Abstract
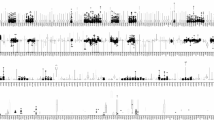
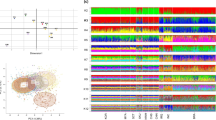
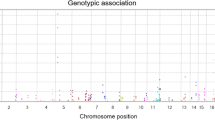
In the present study, 90 autosomal single nucleotide polymorphisms (SNPs) and 34 Y chromosomal SNPs were sequenced simultaneously using HID-Ion AmpliSeq™ Identity Panel on the Ion PGM™ platform for 125 samples in a southern Chinese population. Raw data were analyzed and forensic parameters were calculated. Haplogrouping concordance was also assessed using alternative methods based on Y-SNP haplotypes and Y-STR haplotypes. The results showed that allelic imbalance occurred more frequently with low coverage while several SNPs with high coverage were also observed with poor allelic balance, including rs214955, rs430046, rs7520386, rs876724, rs9171188, rs16981290, and rs2032631. Totally, 21,261 miscalled reads (0.28%) were observed. The rate of allele-specific miscalled reads (ASMRs) was higher than that of allele nonspecific miscalled reads (ANMRs) and associated with genetic diversity of the SNP. The ASMRs of major allele were lower than that of minor allele while there was no difference for ANMRs. The combined discrimination power (CDP) was 1–4.81 × 10−34 and the combined power of exclusion (CPE) was 0.99989 and 0.99999992 for duo and trio paternity testing, respectively. No significant genetic difference was detected between southern and northern Chinese populations. For haplogroup study, O2 was the predominant haplogroup and 97.01% of samples were assigned consistent haplogoups with Y-SNP and Y-STR haplotypes. In conclusion, the AmpliSeq™ Identity Panel was powerful for individual identification and trio paternity testing. ASMRs were associated with the genetic diversity and allele frequency while neither was related for ANMRs. High concordance of haplogrouping assignment can be obtained with Y-STR and Y-SNP haplotypes.





Similar content being viewed by others
References
Kidd KK, Pakstis AJ, Speed WC, Grigorenko EL, Kajuna SLB, Karoma NJ, Kungulilo S, Kim J, Lu R, Odunsi A, Okonofua F, Parnas J, Schulz LO, Zhukova OV, Kidd JR (2006) Developing a SNP panel for forensic identification of individuals. Forensic Sci Int 164:20–32
Amorim A, Pereira L (2005) Pros and cons in the use of SNPs in forensic kinship investigation: a comparative analysis with STRs. Forensic Sci Int 150:17–21
Mo S, Liu Y, Wang S, Bo X, Li Z, Chen Y, Ni M (2016) Exploring the efficacy of paternity and kinship testing based on single nucleotide polymorphisms. Forensic Sci Int Genet 22:161–168
Sanchez JJ, Phillips C, Borsting C, Balogh K, Bogus M, Fondevila M, Harrison CD, Musgrave-Brown E, Salas A, Syndercombe-Court D, Schneider PM, Carracedo A, Morling N (2006) A multiplex assay with 52 single nucleotide polymorphisms for human identification. Electrophoresis 27:1713–1724
Pakstis AJ, Speed WC, Fang R, Hyland FCL, Furtado MR, Kidd JR, Kidd KK (2010) SNPs for a universal individual identification panel. Hum Genet 127:315–324
Sobrino B, Brión M, Carracedo A (2005) SNPs in forensic genetics: a review on SNP typing methodologies. Forensic Sci Int 154:181–194
Seo SB, King JL, Warshauer DH, Davis CP, Ge J, Budowle B (2013) Single nucleotide polymorphism typing with massively parallel sequencing for human identification. Int J Legal Med 127:1079–1086
Guo F, Zhou Y, Song H, Zhao J, Shen H, Zhao B, Liu F, Jiang X (2016) Next generation sequencing of SNPs using the HID-Ion AmpliSeq™ Identity Panel on the Ion Torrent PGM™ platform. Forensic Sci Int Genet 25:73–84
Børsting C, Fordyce SL, Olofsson J, Mogensen HS, Morling N (2014) Evaluation of the Ion Torrent™ HID SNP 169-plex: a SNP typing assay developed for human identification by second generation sequencing. Forensic Sci Int Genet 12:144–154
Loman NJ, Misra RV, Dallman TJ, Constantinidou C, Gharbia SE, Wain J, Pallen MJ (2012) Performance comparison of benchtop high-throughput sequencing platforms. Nat Biotechnol 30:434–439
Eduardoff M, Santos C, de la Puente M, Gross TE, Fondevila M, Strobl C, Sobrino B, Ballard D, Schneider PM, Carracedo Á, Lareu MV, Parson W, Phillips C (2015) Inter-laboratory evaluation of SNP-based forensic identification by massively parallel sequencing using the Ion PGM™. Forensic Sci Int Genet 17:110–121
