Abstract
Background
Recent evidence points toward a role of the small ubiquitin-like modifier (SUMO) system, including SUMO4, in protecting from stress insults and neurodegeneration, such as the progressive motor neuron disease amyotrophic lateral sclerosis (ALS), e.g., by regulating stress granule (SG) dynamics. Here, we investigated whether SUMO4 variants play a role in ALS pathogenesis.
Methods
Whole-exome or targeted SUMO4 sequencing was done in 222 unrelated European ALS patients. The consequences of the identified initiator codon variant were analyzed at the mRNA, protein and cellular level. SUMO4 expression was quantified in human tissues. All patients were subjected to clinical, electrophysiological, and neuroradiological characterization.
Results
A rare heterozygous SUMO4 variant, i.e., SUMO4:c.2T>C p.Met1?, was detected in four of 222 (1.8%) ALS patients, significantly more frequently than in two control cohorts (0.3% each). SUMO4 mRNA and protein expression was diminished in whole blood or fibroblasts of a SUMO4 variant carrier versus controls. Pertinent stress factors, i.e., head trauma or cancer (treated by radiochemotherapy), were significantly more frequent in SUMO4 variant carrier versus non-carrier ALS patients. The mean number of SGs per cell was significantly higher in fibroblasts of a SUMO4 variant carrier compared to controls at baseline, upon oxidative stress, and after recovery, and SUMOylation of ALS-associated valosin-containing protein by SUMO4 was decreased. SUMO4 mRNA expression was highest in brain of all human tissues analyzed.
Conclusions
Our results are consistent with SUMO4 haploinsufficiency as a contributor to ALS pathogenesis impacting SG dynamics and possibly acting in conjunction with environmental oxidative stress-related factors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Amyotrophic lateral sclerosis (ALS) is a progressive fatal neurodegenerative disease characterized by the loss of motor neurons. Mutations in ALS-associated genes, such as FUS (FUS RNA binding protein) and VCP (valosin-containing protein), as well as environmental stress and lifestyle factors are involved in ALS pathogenesis. In response to various stress conditions, stress granules (SGs), cytosolic membrane-less organelles consisting of untranslated mRNAs and RNA binding proteins (RBPs), form. Evidence is emerging that pathological aggregation of RBPs and persistent SGs play a role in neurodegeneration [1], and represent promising therapeutic targets [2].
In a mouse model recently reported by Zhang et al., an ALS-associated FUS mutation was linked to SG misprocessing and motor performance decline aggravated by stress [3]. FUS encodes a nuclear RBP that, during the SG cycle, translocates to the cytoplasm where SGs nucleate, mature, and disperse [1]. Among other things, Zhang et al. reported that cultured motor neurons from FUS knock-in mice were prone to form SGs in response to oxidative stress, and showed decreased disassembly of SGs after recovery from stress challenge. These data corroborate and extend earlier findings in HEK293 cells and the zebrafish spinal cord that expression of a FUS nonsense mutation, known to cause ALS, resulted in cytoplasmic accumulation of mutant FUS, and assembly into SGs upon oxidative stress [4]. Furthermore, mutations in VCP and other ALS-associated genes have been shown to affect SGs under stress conditions [5, 6], suggesting that yet other SG-related proteins may be involved in ALS pathogenesis.
The distribution of VCP to SGs required for SG clearance [7] is facilitated by SUMOylation of VCP [5]. SUMOylation, a ubiquitination-like post-translational modification, involves the reversible covalent conjugation of a small ubiquitin-like modifier (SUMO) to lysine residues of a multitude of cellular proteins [8]. Recent evidence points toward a role of the SUMO system in protecting from ALS pathology, e.g., by regulating SG dynamics [5]. For instance, SG disassembly-engaged proteins include SUMO ligases that are recruited during normal clearance and dysregulated in ALS-like conditions [9]. Interestingly, SUMO E3 ligase activity has been described for FUS [10]. To our knowledge, no ALS-associated mutations have been reported in genes encoding SUMO isoforms 1–4. Here, we provide evidence that not only a FUS mutation, as described in the ALS mouse model by Zhang et al. [3], but also a specific variant in the SUMO4 gene detected in ALS patients affects SG dynamics as an underlying pathomechanism of ALS.
