Abstract
Background
Robust predictive models of clinical impairment and worsening in multiple sclerosis (MS) are needed to identify patients at risk and optimize treatment strategies.
Objective
To evaluate whether machine learning (ML) methods can classify clinical impairment and predict worsening in people with MS (pwMS) and, if so, which combination of clinical and magnetic resonance imaging (MRI) features and ML algorithm is optimal.
Methods
We used baseline clinical and structural MRI data from two MS cohorts (Berlin: n = 125, Amsterdam: n = 330) to evaluate the capability of five ML models in classifying clinical impairment at baseline and predicting future clinical worsening over a follow-up of 2 and 5 years. Clinical worsening was defined by increases in the Expanded Disability Status Scale (EDSS), Timed 25-Foot Walk Test (T25FW), 9-Hole Peg Test (9HPT), or Symbol Digit Modalities Test (SDMT). Different combinations of clinical and volumetric MRI measures were systematically assessed in predicting clinical outcomes. ML models were evaluated using Monte Carlo cross-validation, area under the curve (AUC), and permutation testing to assess significance.
Results
The ML models significantly determined clinical impairment at baseline for the Amsterdam cohort, but did not reach significance for predicting clinical worsening over a follow-up of 2 and 5 years. High disability (EDSS ≥ 4) was best determined by a support vector machine (SVM) classifier using clinical and global MRI volumes (AUC = 0.83 ± 0.07, p = 0.015). Impaired cognition (SDMT Z-score ≤ −1.5) was best determined by a SVM using regional MRI volumes (thalamus, ventricles, lesions, and hippocampus), reaching an AUC of 0.73 ± 0.04 (p = 0.008).
Conclusion
ML models could aid in classifying pwMS with clinical impairment and identify relevant biomarkers, but prediction of clinical worsening is an unmet need.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Multiple sclerosis (MS) is a chronic inflammatory, demyelinating, and neurodegenerative disease with a heterogeneous and unpredictable disease course [1]. Prognostic biomarkers are urgently needed for monitoring disease progression and optimizing therapeutic strategies [2]. The clinical relevance of magnetic resonance imaging (MRI) for diagnosing and monitoring MS by using inflammatory markers (e.g., white matter (WM) lesion counts) is well established [3, 4]. However, these inflammatory markers have limited explanatory value for determining the severity of symptoms and predicting clinical progression [5, 6]. MRI markers of neurodegeneration, instead, are more closely related to clinical outcomes and thought to be the main driver of irreversible disability [7, 8]. Regional volumetric MRI measures such as deep gray matter (DGM) and cortical gray matter (CGM) volumes have shown the closest associations to motor dysfunctions and cognitive decline [9, 10]. However, the use of regional MRI volumes as predictors of disease progression remains largely unexplored.
Machine learning (ML) strategies have been increasingly applied for prediction in medicine and identifying patients at risk [11]. While traditional statistical techniques can typically handle only a few input variables and are often based on strict assumptions, ML is able to derive complex hidden patterns in high-dimensional data [12]. In MS, various ML approaches have been applied, but predicting disability progression with high accuracy remains challenging [13,14,15]. These studies used progression on the Expanded Disability Status Scale (EDSS) as a prediction target, since it is the predominant outcome measure for defining disability accumulation and progression in MS [16, 17]. However, the reliability of the EDSS is compromised by a significant measurement error and interrater variability [18]. In addition, it is heavily influenced by ambulatory functioning, while upper-extremity dysfunction and cognitive dysfunction are not adequately measured resulting in a low sensitivity in identifying crucial factors for disease progression [19, 20]. Recent evidence suggests that other outcome measures, such as the Timed 25-Foot Walk Test (T25FW) and the 9-Hole Peg Test (9HPT), as well as composite scores of EDSS, T25FW, and 9HPT (EDSS +), may be more sensitive in capturing disease progression [21, 22]. For cognitive functioning, the Symbol Digit Modalities Test (SDMT) is widely employed in clinical trials, because it is highly sensitive for measuring information processing speed (IPS), the most affected cognitive function in MS [23, 24].
In this study, we aimed to systematically compare the performance of ML approaches for classifying clinical impairment and predicting disease progression in people with MS (pwMS), based on a range of (composite) clinical outcomes (EDSS, T25FW, 9HPT, EDSS + , and SDMT). Our secondary aim was to identify which clinical and MRI markers were most important in determining clinical impairment and predicting worsening defined for each outcome. ML approaches included logistic regression, support vectors machine, gradient boosting, and random forest classifiers. Models were trained on two clinical data sets, one early MS cohort from Berlin with a follow-up after 2 years and a long-standing MS cohort from Amsterdam with a follow-up after 5 years.
Methods
Study population
Data were retrospectively collected from the early MS cohort of Berlin, Germany (32 people diagnosed with clinically isolated syndrome (CIS) and 93 pwMS) [25, 26] and people with clinically definite MS from the Amsterdam MS cohort, the Netherlands (330 pwMS) [27, 28]. All included subjects were over the age of 18 and had a clinical assessment and a structural MRI examination available at baseline. Clinical measurements included EDSS, 9HPT, T25FW, and SDMT score. The early MS cohort of Berlin had a 2-year clinical follow-up available for all 125 included subjects. For Amsterdam, a 5-year clinical follow-up was available for 225/330 included subjects. The institutional ethics review boards of both institutions (Amsterdam UMC, Amsterdam and Charité, Berlin) approved the study protocol and subjects gave written informed consent prior to participation.
