Abstract
Background
Forensic DNA analysis has seen remarkable advancements with the advent of Next Generation Sequencing (NGS). In particular, NGS analysis of single nucleotide polymorphisms (SNPs) offers significant advantages in the analysis of challenging samples compared to conventional STR analysis.
Objective
This study aimed to investigate the SNPs of the Precision ID Identity Panel, a commercially available NGS panel for personal identification, by generating genetic profiles of 298 Koreans and comparing them with other global populations.
Methods
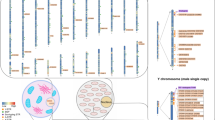
A total of 124 SNPs, including 90 autosomal and 34 Y-SNPs, were analyzed using the Precision ID Identity Panel, and forensic parameters, microhaplotypes, and population differences were investigated.
Results
The NGS data were successfully obtained from 298 Koreans. The analysis of forensic parameters exhibited a low combined match probability of 1.532 × 10− 34, which is comparable to that obtained from commonly used STR analysis. Additionally, the microhaplotype analysis revealed that the use of 16 microhaplotypes provided higher discriminatory power compared to single target SNPs. Furthermore, the adoption of microhaplotype data resulted in an increase of over 20% in expected heterozygosity at five loci. Inter-population analysis showed a close genetic relationship between Koreans and individuals from China and Myanmar in East and Southeast Asia, which are geographically adjacent to Korea.
Conclusions
The results of this study show that the Precision ID Identity panel can be a useful alternative where traditional STR typing is not feasible. Also, the data from our study will be useful as a reference for Koreans in forensic investigations and the prosecution of criminal justice.


Similar content being viewed by others
Availability of data and materials
All data and materials described in the manuscript are available.
References
Avent I, Kinnane AG, Jones N, Petermann I, Daniel R, Gahan ME, McNevin D (2019) The QIAGEN 140-locus single-nucleotide polymorphism (SNP) panel for forensic identification using massively parallel sequencing (MPS): an evaluation and a direct-to-PCR trial. Int J Legal Med 133:677–688
Avila E, Felkl AB, Graebin P, Nunes CP, Alho CS (2019) Forensic characterization of brazilian regional populations through massive parallel sequencing of 124 SNPs included in HID ion Ampliseq Identity Panel. Forensic Sci Int Genet 40:74–84
Borsting C, Fordyce SL, Olofsson J, Mogensen HS, Morling N (2014) Evaluation of the Ion Torrent HID SNP 169-plex: a SNP typing assay developed for human identification by second generation sequencing. Forensic Sci Int Genet 12:144–154
Buchard A, Kampmann ML, Poulsen L, Borsting C, Morling N (2016) ISO 17025 validation of a next-generation sequencing assay for relationship testing. Electrophoresis 37:2822–2831
Budowle B, van Daal A (2008) Forensically relevant SNP classes. Biotechniques 44:603–608
Dash HR, Avila E, Jena SR, Kaitholia K, Agarwal R, Alho CS, Srivastava A, Singh AK (2022) Forensic characterization of 124 SNPs in the central indian population using precision ID identity panel through next-generation sequencing. Int J Legal Med 136:465–473
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Guo F, Zhou Y, Song H, Zhao J, Shen H, Zhao B, Liu F, Jiang X (2016) Next generation sequencing of SNPs using the HID-Ion AmpliSeq Identity Panel on the Ion Torrent PGM platform. Forensic Sci Int Genet 25:73–84
Joo SM, Kwon Y-L, Moon MH, Shin K-J (2022) Investigation of SNPs in the Precision ID Identity Panel using next-generation sequencing in a Myanmar population. Research Square
Karafet TM, Mendez FL, Meilerman MB, Underhill PA, Zegura SL, Hammer MF (2008) New binary polymorphisms reshape and increase resolution of the human Y chromosomal haplogroup tree. Genome Res 18:830–838
Kidd KK, Pakstis AJ (2022) State of the art for Microhaplotypes. Genes (Basel, p 13
Kidd KK, Speed WC (2015) Criteria for selecting microhaplotypes: mixture detection and deconvolution. Investig Genet 6:1
Kidd KK, Pakstis AJ, Speed WC, Lagace R, Chang J, Wootton S, Haigh E, Kidd JR (2014) Current sequencing technology makes microhaplotypes a powerful new type of genetic marker for forensics. Forensic Sci Int Genet 12:215–224
Kim SH, Han MS, Kim W, Kim W (2010) Y chromosome homogeneity in the korean population. Int J Legal Med 124:653–657
Koch E, Ristroph M, Kirkpatrick M (2013) Long range linkage disequilibrium across the human genome. PLoS ONE 8:e80754
