesting a probable activation of DNA repair mechanisms.
Overall from all these results we may assert that the three cell lines respond differently in terms of DNA fragmentation, after RSV and DOXO treatments and the RSV–DOXO cotreatment, probably based on their biological status.
By applying, for the first time, the well-known TUNEL test on nuclei and metaphases, we obtained data relating to DNA fragmentation that can significantly contribute to characterizing the tumor phenotype of a cell type, either in a clinical setting or to test the effect of candidate molecules such as new anticancer drugs.
Data availability
The data and material supporting the findings of this study are available from the corresponding author, Fabio Caradonna, upon request.
Abbreviations
- TUNEL:
-
Terminal deoxynucleotidyl transferase dUTP nick end labeling
- RSV:
-
Resveratrol
- Doxo:
-
Doxorubicin
- GI:
-
Genomic instability
- NGS:
-
Next-generation sequencing
- DMEM:
-
Dulbecco’s modified eagle’s medium
- DMSO:
-
Dimethyl sulfoxide
- IC50:
-
Half maximal inhibitory concentration
- PBS:
-
Phosphate-buffered saline
- rTdT:
-
Enzyme terminal deoxynucleotidyl transferase, recombinant
- SSC:
-
Saline-sodium citrate
- DABCO:
-
1,4-DiAzaBicyclo ( 2–2-2) octane
- SD:
-
Standard deviation
References
Agnello M, Chiarelli R, Martino C et al (2016) Autophagy is required for sea urchin oogenesis and early development. Zygote 24:918–926. https://doi.org/10.1017/S0967199416000253
Atkin-Smith GK, Poon IKH (2017) Disassembly of the dying: mechanisms and functions. Trends Cell Biol 27:151–162. https://doi.org/10.1016/j.tcb.2016.08.011
Battistelli M, Falcieri E (2020) Apoptotic bodies: particular extracellular vesicles involved in intercellular communication. Biology (basel) 9:21. https://doi.org/10.3390/biology9010021
Bong AHL, Bassett JJ, Roberts-Thomson SJ, Monteith GR (2020) Assessment of doxorubicin-induced remodeling of Ca2+ signaling and associated Ca2+ regulating proteins in MDA-MB-231 breast cancer cells. Biochem Biophys Res Commun 522:532–538. https://doi.org/10.1016/j.bbrc.2019.11.136
Bouwman P, Jonkers J (2012) The effects of deregulated DNA damage signalling on cancer chemotherapy response and resistance. Nat Rev Cancer 12:587–598. https://doi.org/10.1038/nrc3342
Caradonna F (2015) Nucleoplasmic bridges and acrocentric chromosome associations as early markers of exposure to low levels of ionising radiation in occupationally exposed hospital workers. Mutagenesis 30:269–275. https://doi.org/10.1093/mutage/geu068
Caradonna F, Schiera G, Di Liegro CM et al (2020) Establishment and preliminary characterization of three astrocytic cells lines obtained from primary rat astrocytes by sub-cloning. Genes (basel) 11:1502. https://doi.org/10.3390/genes11121502
Carusillo A, Mussolino C (2020) DNA damage: from threat to treatment. Cells 9:1665. https://doi.org/10.3390/cells9071665
Chatterjee N, Walker GC (2017) Mechanisms of DNA damage, repair, and mutagenesis. Environ Mol Mutagen 58:235–263. https://doi.org/10.1002/em.22087
Chiarelli R, Roccheri MC (2012) Heavy metals and metalloids as autophagy inducing agents: focus on cadmium and arsenic. Cells 1:597–616. https://doi.org/10.3390/cells1030597
Chiarelli R, Agnello M, Bosco L, Roccheri MC (2014) Sea urchin embryos exposed to cadmium as an experimental model for studying the relationship between autophagy and apoptosis. Mar Environ Res 93:47–55. https://doi.org/10.1016/j.marenvres.2013.06.001