MRI acquisition
All subjects underwent a 3T MRI examination including the following pulse sequences: 3D T1-weighted (3D-T1) and 3D fluid-attenuated inversion recovery (3D-FLAIR). The scanning protocol in Berlin included a 3D-T1 magnetization prepared rapid acquisition gradient echo sequence (1.0 × 1.0 × 1.0 mm resolution, repetition time (TR) = 1900 ms, echo time (TE) = 3.03 ms, inversion time (TI) = 900 ms, flip angle = 9°) and a 3D-FLAIR sequence (1.0 × 1.0 × 1.0 mm resolution, TR = 6000 ms, TE = 388 ms, TI = 2100 ms), using a Tim Trio scanner (Siemens Medical Systems, Erlangen, Germany). The scanning protocol in Amsterdam included a 3D-T1 fast-spoiled gradient-echo sequence (1 × 0.9 × 0.9 mm resolution, TR = 7.8 ms, TE = 3 ms, TI = 450 ms, flip angle = 12°) and a 3D-FLAIR sequence (1.2 × 1.0 × 1.0 mm resolution, TR = 8000 ms, TE = 125 ms, TI = 2350 ms), using a GE Signa HDxt scanner (Milwaukee, WI).
MRI processing
T2-lesion volumes (T2LV) were determined on 3D-FLAIR. In Berlin, lesions were manually segmented using ITK-SNAP (www.itksnap.org) by two expert MRI technicians [26]. In Amsterdam, lesions were automatically segmented using a k-nearest neighbor algorithm and visually checked [28,29,30]. To reduce lesion-associated brain tissue segmentation bias, lesions were filled with values approximating normal WM on 3D-T1 [31]. Whole-brain, CGM, and DGM segmentations were derived for both centers using the FreeSurfer 7.1.1 (http://surfer.nmr.mgh.harvard.edu/) recon-all pipeline on lesion filled 3D-T1. Subsequently, the cortical surface of each subject was parcellated into 210 regions using the Brainnetome Atlas (BNA) [32]. The volumes of the left and right regions were averaged to decrease the number of input features without losing too much anatomical information, resulting in 5 global volumes (whole-brain volume (WBV), CGM volume, DGM volume, lateral ventricular volume (LVV), cortical cerebellum volume), 105 CGM regional volumes, and seven DGM regional volumes (thalamus, accumbens, putamen, caudate, pallidum, amygdala, hippocampus). All volumes, except for T2LV, were corrected for head size by dividing the volume by the estimated total intracranial volume (eTIV).
Overview of machine learning approach
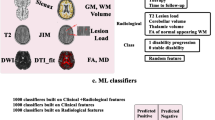
Based on different sets of clinical and MRI features, we trained different ML algorithms for classifying clinical impairment and predicting clinical worsening. The input features included demographic (age, sex) and clinical information (symptom duration, MS subtype, use of disease-modifying therapy (DMT)) as well as structural MRI volumes. The outcome measures for classifying clinical impairment included EDSS (cutoff ≥ 4) to define high disability or SDMT (Z-score ≤ −1.5) to define cognitive impairment. Clinical worsening was evaluated over a follow-up of 2 years (Berlin) and 5 years (Amsterdam), based on EDSS, 9HPT, T25FW, or SDMT scores. The significance of classification and prediction models was assessed with permutation testing and the most important clinical and MRI features were determined using Shapley additive explanations (SHAP). See Fig. 1 for an overview of the machine learning approach.
Overview of the machine learning (ML) approach. Input features for ML included different sets of clinical variables and MRI volumes. Different ML approaches were evaluated to classify clinical impairment and predict clinical worsening. Shapley additive explanations were applied to identify the most important clinical and MRI features for classification and prediction. DGM deep gray matter; BNA Brainnetome Atlas; LR logistic regression; SVM support vector machine; RF random forest; XGBoost eXtreme Gradient Boosting; EDSS Expanded Disability Status Scale; SDMT Symbol Digit Modalities Test; 9HPT 9-Hole Peg Test; T25FW Timed 25-Foot Walk Test
Clinical impairment
Clinical impairment at baseline was defined using the EDSS or SDMT. PwMS were classified as having a low or high disability based on an EDSS ≥ 4 cutoff. For classifying preserved or impaired cognition, SDMT standardized Z-scores were calculated based on German normative healthy control data for the Berlin cohort [33] and norm scores of matched healthy controls for the Amsterdam cohort [28]. PwMS reporting an SDMT Z-score below −1.5 were considered as cognitively impaired.
Clinical worsening
Clinical worsening was assessed using the EDSS, 9HPT, T25FW, a combination of EDSS, 9HPT, and T25FW (EDSS +), or SDMT. EDSS-based worsening was defined as an increase in EDSS of ≥ 1.5 points for a baseline score of 0, an increase of ≥ 1.0 for a baseline score between 1.0 and 5.5, or an increase of ≥ 0.5 for a baseline EDSS score of ≥ 6.0 [34]. For 9HPT and T25FW, clinically meaningful worsening was defined as a 20% increase in the time required to finish the test compared to the baseline measurement [35, 36]. For 9HPT, worsening in the non-dominant hand and dominant hand was assessed separately. Worsening on EDSS + was defined as worsening on ≥ 1 of the three components (EDSS, T25FW, or 9HPT (dominant or non-dominant hand)) [21]. Lastly, SDMT worsening was defined as an increase of ≥ 4 points compared to the baseline measure [37].
