Abstract
Background
Down Syndrome Regression Disorder (DSRD) is a rare and poorly understood disorder of the central nervous system, characterized by acute or subacute neuropsychiatric symptoms in previously healthy individuals with Down syndrome (DS). Many patients exhibit immunotherapy-responsiveness, indicative of immune dysregulation as a potential underlying etiology. While hypotheses are emerging regarding the role of interferon signaling in DSRD and other autoimmune conditions associated with DS, it is unclear why a small subset of individuals with DS develop DSRD. The aim of this study was to investigate genes of immune regulation in persons with DSRD.
Methods
This study included individuals with DSRD aged 10–30 years with trio exome sequencing performed during the diagnostic work up. Descriptive statistics and univariate analysis (Chi-square and Fisher’s exact test) were used to describe and compare the characteristics of individuals with and without variants.
Results
Forty-one individuals with DSRD had trio exome sequencing results. Eight (20%) had heterozygous de novo variants of immune regulatory genes, with four variants being pathogenic or likely pathogenic (UNC13D, XIAP, RNASEH2A, and DNASE1L3). All genes harboring pathogenic variants were associated with interferon type-1 inflammatory response. Individuals harboring variants were more likely to have a preceding trigger (p = 0.03, 95% CI 1.21–97.06), rapid clinical decline in less than 1 month (p = 0.01, 95% CI 1.67–52.06), and MRI abnormalities (p < 0.001, 95% CI 4.89–527.71).
Discussion
A distinct subset of individuals with DSRD exhibited pathogenic variants in immune regulation genes associated with interferon-mediated inflammatory response, coinciding with previously established links between these genes and interferonopathies such as Aicardi-Goutieres syndrome. Our observations suggest that these variants might potentially contribute to the development of DSRD in individuals with DS.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Down syndrome (DS) is one of the most common genetic disorders, with an incidence of 1 in every 800 live births in the United States [1]. An emerging condition, Down Syndrome Regression Disorder (DSRD), has been reported with increasing frequency over the last 2 decades [2,3,4]. DSRD is an acute or subacute neuropsychiatric disorder which primarily occurs in individuals with DS between ages 10 and 30 years [5]. It includes symptoms of bradykinesia, catatonia, developmental regression, encephalopathy, insomnia, mutism, and loss of ability to perform activities of daily living [2,3,4, 6,7,8]. This condition is often severe and can significantly impact the quality of life and autonomy of persons with DS.
The etiology of DSRD is not clear. According to several multi-center studies, as high as 85% of individuals with DSRD have been reported to respond to immunotherapy [5,6,7, 9]. There is a higher likelihood of immunotherapy responsiveness in individuals with neurodiagnostic abnormalities (e.g., abnormal MRI or cerebrospinal fluid studies) though the precise origin of these abnormalities and their link to immunotherapy response remain unclear [6]. Individuals with DS are reported to have other co-occurring genetic and autoimmune conditions [10]. Several studies have examined downstream immune dysregulation in individuals with DS and have hypothesized that amplified interferon signaling secondary to increased interferon receptor gene dosage (due to trisomy of chromosome 21) as a major contributor [11,12,13]. While hypotheses are emerging regarding the role of interferon signaling in DSRD and other autoimmune conditions associated with DS, it is unclear why a small subset of individuals with DS develop DSRD.
Susceptibility to polygenic diseases stem partially from variations in environmental stressors and socioeconomic factors. However, genetic variations also play a significant role. Traditionally, the quest to identify genetic variants linked to complex or polygenic traits and diseases has relied on genome-wide association studies (GWAS), which identify common variants (i.e., single nucleotide polymorphisms, SNP). However, rare variants with potentially larger effect-size are not identified through GWAS. In addition, variants identified through GWAS often localize to non-coding regions, complicating results interpretation [14]. Identification of rare variants in the coding regions could provide insights into the underlying pathophysiological mechanisms, particularly for conditions where a clear etiology has yet to be established.
This study sought to evaluate contribution of rare variants within coding regions of genes related to immune regulation to DSRD. Additionally, we aimed to investigate whether these variants are associated with specific clinical characteristics such as disease severity, abnormality in neurodiagnostic studies, or responsiveness to immunotherapy.
