Abstract
In many organisms, telomere DNA consists of simple sequence repeat tracts that are required to protect the chromosome end. In the yeast Saccharomyces cerevisiae, tract maintenance requires two checkpoint kinases of the ATM family, Tel1p and Mec1p. Previous work has shown that Tel1p is recruited to functional telomeres with shorter repeat tracts to promote telomerase-mediated repeat addition, but the role of Mec1p is unknown. We found that Mec1p telomere association was detected as cells senesced when telomere function was compromised by extreme shortening due to either the loss of telomerase or the double-strand break binding protein Ku. Exonuclease I effects the removal of the 5' telomeric strand, and eliminating it prevented both senescence and Mec1p telomere association. Thus, in contrast to Tel1p, Mec1p associates with short, functionally compromised telomeres.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In many eukaryotes, telomeres consist of specialized proteins bound to repeated DNA sequences (TTAGGG in vertebrates, TG1–3 in yeast), and the length of these repeat tracts is commonly referred to as telomere length. Telomeres are dynamic structures whose lengths are shortened due to nuclease action or incomplete replication of the chromosome end and are normally elongated by telomerase. In cells that lack telomerase, such as human somatic cells or various yeast mutants, telomere repeat tracts gradually shorten until cells stop growing. As telomeres shorten, the reduction in the number of telomere repeat binding proteins is associated with telomere dysfunction, i.e., the loss of capping functions that distinguish telomeres from double-strand breaks (DSBs), which results in the activation of DNA damage checkpoints and senescence in both yeast and mammalian cells (d'Adda di Fagagna et al. 2003; Herbig et al. 2004; Livengood et al. 2002; Smogorzewska and de Lange 2004).
Several lines of genetic evidence indicate that two paralogous DNA damage checkpoint kinases of the ATM family, yeast Tel1p and Mec1p, are trans-acting factors that regulate telomere length by controlling telomerase access to the 3' end of the chromosome (reviewed in (Bianchi and Shore 2008)). Cells lacking Tel1p (the ortholog of human ATM) reduce their telomere lengths to about 30% of wild-type lengths, while cells lacking Mec1p (the ortholog of human ATR) show a smaller reduction to 85% of wild-type lengths (Lustig and Petes 1986; Ritchie et al. 1999). Simultaneous loss of both Tel1p and Mec1p (tel1 mec1 cells) causes cells to behave as if they have no telomerase, even though active telomerase can be isolated from these cells (Chan et al. 2001; Ritchie et al. 1999). Thus, both Tel1p and Mec1p play a role in maintaining telomere repeat tracts.
More is known about the function of Tel1p at telomeres than Mec1p. Tel1p appears to be required for normal telomere elongation as telomeres in cells lacking Tel1p undergo drastic shortening before being elongated (Ray and Runge 2001) and recruit reduced levels of telomerase components to the telomere (Goudsouzian et al. 2006). Tel1p preferentially associates with short telomeres, and this association increases telomerase recruitment and telomere elongation (Bianchi and Shore 2007; Hector et al. 2007; Sabourin et al. 2007). Importantly, Tel1p associates with telomeres whose tract lengths can support chromosomal end replication and maintenance (Hector et al. 2007; Sabourin et al. 2007; Zakian 1996), indicating that Tel1p associates with functional telomeres. Mutant Tel1p proteins that lack kinase activity also associate with telomeres but do not induce normal telomere elongation, indicating that Tel1p kinase activity plays a role in telomere elongation (Hector et al. 2007; Ma and Greider 2009). These data suggest that Tel1p associates with telomeres to phosphorylate telomere-associated proteins, facilitating telomere elongation and maintaining telomere length within its normal range. However, the Telomerase- phenotype of the tel1 mec1 double mutant cells indicates that the Mec1p kinase also plays a role in telomere function. The role of Mec1p is unclear as its association with telomeres is not seen under conditions where Tel1p telomere association is clearly detectable (McGee et al. 2010).
Like their human orthologs ATM and ATR, Tel1p and Mec1p also have important roles in sensing and halting the cell cycle in response to DSBs (Cimprich and Cortez 2008; Lavin 2007; Morrow et al. 1995; Usui et al. 2001), and both proteins associate with DSBs (Lisby et al. 2004; Nakada et al. 2003). Telomere dysfunction also signals cell cycle arrest through Mec1p as yeast lacking telomerase that acquire drastically shortened telomeres are arrested through a Mec1p-dependent pathway (Enomoto et al. 2002; IJpma and Greider 2003). These short telomeres can undergo recombination events similar to DSBs to generate “survivors” that maintain a functional telomere through a telomerase-independent pathway (reviewed in (McEachern and Haber 2006; Runge 2006)). Thus, Mec1p activity can initiate a cell cycle checkpoint delay and recombination pathways both at DSBs and extremely shortened telomeres.
The structure of the extremely short telomeric DNA that activates the Mec1p-dependent checkpoint is not well understood. In the case of DSBs, Mec1p associates only after exonucleolytic resection of the 5' DNA strand of a DSB (Nakada et al. 2004; Tsubouchi and Ogawa 2000). Whether a similar exonucleolytic activity is required at telomeres to promote Mec1p telomere association is unknown. Telomeric Mec1p association has only been detected under conditions where the 5' strand of the telomere has undergone extensive resection (Rouse and Jackson 2002b), and whether Mec1p associates with the non-TG1–3 DNA or the single-stranded TG1–3 DNA is not known. Mec1p association with an artificial telomere, consisting of a selectable marker followed by a short tract of telomere repeats, has been detected, but whether Mec1p associated with telomeres lacking or retaining TG1–3 repeats was not addressed (Abdallah et al. 2009; Khadaroo et al. 2009). In addition, Mec1p association with natural chromosome ends may be different as artificial telomeres do not always behave the same as natural telomeres in response to mutations that alter tract length (Arneric and Lingner 2007; Berthiau et al. 2006; Craven and Petes 1999; Ray and Runge 1999a; Sabourin et al. 2007). Thus, the DNA structure that recruits Mec1p to natural telomeres and whether these telomeres are functionally compromised or have been converted to DSBs that lack TG1–3 repeats are important open questions.
In this study, we investigated the physical association of Mec1p and Tel1p in yeast strains where telomeres progressively shorten until cells senesced. We found that Mec1p associated with natural telomeres when cells had senesced due to telomere dysfunction, that these Mec1p-associated telomeres retained short TG1–3 repeat tracts, and that preventing telomere dysfunction prevented detectable Mec1p telomere association and senescence. Thus, Mec1p associates with telomeres when their function is compromised.