Input features
To compare the performance of clinical and MRI-derived features for classifying clinical impairment and predicting clinical worsening, and assess which combination would result in the highest performing models, five different feature sets were defined: (1) clinical data, (2) global MRI volumes, (3) clinical data + global MRI volumes, (4) regional MRI volumes, (5) clinical data + regional MRI volumes. See Table 1 for an overview of included features in each feature set. Clinical data included age, sex, symptom duration, MS subtype (CIS, RRMS, or progressive MS), and use of DMT (yes or no).
Machine learning model training
Five ML algorithms were compared for classification of clinical impairment and prediction of worsening for the different clinical outcomes. A comparison of multiple ML models was conducted because these models have varying abilities to capture linear and non-linear relationships between input features [38]. As linear classifiers, logistic regression (LR) and support vector machine with a linear kernel (SVM-lin) [39] were selected due to their robust performance demonstrated in prior structural neuroimaging studies [40, 41]. Their performance was compared to three non-linear models that have been successfully applied in other studies [42]: SVM with a radial basis function kernel (SVM-RBF) [43], eXtreme Gradient Boosting Classifier (XGBoost) [44], and random forest (RF) [45]. The ML pipelines were implemented in Python 3.9.15 using the scikit-learn [46] and xgboost [47] software packages. Due to variations in demographics, follow-up times, and the use of different MRI scanners and protocols between the cohorts, it was unfavorable to employ one cohort as a training set and the other as a validation set. Consequently, machine learning models were independently trained on the data from each center. Preprocessing of all input features included standardization by removing the mean and scaling to unit variance [48]. In addition, random oversampling of the minority class was used in the preprocessing pipeline to account for class imbalance in model training. Due to the relatively low sample size of our data, stratified Monte–Carlo cross-validation with ten repetitions (i.e., 10 randomly selected test sets) was performed using an 80%/20% train/test split ratio to avoid evaluation bias resulting from sampling effects [49, 50]. A stratified 5-fold cross-validation was applied within the training set of each repetition for hyperparameter optimization [51]. Since not all clinical outcome variables were available for each participant, train and test sets were different for each clinical outcome due to random selection. Model performance was assessed using several metrics: area under the curve (AUC) of the receiver operating characteristics curve, balanced accuracy (BA), precision, and recall (or sensitivity). AUC assesses the model's ability to distinguish between classes by plotting the true positive rate against the false positive rate across all possible classification thresholds. Balanced accuracy represents the average of recall and specificity, providing a balanced view of the model's performance across both classes. Precision quantifies the proportion of true positives among all positive predictions, highlighting the model's accuracy in predicting positive instances. Recall, also known as sensitivity, measures the proportion of actual positives correctly identified by the model, indicating its effectiveness in detecting positive instances. The final performance of the models was ranked based on the average and standard deviation of the AUC across the 10 repetitions. Statistical significance of the best performing ML model for each outcome measure was determined using permutation testing [52]. To reduce computational time, the p value was derived based on 100 permutations.
Model explanations
To understand which combination of clinical and MRI features were most relevant for clinical impairment and worsening predictions in pwMS, SHAP values were calculated in a post hoc analysis. SHAP is a local model explanation method aimed to explain the model prediction for each subject by computing the relative importance of every input feature for the final prediction [53]. SHAP values explain the difference between the individual prediction and the average prediction. For an individual, the sum of all SHAP values equals the difference between their prediction and the average probability of clinical impairment or worsening. Global ranking of feature importance was defined as the mean absolute SHAP value of each feature across all subjects and all test sets [54]. SHAP values were calculated with the KernelSHAP method implemented in Python.
Results
Demographics
A total of 125 participants from Berlin and 330 participants from Amsterdam were included. Demographic, clinical, and global MRI variables of the cohorts are shown in Table 2. The participants from Amsterdam were older compared to those from Berlin (47.7 ± 10.9 vs. 33.2 ± 7.2, p < 0.001), had a longer disease duration (14.8 ± 8.5 vs. 0.6 ± 0.7, p < 0.001), and a higher EDSS at baseline (3.0 [2.0 − 4.0] vs. 1.5 [1.0 − 2.0], p < 0.001). For the Berlin cohort only, 2% (n = 3) of pwMS had a high disability (EDSS ≥ 4) at baseline and 2% (n = 2) had cognitive impairment (SDMT Z ≤ − 1.5). For the Amsterdam cohort, 39% (n = 129) of pwMS had a high disability at baseline and 36% (n = 119) had cognitive impairment. See Fig. 2 for total sample sizes for each longitudinal clinical outcome measure and percentages of patients showing worsening during the follow-up period. For the Berlin cohort, a 2-year clinical follow-up was available for all 125 patients, of whom 18% (n = 22) showed worsening on EDSS and 21% (n = 20) on SDMT. For the Amsterdam cohort, a 5-year follow-up was available for 225 pwMS, of whom 35% (n = 78) showed worsening on EDSS and 29% (n = 66) on SDMT. The lowest progression rates were seen for the 9HPT in both cohorts, with 2% (dominant hand, n = 2) and 4% (non-dominant hand, n = 4) in the Berlin cohort, and 16% (dominant hand, n = 34) and 15% (non-dominant hand, n = 32) in the Amsterdam cohort.