Chiarelli R, Martino C, Agnello M et al (2016) Autophagy as a defense strategy against stress: focus on Paracentrotus lividus sea urchin embryos exposed to cadmium. Cell Stress Chaperones 21:19–27. https://doi.org/10.1007/s12192-015-0639-3
Chiarelli R, Martino C, Roccheri MC (2019) Cadmium stress effects indicating marine pollution in different species of sea urchin employed as environmental bioindicators. Cell Stress Chaperones 24:675–687. https://doi.org/10.1007/s12192-019-01010-1
Chiarelli R, Martino C, Roccheri MC, Cancemi P (2021) Toxic effects induced by vanadium on sea urchin embryos. Chemosphere 274:129843. https://doi.org/10.1016/j.chemosphere.2021.129843
Chiarelli R, Martino C, Roccheri MC, Geraci F (2022a) Vanadium toxicity monitored by fertilization outcomes and metal related proteolytic activities in Paracentrotus lividus embryos. Toxics 10:83. https://doi.org/10.3390/toxics10020083
Chiarelli R, Martino C, Scudiero R, Geraci F (2022b) Vanadium modulates proteolytic activities and MMP-14-like levels during Paracentrotus lividus embryogenesis. Int J Mol Sci 23:14238. https://doi.org/10.3390/ijms232214238
Chiarelli R, Scudiero R, Memoli V et al (2022c) Toxicity of vanadium during development of sea urchin embryos: bioaccumulation, calcium depletion, ERK modulation and cell-selective apoptosis. Int J Mol Sci 23:6239. https://doi.org/10.3390/ijms23116239
Gäberlein K, Schröder SK, Nanda I et al (2023) Genetic characterization of rat hepatic stellate cell line PAV-1. Cells 12:1603. https://doi.org/10.3390/cells12121603
Galvao J, Davis B, Tilley M et al (2014) Unexpected low-dose toxicity of the universal solvent DMSO. FASEB J 28:1317–1330. https://doi.org/10.1096/fj.13-235440
Gavrieli Y, Sherman Y, Ben-Sasson SA (1992) Identification of programmed cell death in situ via specific labeling of nuclear DNA fragmentation. J Cell Biol 119:493–501. https://doi.org/10.1083/jcb.119.3.493
Geraghty RJ, Capes-Davis A, Davis JM et al (2014) Guidelines for the use of cell lines in biomedical research. Br J Cancer 111:1021–1046. https://doi.org/10.1038/bjc.2014.166
Gomes L, Sorgine M, Passos CLA et al (2019) Increase in fatty acids and flotillins upon resveratrol treatment of human breast cancer cells. Sci Rep 9:1–11. https://doi.org/10.1038/s41598-019-50416-5
Gomes L, Viana L, Silva JL et al (2020) Resveratrol modifies lipid composition of two cancer cell lines. Biomed Res Int 2020:5393041. https://doi.org/10.1155/2020/5393041
González-Marín C, Gosálvez J, Roy R (2012) Types, causes, detection and repair of DNA fragmentation in animal and human sperm cells. Int J Mol Sci 13:14026–14052. https://doi.org/10.3390/ijms131114026
Huang R, Zhou PK (2021) DNA damage repair: historical perspectives, mechanistic pathways and clinical translation for targeted cancer therapy. Signal Transduct Target Ther 6:254. https://doi.org/10.1038/s41392-021-00648-7
Huang Z, Yu P, Tang J (2020) Characterization of triple-negative breast cancer MDA-MB-231 cell spheroid model. Onco Targets Ther 13:5395–5405. https://doi.org/10.2147/OTT.S249756
Jackson SP, Bartek J (2009) The DNA-damage response in human biology and disease. Nature 461:1071–1078. https://doi.org/10.1038/nature08467
Jadid MFS, Aghaei E, Taheri E et al (2021) Melatonin increases the anticancer potential of doxorubicin in Caco-2 colorectal cancer cells. Environ Toxicol 36:1061–1069. https://doi.org/10.1002/tox.23105