Materials and methods
Yeast strains
All experiments were performed in the W303 strain background, using the original MEC1-3HA and TEL1-3HA strains of Mallory and Petes that produce functional versions of Mec1p and Tel1p, each bearing three tandem influenza hemaglutinin (HA) epitope tags (Mallory and Petes 2000). Gene deletions in these strains were made by the method of Baudin et al. (1993), using PCR to construct selectable markers flanked by 40 bp of sequence homologous to the yeast genome to disrupt the indicated ORF. All deletions were subsequently confirmed by PCR analysis of the novel sequence junctions created by the appropriate gene disruption, as previously described (Hector et al. 2007). A complete list of yeast strains (Table S1) and the oligonucleotides (Table S2) used are presented in the on-line supplemental material.
Generation of est2Δ and yku70Δ strains with dysfunctional telomeres
To propagate the est2Δ strain, the cells were transformed with a yeast centromere plasmid bearing the EST2 and URA3 genes (pVL291, provided by V. Lundblad) and then grown on medium lacking uracil to select for the plasmid. The est2Δ/pVL291 cells were plated onto non-selective medium (called the stock plate) to allow colony formation and then stored at 4°C. Portions of individual colonies from the stock plate were tested for the presence or absence of plasmid: Cells from each colony were spotted onto plates containing non-selective medium or selective medium, and a colony whose cells could not grow on selective medium was chosen. The remainder of this colony was taken from the stock plate and used to seed a 500-ml culture of non-selective medium (YEPD (Rose et al. 1990)) that was grown for different numbers of population doublings. At each time point, 16 ml was taken to start the next 500-ml culture for the next time point, 50 ml was taken to prepare genomic DNA (as described in (Ray and Runge 1999b)) for telomere length analysis, and the remaining cells were used for ChIP (described in the following discussion).
To generate yku70Δ cells with dysfunctional telomeres, cells were grown at the permissive temperature (25°C) to mid-log phase (~5 × 106 cells/ml) and then diluted into fresh YEPD medium pre-warmed to 37°C, followed by growth at 37°C for different times, as indicated in the “Results” section. The fold dilutions for each culture were chosen to allow cells to reach mid-log phase by each time point and were determined empirically from pilot experiments. The same approach was used for yku70Δ exo1Δ cells.
Chromatin immunoprecipitations
Samples were prepared for ChIP according to published methods (Strahl-Bolsinger et al. 1997). Cells were grown to an OD660 of 0.5 to 0.8 and cross-linked by adding formaldehyde to a final concentration of 1%, swirling the culture at room temperature for 15 min, followed by the addition of glycine to a final concentration of 125 mM to quench formaldehyde cross-linking. After a 5-min incubation at room temperature, the cells were pelleted, resuspended in Tris buffered saline (TBS, 50-mM Tris-HCl pH 7.5, 150-mM NaCl) cooled to 4°C and transferred to a 1.5-ml tube. Cells were pelleted, washed once again in TBS, once with FA buffer (50-mM HEPES, pH 7.5, 150-mM NaCl, 1-mM EDTA, 0.1% sodium deoxycholate), and then resuspended in 500 μl of cold FA lysis buffer (FA buffer also containing 0.1% SDS and 1% Triton X-100) and stored at −80°C. Cells were thawed and then lysed by filling the tube with acid-washed glass beads and shaking for 20 min at 4°C in a Vortex shaker. Both the bottom and the top of the 1.5-ml tube were punctured with a 22 gauge needle; the tube was placed into a 15-ml conical tube and then spun at 1,000 × g for 2 min to separate the lysate from the glass beads. The lysate was transferred to a new 1.5-ml tube and sonicated with a microtip probe at an output of 4.5 W for 10 s, which sheared the DNA to a modal size of 500 bp with a range of 100 to 5,000 bp. Cell debris was removed by centrifugation at 15,000 × g for 20 min at 4°C, and the clarified lysate was transferred to a new tube. The final volume was adjusted to 2.5 ml with FA lysis buffer, and aliquots of 500 μl of this input chromatin were stored at −80°C, with one aliquot used for each chromatin IP.
Immunoprecipitations were performed by incubating the input chromatin with or without 2 μg of antibody to HA (HA-probe (F7)) and 25-μl of protein G-sepharose (both from Santa Cruz Biotechnology) overnight at 4°C. The resin was washed by centrifugation at 1,800 × g for 1 min at 4°C for each wash as follows: two washes with 1.4 ml of FA-lysis buffer, followed by two washes with 1.4 ml of FA-lysis buffer adjusted to 500-mM NaCl, two washes with 1.4 ml of LiCl wash buffer (10-mM Tris-HCl pH 8.0, 0.25-M LiCl, 0.5% NP-40, 0.5% sodium deoxycholate, 1 mM EDTA), and two washes with 1.4 ml of FA buffer. The immunoprecipitated chromatin was released from the beads by heating for 6 h at 65°C, which also reversed the formaldehyde cross-links. DNA was purified as described (Strahl-Bolsinger et al. 1997) and resuspended in 100 μl of 10-mM Tris, 1-mM EDTA pH 7.0.
To determine the level of telomere enrichment, a standard curve was performed using PCR on DNA isolated from the input chromatin (called the input DNA). A 1:3,000 dilution of input DNA and 4 additional 3-fold serial dilutions were used as template in PCR reactions to determine a linear range of amplification. PCR analysis of ChIP samples used 1:50 and 1:250 dilutions of the ChIP DNA to obtain band intensities within the linear range of the input DNA dilutions. Association with the VI-R telomere was measured using the VI-R70S + VI-R70AS primers (Table S2) to produce a 269-bp fragment. The VI-R association was compared to two control loci from the central regions of two different chromosomes, ARO1 and GAL10, using the primers ARO1S + ARO1AS and GAL10S + GAL10AS (Table S2) to yield products of 541 and 420 bp, respectively. Primer concentrations were adjusted to yield the same amount of each amplified product using the input DNA as template, and these concentrations were used for all subsequent PCR analyses. Association with the XV-L telomere was also determined in some experiments using the primers XV-L-26AS + XV-L-383S (Table S2) to produce a 372-bp fragment that was co-amplified with the ARO1 primers. The amplified DNA fragments were separated on 1.4% agarose gels, and the gels were stained with Vistra Green (GE Healthcare) and scanned on a Storm 840 Imager (GE Healthcare). The relative levels of each PCR product were determined using ImageQuant software (version 5.2, GE Healthcare), and telomere enrichment was calculated as the ratio of the yield of the normalized VI-R telomere product over the ratio of the normalized internal control locus product using the formula
where immunoprecipitations were performed with antibody (+Ab) or without antibody (-Ab) prior to decross-linking and PCR. Each ChIP PCR was performed and quantitated at least twice. Normalizations using either the ARO1 or GAL4 control locus gave very similar values for enrichments, and only ARO1-based normalizations are reported.