Methods
Patient population and approvals
A retrospective, chart-based, review was performed of individuals enrolled in the DS Neurology program at the Children’s Hospital Los Angeles (CHLA) between July 1st, 2019, and October 1st, 2023. This included patients physically evaluated at the primary center or those with whom chart review was performed in secondary consultation with an outside physician. Institutional Review Board (IRB) approval was obtained through CHLA and the University of Southern California, and consent and assent were waived for this study as the clinical data reviewed had already been obtained.
Inclusion criteria
All individuals in both cohorts required a genetically confirmed diagnosis of Trisomy 21. Individuals with either possible or probable DSRD per international consensus criteria were included [15]. Further, all individuals had to be aged 10–30 years at the time of symptom onset to qualify for the study. Individuals in the genetic testing cohort required trio testing (biological parents) to assess if variants were de novo.
Exclusion criteria
Individuals who had translocation or mosaic forms of DS were excluded. Further, if an individual was found to have an alternative explanation for their symptoms (e.g., cerebrovascular accident or epilepsy), that participant was excluded.
Data collection
Data were collected through retrospective review of the electronic medical record (EMR). Demographic data, medical and surgical history, along with clinical findings and diagnostic results, were obtained from clinical records. Any potential triggers linked to the onset of symptoms were noted, including infection, change of school or home environment, loss of care giver or friend, death in family, physical or emotional abuse, or any medical events. These factors were considered potentially contributory if they occurred within 12 weeks before the onset of symptoms. All individuals included in this study had received Bush Francis Catatonia Rating Scale (BFCRS) [16] and Neuropsychiatric Inventory-Questionnaire (NPI-Q) [17] at baseline and 24 weeks following immunotherapy initiation. Immunotherapy consisted of intravenous immunoglobulin (IVIg) and/or steroid administration. Immunotherapy-responsiveness was defined as a 50% or more improvement on BFCRS or NPIQ scores.
Neurodiagnostic abnormalities
EEG abnormality was defined as focal or generalized slowing, focal or generalized epileptiform discharges out of any cortex, or seizure. MRI had to be performed with and without contrast administration on a 3 T scanner. All abnormal findings except for structural malformations (e.g., Chiari malformation) were categorized as abnormal. Any of the following cerebrospinal fluid (CSF) findings were considered abnormal: WBC count > 5 cells/mm3, total protein > 60 mg/dL, presence of oligoclonal bands, an IgG index of > 0.66, and/or an elevated neopterin (> 33 nmol/mL). The presence of over 1000 RBC in the CSF indicated hemorrhagic contamination and, therefore, the results were excluded from analysis [6].
Next generation sequencing and bioinformatic analysis
Next-generation sequencing of exomes was performed. Variants of uncertain significance were first analyzed using Varsome Human Genomic Variant Search Engine and the hg38 genome assembly [18]. The Single Nucleotide Polymorphism Database (dbSNP) reference SNP ID number (rs number) was reported when available. Variant frequency was adapted from Varsome gnomAD Exome v2.1.1. Conservation scores were adapted from PhyloP100 scores available on Varsome browser. Genomic location was taken from the Varsome Variant Tab and used to calculate Combined Annotation-Dependent Depletion (CADD) scores using the Single Nucleotide Variant Lookup tool and the GRCh38-v1.6 CADD model [19]. Maximum CADD scores were reported.
For in silico prediction of variant function, we first downloaded FASTA sequences for each protein using the National Library of Medicine’s protein search. FASTA sequences were then analyzed using Polymorphism Phenotyping v2 (PolyPhen-2) and Sorting Intolerant from Tolerant (SIFT) Sequence tool [20, 21]. For meta in silico prediction, we used the meta scores available on Varsome browser, based on the aggregated evidence from different in silico predictors (MetaRNN, REVEL, BayesDel noAF, BayesDel addAF, MetaLR, MetaSVM) [18].
Variant calling
Interpretation of variants were performed according to the joint consensus recommendation of the American College of Medical Genetics (ACMG) and Genomics and the Association for Molecular Pathology (AMP) [22]. Results were interpreted by a board-certified clinical molecular geneticist. For quantitative variant classification, a naturally converted Bayesian formulation point system was used [23].