Number of subjects with clinical worsening based on multiple clinical end points for the A Berlin cohort and B Amsterdam cohort. The percentage of pwMS showing worsening (orange) or being stable (green) on a clinical outcome measure over the follow-up period varies for each outcome measure. The total sample size for each outcome measure (n) is displayed above the graphs. EDSS Expanded Disability Status Scale; 9HPT 9-Hole Peg Test (D dominant hand; ND non-dominant hand); SDMT Symbol Digit Modalities Test; T25FW Timed 25-Foot Walk Test
Classification of clinical impairment
Due to the low proportion of pwMS having clinical impairment in the Berlin cohort (2% of pwMS had a high disability and 2% had cognitive impairment), cross-sectional clinical impairment classification was only performed for the Amsterdam cohort. The best performance for determining high disability at baseline was achieved by the clinical + global MRI feature set (AUC = 0.83 ± 0.07, BA = 0.76 ± 0.09, precision = 0.68 ± 0.11, recall = 0.74 ± 0.11, p = 0.015) using SVM-RBF. As shown in Fig. 3, all models showed good performance for determining high disability at baseline (mean AUC 0.75–0.78 across all feature sets), with the lowest performance for LR (mean AUC = 0.68). The model performance for determining impaired cognition was the highest for SVM-lin with regional MRI volumes as input feature set (AUC = 0.73 ± 0.04, BA = 0.67 ± 0.03, precision = 0.60 ± 0.11, recall = 0.55 ± 0.09, p = 0.008). Clinical features only could not predict cognitive impairment (SVM-RBF: AUC = 0.55 ± 0.05, p = 0.27).
Performance of ML models in classifying clinical impairment at baseline across ten partitions (Amsterdam cohort, n = 330). For determining EDSS ≥ 4, the highest mean AUC was achieved by SVM-RBF with feature set Clinical + global MRI (AUC = 0.83 ± 0.07, p = 0.016), for determining SDMT Z ≤ 1.5 the highest mean AUC was found for SVM-lin with regional MRI as feature set (AUC = 0.73 ± 0.04, p = 0.008). AUC area under the curve; EDSS Expanded Disability Status Scale; SDMT Symbol Digit Modalities Test; LR logistic regression; SVM-lin support vector machine with a linear kernel; SVM-RBF SVM with a radial basis function kernel; XGBoost eXtreme Gradient Boosting Classifier; RF random forest
Important brain regions for determining clinical impairment
We used SHAP values to assess which clinical features and brain regions were most important for classifying high disability and impaired cognition in long-standing MS (Amsterdam cohort). The most informative features were determined for the SVM-RBF model since it had the highest average performance across all feature sets in determining high disability (AUC = 0.78 ± 0.07) and impaired cognition (AUC = 0.67 ± 0.08). Figure 4 presents the 20 most important features from the feature set including all clinical and MRI features (clinical data + regional MRI volumes). Based on all input features, the SVM-RBF model achieved an AUC of 0.77 ± 0.05 (p = 0.035) in determining high disability and an AUC of 0.72 ± 0.04 (p = 0.008) in determining cognitive impairment. The feature importance for the other feature sets are shown in Supplementary Figs. 1 and 2. Most relevant features for determining high disability were age, disease duration, LVV, DGM volumes (thalamus, hippocampus, caudate, and accumbens) and CGM regions in the middle frontal gyrus (BNA: A46, A46v, A6vl, A10l) and inferior temporal gyrus (BNA: A20cv, A20il), see Fig. 5 and Table 3. For determining cognitive impairment, the most relevant features were regional MRI volumes, including DGM volumes (thalamus, hippocampus, putamen, accumbens), LVV, T2LV, and volumes of the superior frontal gyrus (BNA: A8dl, A8m), orbital gyrus (BNA: A11m), and parahippocampal gyrus (BNA: TI).
Distributions of SHAP feature importance for the 20 most important features using SVM-RBF in determining high disability (left, AUC = 0.77 ± 0.05, p = 0.035) and cognitive impairment (right, AUC = 0.72 ± 0.04, p = 0.008) at baseline (Amsterdam cohort, n = 330). The mean of the distributions is shown in red. The Brainnetome Atlas (BNA) cortical regions are plotted in Fig. 5 and described in Table 3. EDSS Expanded Disability Status Scale; LVV lateral ventricular volume; SDMT Symbol Digit Modalities Test; T2LV T2-lesion volume; WBV whole-brain volume
Most important brain regions for classifying clinical impairment using SVM-RBF for determining high disability (left) and cognitive impairment (right) at baseline (Amsterdam cohort, n = 330). Intensity represents the median SHAP value across all ten partitions. EDSS Expanded Disability Status Scale; SDMT Symbol Digit Modalities Test
Prediction of clinical worsening
The performance of the ML models in predicting clinical worsening was evaluated based on the mean AUC, BA, precision, and recall for each longitudinal outcome measure across all test sets. The combination of the best ML models and feature sets for each prediction task are presented in Table 4. For the Berlin cohort, prediction of 9HPT 20% worsening could not be assessed due to a large class imbalance (2% worsening on dominant hand and 4% worsening on non-dominant hand). Although the prediction of EDSS worsening over a 2-year follow-up showed the highest AUC using SVM-RBF and global MRI volumes as input features (AUC = 0.73 ± 0.13), precision was low (0.24 ± 0.09) and the prediction was not significant after permutation testing (p = 0.163). Other measures of clinical worsening also did not reach significance after permutation testing, with mean AUC varying from 0.53 to 0.73 and mean BA between 0.51 and 0.63. For the Amsterdam cohort, clinical worsening could not be predicted over the 5-year follow-up, as none of the ML models reached significance after permutation testing. The highest prediction performance was achieved for 9HPT 20% worsening in the dominant hand (AUC = 0.63 ± 0.11, BA = 0.53 ± 0.08), with low precision (0.21 ± 0.12) and recall (0.23 ± 0.17) values. The ML models for predicting other clinical worsening outcomes reached a mean AUC between 0.54 and 0.63, with a mean BA between 0.50 and 0.59.