Kim BS, Park JJ, Edler L et al (2002) New measure of DNA repair in the single-cell gel electrophoresis (comet) assay. Environ Mol Mutagen 40:50–56. https://doi.org/10.1002/em.10090
Kitazumi I, Tsukahara M (2011) Regulation of DNA fragmentation: the role of caspases and phosphorylation. FEBS J 278:427–441. https://doi.org/10.1111/j.1742-4658.2010.07975.x
Kroemer G, Galluzzi L, Vandenabeele P et al (2009) Classification of cell death: recommendations of the nomenclature committee on cell death 2009. Cell Death Differ 16:3–11. https://doi.org/10.1038/cdd.2008.150
Lin CJ, Chen TH, Yang LY, Shih CM (2014) Resveratrol protects astrocytes against traumatic brain injury through inhibiting apoptotic and autophagic cell death. Cell Death Dis 5:e1147. https://doi.org/10.1038/cddis.2014.123
Liu B, Zhou Z, Zhou W et al (2014) Resveratrol inhibits proliferation in human colorectal carcinoma cells by inducing G1/S-phase cell cycle arrest and apoptosis through caspase/cyclin-CDK pathways. Mol Med Rep 10:1697–1702. https://doi.org/10.3892/mmr.2014.2406
Luparello C, Asaro DML, Cruciata I et al (2019) Cytotoxic activity of the histone deacetylase 3-selective inhibitor Pojamide on MDA-MB-231 triple-negative breast cancer cells. Int J Mol Sci 20:804. https://doi.org/10.3390/ijms2004080410.3390/ijms20040804
Luparello C, Cruciata I, Joerger AC et al (2021) Genotoxicity and epigenotoxicity of carbazole-derived molecules on MCF-7 breast cancer cells. Int J Mol Sci 22:3410. https://doi.org/10.3390/ijms22073410
Majtnerová P, Roušar T (2018) An overview of apoptosis assays detecting DNA fragmentation. Mol Biol Rep 45:1469–1478. https://doi.org/10.1007/s11033-018-4258-9
Mannino G, Caradonna F, Cruciata I et al (2019) Melatonin reduces inflammatory response in human intestinal epithelial cells stimulated by interleukin-1β. J Pineal Res 67:e12598. https://doi.org/10.1111/jpi.12598
Martino C, Byrne M, Roccheri MC, Chiarelli R (2021) Interactive effects of increased temperature and gadolinium pollution in Paracentrotus lividus sea urchin embryos: a climate change perspective. Aquat Toxicol 232:105750. https://doi.org/10.1016/j.aquatox.2021.105750
Maszczyk M, Banach K, Karkoszka M et al (2022) Chemosensitization of U-87 MG glioblastoma cells by neobavaisoflavone towards doxorubicin and etoposide. Int J Mol Sci 23:5621. https://doi.org/10.3390/ijms23105621
Moore CL, Savenka AV, Basnakian AG (2021) Tunel assay: a powerful tool for kidney injury evaluation. Int J Mol Sci 22:412. https://doi.org/10.3390/ijms22010412
Naselli F, Belshaw NJ, Gentile C et al (2015) Phytochemical indicaxanthin inhibits colon cancer cell growth and affects the DNA methylation status by influencing epigenetically modifying enzyme expression and activity. J Nutrigenet Nutrigenomics 8:114–127. https://doi.org/10.1159/000439382
Ou HL, Schumacher B (2018) DNA damage responses and p53 in the aging process. Blood 131:488–495. https://doi.org/10.1182/blood-2017-07-746396
Razak NA, Abu N, Ho WY et al (2019) Cytotoxicity of eupatorin in MCF-7 and MDA-MB-231 human breast cancer cells via cell cycle arrest, anti-angiogenesis and induction of apoptosis. Sci Rep 9:1514. https://doi.org/10.1038/s41598-018-37796-w
Rosochowicz MA, Lach MS, Richter M et al (2023) Conditioned medium—is it an undervalued lab waste with the potential for osteoarthritis management? Stem Cell Rev Reports 19:1185–1213. https://doi.org/10.1007/s12015-023-10517-1