Pathway analysis
To perform Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis using clinical diagnosis gene lists, lists of gene names were first imported into the NIAID/NIH Database for Annotation, Visualization and Integrated Discovery (DAVID) Bioinformatics Resources v.6.8 Analysis Wizard Tool. “OFFICIAL_GENE_SYMBOL” was selected in the Identifier field, Homo sapiens was inputted within the Species field, and “Gene List” was selected under List Type. Next, the imported gene list was analyzed using the DAVID Functional Annotation Tool set, specifically looking within the “Pathway” and “KEGG_Pathway” tools [24, 25].
Statistical analysis
Descriptive statistics were used to summarize the characteristics of patients included in this study. Univariate analyses (Chi square and Fisher’s exact test) were used to compare the characteristics of individuals with and without variants. For KEGG pathway analysis, p values and Benjamini corrections were calculated. Benjamini values of < 0.05 were considered statistically significant. Analyses were performed using DAVID Bioinformatics Resources 6.8.
Results
Of the 347 individuals with DSRD identified for inclusion, 41 (12%) had trio exome sequencing performed during their diagnostic work up. Demographic and clinical features of the cohort are reported in Table 1. A high percentage of individuals with exome sequencing testing had commercial insurance (35/41, 85%) which was significantly higher than the rate of commercial insurance in our comparator DSRD cohort (159/306, 52%) (p ≤ 0.001, 95% CI 2.20–13.19). There were no other statistically significant differences with regards to the DSRD cohort without exome sequencing.
Of all individuals with exome sequencing testing, eight (20%) were identified as having de novo heterozygous variants classified as pathogenic, likely pathogenic or uncertain clinical significance. Variants of eight immune regulatory genes were detected (Table 2). Of the eight variants identified, four variants were classified as pathogenic (RNASEH2A: NM_006397.3:c.557G>A) or likely pathogenic (UNC13D: NM_199242.3:c.652G>T, XIAP: NM_001378592.1:c.655G>A, and DNASE1L3: NM_004944.4:c.581G>A). Tables S1 and S2 summarize clinical features and neurodiagnostic findings in individuals with variants. On KEGG pathway analysis, the identified enriched pathway (NOD2 receptor signaling) did not reach statistical significance.
Across all gene variants identified, no individual harboring a variant exhibited a clinical phenotype consistent with a non-DSRD clinical disorder associated with the gene of concern (e.g., Aicardi-Goutières syndrome). However, individuals with identified variants were noted to have differences in clinical characteristics compared to individuals with no variants (Table 3). Notably, individuals with variants were more likely to have a preceding trigger (p = 0.03, 95% CI 1.21–97.06), rapid clinical decline with nadir in less than 1 month (p = 0.01, 95% CI 1.67–52.06), and MRI abnormalities (p < 0.001, 95% CI 4.89–527.71). The type of trigger (e.g., temporally associated environmental, social, or infectious event) was not associated with a higher likelihood of harboring a variant (p = 0.63, 95% CI 0.37–1.96). Other clinical factors such as abnormality of EEG, CSF or serum cytokine panel were not statistically significant nor was the presence of catatonia, BFCRS > 20 or immunotherapy-responsiveness.
Discussion
This study represents the first extensive examination of exome sequencing in individuals with DSRD. The authors identified heterozygous de novo variants in immune regulation genes in 20% of individuals with DSRD, including pathogenic or likely pathogenic variants in DNASE1L3, RNASEH2A, UNC13D, and XIAP in 10% of cases. Among those with variants, a notable pattern emerged, indicating a higher likelihood of preceding triggers, a more rapid clinical decline, and higher rate of neuroimaging abnormalities. Individuals with these gene variants were previously healthy and did not exhibit the previously recognized conditions associated with pathogenic variants in RNASEH2A, UNC13D, XIAP, or DNASE1L3. There was no significant difference in immune therapy responsiveness between individuals with and without variants, although all individuals with variants were immune therapy responsive.