Patients and methods
Subjects and their clinical characterization
The study was approved by the Ethics Board of Hannover Medical School (ID # 6269). Written informed consent was obtained from all individuals. At the ALS/motor neuron disease clinic of the Department of Neurology, Hannover Medical School, Hannover, Germany, 222 central European ALS patients (11 with familial ALS and 211 with sporadic ALS), and in some cases, their parents or siblings were recruited to the study. The median age at onset of patients was 61 years, and the male to female ratio was 1.4:1. Regarding the site of symptom onset, the proportions of subjects with initial bulbar and spinal symptoms were 23.4 and 76.6%, respectively. Patients were subdivided into eight clinical subtypes (upper motor neuron dominant ALS, bulbar phenotype, flail arm syndrome, flail leg syndrome, respiratory phenotype, progressive muscular atrophy, lower motor neuron dominant ALS, and classic (Charcot) ALS) as described previously [11]. Disease status was evaluated using the revised ALS functional rating scale (ALSFRS-R). At first presentation, the progression rate was calculated using the following formula: 48–total ALSFRS-R/symptom duration in months [12]. Of the proposed environmental stress and lifestyle factors increasing the risk of ALS [13], smoking habits, occupation, chemical exposure, and physical exercise were not assessed routinely, whereas history of head trauma and tumors with associated treatments, such as chemotherapy or radiation, could be retrieved from patients’ past medical records. Detailed information on the severity of traumatic head injury or the specific radiotherapy/chemotherapeutic regimens used was not available.
Whole-exome/targeted sequencing and data analysis
DNA was extracted from whole blood samples using the QIAamp DNA Blood Maxi Kit (Qiagen, Hilden, Germany). Whole-exome sequencing (WES) was performed on leukocyte DNA of 24 ALS patients and 1787 in-house individuals. Target enrichment was done using the Agilent SureSelect Human All Exon v4 Target Enrichment System (Agilent Technologies, Inc., Santa Clara, CA, U.S.A.) or the xGen® Exome Research Panel (Integrated DNA Technologies, Inc., Coralville, U.S.A.). All samples were sequenced to a mean target coverage of >50× on an Illumina HiSeq 2000 or an Illumina NextSeq 500 using the NextSeq 500/550 High Output v2 kit (all Illumina, San Diego, CA, U.S.A.). Data were processed and aligned to the GRCh37/hg19 reference human genome build using the Biomedical Genomics Workbench (version 5.0; Qiagen) or megSAP, version 0.1-710-g52d2b0c (https://github.com/imgag/megSAP). Variant prioritization and visualization were performed using Ingenuity Variant Analysis (Qiagen), GSvar version 2018_04 (https://github.com/imgag/ngs-bits), IGV version 2.4.14 (https://software.broadinstitute.org/software/igv/), and Alamut® visual version 2.11 (Interactive Biosoftware, Rouen, France). Targeted sequencing of the coding and flanking intronic sequences of the SUMO4 gene (NM_001002255) by a conventional chain termination protocol was used to verify SUMO4 variants identified by WES, and to do mutational analysis of 198 additional ALS patients. Minor allele frequencies (MAF) of genetic variants were extracted from the Genome Aggregation Database (gnomAD) Browser v2.1.1 (https://gnomad.broadinstitute.org). Variant pathogenicity was predicted using SIFT (http://sift.jcvi.org/), PolyPhen-2 (http://genetics.bwh.harvard.edu/pph2/), and MutationTaster (http://www.mutationtaster.org).
Quantitative SUMO4 mRNA expression analysis in ALS patients and human tissues
RNA was isolated from whole blood of three ALS patients, i.e., SUMO4 variant carrier TALS004-01 and two non-carriers, using the RNeasy Mini Kit (Qiagen). To eliminate DNA contamination, RNA samples were treated with RNase-free DNase I (Qiagen). cDNA synthesis was performed using the SuperScript III First-Strand Synthesis System (Thermo Fisher Scientific, Waltham, MA, U.S.A.). For quantitative SUMO4 mRNA expression analysis, the TaqMan Universal PCR Master Mix and a TaqMan gene expression assay for SUMO4 (Hs01940570_g1; Thermo Fisher Scientific) were used to analyze cDNA of ALS patients and Human Multiple Tissue cDNA (MTC) Panel I (#636742; Clontech-Takara Bio Europe, Saint Germain-en-Laye, France) in three and two independent experiments, respectively. Target gene expression levels were normalized to expression levels of B2M (Hs00187842_m1; Thermo Fisher Scientific), and comparative Ct quantification was applied.
Cell culture of fibroblasts of patient TALS004-01 and controls
Skin biopsy-derived fibroblasts of patient TALS004-01 and of two age- and sex-matched non-ALS controls were cultured in Dulbecco’s Modified Eagle Medium (DMEM, Merck, Darmstadt, Germany) supplemented with 10% fetal bovine serum, 2 mM l-glutamine, and 1% penicillin/streptomycin (all Thermo Fisher Scientific).