Telomere PCR
The sequence of the Mec1-HAp-associated DNA was determined using a modified version of the telomere PCR protocol (Förstemann et al. 2000). Genomic DNA (100–200 ng, usually 1.0 μl, prepared as described (Ray and Runge 1999b)) or ChIP DNA (4.0 μl, prepared as described previously) was suspended in a total of 10 μl of One-phor-all buffer (20-mM Tris-HCl pH 7.8, 50-mM potassium acetate, 10-mM magnesium acetate) and incubated at 96°C for 10 min and then shifted to 4°C for 20 min. Tailing was performed by adjusting the reaction to 1-mM dCTP and adding 1 unit of terminal deoxynucleotide transferase (New England Biolabs), incubating for 30 min at 37°C, followed by an incubation at 65°C for 10 min to stop the reaction. The poly-dC-tailed DNA was then PCR amplified using Taq polymerase with a primer specific for the subtelomeric region of chromosome VI-R (TELVIR-88, Table S2) and a primer complimentary to the poly-dC-tail (dG18-BamHI, Table S2). Samples were denatured for 3 min at 94°C, then cycled 45 times with 30 s of denaturation (94°C), 15 s of annealing (63°C), and 30 s of extension (72°C), followed by a final extension step of 5 min at 72°C. The PCR products were cloned into the plasmid pCR2.1-Topo (Invitrogen) and sequenced.
Telomere length analysis
For analysis of the VI-R telomere length, ~10 μg of yeast genomic DNA was digested with 20 units of EcoR V (New England Biolabs) and analyzed by Southern blotting using a 32P-labeled VI-R PCR product made from primers VI-R70S + VI-R70AS (Table S2) as probe. Pre-hybridization and hybridization using PerfectHyb (Sigma), as well as washing of the blots, were performed according to the instructions of the manufacturer. Fragment sizes were determined using Fragment NT software, and mean TG1–3 lengths were determined by subtracting the known lengths of the non-telomeric DNA sequences from the size of the most intense part of the band.
Analysis of Mec1-HAp association in synchronized cells
Alpha factor was used to synchronize cells in G1 following the methods described in Hector et al. (2007), based on previous methods (Spellman et al. 1998). Briefly, 1.8 l of MEC1-HA bar1Δ or MEC1-HA yku70Δ bar1Δ cells was grown in YEPD pH 5.5 at 20°C to an OD660 of 0.2. Alpha factor peptide (Sigma) was then added to 12 ng/ml, and the cells were grown for an additional 3 h. Aliquots of cells were removed to measure Mec1-HAp association by ChIP (200 ml) and DNA content by FACS (15 ml). Cells were then released into the cell cycle by pelleting the cells, washing them once with 200 ml of 20°C YEPD medium and resuspending the washed pellets in 1.6 l of 20°C YEPD pH 5.5. Aliquots were taken at the time points indicated in the “Results” section and processed as previously discussed. Time points were chosen based on a pilot experiment with a 50-ml culture and performing FACS analysis on aliquots taken every 5 min. Cells were either followed until all cells were 2 N, consistent with G2/M phase, or were allowed to enter the next G1 phase to analyze Mec1-HAp telomere association during this transition. Processing for ChIP followed the protocol described previously. FACS analysis was performed by immediately fixing the culture aliquot by the addition of 100% EtOH to a final concentration of 70% and then processing cells for propidium iodide staining at the Lerner Research Institute Cell Sorting Core Facility.
Results
Mec1-HAp associates with short telomeres as cells escape from senescence
The telomerase- phenotype of the tel1Δ mec1Δ double mutant (Ritchie et al. 1999) suggests a role for Mec1p in the maintenance of telomere repeat tracts, possibly by directly associating with the telomere as occurs with Tel1p (Bianchi and Shore 2007; Hector et al. 2007; Sabourin et al. 2007). Given that cells lacking telomerase eventually shorten their telomeres to the point when they activate a Mec1p-dependent cell cycle arrest (Enomoto et al. 2002; IJpma and Greider 2003), the possible association of Mec1p with the telomere was examined in cells whose telomeres shortened until the cells entered crisis. These experiments used an epitope-tagged Mec1p protein (Mec1-HAp) that was previously shown to retain DNA damage checkpoint activity in vivo and kinase activity in vitro with no obvious effect on telomere length (Mallory and Petes 2000). We verified that Mec1-HAp protein also associated with an induced DSB in ChIP experiments (Fig. S1), as shown for other tagged Mec1 variants (Lisby et al. 2004; Nakada et al. 2005; Rouse and Jackson 2002b; Zhang and Durocher 2010). Thus, the HA-tagged protein retained the functions of normal Mec1p.
Mec1-HAp association with telomeres of different lengths was first assayed in cultures of cells in which telomeres were steadily shortening. Telomerase was inactivated in Mec1-HAp cells by inducing the loss of the gene encoding the telomerase catalytic subunit, Est2p (“Materials and methods”). The Telomerase- cells were then kept in a state of continuous growth, which allowed telomeres to gradually shorten over many population doublings. Samples were taken over the course of 70 population doublings to monitor cell growth rate, the length of a unique chromosomal telomere, and Mec1-HAp association with this telomere.
During the course of the experiment, the cells followed the well-established phenotype of cells lacking telomerase (Lundblad and Blackburn 1993). As the population doublings increased, telomeres shortened, the culture took longer to double, and rearranged telomeres were detected at a point coincident with a decrease in population doubling time. Thus, as the cells lacking telomerase grew from population doubling 0 to 60 (PD 0 to PD 60), the size of the restriction fragment for the unique telomere on the right arm of chromosome VI (VI-R) gradually decreased (Fig. 1a), and population doubling time increased to a maximum at PD 60 (Fig. 1d). At PD 70, new telomeric restriction fragments appeared (Fig. 1a), and the population doubling time decreased (Fig. 1d), indicating the appearance of telomerase-independent survivor cells between PD 60 and PD 70. An assessment of several other chromosomal telomeres that bear the middle-repetitive element Y' adjacent to the terminal TG1–3 tracts confirmed that cells in PD 70 contained recombined telomeres normally associated with the survivor pathway (Fig. S2). The combined data of the culture growth rates (Fig. 1d), telomere shortening, and appearance of recombined telomeres (Figs. 1a and S 2) indicate that some of the cells in the PD 70 culture were escaping the senescence caused by extreme telomere shortening through the activation of recombination pathways that allow telomeres to be elongated in the absence of telomerase (Chen et al. 2001; Le et al. 1999; Lundblad and Blackburn 1993; McEachern and Haber 2006; Teng et al. 2000; Teng and Zakian 1999).
