Abstract
Parkinson’s disease (PD) may be misdiagnosed due to the clinical overlap between PD and atypical parkinsonism. The utility of α-Synuclein (αSyn) Seed Amplification Assay (SAA) as a diagnostic indicator for PD has been reported in numerous studies, but never when administered as a validated clinical laboratory test. This study compares results from αSyn-SAA validation testing performed using well-characterized cohorts from two biorepositories to better understand the accuracy of PD clinical diagnosis. Blinded cerebrospinal fluid (CSF) specimens from a repository that included cohorts of subjects clinically diagnosed as PD or healthy controls, both with confirmatory dopamine transporter single-photon emission computed tomography (DAT SPECT) imaging, and blinded CSF specimens from a repository that included cohorts of subjects clinically diagnosed as PD or healthy controls based on clinical diagnosis alone, were tested as part of the validation studies for the diagnostic αSyn-SAA test (SYNTap® Biomarker Test). Measured αSyn-SAA test accuracy was 83.9% using clinical diagnosis as comparator, and 93.6% using clinical diagnosis with confirmatory DAT- SPECT imaging as comparator. The statistically significant discordance between accuracy determinations using specimens classified using different diagnostic inclusion criteria indicates that there is some symbiosis between dopamine-weighted imaging and αSyn-SAA results, both of which are associated with higher accuracy compared with the clinical diagnosis alone.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Approximately 20% of Parkinson’s disease (PD) may be misdiagnosed due to the clinical overlap between PD and atypical Parkinsonism [1]. Non-PD Parkinsonism disorders confound the diagnosis of PD and may have different pathological mechanisms (e.g., tauopathy, TDP-43 aggregation, etc.), and management [2]. Dopamine transporter single-photon emission computed tomography (DAT-SPECT) identifies deficits in presynaptic domine levels and is useful for demonstrating nigrostriatal degeneration occurring in PD and other diagnoses involving dopamine deficit [3, 4]. DAT-SPECT findings reflect dopaminergic function and do not necessarily identify the exact underlying etiology when a deficit is identified. However, when used for the indication of aiding PD diagnosis in cohorts with suspected PD (high pre-test probability), DAT-SPECT strengthens the accuracy of diagnosis [5]. The Parkinson’s Progressive Markers Initiative (PPMI) study included DAT-SPECT imaging to enhance the accuracy of diagnosis for PD and control cohorts with the rationale that individuals with “scans without DAT deficit are unlikely to have PD.” [5]. An accurate PD biomarker test is therefore expected to have a high correlation to PPMI cohort assignment, since there is high confidence in the accuracy of clinical diagnosis associated with these specimens.
The α-Synuclein (αSyn) Seed Amplification Assay (SAA) is a novel biomarker test that has significant support in the clinical research community. The assay utilizes in vitro propagation of minute amounts of aggregated misfolded αSyn to achieve levels detectable by simple fluorescence measurements of the amyloid-specific dye, Thioflavin T (ThT) [6,7,8,9]. αSyn‑SAA reproducibility across laboratories and methodological variations has been previously demonstrated; however, no study has been performed in a CLIA (Clinical Laboratory Improvement Amendments)/CAP (College of American Pathologists) accredited laboratory, and no study has compared cohorts selected with and without diagnostic imaging using SAA as the comparator [10]. To this end, we compared assay results between two cerebrospinal fluid (CSF) repositories using a CLIA/CAP validated αSyn-SAA, and compared test performance to repository inclusion diagnostic criteria.
Materials and methods
All patient specimens were collected under institutional review board approved protocols and with informed consent.
Study populations
For the first study, clinical data and frozen CSF from 98 participants were obtained from the Parkinson's Disease Biomarkers Program (PDBP) biorepository [11]. PDBP is a consortium in which participants are assessed longitudinally using standardized sample collection protocols and clinical assessments using Movement Disorder Society or UK Brain Bank criteria [12]. We initially identified case samples from subjects with PD (n = 41) and then identified control samples age and sex-matched to case samples to the extent possible (n = 57).