Targeted SUMO4 sequencing on fibroblast DNA of patient TALS004-01 and controls
DNA was isolated from fibroblasts of patient TALS004-01 and the non-ALS controls using the innuPREP DNA Mini Kit (Analytik Jena, Jena, Germany). The coding sequence and parts of the flanking untranslated regions of SUMO4 were amplified and sequenced using a conventional chain termination protocol. Sequence data were analyzed using the SeqPilot software v4.3.1 (JSI Medical Systems GmbH, Ettenheim, Germany).
Western blot analysis of fibroblast lysates of patient TALS004-01 and controls
Fibroblasts were homogenized in lysis buffer (20 mM Tris–HCl, pH 8.0, 50 mM sodium fluoride, 1 mM sodium orthovanadate, 1% Nonidet P40, supplemented with protease inhibitors (Roche Diagnostics, Mannheim, Germany)), and protein concentration was determined using the Pierce BCA Protein Assay Kit (Thermo Fisher Scientific). Cell lysates were adjusted to equal protein concentrations, mixed with 4 × Laemmli buffer (250 mM Tris–HCl, pH 6.8, 40% glycerol, 8% sodium dodecyl sulfate (SDS), 20% 2-mercaptoethanol, 4 mM ethylenediaminetetraacetic acid (EDTA), 0.04% bromophenol blue (w/v)), and used in equal amounts for SDS–polyacrylamide gel electrophoresis and semidry electroblotting. After incubating polyvinylidene difluoride or nitrocellulose membranes (GE Healthcare, Chicago, IL, U.S.A.) in 5% fat-free milk powder dissolved in phosphate-buffered saline (PBS) with 0.05% Tween 20 (PBST) to block unspecific binding, the following primary antibodies were diluted in 5% bovine serum albumin (BSA) in PBST and incubated at 4 °C overnight for immunodetection: anti-GAPDH (#MAB374, Merck; dilution 1:3000), anti-pan-SUMO detecting SUMO1-4 (#A-714, R&D Systems, Minneapolis, MN, U.S.A.; dilution 1:1000), anti-SUMO4 [EPR7163], lot: GR155829-1 (#ab126606, Abcam, Cambridge, U.K.; dilution 1:1000), anti-VCP (#MA3-004; Thermo Fisher Scientific; dilution 1:1000). All membranes were exposed to the corresponding horseradish peroxidase-conjugated secondary antibody (Thermo Fisher Scientific) diluted at 1:3000 in 5% fat-free milk powder dissolved in PBST, and developed using the Pierce SuperSignal West Dura detection kit (Thermo Fisher Scientific). Images were acquired using the Fusion-FX7 gel documentation system (Vilber, Collégien, France). Protein bands were quantified using the ImageJ software (https://imagej.nih.gov/ij/). SUMO4, pan-SUMO and GAPDH protein levels were calculated from three independent experiments.
Immunofluorescence of fibroblasts of patient TALS004-01 and controls to visualize stress granules
Fibroblasts (5.0 × 104 cells) were seeded on fibronectin-coated glass coverslips. After 24 h, cells were treated with 0.5 mM sodium arsenite (NaAsO2) solution (Sigma-Aldrich, St. Louis, MO, U.S.A.) for 45 min at 37 °C (oxidative stress). After removal of NaAsO2, cells were incubated in DMEM with supplements for 60 min at 37 °C (recovery). Cells were fixed with 4% paraformaldehyde for 15 min at 4 °C, permeabilized with 0.25% Triton X-100 in PBS for 25 min at room temperature (RT), and incubated with 0.1% Triton X-100, 6% BSA in PBS for 1 h at RT to block unspecific binding. The primary anti-TIAR rabbit antibody (D32D3, #8509, Cell Signaling Technology, Danvers, MA, U.S.A.) was diluted at 1:1600 in 0.1% Triton X-100, 1% BSA in PBS and incubated overnight at 4 °C. After washing three times in PBS, the Alexa Fluor 488-conjugated anti-rabbit secondary antibody (Thermo Fisher Scientific; dilution 1:500) was incubated for 1 h at RT. After counterstaining with 4′,6-diamidino-2-phenylindole (DAPI), cells were mounted in Mowiol mounting medium. Images were captured using an epifluorescence microscope (Leica DM RXA2) equipped with a cooled charge-coupled device camera (SenSys, Photometrics, Tucson, AZ, U.S.A.). Quantification of TIAR-positive stress granules was performed using Fiji software [14]. Stress granules were arbitrarily defined as cytoplasmic foci of ≥ 0.5 µm in diameter staining positive for TIAR. At least 30 cells were analyzed per condition and individual in each of three independent experiments.