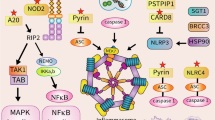
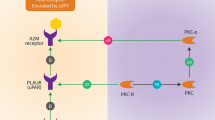
Trisomy 21 is known to cause an amplified interferon response [11]. The chromosome 21 carries genes encoding four out of the six interferon receptor subunits. The presence of an extra copy of chromosome 21 leads to increased gene dosage, thereby elevating the expression of interferon receptors [11]. The increased expression amplifies the JAK/STAT signaling pathway, culminating in the upregulation of interferon-stimulated gene (ISG) expression in response to various triggers individuals with DS (Fig. 1) [26]. Our study did not find any one specific type of trigger (e.g., infectious) being preferentially associated with the harboring of gene variants although it is possible that the low total number of individuals with triggers was not sufficient to detect any effect. Alternatively, this result could also be interpreted as the etiology of the biological stress to the system may be irrelevant and that any activation of the inflammatory response may be sufficient to cause an overactive signaling pathway to cause pathology. This will be an important area of future investigation.
Interferon type-1 response. (1) Pathogens and damaged cells release nucleic acids. (2) The cytosolic DNA activates the cGAS-STING pathway, resulting in phosphorylation of interferon regulatory factor 3 (IRF3). (3) The activated IRF3 induces transcription of interferons in the nucleus. (4) Interferon type-1 binds to the heterodimeric IFNα/β receptor (IFNAR), activating the associated tyrosine kinases JAK1 and TYK2, which in-turn activate STAT1 and STAT2 transcription factors. (5) STAT homo/heterodimers, combined with IRF-9, activate the transcription of interferon-stimulated genes (ISGs), resulting in inflammation. (6) Chromosome 21 encodes four of the INFAR subunits. Trisomy of chromosome 21 results in increased expression of INFAR, resulting in an amplified JAK/STAT response to triggers. *Proteins encoded by genes harboring a pathogenic variant identified in DSRD are highlighted with a red star. UNC13D regulates the inflammatory response by inhibiting the oligomerization of STING. Deoxyribonuclease 1 like 3 (DNase1L3) degrades cell-free DNA, and Ribonuclease H2 (RNase H2) cleaves RNA-DNA hybrids, preventing cytosolic entry and accumulation of unprocessed nucleic acids
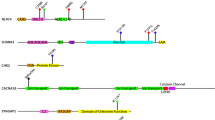
It is important to note that all genes harboring pathogenic variants identified in our cohort encode proteins associated with interferon type-1 inflammatory response. The Unc-13 homolog D (UNC13D) protein regulates the inflammatory response by inhibiting the oligomerization of Stimulator of interferon genes (STING) [27]. Deoxy- ribonuclease 1 like 3 (DNase1L3) degrades cell-free DNA [28], and Ribonuclease H2 (RNase H2) cleaves RNA–DNA hybrids, preventing the inflammatory response triggered by cytosolic entry and accumulation of unprocessed nucleic acids [29]. X-linked inhibitor of apoptosis protein (XIAP) increases the stability of Interferon regulatory factor 7 (IRF7), a transcription factor that drives expression of type 1 interferons (Fig. 1) [30].
Furthermore, pathogenic variants in the identified genes are linked to interferonopathies, autoimmune, and autoinflammatory disorders such as Aicardi-Goutieres syndrome (AGS), familial hemophagocytic lymphohistiocytosis (HLH), systemic lupus erythematous (SLE), and inflammatory bowel disease (IBD). For instance, biallelic variants of UNC13D are associated with familial HLH type 3 [31,32,33]. Pathogenic variants in genes encoding RNase H2 subunits (RNASEH2A, RNASEH2B, and RNASEH2C) are known causes of AGS [34, 35]. In addition, heterozygous variants of RNASEH2 are shown to be associated with SLE and increased risk of systemic autoimmunity [36]. Pathogenic variants of DNASE1L3 are associated with familial SLE, marked by childhood-onset of anti-ds DNA positivity [37,38,39]. XIAP deficiency is marked by immune system dysfunction and a wide range autoinflammatory manifestations, including HLH and IBD [40]. Interestingly, serum analysis of individuals in our cohort with identified pathogenic variants was unremarkable and not consistent with any of the mentioned inflammatory conditions as noted in prior reports [6].