Rothberg JM, Hinz W, Rearick TM, Schultz J, Mileski W, Davey M, Leamon JH, Johnson K, Milgrew MJ, Edwards M, Hoon J, Simons JF, Marran D, Myers JW, Davidson JF, Branting A, Nobile JR, Puc BP, Light D, Clark TA, Huber M, Branciforte JT, Stoner IB, Cawley SE, Lyons M, Fu Y, Homer N, Sedova M, Miao X, Reed B, Sabina J, Feierstein E, Schorn M, Alanjary M, Dimalanta E, Dressman D, Kasinskas R, Sokolsky T, Fidanza JA, Namsaraev E, McKernan KJ, Williams A, Roth GT, Bustillo J (2011) An integrated semiconductor device enabling non-optical genome sequencing. Nature 475:348–352
Elena S, Alessandro A, Ignazio C, Sharon W, Luigi R, Andrea B (2016) Revealing the challenges of low template DNA analysis with the prototype Ion AmpliSeq™ Identity panel v2.3 on the PGM™ Sequencer. Forensic Sci Int Genet 22:25–36
Karafet TM, Mendez FL, Meilerman MB, Underhill PA, Zegura SL, Hammer MF (2008) New binary polymorphisms reshape and increase resolution of the human Y chromosomal haplogroup tree. Genome Res 18:830–838
Zhang S, Bian Y, Zhang Z, Zheng H, Wang Z, Zha L, Cai J, Gao Y, Ji C, Hou Y, Li C (2015) Parallel analysis of 124 universal SNPs for human identification by targeted semiconductor sequencing. Sci Rep-UK 5:18683
Athey TW (2006) Haplogroup prediction from Y-STR values using a Bayesian-allele-frequency approach. J Genet Geneal 2:34–39
Athey TW (2005) Haplogroup prediction from Y-STR values using an allele-frequency approach. J Genet Geneal 1:1–7
Kalinowski ST, Taper ML, Marshall TC (2007) Revising how the computer program CERVUS accommodates genotyping error increases success in paternity assignment. Mol Ecol 16:1099–1106
Excoffier L, Lischer HE (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Amigo J, Salas A, Phillips C, Carracedo A (2008) SPSmart: adapting population based SNP genotype databases for fast and comprehensive web access. BMC Bioinformatics 9:428
International Society of Genetic Genealogy (ISOGG): Y-DNA Haplogroup Tree 2017, Version: 12.128. In, 2017. Available at: http://www.isogg.org/tree/
Wright S (1978) Evolution and the genetics of populations. Vol. 4. Variability within and among natural populations. University of Chicago Press, Chicago
Phillips C, García-Magariños M, Salas A, Carracedo Á, Lareu MV (2012) SNPs as supplements in simple kinship analysis or as core markers in distant pairwise relationship tests: when do SNPs add value or replace well-established and powerful STR tests? Transfus Med Hemoth 39:202–210
Wang Y, Liu C, Zhang CC, Li R, Li Y, XL O, Sun HY (2015) Analysis of 17 Y-STR loci haplotype and Y-chromosome haplogroup distribution in five Chinese ethnic groups. Electrophoresis 36:2546–2552
Muzzio M, Ramallo V, Motti JMB, Santos MR, López Camelo JS, Bailliet G (2011) Software for Y-haplogroup predictions: a word of caution. Int J Legal Med 125:143–147
Petrejcikova E, Carnogurska J, Hronska D, Bernasovska J, Boronova I, Gabrikova D, Bozikova A, Macekova S (2014) Y-SNP analysis versus Y-haplogroup predictor in the Slovak population. Anthropol Anz 71:275–285
Dogan S, Babic N, Gurkan C, Goksu A, Marjanovic D, Hadziavdic V (2016) Y-chromosomal haplogroup distribution in the Tuzla Canton of Bosnia and Herzegovina: a concordance study using four different in silico assignment algorithms based on Y-STR data. Homo 67:471–483
Nunez C, Geppert M, Baeta M, Roewer L, Martinez-Jarreta B (2012) Y chromosome haplogroup diversity in a Mestizo population of Nicaragua. Forensic Sci Int Genet 6:e192–e195
Athey W (2011) Comments on the article, “Software for Y haplogroup predictions, a word of caution”. Int J Legal Med 125(901–903):905–906
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Rights and permissions
About this article
Cite this article
Li, R., Zhang, C., Li, H. et al. SNP typing using the HID-Ion AmpliSeq™ Identity Panel in a southern Chinese population. Int J Legal Med 132, 997–1006 (2018). https://doi.org/10.1007/s00414-017-1706-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-017-1706-3