Enrichment of VCP from fibroblast lysates of patient TALS004-01 and controls by immunoprecipitation
Fibroblasts were washed with ice-cold PBS and lysed for 30 min on ice in 1 ml of immunoprecipitation (IP) lysis buffer (20 mM Tris–HCl, pH 8.0, 50 mM sodium fluoride, 1 mM sodium orthovanadate, 1% Nonidet P40, supplemented with protease and phosphatase inhibitors; Roche Diagnostics). Cell lysates were incubated with 1 µg anti-VCP antibody (#MA3-004; Thermo Fisher Scientific) by tumbling over night at 4 °C. Protein G Sepharose beads (GE Healthcare) were equilibrated in IP lysis buffer and incubated with the cell lysates for 4 h at 4 °C. After six washing steps with IP lysis buffer, bound protein was eluted from the beads using 1 × Laemmli buffer (62.5 mM Tris–HCl, pH 6.8, 10% glycerol, 2% SDS, 5% 2-mercaptoethanol, 1 mM EDTA, 0.01% bromophenol blue). VCP and SUMO4-VCP were detected by Western blot analysis using anti-VCP and anti-SUMO4 antibodies, as described above. Protein bands were quantified using the ImageJ software (https://imagej.nih.gov/ij/). Levels of SUMO4-VCP and VCP immunoprecipitates were calculated from two independent experiments.
Statistics
Statistical comparisons were made using two-tailed Fisher’s exact test, Mann–Whitney U Test or Chi-square test, as appropriate and indicated in table legends. Statistical significance of experiments was calculated using two-tailed Student’s t test or one-way ANOVA as indicated in the figure legend. The analysis was carried out using MATLAB (R2017a; The MathWorks, Inc., Natick, MA, U.S.A.), PRISM software (v.5; GraphPad software, San Diego, CA, U.S.A.) or SPSS (IBM® Statistical Software Package of Social Science, Chicago, IL, U.S.A.) version 26, p-values of ≤ 0.05 (*), ≤ 0.01 (**), and ≤ 0.001 (***) were considered statistically significant.
Results
With the aim to identify new ALS-associated genes and to investigate whether rare SUMO4 variants play a role in ALS pathogenesis, we analyzed leukocyte DNA of 222 unrelated central European ALS patients using whole-exome or targeted SUMO4 sequencing. We detected an identical initiator codon variant in the SUMO4 gene, NM_001002255(SUMO4):c.2T>C p.Met1?, in four of the 222 ALS patients, verified to be heterozygous by Sanger sequencing (Table 1; Fig. 1a). The four ALS patients carrying the SUMO4:c.2T>C variant were of German origin, but from different German regions/federal states, and, thus, do not belong to a specific subpopulation. The SUMO4:c.2T>C variant (with a MAF < 0.2%, Table 1) as well as all rare (MAF < 0.2%) SUMO4 variants predicted to be deleterious by SIFT and/or PolyPhen-2 (according to the Genome Aggregation Database, v2.1.1) taken together were significantly more frequent in central European ALS patients (4/222 (1.8%), both comparisons) compared to in-house individuals mostly of German origin (5/1786 (0.3%), p = 0.0117 and 7/1786 (0.4%), p = 0.0256, two-tailed Fisher’s exact test) and non-Finnish European controls from the Genome Aggregation Database, v2.1.1, controls (74/24,101 (0.3%), p < 0.0001 and 90/20,432 (0.4%), p = 0.0027, Chi-square test). None of the four SUMO4:c.2T>C variant carriers harbored a hexanucleotide repeat expansion or pathogenic variant in the ALS-associated genes C9orf72 (not analyzed in VALS051), FUS, SOD1, TARDBP, or VCP, or had a family history of ALS. None of the 24 ALS patients analyzed by WES carried a rare (MAF < 0.2%) variant in the SUMO1, SUMO2 or SUMO3 gene. The SUMO4:c.2T>C variant was also identified in the healthy father of variant carrier TALS004-01, the only parent available for genetic testing, suggesting either reduced penetrance of the variant, later onset of ALS in the father, or combined genetic and environmental causes of ALS in the patient. The latter assumption is corroborated by the fact that exposure to certain potentially pertinent oxidative stress-related factors, i.e., head trauma or tumor with and without treatment by radiation or chemotherapy, were significantly more frequent in patients carrying the SUMO4:c.2T>C variant compared to non-carriers in our ALS cohort (p = 0.012; Tables 2, 3). Patient TALS004-01 presented with a history of head trauma at 6 years of age (Table 3), whereas no stress factor has been reported in his father. Other characteristics of SUMO4:c.2T>C variant carriers versus non-carriers were not significantly different in our ALS cohort (Table 2).