Mec1-HAp associates with telomeres as cells escape the cell cycle arrest that occurs due to telomere shortening. a Telomere shortening in Mec1-HAp cells lacking the telomerase catalytic subunit Est2p. Cells lacking telomerase were generated by deleting the genomic copy of EST2, covering the deletion with a plasmid bearing the EST2 gene, and then isolating cells that had lost the plasmid. Cells lacking telomerase were then cultured for the number of population doublings (PD) shown, where the 0 PD sample contains cells that retain the EST2 plasmid and the other PD samples contain cells that lack telomerase. Genomic DNA from each PD was digested with EcoR V and analyzed by Southern blotting using the VI-R PCR product from the primers shown in panel b as probe. Telomeres gradually shortened with increasing PD, allowing the isolation of cells with different telomere lengths. The black arrow designates a background band that hybridizes to probe. The white arrowheads designate rearranged telomeres in PD 70 that are characteristic of the survivor cells that escape the short telomere cell cycle arrest. b A schematic of the loci and primers used to monitor telomere association. The telomeres from the right arm of chromosome VI (VI-R) and two internal loci from separate chromosomes (GAL10 and ARO1) were PCR amplified in the ChIP experiments in this study. c Analysis of the Mec1-HAp immunoprecipitates using the PCR primers indicated in panel b. Telomere enrichment values are calculated as the ratio of the intensities of the VI-R telomere band to the control ARO1 chromosomal band as described in the “Materials and methods.” The values shown are averages of multiple determinations. The input dilution is the input DNA that was used to determine the linear range of PCR amplification (see “Materials and methods”). d Comparison of cell doubling times and Mec1-HAp telomere enrichments. Mec1-HAp association increases at PD 70 just as survivors are beginning to arise in the culture, as indicated by the simultaneous decrease in doubling time and appearance of rearranged telomeres (panel a)
Mec1-HAp association with the VI-R telomere was monitored by ChIP. At each time point, aliquots of cells were treated with formaldehyde, and cross-linked chromatin fragments were prepared. The chromatin was immunoprecipitated with antibody against the HA epitope, and the immunoprecipitates were then analyzed by quantitative PCR for the enrichment of the VI-R telomere compared to control loci from the middle of chromosome IV or II (Fig. 1b). Mec1-HAp telomere association was most clearly detectable at PD 70 (Fig. 1c), the same point in the experiment when rearranged telomeres appeared and the culture growth rate increased (Fig. 1a, d). These data suggest that Mec1-HAp associates with short telomeres that are losing telomere function and entering a recombination-based survivor pathway for telomere maintenance.
The structure of the DNA bound by Mec1-HAp was investigated. Thus, the increased Mec1-HAp enrichment detected at PD 70 could arise from Mec1-HAp associating with DNA that had lost all of the TG1–3 telomeric repeats in which the degraded telomere is equivalent to one end of a DSB. Alternatively, Mec1-HAp may have associated with telomeres that retained short tracts of telomere repeats, but became dysfunctional due to the loss of DNA binding proteins that effect telomere function. To distinguish between these two possibilities, VI-R telomeres from the Mec1-HAp ChIP sample from PD 70 were cloned by telomere PCR and sequenced. Briefly, a poly-dC tail was added to the DNA fragments in the Mec1-HAp ChIP sample, and the VI-R telomeres were amplified with a unique primer and a poly-dG primer (Fig. 2a, described in (Förstemann et al. 2000; Hector et al. 2007)). Cloning and sequencing of the products from the Mec1-HAp ChIP revealed several VI-R telomeres; all of which had TG1–3 repeats of 36 to 55 nucleotides (Fig. 2b). These Mec1-HAp-associated telomeres bearing short tract of TG1–3 repeats appeared as cells escaping senescence due to telomeres entering a recombination pathway. These data strongly suggest that Mec1-HAp associates with dysfunctional telomeres.
The VI-R telomere sequences isolated from the Mec1-HAp ChIP are capped by short stretches of TG1–3 sequences. a The telomere PCR method (adapted from the method of Förstemann et al. (2000)). The Mec1-HAp ChIP is heated to reverse the formaldehyde cross-links, and DNA is subsequently purified and treated with dCTP and lambda terminal transferase to add an oligo-dC 3' extension. PCR is then performed with a specific primer to the VI-R telomere (TELVIR-88) and an oligo-dG primer (dG18-BamHI) to amplify a fragment containing the TG1–3 sequences (represented by the red box). PCR products were then cloned and sequenced. b Telomere sequences from the Mec1-HAp ChIP from population doubling 70 (Fig. 1). The VI-R reference sequence is the published genomic sequence that starts at chromosome VI base number 270,089 (Engel et al. 2010)
The average telomere repeat tract sizes at PD 50 and PD 70 differed by ~40 bp by Southern blotting (Fig. 1a), while the tract sizes of the sequences in the Mec1-HAp ChIP were even smaller (Fig. 2b). These differences in tract size combined with the increase in Mec1-HAp association as survivors appeared in the culture raised the possibility that the structure of the double-stranded repeat tracts may be important for recruiting Mec1-HAp. To address this question, the association of Mec1-HAp with the telomeres of cells lacking Yku70p was examined.
Mec1-HAp telomere association occurs when telomere dysfunction is induced in cells lacking Yku70p
As an independent test for the association of Mec1-HAp with dysfunctional telomeres, a system in which telomere dysfunction can be synchronously induced in a population of cells was used, i.e., yku70Δ cells that lack the 70-kD subunit of the Ku heterodimer and maintain short telomeres with 100–150 bp of TG1–3. This telomere shortening is independent of MEC1 as yku70Δ and yku70Δ mec1Δ cells have short telomeres of the same lengths (Corda et al. 2005). The telomeres of yku70Δ cells undergo a slow degradation of their 5' strands over a 24-h period when shifted from the permissive temperature (25°C) to the non-permissive temperature (37°C) (Fig. 3a), and a short telomere checkpoint is activated, which causes cell cycle arrest (Maringele and Lydall 2002). The yku70Δ cells can resume growth by means of recombination-based rearrangements of their telomeres similar to survivors seen in cells that lack telomerase activity (Chen et al. 2001; Fellerhoff et al. 2000; Huang et al. 2001; Maringele and Lydall 2002; Teng and Zakian 1999). Synchronous induction of telomere dysfunction is more readily controlled in the yku70Δ system, which has the added advantage of allowing the introduction of additional mutations into the yku70Δ strain background. We therefore examined cell doubling time and Mec1-HAp telomere association in yku70Δ cells grown at the non-permissive temperature.