Discussion
In this study, five different ML models were applied to determine the clinical impairment and predict clinical worsening in pwMS based on clinical and structural MRI input features. For baseline models, the classification of high disability and cognitive impairment exhibited commendable accuracy when relying on a combination of clinical and regional MRI input features. Nevertheless, forecasting disease worsening in pwMS over a longitudinal period of 2–5 years, utilizing baseline clinical and structural MRI features, did not yield significant prediction accuracies.
In the classification of high versus low disability at baseline, both clinical characteristics and MRI volumes played a crucial role in the prediction, achieving an AUC of 0.83. However, for cognitive impairment, regional MRI volumes were most important, as clinical features alone offered little explanatory value. The performance of the cognitive classification task (AUC = 0.73) was highly comparable to another study predicting cognitive impairment in a large cohort (n = 540), achieving an AUC of 0.74 using MRI features as input only [55]. From all MRI features, the thalamus volume was the most important regional MRI feature associated with both disability and cognitive impairment, which is in line with previous findings [9, 56, 57]. Regarding other DGM volumes, atrophy of the hippocampus and accumbens was important for determining both high disability and cognitive impairment, while caudate volume seemed more predictive for disability, and the volume of the amygdala and putamen for cognitive impairment. The classification tasks demonstrated comparable performance whether utilizing global MRI volumes or intricate parcellations of the CGM with the BNA. The advantage of the latter approach is that distinct CGM regions could be identified that were important for high disability and cognitive impairment. Consistent with previous work using other statistical methods, mostly regions in the temporal and frontal gyrus were important for determining high disability and cognitive impairment [58, 59]. From the temporal lobe, specific areas in the inferior and middle temporal gyrus were most important for EDSS, while the parahippocampal gyrus was more important for determining cognitive impairment. When comparing the stability of feature importance across tasks, it is important to highlight that the feature importance for predicting cognitive impairment were less consistent across various test splits. This reduced stability might be attributed to the lower task performance (0.73 compared to 0.83). However, it could also suggest that the model captures heterogeneous cognitive profiles in pwMS, since the SDMT assesses various cognitive processes [60].
Despite achieving good performance in cross-sectional classification tasks, predicting future disease worsening over 2 and 5 years proved to be unattainable using the same ML models and input variables. While two earlier ML studies reported more promising results in predicting MS disease worsening, these results were limited by relying on relatively small sample sizes and model evaluation on only one small test set, possibly reporting overly optimistic results [15, 61]. In larger cohort studies using similar ML approaches, baseline data had limited predictive value for disease worsening over 5 years, while incorporating longitudinal observations of clinical and brain MRI changes in the first two years improved prediction performance (AUC = 0.75–0.83) [13, 62]. Furthermore, incorporating standardized disease history from electronic health records in combination with MRI might be a promising direction to increase data availability and information content with higher frequency for improving ML prediction models [63]. In addition, all previous ML studies investigating disease worsening in MS used EDSS as the clinical end point. While EDSS-based definitions are still considered the gold standard in clinical research, they are often criticized for low sensitivity and reliability [64]. To address the challenge of heterogeneous subjective scoring in clinical assessments, one promising approach involves leveraging ML models capable of accommodating label uncertainty [65]. In our study, we took a different approach by exploring a range of other widely used clinical end points, including assessments of hand dysfunction (9HPT), walking dysfunction (T25FW), and a combination of EDSS, 9HPT, and T25FW (EDSS +). However, we did not observe an improvement in the prediction performance for all these outcomes. While 9HPT and T25FW have a lower measurement error compared to the EDSS, worsening could still not be predicted by baseline MRI measures [22]. These findings highlight the need for defining accurate clinical end points in addition to the exploration of better predictors [66]. In addition to predicting disability worsening, SDMT worsening of at least 4 points over 5 years was defined as a cutoff for cognitive worsening, but also did not achieve significant prediction performance. This is in contrast to an earlier investigation of the Amsterdam MS cohort, which showed that cortical atrophy was predictive of cognitive decline, using traditional logistic regression and an extensive cognitive test battery to define cognitive worsening [30]. The difference in predictive performance may be explained by the lower sensitivity of the SDMT as a standalone measure to capture cognitive decline [67, 68].