We speculate that deleterious variations in genes associated with interferon response may increase the likelihood of abnormal responses to common pathogens or stressors. Instances of rapid deterioration in response to common infections have been described in individuals with AGS and HLH, conditions arising from biallelic pathogenic variants in genes identified in this study [32, 34, 41]. This suggests that heterozygous variants in these genes might exacerbate the inflammatory response to various triggers, potentially contributing to the development of DSRD. This hypothesis is supported by the observation of a higher likelihood of preceding triggers and a more rapid decline to nadir of DSRD symptoms among individuals with variants.
Additional evidence comes from the neuroimaging abnormalities in DSRD, including susceptibility weighted imaging (SWI) findings of dystrophic mineralization of the basal ganglia, which resemble findings in AGS and HLH, albeit often more diffuse and extensive in the latter conditions (Fig. 2) [42, 43]. Individuals with variants were more likely to demonstrate abnormal neuroimaging findings compared to those without identified variants, although the total number compared was relatively small.
MRI images in a 4-year-old male with Aicardi-Goutieres syndrome (A–E) and an 18-year-old female with Down syndrome regression disorder (E–I). Sagittal T1-weighted (A, E), axial T2-weighted (B, F), and axial SWI (C–E, G–I) are shown. In AGS, there is diffuse cerebral and cerebellar atrophy with ex-vacuo ventriculomegaly (A, B), diffuse dystrophic calcification involving cerebellum (C), basal ganglia (D), and supratentorial cortex and white matter (E, arrowheads). In DSRD, there is mild cerebral atrophy (F, G). Dystrophic mineralization demonstrated in dentate nuclei of the cerebellum (H), basal ganglia (I, J, arrowheads)
Interestingly, harboring variants did not correlate with an increased rate of immunotherapy responsiveness in this cohort. This may be secondary to the small sample size and the already notable rate of immunotherapy responsiveness in individuals with DSRD. However, it is noteworthy that all individuals with a pathogenic or likely pathogenic variant were immunotherapy responsive.
This study is not without limitation. Firstly, our analysis was retrospective in nature and is influenced by both selection and severity bias. Individuals evaluated or seeking second opinions at the host institution would be presumed to have more severe phenotypes. Nevertheless, the inclusion of individuals with severe phenotypes offers a unique opportunity to detect rare variants with potentially significant effect sizes [44]. Individuals who had exome sequencing were more likely to have private insurance, further introducing selection bias towards individuals with higher socioeconomic status. Compounded with selection and severity bias, generalizability of these findings is limited and should be interpreted with reservation. The rare nature of DSRD and financial limitations on obtaining exome sequencing were factors in lowering the power of this study as well.
The causality of identified variants requires validation through functional studies. The interface between phenotype and genotype is complicated and influenced by a variety of environmental, social and health factors, and while these genes are of interest, they may not be sufficient to cause DSRD. Further exploration of this subset of individuals may provide crucial insights into DSRD's pathophysiology. Further large-scale studies comparing whole genome sequencing and genome-wide methylation analysis in DSRD and DS controls are warranted to better understand the genetic and epigenetic changes associated with DSRD.
Conclusion
A small subset of individuals with DSRD were observed to have pathogenic variants in genes of immune regulation. The significance of this finding is underscored by the established links between these genes and interferon-mediated disorders such as AGS and SLE. These results mark a promising area for future investigations into the intricate mechanisms underlying DSRD and may lead to new immune targeting treatments.
Data availability
Additional data regarding patients is available to qualified investigators pending IRB approval.