Mec1-HAp and Tel1-HAp telomere association during induced telomere dysfunction in yku70Δ cells. a Telomere dysfunction is inducible in yku70Δ cells. A schematic of chromosomal ends where the telomere repeats are shown as curved lines. At 25°C, yku70Δ cell telomere repeats are maintained at a constant length. Previous work in the W303 strain, the same strain used here, showed that following prolonged incubation at 37°C, the 5' strand of the chromosome is degraded and a DNA damage checkpoint-mediated arrest ensues (Maringele and Lydall 2002). b A Southern blot for VI-R telomere lengths of cells grown at 25°C or shifted to 37°C and grown for the times indicated. The blot was probed and processed as in Fig. 1a. The 0-h time point corresponds to cells grown at 25°C. WT indicates samples from MEC1-HA cells or TEL1-HA cells with a wild-type YKU70 gene. This Southern analysis showed no evidence of survivor formation in the cells grown for 24 h at 37°C. c Representative Mec1-HAp ChIP results for wild-type and yku70Δ cells grown under the conditions shown in panel b. The ChIP was performed and analyzed as in Fig. 1c. The levels of VI-R telomere enrichment show the results of two independent ChIP experiments. d Culture doubling times of yku70Δ Mec1-HAp cells are plotted along with the VI-R enrichment values from panel c. The 70-h doubling time between the 12- and 24-h time points is indistinguishable from cessation of cell growth. e Representative Tel1-HAp ChIP results for wild-type and yku70Δ cells grown under the conditions shown in panel b. The levels of VI-R telomere enrichment show the results of two independent ChIP experiments. f Culture doubling times of yku70Δ Tel1-HAp cells are plotted along with the VI-R enrichment values from panel e. The telomere lengths and ChIP enrichments for the Tel1-HAp wild-type and yku70Δ samples at 0 and 8 h in panels b, e, and f have been published previously ((Hector et al. 2007) used with permission) and are shown as part of the larger time course to allow a direct comparison of Mec1-HAp telomere association assayed under the same conditions
After shifting yku70Δ cells from 25°C to 37°C, cell doubling time increased more than 20-fold over 24 h, and the VI-R telomere shortened slightly (Fig. 3b, d). Mec1-HAp association was not detectable at the beginning of the experiment but increased the longer cells were held at 37°C and the slower the cells grew (Fig. 3c, d). Interestingly, this increase in association paralleled the known increase in telomere strand degradation and dysfunction in yku70Δ cells that occurs over 24 h at 37°C (Maringele and Lydall 2002). These data are consistent with the hypothesis that Mec1-HAp associates with telomeres that are losing their function as yku70Δ cells begin to senesce at the non-permissive temperature. This Mec1-HAp pattern of association was distinct from that of Tel1-HAp as Tel1-HAp association was detected at all time points in yku70Δ cells (Fig. 3e, f). Thus, Mec1-HAp and Tel1-HAp displayed distinct patterns of telomere association as cell growth ceased.
Suppressing telomere dysfunction decreases Mec1-HAp telomere association
The impact of telomere dysfunction on Mec1-HAp association was further investigated by suppressing telomere resection in yku70Δ cells through deletion of the gene for Exo1p, an exonuclease known to degrade the 5' to 3' strand of DSBs (Fiorentini et al. 1997; Maringele and Lydall 2002; Tran et al. 2002). Exo1p appears to also act at telomeres in a similar manner to DSBs because cells bearing the yku70Δ exo1Δ mutations can be grown at 37°C for 24 h without senescing and show greatly reduced telomere degradation (Maringele and Lydall 2002). If Mec1-HAp associates with telomeres that are being degraded and losing their function in yku70Δ cells grown at 37°C, then blocking telomere degradation and dysfunction in yku70Δ exo1Δ cells should block Mec1-HAp association. We therefore examined whether loss of Exo1p altered Mec1-HAp telomere association and also analyzed Tel1-HAp in this system.
Eliminating Exo1p from yku70Δ cells suppressed the yku70Δ temperature-sensitive growth phenotype (Fig. 4a), while telomeres remained short (Fig. 4b). Thus, our system recapitulated both the kinetics of senescence and its suppression by loss of Exo1p previously shown by others (Maringele and Lydall 2002). Mec1-HAp VI-R telomere association was reproducibly reduced in yku70Δ exo1Δ cells grown for 24 h at 37°C (Fig. 4c, e). A similar reduction in Mec1-HAp association with the XV-L telomere was also observed (Fig. 5). Thus, suppressing the arrest due to telomere dysfunction abrogated Mec1-HAp telomere association.
Suppressing telomere dysfunction in yku70Δ cells abrogates Mec1-HAp telomere association. a Temperature sensitive growth is suppressed in yku70Δ exo1Δ cells. Aliquots (5 μl) of growing cells were spotted onto plates pre-warmed to 25°C or 37°C and then grown for 24 h at the indicated temperatures. b A Southern blot for VI-R telomere lengths of cells grown as indicated, where the 0-h time point corresponds to cells grown continuously at 25°C, probed and processed as in Fig. 1a. c Representative Mec1-HAp ChIP results for cells grown under the conditions shown in panel b. The ChIP was performed and analyzed as in Fig. 1c. The levels of VI-R telomere enrichment show the results of two independent ChIP experiments. The wild-type and yku70Δ cell samples from Fig. 3c were processed alongside the other samples for comparison. d Representative Tel1-HAp ChIP results for cells grown under the conditions shown in panel b, where the enrichment values are the results of two independent experiments. The wild-type and yku70Δ cell samples from Fig. 3e were processed alongside the other samples for comparison. e Graphical summary of the ChIP results of different Mec1-HAp and Tel1-HAp cells grown at 25°C or shifted to 37°C for 24 h. The Mec1-HAp telomere association observed in yku70Δ cells at 37°C is absent in yku70Δ exo1Δ cells
Mec1-HAp and Tel1-HAp association with the XV-L telomere parallels the results with the VI-R telomere. a A schematic showing the loci and primers used to monitor Mec1-HAp association with the XV-L telomere and the negative control internal locus ARO1, with the PCR primers indicated by arrows. b Representative Mec1-HAp ChIP results from the same strains analyzed for VI-R telomere association in Fig. 4c for cells grown continuously at 25°C or shifted to 37°C for 24 h. The quantitations shown are the results of two independent ChIP experiments. c Representative Tel1-HAp ChIP results for the same strains analyzed in Fig. 4d. The quantitations shown are the results of two independent ChIP experiments. d Graphical summary of the ChIP results. The magnitude of Tel1-HAp association with the XV-L and VI-R (Fig. 4) telomeres in yku70Δ cells was similar at 37°C but different at 25°C, possibly due to the different DNA sequences adjacent to the TG1–3 sequences at each locus
Tel1-HAp gave a different response than Mec1-HAp as Tel1-HAp associated with the VI-R and XV-L telomeres in yku70Δ exo1Δ cells grown for 24 h at 37°C (Figs. 4d, e and 5). Thus, suppressing the temperature-sensitive telomere dysfunction of yku70Δ cells still allowed Tel1-HAp telomere association. Intriguingly, Tel1-HAp telomere association in yku70Δ exo1Δ cells was reduced at 25°C, even though Tel1-HAp normally associated with the short yku70Δ telomeres at that temperature (Figs. 4e and 5d). The reason for this reduction and the role of Exo1p in Tel1p telomere association is not currently understood. The sum of these data indicates that Tel1-HAp associates with short functional telomeres while Mec1-HAp is recruited to dysfunctional telomeres.