The main limitation of our study is the relatively small sample size of our data sets for an ML-based analysis, along with the lack of evaluation on an additional external test set. Despite the application of our approach across two diverse cohorts that span early to late stages of MS, the divergent data collection and follow-up times (2 vs 5 years) prevented the merging of cohorts for analysis and the utilization of one cohort as an independent test set. Future ML studies with larger datasets are required to explore individualized prediction of disease course both in early and later MS phases, facilitating early detection of high-risk patients and enabling personalized treatment strategies. Predictive variables for disease progression may differ between MS disease stages, as there are some indications that some predictors such as white matter damage are predictive for worsening in early stages and other measures such as cortical atrophy are specific for later, more progressive, stages of MS [30]. Furthermore, considering the predictive value of spinal cord lesions and atrophy is crucial, as these factors significantly contribute to walking dysfunction in long-standing disease [69]. In addition, the limited sample size resulted in a low stability of feature importance across the evaluated test sets, especially for classifying cognitive impairment. To establish a more robust understanding of the relationship between clinical and structural brain information and clinical end points related to disease severity and progression, further investigations with larger sample sizes and extended follow-up durations are essential. Lastly, the currently used clinical end points may not be specific enough to capture disease progression in MS. Development of new outcome measures, such as frequent smartphone-based assessments, may be more sensitive in capturing worsening in various symptoms of MS [70].
Conclusion
While ML models could accurately identify patients with clinical impairment based on clinical data and regional MRI volumes, accurate prediction of MS disease worsening remains an unmet need. Improving clinical end point metrics and using larger standardized datasets representative of the heterogeneity in MS can be a promising direction for future ML research.
Data availability
Anonymized data can be shared upon reasonable request from a qualified investigator.
References
Confavreux C, Vukusic S (2014) The clinical course of multiple sclerosis. Handb Clin Neurol 122:343–369
Tavazzi E et al (2020) MRI biomarkers of disease progression and conversion to secondary-progressive multiple sclerosis. Expert Rev Neurother 20(8):821–834
Filippi M et al (2019) Assessment of lesions on magnetic resonance imaging in multiple sclerosis: practical guidelines. Brain 142(7):1858–1875
Tomassini V et al (2020) Diagnosis and management of multiple sclerosis: MRI in clinical practice. J Neurol 267:2917–2925
Barkhof F (2002) The clinico-radiological paradox in multiple sclerosis revisited. Curr Opin Neurol 15(3):239–245
Brex PA et al (2002) A longitudinal study of abnormalities on MRI and disability from multiple sclerosis. N Engl J Med 346(3):158–164
Horakova D et al (2009) Gray matter atrophy and disability progression in patients with early relapsing–remitting multiple sclerosis: a 5-year longitudinal study. J Neurol Sci 282(1–2):112–119
Dekker I et al (2019) Predicting clinical progression in multiple sclerosis after 6 and 12 years. Eur J Neurol 26(6):893–902
Eshaghi A et al (2018) Deep gray matter volume loss drives disability worsening in multiple sclerosis. Ann Neurol 83(2):210–222
Eijlers AJC et al (2019) Cortical atrophy accelerates as cognitive decline worsens in multiple sclerosis. Neurology 93(14):e1348–e1359
Chen JH, Asch SM (2017) Machine learning and prediction in medicine—beyond the peak of inflated expectations. N Engl J Med 376(26):2507
Bzdok D, Altman N, Krzywinski M (2018) Statistics versus machine learning. Nat Methods 15(4):233–234
Zhao Y et al (2017) Exploration of machine learning techniques in predicting multiple sclerosis disease course. PLoS ONE 12(4):e0174866
Law MTK et al (2019) Machine learning in secondary progressive multiple sclerosis: an improved predictive model for short-term disability progression. Mult Scler J Exp Transl Clin 5(4):2055217319885983
Pinto MF et al (2020) Prediction of disease progression and outcomes in multiple sclerosis with machine learning. Sci Rep 10(1):1–13
Kurtzke JF (1983) Rating neurologic impairment in multiple sclerosis: an expanded disability status scale (EDSS). Neurology 33(11):1444–1444
Meyer-Moock S et al (2014) Systematic literature review and validity evaluation of the Expanded Disability Status Scale (EDSS) and the Multiple Sclerosis Functional Composite (MSFC) in patients with multiple sclerosis. BMC Neurol 14(1):1–10
Koch MW et al (2021) Reliability of outcome measures in clinical trials in secondary progressive multiple sclerosis. Neurology 96(1):e111–e120
Rudick RA et al (2009) Assessing disability progression with the multiple sclerosis functional composite. Mult Scler J 15(8):984–997
Balabanov P et al (2014) Addressing the regulatory and scientific challenges in multiple sclerosis—a statement from the EU regulators. Mult Scler J 20(10):1282–1287
Cadavid D et al (2017) The EDSS-Plus, an improved endpoint for disability progression in secondary progressive multiple sclerosis. Mult Scler J 23(1):94–105