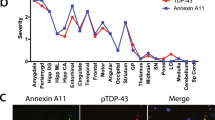
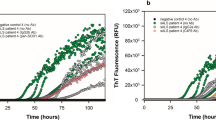
A rare heterozygous SUMO4:c.2T>C initiator codon variant detected in 1.8% of ALS patients leads to reduced SUMO4 expression, altered stress granule dynamics, and reduced SUMOylation of VCP. a Electropherograms demonstrating the rare heterozygous SUMO4:c.2T>C p.Met1? variant in leukocyte DNA of four sporadic ALS patients. b SUMO4 mRNA expression in whole blood of SUMO4:c.2T>C variant carrier TALS004-01 and two ALS control patients without the SUMO4:c.2T>C variant was quantified by real-time RT-PCR and normalized to B2M mRNA. SUMO4 mRNA levels were significantly reduced in whole blood of patient TALS004-01 compared to ALS controls (mean ± SD of triplicate samples from three independent experiments). c–e Detection (d) and densitometric quantification (e) of unconjugated SUMO4 protein and all unconjugated SUMO isoforms 1–4 by Western blot analysis of lysates of fibroblasts of patient TALS004-01 harboring the SUMO4:c.2T>C variant (c) and of two sex- and age-matched non-ALS controls without rare SUMO4 variants using anti-SUMO4 and anti-pan-SUMO (detecting SUMO isoforms 1–4) antibodies, showing that SUMO4 protein levels were significantly reduced and levels of all SUMO isoforms were reduced in fibroblasts of patient TALS004-01 compared to controls (mean ± SD from three independent experiments). f, g Detection (f) of stress granules (SGs) in fibroblasts of patient TALS004-01 and two non-ALS controls that were untreated (baseline), treated for 45 min with 0.5 mM sodium arsenite (NaAsO2) or had recovered for 60 min from NaAsO2 treatment (recovery) by immunostaining of the SG marker protein nucleolysin TIAR; nuclei were counterstained with DAPI; scale bar 10 µm. Quantification (g) of SGs from (f), showing significantly higher numbers of SGs in fibroblasts of patient TALS004-01 compared to fibroblasts of non-ALS controls at baseline, upon NaAsO2, and after recovery (mean ± SEM of ≥ 95 cells per individual and condition from three independent experiments with at least 30 cells per individual, condition and experiment). h, i Immunoprecipitation (IP) of VCP, and detection (h) and densitometric quantification (i) of SUMO4-conjugated VCP by Western blot analysis of fibroblast lysates of patient TALS004-01 and non-ALS controls using an anti-SUMO4 antibody, showing that VCP is SUMOylated by SUMO4 and that SUMOylation of VCP by SUMO4 is significantly reduced in fibroblasts of patient TALS004-01 compared to controls (mean ± SD from two independent experiments). j SUMO4 mRNA expression in a human adult multiple tissue cDNA panel was quantified by real-time PCR, normalized to B2M mRNA, and displayed relative to mRNA levels in brain (†). SUMO4 mRNA levels were highest in brain followed by placenta and skeletal (sk.) muscle (mean ± SD of duplicate samples from two independent experiments). *p < 0.05; **p ≤ 0.01; ***p ≤ 0.001; n.s. not significant; according to two-tailed Student’s t test (b, e, i) or one-way ANOVA (g)
To analyze the consequences of the heterozygous SUMO4:c.2T>C initiator codon variant, RNA was extracted from whole blood of patient TALS004-01, who was still alive, and two ALS patients not carrying the SUMO4 variant. By real-time reverse transcription PCR, SUMO4 mRNA levels of SUMO4:c.2T>C variant carrier TALS004-01 were significantly lower than mean levels of the two non-carriers (Fig. 1b). These results were confirmed on the protein level by Western blot analysis of lysates of skin biopsy-derived fibroblasts of patient TALS004-01, which were also shown to harbor the SUMO4:c.2T>C variant (Fig. 1c) and to contain significantly decreased unconjugated SUMO4 protein levels compared to fibroblasts of two sex- and age-matched non-ALS controls without rare SUMO4 variants (Fig. 1d, e). Similar results were obtained using a pan-SUMO antibody detecting all SUMO isoforms, i.e., SUMO1-4, although not statistically significant (Fig. 1d, e). Taken together, we show diminished SUMO4 mRNA and protein expression due to the heterozygous SUMO4:c.2T>C variant, apparently not fully compensated by an increased expression of other SUMO isoforms.