Sadeghi-Aliabadi H, Mosavi H, Mirian M et al (2012) The cytotoxic and synergistic effects of flavonoid derivatives on doxorubicin cytotoxicity in hela, MDA-MB-231, and HT-29. Iran J Toxicol 5:558–564
Schiera G, Cancemi P, Di Liegro CM et al (2023) An in vitro model of glioma development. Genes (basel) 14:990. https://doi.org/10.3390/genes14050990
Sirchia R, Luparello C (2007) Mid-region parathyroid hormone-related protein (PTHrP) and gene expression of MDA-MB231 breast cancer cells. Biol Chem 388:457–465. https://doi.org/10.1515/BC.2007.059
Tsukada T, Watanabe M, Yamashima T (2001) Implications of CAD and DNase II in ischemic neuronal necrosis specific for the primate hippocampus. J Neurochem 79:1196–1206. https://doi.org/10.1046/j.1471-4159.2001.00679.x
van Dijk EL, Jaszczyszyn Y, Naquin D, Thermes C (2018) The third revolution in sequencing technology. Trends Genet 34:666–681. https://doi.org/10.1016/j.tig.2018.05.008
Volpes S, Cruciata I, Ceraulo F et al (2023) Nutritional epigenomic and DNA-damage modulation effect of natural stilbenoids. Sci Rep 13:1–12. https://doi.org/10.1038/s41598-022-27260-1
Xu X, Lai Y, Hua ZC (2019) Apoptosis and apoptotic body: disease message and therapeutic target potentials. Biosci Rep 39:BSR20180992. https://doi.org/10.1042/BSR20180992
Yoshida A, Pommier Y, Ueda T (2006) Endonuclease activation and chromosomal DNA fragmentation during apoptosis in leukemia cells. Int J Hematol 84:31–37. https://doi.org/10.1007/BF03342699
Zhang JH, Xu M (2000) DNA fragmentation in apoptosis. Cell Res 10:205–211. https://doi.org/10.1038/sj.cr.7290049
Acknowledgements
The authors are grateful to Prof. M.C. Roccheri for precious technical scientific collaboration.
Funding
Open access funding provided by Università degli Studi di Palermo within the CRUI-CARE Agreement. This research was partially funded by (i) grants from University of Palermo (FFR 2021–2023 to R.C., F.C., and F.N); (ii) National Biodiversity Future Center (identification code CN00000033, CUP B73C22000790001) on “Biodiversity,” financed under the National Recovery and Resilience Plan (NRRP), Mission 4, Component 2, Investment 1.4 “Strengthening of research structures and creation of R and D “national champions” on some “Key Enabling Technologies”—call for tender no. 3138, 16 December 2021, rectified by Decree no. 3175, 18 December 2021, of the Italian Ministry of University and Research funded by the European Union—NextGenerationEU. Award number: project code CN_00000033, concession decree no. 1034 of 17 June 2022 adopted by the Italian Ministry of University and Research, CUP B73C22000790001, project title “National Biodiversity Future Center—NBFC” (F.C. and C.L.).
Author information
Authors and Affiliations
Contributions
R.C. and F.C. designed the study. R.C., P.S.C., and F.N. performed the cytogenetic experiments. C.L. and C.M. interpreted the data. R.C., P.S.C., S.V., I.C., and R.V. wrote the draft. All authors commented on the manuscript and contributed to the final version.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Naselli, F., Cardinale, P.S., Volpes, S. et al. An alternative approach of TUNEL assay to specifically characterize DNA fragmentation in cell model systems. Histochem Cell Biol (2024). https://doi.org/10.1007/s00418-024-02306-9
Accepted:
Published:
DOI: https://doi.org/10.1007/s00418-024-02306-9