Koch MW et al (2022) The timed 25-foot walk is a more sensitive outcome measure than the EDSS for PPMS trials: an analysis of the PROMISE clinical trial dataset. J Neurol 269(10):5319–5327
Smith A (1973) Symbol digit modalities test. Western Psychological Services, USA
Benedict RHB, Amato MP, DeLuca J, Geurts JJG (2020) Cognitive impairment in multiple sclerosis: clinical management, MRI, and therapeutic avenues. Lancet Neurol 19(10):860–871
Thompson AJ et al (2018) Diagnosis of multiple sclerosis: 2017 revisions of the McDonald criteria. Lancet Neurol 17(2):162–173
Chien C et al (2022) Prediction of high and low disease activity in early MS patients using multiple kernel learning identifies importance of lateral ventricle intensity. Mult Scler J Exp Transl Clin 8(3):20552173221109770
Polman CH et al (2011) Diagnostic criteria for multiple sclerosis: 2010 revisions to the McDonald criteria. Ann Neurol 69(2):292–302
Eijlers AJC et al (2017) Increased default-mode network centrality in cognitively impaired multiple sclerosis patients. Neurology 88(10):952–960
Steenwijk MD et al (2013) Accurate white matter lesion segmentation by k nearest neighbor classification with tissue type priors (kNN-TTPs). NeuroImage Clin 3:462–469
Eijlers AJC et al (2018) Predicting cognitive decline in multiple sclerosis: a 5-year follow-up study. Brain 141(9):2605–2618
Valverde S, Oliver A (2014) Llad X, A white matter lesion-filling approach to improve brain tissue volume measurements. NeuroImage Clin 6:86–92
Fan L et al (2016) The human Brainnetome Atlas: a new brain atlas based on connectional architecture. Cereb Cortex 26(8):3508–3526
Scherer P et al (2004) Normierung der brief repeatable battery of neuropsychological tests (BRB-N) fur den deutschsprachigen Raum. Nervenarzt 10(75):984–990
Kalincik T et al (2015) Defining reliable disability outcomes in multiple sclerosis. Brain 138(11):3287–3298
Kaufman M, Moyer D, Norton J (2000) The significant change for the Timed 25-foot Walk in the multiple sclerosis functional composite. Mult Scler J 6(4):286–290
Feys P et al (2017) The Nine-Hole Peg test as a manual dexterity performance measure for multiple sclerosis. Mult Scler J 23(5):711–720
Strober L et al (2019) Symbol digit modalities test: a valid clinical trial endpoint for measuring cognition in multiple sclerosis. Mult Scler J 25(13):1781–1790
Lemm S, Blankertz B, Dickhaus T, Mller K-R (2011) Introduction to machine learning for brain imaging. Neuroimage 56(2):387–399
Boser BE, Guyon IM, Vapnik VN (1992) A training algorithm for optimal margin classifiers. Proc fifth annual workshop Comput Learn theory:144–152
Desikan RS et al (2009) Automated MRI measures identify individuals with mild cognitive impairment and Alzheimer’s disease. Brain 132(8):2048–2057
Zurita M, Montalba C, Labb T (2018) Characterization of relapsing-remitting multiple sclerosis patients using support vector machine classifications of functional and diffusion MRI data. NeuroImage Clin 20:724–730
Mateos-Prez JM et al (2018) Structural neuroimaging as clinical predictor: a review of machine learning applications. NeuroImage Clin 20:506–522
Chapelle O, Vapnik V, Bousquet O, Mukherjee S (2002) Choosing multiple parameters for support vector machines. Mach Learn 46:131–159
Friedman JH (2001) Greedy function approximation: a gradient boosting machine. Ann Stat 29:1189–1232
Breiman L (1996) Bagging predictors. Mach Learn 24:123–140
Pedregosa F et al (2011) Scikit-learn: machine learning in Python. J Mach Learn Res 12:2825–2830
Chen T, Guestrin C (2016) Xgboost: a scalable tree boosting system. Proc 22nd acm sigkdd Int Conf Knowl Disc data Min:785–794
Luor D-C (2015) A comparative assessment of data standardization on support vector machine for classification problems. Intell Data Anal 19(3):529–546
Xu QS, Liang YZ, Du YP (2004) Monte Carlo cross-validation for selecting a model and estimating the prediction error in multivariate calibration. J Chemometr Soc 18(2):112–120
Shan G (2022) Monte Carlo cross-validation for a study with binary outcome and limited sample size. BMC Med Inform Decis Mak 22(1):270
Varoquaux G et al (2017) Assessing and tuning brain decoders: cross-validation, caveats, and guidelines. Neuroimage 145:166–179
Ojala M, Garriga GC (2010) Permutation tests for studying classifier performance. J Mach Learn Res 11(6):1833–1863
Lundberg SM, Lee S-I (2017) A unified approach to interpreting model predictions. Adv Neural Inf Process Syst 30:22397
Lundberg SM et al (2020) From local explanations to global understanding with explainable AI for trees. Nat Mach Intell 2(1):56–67
Marzi C et al (2023) Prediction of the information processing speed performance in multiple sclerosis using a machine learning approach in a large multicenter magnetic resonance imaging data set. Hum Brain Mapp 44(1):186–202
Rocca MA et al (2010) Thalamic damage and long-term progression of disability in multiple sclerosis. Radiology 257(2):463–469
Bisecco A et al (2018) Attention and processing speed performance in multiple sclerosis is mostly related to thalamic volume. Brain Imaging Behav 12:20–28
Tsagkas C et al (2020) Longitudinal patterns of cortical thinning in multiple sclerosis. Hum Brain Mapp 41(8):2198–2215
Colato E et al (2021) Predicting disability progression and cognitive worsening in multiple sclerosis using patterns of grey matter volumes. J Neurol Neurosurg Psychiatry 92(9):995–1006