In three of four SUMO4 variant carriers, environmental stress factors were identified as potential additional triggers of ALS pathogenesis (Tables 2, 3). Therefore, we investigated whether the cellular response to stress was altered in fibroblasts of SUMO4 variant carrier TALS004-01. Using immunofluorescence and the SG marker nucleolysin TIAR, SGs were quantified in fibroblasts of patient TALS004-01 and sex- and age-matched non-ALS controls at baseline, upon arsenite-induced oxidative stress, and after recovery (Fig. 1f). The mean number of TIAR-positive SGs per cell was significantly higher in fibroblasts of patient TALS004-01 compared to control fibroblasts in all three conditions, i.e., baseline, arsenite treatment, and recovery (Fig. 1g). These data demonstrate persistent SGs, increased SG accumulation upon stress exposure, and reduced SG dynamics after stress challenge in fibroblasts of an ALS patient carrying the SUMO4:c.2T>C variant.
As SUMOylation of the ALS-associated protein VCP by SUMO1 was previously reported [5], we analyzed whether SUMO4 is also conjugated to VCP. Our analysis of VCP-immunoprecipitates from fibroblast lysates demonstrates for the first time that VCP is also SUMOylated by SUMO4 (Fig. 1h). Consistent with the lower unconjugated SUMO4 levels in fibroblasts of SUMO4 variant carrier TALS004-01, SUMOylation of VCP by SUMO4 was significantly reduced in these cells compared to sex- and age-matched non-ALS control fibroblasts (Fig. 1h, i).
To further establish whether SUMO4 variants may play a role in a neurodegenerative disease such as ALS, SUMO4 mRNA expression was quantified in different human tissues. By real-time PCR, the highest human SUMO4 mRNA levels were observed in brain, followed by placenta and skeletal muscle (Fig. 1j).
Discussion
Here, we demonstrate that a heterozygous initiator codon variant in the SUMO4 gene is significantly more frequent in 222 European ALS patients than in two control cohorts from Europe. SUMO4 variants have previously been associated with human disease in that the SUMO4 Met55Val variant was implicated in the susceptibility to type I diabetes mellitus [15, 16]. SUMO4 encodes one of four human SUMO isoforms that are attached to more than a thousand proteins in human cells and are essential for the regulation of several cellular processes such as transcription and mRNA processing [6, 8]. As may be expected from a disrupted initiation codon shown to diminish gene expression in the case of an ATG to ATA conversion at the Fgf5 locus [17], the consequences of the heterozygous SUMO4:c.2T>C variant were reduced SUMO4 mRNA and protein levels. This variant effect does not seem to be fully compensated by an increased expression of other SUMO isoforms consistent with SUMO4 haploinsufficiency as a contributor to ALS pathogenesis. In addition, we identified that the stress factors head trauma or cancer treated by radiochemotherapy, which are discussed as potential additional triggers of ALS pathogenesis via an increased generation of free radicals [13], occurred in carriers of the SUMO4 initiator codon variant prior to ALS onset. Therefore, the combination of this genetic alteration with environmental factors contributing to oxidative stress may cause ALS in these patients.
SGs that form in response to various cellular stress conditions, including heat or oxidative stress, are increasingly associated with neurodegenerative diseases, particularly ALS [1]. Interestingly, the SUMO system may control both the assembly and dissolution of SGs [6]. In line with this notion, our data link SUMO4 haploinsufficiency in an ALS patient to persistent SGs, excessive SG accumulation upon stress, and reduced dynamics after stress challenge. Similarly, altered SG dynamics were shown to be a consequence of ALS-associated mutations in another gene, e.g., in FUS [3, 4]. Moreover, SG clearance by VCP was reduced by ALS-associated VCP mutations [7], which also inhibited the SUMOylation of VCP by SUMO1 [5]. Here, SUMOylation of VCP by SUMO4 was reported and found to be diminished in ALS patient fibroblasts harboring the SUMO4 initiator codon variant, suggesting that this variant may similarly affect SUMOylation of VCP with relevance for SG disassembly, as do ALS-associated VCP mutations.