Mec1-HAp shows modest association with short yku70Δ cell telomeres in late S phase
Mec1-HAp telomere association was also monitored during the cell cycle because yeast telomeres undergo a telomerase-independent resection of the 5' telomere strand in late S-phase to produce a transient 50- to 100-bp single-stranded overhang (Wellinger et al. 1996). In the case of yku70Δ cells bearing telomere repeat tracts that are ~200 bp shorter than those of wild-type cells, we hypothesized that this resection may reduce these tracts to the point where so few telomere binding proteins are present that the telomere becomes dysfunctional and Mec1-HAp is recruited. We therefore examined Mec1-HAp telomere association during the cell cycle in yku70Δ cells at the permissive temperature and wild-type cells.
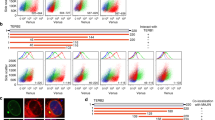
Mec1-HAp association with the VI-R telomere in yku70Δ cells showed a modest but detectable increase in Mec1-HAp association as cells entered late S-phase, with a subsequent reduction as the cells entered G2/M (Fig. 6a–c). These results are consistent with the hypothesis that the reduction of double-stranded TG1–3 repeats by degradation of the 5' telomere strand during late S-phase on short telomeres can promote Mec1-HAp telomere association. In contrast, Mec1-HAp association with the VI-R telomere did not significantly change during the cell cycle in wild-type cells, even when cells were allowed to progress into the next G1 phase (Fig. 6d–f). Thus, the progression of wild-type cells with normal length telomeres through S-phase did not cause an alteration in telomere structure to result in detectable Mec1-HAp telomere association.
Cell cycle analysis of Mec1-HAp telomere association in yku70Δ cells (a–c) and wild-type cells (d–f). Cells were grown at the permissive temperature (20°C), arrested with α-factor and released into the cell cycle until all of the cells were in G2/M phase, and samples were taken for FACS and ChIP. Both strains bear the bar1Δ mutation, which eliminates a secreted protein that degrades α-factor and facilitates the synchronization and release. a FACS analysis showing DNA content of the yku70Δ cells analyzed. b Representative ChIP results for the yku70Δ VI-R telomere. Values below each lane are the quantitations of all enrichment determinations, each analyzed in triplicate. c Graphical presentation of the fold enrichments in panel b, where the error bars represent the standard deviations of the analysis of each time point. d FACS analysis showing DNA content of the wild-type cells analyzed. This synchrony was allowed to proceed until some of the cells began entering the next G1 phase. e Representative ChIP results for the wild-type VI-R telomere. Quantitations shown below the gel are the average of two separate experiments, where each ChIP experiment was analyzed in triplicate. f Graphical presentation of a representative cell cycle synchrony, where the error bars represent the standard deviations of the analysis of each sample. The cell cycle phases from panel d are indicated, including the portion of cells that re-entered G1 phase. The entire experiment was done twice
Discussion
We have shown that Mec1-HAp associates with dysfunctional telomeres in two separate cellular conditions: extreme telomere shortening in cells lacking telomerase and senescence induced in yku70Δ cells grown at the non-permissive temperature. In the case of cells lacking telomerase, Mec1-HAp telomere association was not detectable as telomeres gradually shortened or as cell growth dramatically slowed (Fig. 1d). Only when telomerase-independent survivors had begun to appear in the culture was Mec1-HAp telomere association observed (Fig. 1a, c, d). As survivor formation is a process that requires telomere recombination and DSB recombination proteins (reviewed in (McEachern and Haber 2006; Runge 2006)), these data indicate that Mec1-HAp telomere association occurs when cells no longer distinguish the chromosome end from a DSB. In the case of yku70Δ cells grown at the non-permissive temperature, Mec1-HAp telomere association was highest when cells had stopped growing (Fig. 3), coincident with the time when degradation of the 5' strand of the telomere is known to occur and cell cycle arrest ensues (Maringele and Lydall 2002), suggesting that Mec1-HA associated with the resected telomeres. Consistent with this interpretation, deletion of the gene for EXO1, which has been shown to delay degradation of the 5' strand of the telomere (Maringele and Lydall 2002), prevented cell senescence and Mec1-HAp telomere association (Figs. 4 and 5). In contrast, the Tel1-HAp pattern of telomere association was different as Tel1p remained bound in both growing and senescent yku70Δ cells and in yku70Δ exo1Δ cells (Figs. 3, 4, and 5), consistent with previous results showing that Tel1-HAp associates with short telomeres to stimulate their elongation (Bianchi and Shore 2007; Hector et al. 2007; Sabourin et al. 2007). These data highlight the different cellular functions of these two ATM family member kinases and indicate that Mec1-HAp associates with telomeres that no longer act as stable chromosome ends.