References
de Graaf G, Buckley F, Skotko BG (2016) Live births, natural losses, and elective terminations with Down syndrome in Massachusetts. Genet Med 18:459–466
Mircher C, Cieuta-Walti C, Marey I et al (2017) Acute regression in young people with Down Syndrome. Brain Sci 7:57
Rosso M, Fremion E, Santoro SL et al (2020) Down syndrome disintegrative disorder: a clinical regression syndrome of increasing importance. Pediatrics 145:e20192939
Worley G, Crissman BG, Cadogan E, Milleson C, Adkins DW, Kishnani PS (2015) Down syndrome disintegrative disorder: new-onset autistic regression, dementia, and insomnia in older children and adolescents with down syndrome. J Child Neurol 30:1147–1152
Santoro JD, Spinazzi NA, Filipink RA et al (2023) Immunotherapy responsiveness and risk of relapse in Down syndrome regression disorder. Transl Psychiatry 13:276
Santoro JD, Partridge R, Tanna R et al (2022) Evidence of neuroinflammation and immunotherapy responsiveness in individuals with down syndrome regression disorder. J Neurodev Disord 14:35
Santoro SL, Baumer NT, Cornacchia M et al (2022) Unexplained regression in Down syndrome: management of 51 patients in an international patient database. Am J Med Genet A 188:3049–3062
Santoro SL, Cannon S, Capone G et al (2020) Unexplained regression in Down syndrome: 35 cases from an international Down syndrome database. Genet Med 22:767–776
Cardinale KM, Bocharnikov A, Hart SJ et al (2019) Immunotherapy in selected patients with Down syndrome disintegrative disorder. Dev Med Child Neurol 61:847–851
Harisinghani A, Raffaele G, Zawatsky CB, Santoro SL (2023) Beyond chromosome analysis: additional genetic testing practice in a Down syndrome clinic. Am J Med Genet C Semin Med Genet 193:e32063
Sullivan KD, Lewis HC, Hill AA et al (2016) Trisomy 21 consistently activates the interferon response. Elife 5:e16220
Powers RK, Culp-Hill R, Ludwig MP et al (2019) Trisomy 21 activates the kynurenine pathway via increased dosage of interferon receptors. Nat Commun 10:4766
Waugh KA, Araya P, Pandey A et al (2019) Mass cytometry reveals global immune remodeling with multi-lineage hypersensitivity to type i interferon in Down syndrome. Cell Rep 29:1893-1908.e1894
Mitrovič M, Patsopoulos NA, Beecham AH et al (2018) Low-frequency and rare-coding variation contributes to multiple sclerosis risk. Cell 175:1679-1687.e1677
Santoro JD, Patel L, Kammeyer R et al (2022) Assessment and diagnosis of down syndrome regression disorder: international expert consensus. Front Neurol 13:940175
Bush G, Fink M, Petrides G, Dowling F, Francis A (1996) Catatonia. I. Rating scale and standardized examination. Acta Psychiatr Scand 93:129–136
Cummings JL, Mega M, Gray K, Rosenberg-Thompson S, Carusi DA, Gornbein J (1994) The Neuropsychiatric Inventory: comprehensive assessment of psychopathology in dementia. Neurology 44:2308–2314
Kopanos C, Tsiolkas V, Kouris A et al (2019) VarSome: the human genomic variant search engine. Bioinformatics 35:1978–1980
Rentzsch P, Witten D, Cooper GM, Shendure J, Kircher M (2019) CADD: predicting the deleteriousness of variants throughout the human genome. Nucl Acids Res 47:D886-d894
Adzhubei IA, Schmidt S, Peshkin L et al (2010) A method and server for predicting damaging missense mutations. Nat Methods 7:248–249
Ng PC, Henikoff S (2001) Predicting deleterious amino acid substitutions. Genome Res 11:863–874
Richards S, Aziz N, Bale S et al (2015) Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 17:405–424
Tavtigian SV, Harrison SM, Boucher KM, Biesecker LG (2020) Fitting a naturally scaled point system to the ACMG/AMP variant classification guidelines. Hum Mutat 41:1734–1737
da Huang W, Sherman BT, Lempicki RA (2009) Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucl Acids Res 37:1–13
da Huang W, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4:44–57
Chung H, Green PHR, Wang TC, Kong XF (2021) Interferon-driven immune dysregulation in down syndrome: a review of the evidence. J Inflamm Res 14:5187–5200