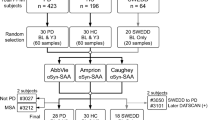
For the second study, different specimens from different individuals were obtained from the Parkinson’s Progression Markers Initiative (PPMI) repository [5, 10]. Unlike the PDBP repository, the PPMI enrollees require an abnormal DAT-SPECT test in addition to clinical symptoms to be included in the PD cohort and a normal DAT-SPECT test to be included in the control cohort. A total of 343 blinded samples from the PPMI biorepository were tested. The 343 samples were composed of 250 specimens from 109 individuals classified as controls, and 93 specimens collected from 55 individuals classified as PD. Specimens from participants who were clinically diagnosed with PD but had a normal DAT-SPECT on visual inspection (Scans Without Evidence of Dopaminergic Deficit; SWEDD) were not included in this study [5, 13]. In most instances where multiple samples were tested for one subject, these were collected from different visits. To appropriately weigh the influence of individual subjects on the statistics, results within the subject were randomly selected through computer-generated modeling to determine the most likely accuracy metrics for reporting. (Refer to Online Resource for additional information.) Fig. 1 depicts the study populations’ testing for both cohorts.
SAA procedure
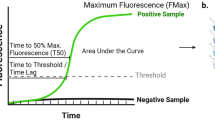
Briefly, a 2.45 mm diameter borosilicate glass bead (Sigmund Lindner GmbH, cat# 55-02450-89RTS), 160 µL of the reaction mixture, and 40µL of test sample were combined in each test well of a 96-well plate, and the plate was sealed with an optical adhesive film (Applied Biosystems, cat# 4311971). The reaction mixture used throughout the validation studies was similar to formulations described in previous studies [10, 14] and was composed of 100 mM PIPES (MilliporeSigma, cat#80635) pH 6.5, 10 µM ThT (MilliporeSigma, cat#T3516-25G), nuclease-free water (Growcells, cat#NUPW-0500), 500 mM NaCl (Lonza, cat#51202), and 0.3 mg/mL recombinant monomeric human αSyn (Amprion, cat#S2020). Three test wells are used for each patient assessment. A BMG LABTECH FLUOStar Ω Microplate Reader (excitation wavelength, 440 nm; emission wavelength, 490 nm) was used to measure ThT fluorescence in relative fluorescence units (RFU). Following a baseline reading, the assay plate was incubated at 37 °C and subjected to cycles of orbital shaking at 800 rpm for 1 min followed by a pause for 29 min. Fluorescence readings were taken once per day for 7 days with the exception of infrequent extended cases requiring 10 days. Extension occurs when a partial fluorescence signal has evolved near the end of the 7-day read window. Following the final measurement, the maximum relative fluorescence units (RFU) of each well is determined and the median of the 3 wells for each sample is calculated. Samples with median signal values greater than or equal to 25,000 RFU are classified as “Detected” (positive for aggregates of misfolded αSyn), and samples with median signal values < 25,000 RFU are classified as “Not Detected” (negative for aggregates of misfolded αSyn).
Statistics
The positive predictive value, negative predictive value, sensitivity, specificity, accuracy, and error rate (false results divided by the total number of results) are reported. 95% confidence intervals were calculated using the binomial confidence interval calculator in https://statpages.info/confint.html. Graphs and tables were generated using Microsoft Excel. Statistics were performed using R version 4.1.1 with statistical significance set to 0.05.
Results
We first looked at potential demographic differences between PPMI and PDBP repositories and found no statistical differences in age, race, or ethnicity apart from race and gender for the control cohort comparison (Table 1). To verify the PDBP and PPMI cohorts’ comparability in terms of symptomology, United Parkinson’s Disease Rating Scale (UPDRS) data were analyzed; there were no statistical differences between repositories (Table 1, Fig. 2).