Li R, Zhang C, Li H, Wu R, Li H, Tang Z, Zhen C, Ge J, Peng D, Wang Y et al (2018) SNP typing using the HID-Ion AmpliSeq Identity Panel in a southern chinese population. Int J Legal Med 132:997–1006
Liu J, Wang Z, He G, Zhao X, Wang M, Luo T, Li C, Hou Y (2018) Massively parallel sequencing of 124 SNPs included in the precision ID identity panel in three east asian minority ethnicities. Forensic Sci Int Genet 35:141–148
Mehta B, Daniel R, Phillips C, Doyle S, Elvidge G, McNevin D (2016) Massively parallel sequencing of customised forensically informative SNP panels on the MiSeq. Electrophoresis 37:2832–2840
Pakstis AJ, Speed WC, Fang R, Hyland FC, Furtado MR, Kidd JR, Kidd KK (2010) SNPs for a universal individual identification panel. Hum Genet 127:315–324
Pang JB, Rao M, Chen QF, Ji AQ, Zhang C, Kang KL, Wu H, Ye J, Nie SJ, Wang L (2020) A 124-plex Microhaplotype Panel based on next-generation sequencing developed for forensic applications. Sci Rep 10:1945
Park JH, Hong SB, Kim JY, Chong Y, Han S, Jeon CH, Ahn HJ (2013) Genetic variation of 23 autosomal STR loci in korean population. Forensic Sci Int Genet 7:e76–77
Phillips C, Fang R, Ballard D, Fondevila M, Harrison C, Hyland F, Musgrave-Brown E, Proff C, Ramos-Luis E, Sobrino B et al (2007) Evaluation of the Genplex SNP typing system and a 49plex forensic marker panel. Forensic Sci Int Genet 1:180–185
Phillips C, Manzo L, de la Puente M, Fondevila M, Lareu MV (2020) The MASTiFF panel-a versatile multiple-allele SNP test for forensics. Int J Legal Med 134:441–450
Shi H, Zhong H, Peng Y, Dong YL, Qi XB, Zhang F, Liu LF, Tan SJ, Ma RZ, Xiao CJ et al (2008) Y chromosome evidence of earliest modern human settlement in East Asia and multiple origins of tibetan and japanese populations. BMC Biol 6:45
Standage DS, Mitchell RN (2020) MicroHapDB: a portable and extensible database of all published microhaplotype marker and frequency data. Front Genet 11:781
Sun L, Fu L, Liu Q, Zhou J, Ma C, Cong B, Li S (2019) Population data using Precision ID Identity Panel in a chinese Han population from Hebei Province. Forensic Sci Int Genet 42:e27–e29
Tillmar A, Sturk-Andreaggi K, Daniels-Higginbotham J, Thomas JT, Marshall C (2021) The FORCE Panel: An All-in-One SNP Marker Set for Confirming Investigative Genetic Genealogy Leads and for General Forensic Applications. Genes (Basel) 12
van der Heijden S, de Oliveira SJ, Kampmann ML, Borsting C, Morling N (2017) Comparison of manual and automated AmpliSeq workflows in the typing of a somali population with the Precision ID Identity Panel. Forensic Sci Int Genet 31:118–125
Wang CC, Li H (2013) Inferring human history in East Asia from Y chromosomes. Investig Genet 4:11
Wendt FR, Novroski NM (2019) Identity informative SNP associations in the UK Biobank. Forensic Sci Int Genet 42:45–48
Zhang S, Bian Y, Zhang Z, Zheng H, Wang Z, Zha L, Cai J, Gao Y, Ji C, Hou Y et al (2015) Parallel analysis of 124 Universal SNPs for human identification by targeted Semiconductor sequencing. Sci Rep 5:18683
Zhang S, Bian Y, Chen A, Zheng H, Gao Y, Hou Y, Li C (2017) Developmental validation of a custom panel including 273 SNPs for forensic application using Ion Torrent PGM. Forensic Sci Int Genet 27:50–57
Acknowledgements
This work was supported by the grant from the National Research Foundation of Korea (NRF-2019R1A2C1086423). This study was conducted with bioresources from National Biobank of Korea, the Centers for Disease Control and Prevention, Republic of Korea (KBN-2020-009).
Funding
This work was supported by the grant from the National Research Foundation of Korea (NRF-2019R1A2C1086423).
Author information
Authors and Affiliations
Contributions
Soo-Bin Yang: conceptualization, methodology, validation, investigation, visualization and writing of original draft; Ji Eun Lee: methodology, partial investigation and resources; Hwan Young Lee: conceptualization, project administration, supervision, reviewing and editing, and funding acquisition. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Ethics approval
This study had been approved by the Seoul National University Hospital IRB (No. 1912-053-1087). All human samples used in the experiments were conducted in compliance with the guidelines approved by the Committee of Seoul National University Hospital.
Conflict of interest
All authors declare no conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, SB., Lee, J.E. & Lee, H.Y. Forensic genetic analysis of single-nucleotide polymorphisms and microhaplotypes in Koreans through next-generation sequencing using precision ID identity panel. Genes Genom 45, 1281–1293 (2023). https://doi.org/10.1007/s13258-023-01424-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-023-01424-3