SUMO4 expression was previously detected in human kidney [15], immune-related tissues [16], and placenta [18]. The fact that the brain showed the highest SUMO4 mRNA levels of all human tissues analyzed in our study establishes that SUMO4 variants may impact the brain, and that SUMO4 haploinsufficiency may cause neurological symptoms.
In summary, we identified the rare initiator codon variant SUMO4:c.2T>C at a significantly higher frequency in a European cohort of ALS patients than in controls. Our findings are based on a limited number of ALS patients and require confirmation in larger cohorts. Here, we introduce the SUMO4:c.2T>C variant leading to SUMO4 haploinsufficiency as a novel potential genetic risk factor for ALS that possibly acts in conjunction with the oxidative stress-related factors trauma and cancer (plus radiochemotherapy), and provide evidence of an impact on SG dynamics and SUMOylation of VCP as similarly shown for mutations in the ALS-linked genes FUS and VCP [3,4,5, 7]. Our data corroborate the notion that the SUMO system may protect from stress insults potentially reducing ALS risk.
Data availability
Anonymized data from the present study can be made available upon reasonable request after contact with the corresponding authors.
References
Wolozin B, Ivanov P (2019) Stress granules and neurodegeneration. Nat Rev Neurosci 20(11):649–666. https://doi.org/10.1038/s41583-019-0222-5
Becker LA, Huang B, Bieri G, Ma R, Knowles DA, Jafar-Nejad P, Messing J, Kim HJ, Soriano A, Auburger G, Pulst SM, Taylor JP, Rigo F, Gitler AD (2017) Therapeutic reduction of ataxin-2 extends lifespan and reduces pathology in TDP-43 mice. Nature 544(7650):367–371. https://doi.org/10.1038/nature22038
Zhang X, Wang F, Hu Y, Chen R, Meng D, Guo L, Lv H, Guan J, Jia Y (2020) In vivo stress granule misprocessing evidenced in a FUS knock-in ALS mouse model. Brain 143(5):1350–1367. https://doi.org/10.1093/brain/awaa076
Bosco DA, Lemay N, Ko HK, Zhou H, Burke C, Kwiatkowski TJ Jr, Sapp P, McKenna-Yasek D, Brown RH Jr, Hayward LJ (2010) Mutant FUS proteins that cause amyotrophic lateral sclerosis incorporate into stress granules. Hum Mol Genet 19(21):4160–4175. https://doi.org/10.1093/hmg/ddq335
Wang T, Xu W, Qin M, Yang Y, Bao P, Shen F, Zhang Z, Xu J (2016) Pathogenic Mutations in the Valosin-containing Protein/p97(VCP) N-domain inhibit the SUMOylation of VCP and lead to impaired stress response. J Biol Chem 291(27):14373–14384. https://doi.org/10.1074/jbc.M116.729343
Keiten-Schmitz J, Roder L, Hornstein E, Muller-McNicoll M, Muller S (2021) SUMO: glue or solvent for phase-separated ribonucleoprotein complexes and molecular condensates? Front Mol Biosci 8:673038. https://doi.org/10.3389/fmolb.2021.673038
Buchan JR, Kolaitis RM, Taylor JP, Parker R (2013) Eukaryotic stress granules are cleared by autophagy and Cdc48/VCP function. Cell 153(7):1461–1474. https://doi.org/10.1016/j.cell.2013.05.037
Hendriks IA, Vertegaal AC (2016) A comprehensive compilation of SUMO proteomics. Nat Rev Mol Cell Biol 17(9):581–595. https://doi.org/10.1038/nrm.2016.81
Marmor-Kollet H, Siany A, Kedersha N, Knafo N, Rivkin N, Danino YM, Moens TG, Olender T, Sheban D, Cohen N, Dadosh T, Addadi Y, Ravid R, Eitan C, Toth Cohen B, Hofmann S, Riggs CL, Advani VM, Higginbottom A, Cooper-Knock J, Hanna JH, Merbl Y, Van Den Bosch L, Anderson P, Ivanov P, Geiger T, Hornstein E (2020) Spatiotemporal proteomic analysis of stress granule disassembly using APEX reveals regulation by SUMOylation and links to ALS pathogenesis. Mol Cell 80(5):876-891.e6. https://doi.org/10.1016/j.molcel.2020.10.032
Oh SM, Liu Z, Okada M, Jang SW, Liu X, Chan CB, Luo H, Ye K (2010) Ebp1 sumoylation, regulated by TLS/FUS E3 ligase, is required for its anti-proliferative activity. Oncogene 29(7):1017–1030. https://doi.org/10.1038/onc.2009.411