Box and whisker plots of the total UPDRS Scores broken down by repository and case/control cohorts. The median UPDRS scores for the PDBP and the PPMI control and case subjects were 4 and 5 and 48 and 50, respectively. p values from robust T tests (Yuen) were 0.37 and 0.73, respectively, when comparing control and case scores between repositories
An important difference between these repositories is the inclusion criteria, which includes DAT-SPECT in the PPMI cohort. Considering repository diagnosis as the comparator, αSyn-SAA accuracy was higher in the PPMI cohort (93.6%) than in the PDBP cohort (83.9%) (Table 2). It is important to note that the error rate for SAA using the PPMI cohort was 6.4% compared to 16.1% in the PDBP cohort (p 0.045) (Table 2). For information, additional validation results, including analytical validation and specimen stability studies, are shown in the Online Resource.
Discussion
The αSyn-SAA results reveal discordance in the accuracy of clinical diagnosis between two repositories of samples collected from donors diagnosed with PD based on different criteria, namely clinical diagnosis with and without ancillary imaging/enrichment by DAT SPECT. Discordance was anticipated since clinical diagnosis of PD based on clinical evaluation alone carries a significant misdiagnosis rate of ~ 20% [1, 15] and is known to be enhanced with a demonstration of the presence or absence of degenerative findings by DAT-SPECT [16].
Because of the relation between higher αSyn-SAA accuracy and enrichment by DAT-SPECT in the inclusion criterion, we endorse that diagnoses associated with specimens obtained from the PPMI are close to clinical truth (precision between Parkinson diagnostic rate and Parkinson pathobiology). Enrichment by αSyn-SAA alone might reach similar or higher PD diagnostic accuracy than clinical diagnosis enhanced by DAT-SPECT, since αSyn-SAA reaches high accuracy in the PPMI cohort (~ 94%) and DAT-SPECT enrichment is not 100% diagnostic [17]. Moreover, αSyn-SAA has been shown to be more sensitive in borderline PD cases of the PPMI cohort [14]. Dopaminergic depletion is hypothesized to occur later in the disease process than αSyn misfolding, so it is possible that the positive αSyn-SAA samples with normal DAT-SPECT are at earlier stages of the disease. In theory, the provider may find utility in either of these methods depending on the clinical presentation since both DAT-SPECT and αSyn-SAA studies have shown utility in refining PD etiology by uncovering information from two discrete areas in the PD landscape.
These hypotheses were confirmed in a recently published research study using > 1100 PPMI CSF specimens and representing the largest analysis to date of αSyn-SAA [18]. Similar to the results in the present study using PPMI specimens, this study showed sensitivity for sporadic PD of 93.3%, and specificity for healthy controls of 96.3%. The larger PPMI study also included specimens from SWEDD, prodromal, and non-manifesting genetic carriers. Sensitivity for prodromal PD (hyposmia or REM behavior sleep disorder) was 86%. Importantly, there was evidence that abnormal αSyn aggregation detectable by αSyn-SAA occurs before other detectable clinical or biomarker changes, including DAT-SPECT. In addition, αSyn-SAA provided information about molecular heterogeneity, with αSyn-SAA positivity lower in LRRK2 PD (67.5%) and higher in GBA PD (95.9%) compared with sporadic PD. Overall, the authors conclude that αSyn-SAA classifies people with PD with high accuracy and can play a crucial role to identify pathologically defined subgroups of people with PD and to establish a biomarker-driven definition of the disease. αSyn-SAA technology has been extensively proven in the research environment, and the test has been officially validated per CLIA and CAP guidelines allowing a significant advancement in diagnostic biochemistry that has the capacity to change the diagnostic approach to neurodegenerative diseases. Accurate “rule-in” and “rule-out” of synucleinopathies with a biochemical test promises utility since diagnosis of synucleinopathies can otherwise consume long periods of time and potentially involve unnecessary drug challenges. This validation also serves as a precedent for a new class of clinical testing using protein aggregation science in a CLIA-certified high-complexity laboratory. Such a test will also aid in clinical research trials requiring a validated test platform for subject screening or monitoring.