Our data showing that the Mec1-HAp ChIP contained fragments with short TG1–3 tracts (Fig. 2) have implications for how the telomere transitions from a functional state to one that recruits Mec1-HAp. While the length of the double-stranded repeats and the number of repeat-binding proteins required to distinguish a telomere from a DSB are unclear, recent experiments suggest that a minimum of ~70 bp is required. Briefly, telomeres possess an “anti-checkpoint” activity that can repress DNA damage signaling when they are adjacent to a DSB (Michelson et al. 2005) and can block the resection of the 5' DNA strand (Hirano and Sugimoto 2007). Arrays of four or more sites for the telomere binding protein Rap1p tethered adjacent to a DSB also block resection and instead support telomere formation (Negrini et al. 2007), implicating bound Rap1p molecules as a key component in distinguishing a nascent telomere from a DSB. Consistent with this idea, reduced levels of cellular Rap1p can cause telomeres to enter recombination pathways at increased frequency (Pardo and Marcand 2005), suggesting that fewer bound Rap1p molecules allow telomeres to behave as DSBs. As the yeast cell appears to equate each bound Rap1p with 18–19 bp of TG1–3 repeats (Ray and Runge 1999b), four Rap1p sites would suggest that a tract of ≥70 bp of TG1–3 sequences is required to elicit an “anti-checkpoint activity” that prevents telomeres from recombining with other DNA molecules similar to DSBs. Our observation that Mec1-HAp was bound to telomeres with less than 60 bases of TG1–3 (Fig. 2b) supports the hypothesis that Mec1-HAp associates with telomeres that have lost their function. Thus, we suggest that as the telomere tracts shorten below a minimal threshold, proteins that bind to double-stranded telomere repeats are lost and the telomere becomes dysfunctional, resulting in recruitment of Mec1p.
Mec1-HAp association with both dysfunctional telomeres and DSBs appears to require resection of the 5' strand. At a specific HO-induced DSB, Mec1p association with sequences adjacent to the break can be detected soon after synthesis of the endonuclease is induced (Nakada et al. 2004) (Fig. S1), and this association depends in part on the gene for EXO1 (Nakada et al. 2004). Similarly, Mec1-HAp association with the yku70Δ cell telomeres at the non-permissive temperature also depends in part upon EXO1 (Figs. 4 and 5). Mec1-HAp association with DNA bearing a resected 5' strand is also indicated at telomeres in a cdc13-1 mutant (Rouse and Jackson 2002b), where growth at the non-permissive temperature results in degradation of the 5' strand of the telomere for scores of kb into the chromosome (Booth et al. 2001; Garvik et al. 1995). Thus, Mec1-HAp associates with resected DSBs and telomeres that lack long tracts of double-stranded TG1–3 repeats, suggesting that the mechanisms that recruit Mec1-HAp to both structures are the same.
The combination of short TG1–3 tracts in the Mec1-HAp ChIP and the requirement of Exo1p for Mec1-HAp telomere association in yku70Δ cells suggest that the number of double-stranded telomere binding proteins present at the chromosome end is what distinguishes functional and dysfunctional telomeres. The Mec1-HAp association results with yku70Δ and wild-type telomeres during the cell cycle are consistent with this idea. In the cell cycle association experiments, wild-type cells have ~350 bp of double-stranded telomere repeats, while yku70Δ cells have only 100–150 bp (Porter et al. 1996). Mec1-HAp telomere association could be detected in yku70Δ cells in late S phase (Fig. 6a–c), when the 5' strand resection of 50–100 bp (Larrivee et al. 2004; Wellinger et al. 1996) would remove most of the double-stranded repeats in these mutant cells. However, association was not detected in wild-type cells (Fig. 6d–f) where ~200 bp of the repeats would be expected to remain double stranded. Our working model for Mec1-HAp recruitment to yku70Δ telomeres then is that the loss of double-stranded repeats would reduce the length of the telomere repeats to less than 70 bp at some chromosome ends, eliminating many of the Rap1p binding sites and removing the chromatin that distinguishes telomeres from DSBs.
If Mec1-HAp telomere association occurs when telomeres become dysfunctional, why does the culture doubling time of the cells lacking telomerase increase before Mec1-HAp recruitment is detected (Fig. 1d)? We propose that this slowed culture doubling time is due to Tel1p activity. Tel1p is known to associate with short telomeres and stimulate their elongation by telomerase (Bianchi and Shore 2007; Hector et al. 2007; Sabourin et al. 2007). In cells lacking telomerase, Tel1p telomere association correlates with the increase in population doubling time (Hector et al. 2007). However, Tel1p telomere association does not appear to cause a permanent arrest as the culture continues growing, which may reflect a transient cell cycle pause that slows culture growth rate. This suggestion is consistent with earlier experiments in cells lacking telomerase in which the final G2 arrest required MEC1, but removal of TEL1 changed the number of population doublings required for this arrest to occur (Enomoto et al. 2002; IJpma and Greider 2003). We therefore hypothesize that the increase in PD time seen in the Mec1-HAp cells lacking telomerase reflects a cell cycle pause caused by Tel1p association with short telomeres, while the final G2 arrest that occurs in the majority of telomerase-null cells is caused by Mec1p association with short, resected telomeres that have lost their telomere function.
While Mec1p associates with DSBs and dysfunctional telomeres, it is not clear if all of the consequences of Mec1p association are the same at both structures. At DSBs, Mec1p association promotes a cell cycle arrest, activation of the DNA damage response, and the initiation of DNA recombination and repair (Longhese 2008; Rouse and Jackson 2002a). A recently discovered function for Mec1p is the suppression of inappropriate telomere formation at DSBs in the middle of the chromosome. Mec1p-dependent phosphorylation of Pif1p (Makovets and Blackburn 2009) and Cdc13 (Zhang and Durocher 2010) blocks the activity of telomerase to prevent telomere formation at DSBs. If a similar process occurred at dysfunctional telomeres, telomerase recruitment would be prevented, and Mec1p binding would induce recombination to produce survivors that escape the cell cycle pause. However, cells could also escape this pause if Mec1p association somehow allowed the addition of more telomere repeats, and the telomerase-null phenotype of mec1Δ tel1Δ cells is consistent with a role for Mec1p in stimulating telomere elongation. As dysfunctional telomeres are distinct from DSBs in that they retain short stretches of TG1–3 tracts (Fig. 2b) and are next to the telomere-associated middle repetitive elements (Zakian 1996), telomere-associated proteins at these sites may potentiate elongation of telomere repeats to repair the telomere and allow cells to re-enter the cell cycle. It will be interesting to determine if cells possess a mechanism that allows very short telomeres bound by Mec1p to become telomerase substrates, in contrast to what occurs at DSBs.