Song P, Yang W, Lou KF et al (2022) UNC13D inhibits STING signaling by attenuating its oligomerization on the endoplasmic reticulum. EMBO Rep 23:e55099
Mathapathi S, Chu CQ (2022) Contribution of impaired DNASE1L3 activity to anti-DNA autoantibody production in systemic lupus erythematosus. Rheumatol Immunol Res 3:17–22
Cristini A, Tellier M, Constantinescu F et al (2022) RNase H2, mutated in Aicardi-Goutières syndrome, resolves co-transcriptional R-loops to prevent DNA breaks and inflammation. Nat Commun 13:2961
Liu BQ, Liu RB, Li WP et al (2023) XAF1 prevents hyperproduction of type I interferon upon viral infection by targeting IRF7. EMBO Rep 24:e55387
Rohr J, Beutel K, Maul-Pavicic A et al (2010) Atypical familial hemophagocytic lymphohistiocytosis due to mutations in UNC13D and STXBP2 overlaps with primary immunodeficiency diseases. Haematologica 95:2080–2087
Meeths M, Chiang SC, Wood SM et al (2011) Familial hemophagocytic lymphohistiocytosis type 3 (FHL3) caused by deep intronic mutation and inversion in UNC13D. Blood 118:5783–5793
Aricò M, Boggio E, Cetica V et al (2013) Variations of the UNC13D gene in patients with autoimmune lymphoproliferative syndrome. PLoS ONE 8:e68045
Mackenzie KJ, Carroll P, Lettice L et al (2016) Ribonuclease H2 mutations induce a cGAS/STING-dependent innate immune response. Embo j 35:831–844
Reijns MA, Bubeck D, Gibson LC et al (2011) The structure of the human RNase H2 complex defines key interaction interfaces relevant to enzyme function and human disease. J Biol Chem 286:10530–10539
Günther C, Kind B, Reijns MA et al (2015) Defective removal of ribonucleotides from DNA promotes systemic autoimmunity. J Clin Invest 125:413–424
Chan RWY, Serpas L, Ni M et al (2020) Plasma DNA profile associated with DNASE1L3 gene mutations: clinical observations, relationships to nuclease substrate preference, and in vivo correction. Am J Hum Genet 107:882–894
Yasutomo K, Horiuchi T, Kagami S et al (2001) Mutation of DNASE1 in people with systemic lupus erythematosus. Nat Genet 28:313–314
Al-Mayouf SM, Sunker A, Abdwani R et al (2011) Loss-of-function variant in DNASE1L3 causes a familial form of systemic lupus erythematosus. Nat Genet 43:1186–1188
Mudde ACA, Booth C, Marsh RA (2021) Evolution of our understanding of XIAP deficiency. Front Pediatr 9:660520
Maakaroun NR, Moanna A, Jacob JT, Albrecht H (2010) Viral infections associated with haemophagocytic syndrome. Rev Med Virol 20:93–105
Livingston J (2014) Common pathways of intracranial calcification and the role of the pericyte: insights from neuropathology. Dev Med Child Neurol 56:924–925
Klok MD, Bakels HS, Postma NL, van Spaendonk RM, van der Knaap MS, Bugiani M (2015) Interferon-α and the calcifying microangiopathy in Aicardi-Goutières syndrome. Ann Clin Transl Neurol 2:774–779
El-Fishawy P, State MW (2010) The genetics of autism: key issues, recent findings, and clinical implications. Psychiatr Clin North Am 33:83–105
Acknowledgements
The authors would like to thank Dr. Matthew Deardorff for his invaluable consultation and advice.
Funding
Open access funding provided by SCELC, Statewide California Electronic Library Consortium. Dr. Santoro receives funding through the National Institutes of Health (NHLBI/NICHD) (Grant no. R611HD109748). He has previously received funding through the National MS Society and Race to Erase MS Foundation. Dr. Santoro receives consulting fees from UCB for unrelated work on myelin oligodendrocyte glycoprotein related disease. He also receives consulting fees from Cycle Pharmaceuticals on unrelated work. No funding was received for conducting this study or preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare no conflicts of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jafarpour, S., Banerjee, A.K., Khoshnood, M.M. et al. De novo variants in immune regulatory genes in Down syndrome regression disorder. J Neurol (2024). https://doi.org/10.1007/s00415-024-12521-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00415-024-12521-y