Osmanovic A, Widjaja M, Förster A, Weder J, Wattjes MP, Lange I, Sarikidi A, Auber B, Raab P, Christians A, Preller M, Petri S, Weber RG (2020) SPG7 mutations in amyotrophic lateral sclerosis: a genetic link to hereditary spastic paraplegia. J Neurol 267(9):2732–2743. https://doi.org/10.1007/s00415-020-09861-w
Labra J, Menon P, Byth K, Morrison S, Vucic S (2016) Rate of disease progression: a prognostic biomarker in ALS. J Neurol Neurosurg Psychiatry 87(6):628–632. https://doi.org/10.1136/jnnp-2015-310998
Oskarsson B, Horton DK, Mitsumoto H (2015) Potential environmental factors in amyotrophic lateral sclerosis. Neurol Clin 33(4):877–888. https://doi.org/10.1016/j.ncl.2015.07.009
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, Preibisch S, Rueden C, Saalfeld S, Schmid B, Tinevez JY, White DJ, Hartenstein V, Eliceiri K, Tomancak P, Cardona A (2012) Fiji: an open-source platform for biological-image analysis. Nat Methods 9(7):676–682. https://doi.org/10.1038/nmeth.2019
Bohren KM, Nadkarni V, Song JH, Gabbay KH, Owerbach D (2004) A M55V polymorphism in a novel SUMO gene (SUMO-4) differentially activates heat shock transcription factors and is associated with susceptibility to type I diabetes mellitus. J Biol Chem 279(26):27233–27238. https://doi.org/10.1074/jbc.M402273200
Guo D, Li M, Zhang Y, Yang P, Eckenrode S, Hopkins D, Zheng W, Purohit S, Podolsky RH, Muir A, Wang J, Dong Z, Brusko T, Atkinson M, Pozzilli P, Zeidler A, Raffel LJ, Jacob CO, Park Y, Serrano-Rios M, Larrad MT, Zhang Z, Garchon HJ, Bach JF, Rotter JI, She JX, Wang CY (2004) A functional variant of SUMO4, a new I kappa B alpha modifier, is associated with type 1 diabetes. Nat Genet 36(8):837–841. https://doi.org/10.1038/ng1391
Chen S, Xie W, Liu Z, Shan H, Chen M, Song Y, Yu H, Lai L, Li Z (2020) CRISPR start-loss: a novel and practical alternative for gene silencing through base-editing-induced start codon mutations. Mol Ther Nucleic Acids 21:1062–1073. https://doi.org/10.1016/j.omtn.2020.07.037
Baczyk D, Audette MC, Drewlo S, Levytska K, Kingdom JC (2017) SUMO-4: A novel functional candidate in the human placental protein SUMOylation machinery. PLoS One 12(5):e0178056. https://doi.org/10.1371/journal.pone.0178056
Acknowledgements
The authors wish to thank all patients for participating in this study. This work was supported by the Deutsche Forschungsgemeinschaft (PRACTIS–Clinician Scientist Program of Hannover Medical School, grant no. ME3696/3-1 to AO; and grant no. KO5614/2-1 to AC), the Else Kröner-Fresenius-Stiftung (scholarship to MW, grant to SP and RGW within the KlinStrucMed Program of Hannover Medical School), Hannover Medical School (Hochschulinterne Leistungsförderung (HiLF), grant to AO), and the Petermax Müller-Stiftung (grant to SP and RGW).
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
AO, SP, and RGW designed the study. AO, AF, MW, BA, AMD, and FB carried out the experiments. AO, AF, MW, BA, AC, FB, SP, and RGW analyzed and interpreted the data. AO, MW, and SP contributed patient material and clinical data. AO, AF, FB, SP, and RGW wrote the manuscript with contributions from all other authors.
Corresponding authors
Ethics declarations
Conflicts of interest
AO received honoraria from Biogen; SP received honoraria from Biogen, Cytokinetics, Inc., Desitin Pharma, Novartis, Roche, and Teva. The other authors report no conflict of interest relevant to this study.
Ethical approval
This study was approved by the Ethics Board of Hannover Medical School (ID # 6269) and was, therefore, performed in accordance with the ethical standards laid down in the 1964 Declaration of Helsinki and its later amendments.
Informed consent
Written informed consent was obtained from all individuals before entering the study.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Osmanovic, A., Förster, A., Widjaja, M. et al. A SUMO4 initiator codon variant in amyotrophic lateral sclerosis reduces SUMO4 expression and alters stress granule dynamics. J Neurol 269, 4863–4871 (2022). https://doi.org/10.1007/s00415-022-11126-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00415-022-11126-7