Sandry J et al (2021) The symbol digit modalities test (SDMT) is sensitive but non-specific in MS: lexical access speed, memory, and information processing speed independently contribute to SDMT performance. Mult Scler Relat Disord 51:102950
Tommasin S, Cocozza S, Taloni A, Giann C (2021) Machine learning classifier to identify clinical and radiological features relevant to disability progression in multiple sclerosis. J Neurol 268(12):4834–4845
Zhao Y et al (2020) Ensemble learning predicts multiple sclerosis disease course in the SUMMIT study. NPJ Digit Med 3(1):135
Zhang K et al (2023) Predicting multiple sclerosis disease severity with multimodal deep neural networks. arXiv preprint arXiv:2304.04062
Cohen M et al (2021) Should we still only rely on EDSS to evaluate disability in multiple sclerosis patients? A study of inter and intra rater reliability. Mult Scler Relat Disord 54:103144
Tayyab M et al (2023) Accounting for uncertainty in training data to improve machine learning performance in predicting new disease activity in early multiple sclerosis. Front Neurol 14:1165267
Pellegrini F et al (2020) Predicting disability progression in multiple sclerosis: Insights from advanced statistical modeling. Mult Scler J 26(14):1828–1836
Sonder JM et al (2014) Comparing long-term results of PASAT and SDMT scores in relation to neuropsychological testing in multiple sclerosis. Mult Scler 20(4):481–488
Castrogiovanni N et al (2023) Longitudinal changes in cognitive test scores in patients with relapsing-remitting multiple sclerosis: an analysis of the DECIDE dataset. Neurology 101(1):e1–e11
Daams M et al (2014) Mean upper cervical cord area (MUCCA) measurement in long-standing multiple sclerosis: relation to brain findings and clinical disability. Mult Scler J 20(14):1860–1865
Montalban X et al (2021) A smartphone sensor-based digital outcome assessment of multiple sclerosis. Mult Scler J 18:654–664
Funding
This work is funded by the DeSBi Research Unit (DFG; KI-FOR 5363; Project ID 459422098), the Deutsche Multiple Sklerose Gesellschaft (DMSG) and the Fondation Eugène Devic EDMUS contre la Sclérose en Plaques & Observatoire Français de la Sclérose en Plaques (grant name DEEP MS), ZonMW (Vidi grant of MMS, project number 09150172010056) and the Dutch MS Research Foundation (08-650, 13-820 and 14-358e).
Author information
Authors and Affiliations
Contributions
SN: conception, execution of the study, analysis of the data, writing of the first draft, and review of the manuscript. MS: conception, execution of the study, analysis of the data, writing of the first draft, and review of the manuscript. CC: conception and review and critique of the manuscript. RPR: execution of the study and review and critique of the manuscript. FB: review and critique of the manuscript. EMMS: conception and review and critique of the manuscript. FP: conception and review and critique of the manuscript. MMS: conception, organization of the study, and review and critique of the manuscript. KR: conception, organization of the study, and review and critique of the manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
SN was supported by research grants from Atara Biotherapeutics, Merck and Biogen. CC reported grants from Novartis and Alexion during the conduct of the study, is part of a consortium project funded by the U.S. Department of Defense, and has a non-financial affiliation as a member from the Canadian Institutes of Health Research Standing Committee on Science outside the submitted work. FB is a steering committee or Data Safety Monitoring Board member for Biogen, Merck, ATRI/ACTC and Prothena; a consultant for Roche, Celltrion, Rewind Therapeutics, Merck, IXICO, Jansen and Combinostics; and has research agreements with Merck, Biogen, GE Healthcare and Roche. EMMS serves on the editorial board of Frontiers in Neurology received speaker fees from Merck and Novartis. FP served on the scientific advisory boards of Novartis and MedImmune; received travel funding and/or speaker honoraria from Bayer, Novartis, Biogen, Teva, Sanofi-Aventis/Genzyme, Merck Serono, Alexion, Chugai, MedImmune, and Shire; is an associate editor of Neurology: Neuroimmunology & Neuroinflammation; is an academic editor of PLoS ONE; consulted for Sanofi Genzyme, Biogen, MedImmune, Shire, and Alexion; received research support from Bayer, Novartis, Biogen, Teva, Sanofi-Aventis/Geynzme, Alexion, and Merck Serono; and received research support from the German Research Council, Werth Stiftung of the City of Cologne, German Ministry of Education and Research, Arthur Arnstein Stiftung Berlin, EU FP7 Framework Program, Arthur Arnstein Foundation Berlin, Guthy-Jackson Charitable Foundation, and NMSS. MMS serves on the editorial board of Neurology and Frontiers in Neurology, receives research support from the Dutch MS Research Foundation, Eurostars-EUREKA, ARSEP, Amsterdam Neuroscience, MAGNIMS and ZonMW (Vidi grant, project number 09150172010056) and has served as a consultant for or received research support from Atara Biotherapeutics. No other disclosures were reported.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Noteboom, S., Seiler, M., Chien, C. et al. Evaluation of machine learning-based classification of clinical impairment and prediction of clinical worsening in multiple sclerosis. J Neurol 271, 5577–5589 (2024). https://doi.org/10.1007/s00415-024-12507-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00415-024-12507-w