Data availability statement
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Rizzo G, Copetti M, Arcuti S et al (2016) Accuracy of clinical diagnosis of Parkinson disease: A systematic review and meta-analysis. Neurology 86(6):566–576
Dickson DW (2012) Parkinson’s disease and parkinsonism: neuropathology. Cold Spring Harb Perspect Med 2(8):009258
Bega D, Kuo PH, Chalkidou A et al (2021) Clinical utility of DaTscan in patients with suspected Parkinsonian syndrome: a systematic review and meta-analysis. NPJ Parkinsons Dis 7(1):43
Tatsch K, Poepperl G (2013) Nigrostriatal dopamine terminal imaging with dopamine transporter SPECT: an update. J Nucl Med 54(8):1331–1338
Parkinson Progression Marker I (2011) The Parkinson Progression Marker Initiative (PPMI). Prog Neurobiol 95(4):629–635
Fairfoul G, McGuire LI, Pal S et al (2016) Alpha-synuclein RT-QuIC in the CSF of patients with alpha-synucleinopathies. Ann Clin Transl Neurol 3(10):812–818
Shahnawaz M, Tokuda T, Waragai M et al (2017) Development of a biochemical diagnosis of Parkinson disease by detection of alpha-synuclein misfolded aggregates in cerebrospinal fluid. JAMA Neurol 74(2):163–172
Groveman BR, Orru CD, Hughson AG et al (2018) Rapid and ultra-sensitive quantitation of disease-associated alpha-synuclein seeds in brain and cerebrospinal fluid by alphaSyn RT-QuIC. Acta Neuropathol Commun 6(1):7
Arnold MR, Coughlin DG, Brumbach BH et al (2022) alpha-synuclein seed amplification in CSF and brain from patients with different brain distributions of pathological alpha-synuclein in the context of co-pathology and non-LBD diagnoses. Ann Neurol 92:650–662
Russo MJ, Orru CD, Concha-Marambio L et al (2021) High diagnostic performance of independent alpha-synuclein seed amplification assays for detection of early Parkinson’s disease. Acta Neuropathol Commun 9(1):179
Gwinn K, David KK, Swanson-Fischer C et al (2017) Parkinson’s disease biomarkers: perspective from the NINDS Parkinson’s Disease Biomarkers Program. Biomark Med 11(6):451–473
Marsili L, Rizzo G, Colosimo C (2018) Diagnostic criteria for Parkinson’s disease: from James Parkinson to the concept of prodromal disease. Front Neurol 9:156
Erro R, Schneider SA, Stamelou M et al (2016) What do patients with scans without evidence of dopaminergic deficit (SWEDD) have? New evidence and continuing controversies. J Neurol Neurosurg Psychiatry 87(3):319–323
Concha-Marambio L, Farris CM, Holguin B et al (2021) Seed amplification assay to diagnose early Parkinson’s and predict dopaminergic deficit progression. Mov Disord 36(10):2444–2446
Tolosa E, Garrido A, Scholz SW et al (2021) Challenges in the diagnosis of Parkinson’s disease. Lancet Neurol 20(5):385–397
Gayed I, Joseph U, Fanous M et al (2015) The impact of DaTscan in the diagnosis of Parkinson disease. Clin Nucl Med 40(5):390–393
Brigo F, Matinella A, Erro R et al (2014) [(1)(2)(3)I]FP-CIT SPECT (DaTSCAN) may be a useful tool to differentiate between Parkinson’s disease and vascular or drug-induced Parkinsonisms: a meta-analysis. Eur J Neurol 21(11):1369-e1390
Siderowf A, Concha-Marambio L, Lafontant DE et al (2023) Assessment of heterogeneity among participants in the Parkinson’s Progression Markers Initiative cohort using alpha-synuclein seed amplification: a cross-sectional study. Lancet Neurol 22(5):407–417
Acknowledgements
Some of the data and biospecimens used in the preparation of this manuscript were obtained from the Parkinson’s Disease Biomarkers Program (PDBP) Consortium, supported by the National Institute of Neurological Disorders and Stroke at the National Institutes of Health. Investigators include Roger Albin, Roy Alcalay, Alberto Ascherio, Thomas Beach, Sarah Berman, Bradley Boeve, F. DuBois Bowman, Shu Chen, Alice Chen-Plotkin, William Dauer, Ted Dawson, Paula Desplats, Richard Dewey, Ray Dorsey, Jori Fleisher, Kirk Frey, Douglas Galasko, James Galvin, Dwight German, Steven Gunzler, Lawrence Honig, Xuemei Huang, David Irwin, Kejal Kantarci, Anumantha Kanthasamy, Daniel Kaufer, Qingzhong Kong, James Leverenz, Carol Lippa, Irene Litvan, Oscar Lopez, Jian Ma, Lara Mangravite, Karen Marder, Nandakumar Narayanan, Laurie Orzelius, Vladislav Petyuk, Judith Potashkin, Liana Rosenthal, Rachel Saunders-Pullman, Clemens Scherzer, Michael Schwarzschild, Tanya Simuni, Andrew Singleton, David Standaert, Debby Tsuang, David Vaillancourt, Jerrold Vitek, David Walt, Andrew West, Cyrus Zabetian, and Jing Zhang. The PDBP Investigators have not participated in reviewing the data analysis or content of the manuscript. Research reported in this document was supported in part by the National Institute of Neurological Disorders and Stroke of the National Institutes of Health under Award Number U44NS111672 and the Michael J. Fox Foundation for Parkinson’s Research grant MJFF-020475. Data used in the preparation of this article were obtained from the Parkinson’s Progression Markers Initiative (PPMI) database (www.ppmi-info.org/data). For up-to-date information on the study, visit www.ppmi-info.org. Data and biospecimens used in the analyses presented in this article were obtained from the Parkinson’s Progression Markers Initiative (PPMI) (www.ppmi-info.org/access-data-specimens/download-data). As such, the investigators within PPMI contributed to the design and implementation of PPMI and/or provided data and collected biospecimens but did not participate in the analysis or writing of this report. PPMI – a public-private partnership – is funded by the Michael J. Fox Foundation for Parkinson’s Research funding partners 4D Pharma, Abbvie, Acurex Therapeutics, Allergan, Amathus Therapeutics, ASAP, Avid Radiopharmaceuticals, Bial Biotech, Biogen, BioLegend, Bristol-Myers Squibb, Calico, Celgene, Dacapo Brain Science, Denali, The Edmond J. Safra Foundation, GE Healthcare, Genentech, GlaxoSmithKline, Golub Capital, Handl Therapeutics, Insitro, Janssen Neuroscience, Lilly, Lundbeck, Merck, Meso Scale Discovery, Neurocrine Biosciences, Pfizer, Piramal, Prevail, Roche, Sanofi Genzyme, Servier, Takeda, Teva, UCB, Verily, and Voyager Therapeutics. The content is solely the responsibility of Amprion and does not necessarily represent the official views of the National Institutes of Health or the Michael J. Fox Foundation. Samples from the NINDS BioSEND, which receives government support under a cooperative agreement grant (U24 NS095871) awarded by the National Institute of Neurological Disorders and Stroke (NINDS), were used in this study. We thank contributors who collected samples used in this study, as well as patients and their families, whose help and participation made this work possible. We would like to acknowledge the PDBP repository staff for generous assistance in the blinding/unblinding of PDBP patient samples.
Funding
This work was supported by the National Institute of Neurological Disorders and Stroke of the National Institutes of Health under Award Number U44NS111672 and the Michael J. Fox Foundation for Parkinson’s Research Grant MJFF-020475.
Author information
Authors and Affiliations
Contributions
Conception, writing, data interpretation: JSM, KJJ and KRM. Data analysis: JSM, HH. Data acquisition: NK, MJM. Manuscript editing: JSM, HLH, KRM, LC-M, KJJ.
Corresponding author
Ethics declarations
Conflicts of interest
Dr. Jensen consults for Prometheus Laboratories, Sharp Healthcare, and Adaptive Biotechnologies, and holds no patent rights and stocks with these entities. Dr. Concha is an inventor on several patents related to SAA technology. All authors are associated with Amprion Inc., a biotech company focused on the development and commercial utilization of SAA technology.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Middleton, J.S., Hovren, H.L., Kha, N. et al. Seed amplification assay results illustrate discrepancy in Parkinson’s disease clinical diagnostic accuracy and error rates. J Neurol 270, 5813–5818 (2023). https://doi.org/10.1007/s00415-023-11810-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00415-023-11810-2