References
Abdallah P, Luciano P, Runge KW, Lisby M, Geli V, Gilson E, Teixeira MT (2009) A two-step model for senescence triggered by a single critically short telomere. Nat Cell Biol 11:988–993
Arneric M, Lingner J (2007) Tel1 kinase and subtelomere-bound Tbf1 mediate preferential elongation of short telomeres by telomerase in yeast. EMBO Rep 8:1080–1085
Baudin A, Ozier-Kalogeropoulos O, Denouel A, Lacroute F, Cullin C (1993) A simple and efficient method for direct gene deletion in Saccharomyces cerevisiae. Nucleic Acids Res 21:3329–3330
Berthiau AS, Yankulov K, Bah A, Revardel E, Luciano P, Wellinger RJ, Geli V, Gilson E (2006) Subtelomeric proteins negatively regulate telomere elongation in budding yeast. EMBO J 25:846–856
Bianchi A, Shore D (2007) Increased association of telomerase with short telomeres in yeast. Genes Dev 21:1726–1730
Bianchi A, Shore D (2008) How telomerase reaches its end: mechanism of telomerase regulation by the telomeric complex. Mol Cell 31:153–165
Booth C, Griffith E, Brady G, Lydall D (2001) Quantitative amplification of single-stranded DNA (QAOS) demonstrates that cdc13-1 mutants generate ssDNA in a telomere to centromere direction. Nucleic Acids Res 29:4414–4422
Chan SW, Chang J, Prescott J, Blackburn EH (2001) Altering telomere structure allows telomerase to act in yeast lacking ATM kinases. Curr Biol 11:1240–1250
Chen Q, Ijpma A, Greider CW (2001) Two survivor pathways that allow growth in the absence of telomerase are generated by distinct telomere recombination events. Mol Cell Biol 21:1819–1827
Cimprich KA, Cortez D (2008) ATR: an essential regulator of genome integrity. Nat Rev Mol Cell Biol 9:616–627
Corda Y, Lee SE, Guillot S, Walther A, Sollier J, Arbel-Eden A, Haber JE, Geli V (2005) Inactivation of Ku-mediated end joining suppresses mec1Delta lethality by depleting the ribonucleotide reductase inhibitor Sml1 through a pathway controlled by Tel1 kinase and the Mre11 complex. Mol Cell Biol 25:10652–10664
Craven RJ, Petes TD (1999) Dependence of the regulation of telomere length on the type of subtelomeric repeat in the yeast Saccharomyces cerevisiae. Genetics 152:1531–1541
d'Adda di Fagagna F, Reaper PM, Clay-Farrace L, Fiegler H, Carr P, von Zglinicki T, Saretzki G, Carter NP, Jackson SP (2003) A DNA damage checkpoint response in telomere-initiated senescence. Nature 426:194–198
Engel SR, Balakrishnan R, Binkley G, Christie KR, Costanzo MC, Dwight SS, Fisk DG, Hirschman JE, Hitz BC, Hong EL, Krieger CJ, Livstone MS, Miyasato SR, Nash R, Oughtred R, Park J, Skrzypek MS, Weng S, Wong ED, Dolinski K, Botstein D, Cherry JM (2010) Saccharomyces genome database provides mutant phenotype data. Nucleic Acids Res 38:D433–D436
Enomoto S, Glowczewski L, Berman J (2002) MEC3, MEC1, and DDC2 are essential components of a telomere checkpoint pathway required for cell cycle arrest during senescence in Saccharomyces cerevisiae. Mol Biol Cell 13:2626–2638
Fellerhoff B, Eckardt-Schupp F, Friedl AA (2000) Subtelomeric repeat amplification is associated with growth at elevated temperature in yku70 mutants of Saccharomyces cerevisiae. Genetics 154:1039–1051
Fiorentini P, Huang KN, Tishkoff DX, Kolodner RD, Symington LS (1997) Exonuclease I of Saccharomyces cerevisiae functions in mitotic recombination in vivo and in vitro. Mol Cell Biol 17:2764–2773
Förstemann K, Hoss M, Lingner J (2000) Telomerase-dependent repeat divergence at the 3' ends of yeast telomeres. Nucleic Acids Res 28:2690–2694
Garvik B, Carson M, Hartwell L (1995) Single-stranded DNA arising at telomeres in cdc13 mutants may constitute a specific signal for the RAD9 checkpoint. Mol Cell Biol 15:6128–6138
Goudsouzian LK, Tuzon CT, Zakian VA (2006) S. cerevisiae Tel1p and Mre11p are required for normal levels of Est1p and Est2p telomere association. Mol Cell 24:603–610
Hector RE, Shtofman RL, Ray A, Chen BR, Nyun T, Berkner KL, Runge KW (2007) Tel1p preferentially associates with short telomeres to stimulate their elongation. Mol Cell 27:851–858
Herbig U, Jobling WA, Chen BP, Chen DJ, Sedivy JM (2004) Telomere shortening triggers senescence of human cells through a pathway involving ATM, p53, and p21(CIP1), but not p16(INK4a). Mol Cell 14:501–513
Hirano Y, Sugimoto K (2007) Cdc13 telomere capping decreases Mec1 association but does not affect Tel1 association with DNA ends. Mol Biol Cell 18:2026–2036
Huang P, Pryde FE, Lester D, Maddison RL, Borts RH, Hickson ID, Louis EJ (2001) SGS1 is required for telomere elongation in the absence of telomerase. Curr Biol 11:125–129
IJpma AS, Greider CW (2003) Short telomeres induce a DNA damage response in Saccharomyces cerevisiae. Mol Biol Cell 14:987–1001
Khadaroo B, Teixeira MT, Luciano P, Eckert-Boulet N, Germann SM, Simon MN, Gallina I, Abdallah P, Gilson E, Geli V, Lisby M (2009) The DNA damage response at eroded telomeres and tethering to the nuclear pore complex. Nat Cell Biol 11:980–987
Larrivee M, LeBel C, Wellinger RJ (2004) The generation of proper constitutive G-tails on yeast telomeres is dependent on the MRX complex. Genes Dev 18:1391–1396
Lavin MF (2007) ATM and the Mre11 complex combine to recognize and signal DNA double-strand breaks. Oncogene 26:7749–7758
Le S, Moore JK, Haber JE, Greider CW (1999) RAD50 and RAD51 define two pathways that collaborate to maintain telomeres in the absence of telomerase. Genetics 152:143–152
Lisby M, Barlow JH, Burgess RC, Rothstein R (2004) Choreography of the DNA damage response: spatiotemporal relationships among checkpoint and repair proteins. Cell 118:699–713
Livengood AJ, Zaug AJ, Cech TR (2002) Essential regions of Saccharomyces cerevisiae telomerase RNA: separate elements for Est1p and Est2p interaction. Mol Cell Biol 22:2366–2374
Longhese MP (2008) DNA damage response at functional and dysfunctional telomeres. Genes Dev 22:125–140
Lundblad V, Blackburn EH (1993) An alternative pathway for yeast telomere maintenance rescues est1 - senescence. Cell 73:347–360
Lustig AJ, Petes TD (1986) Identification of yeast mutants with altered telomere structure. Proc Natl Acad Sci U S A 83:1398–1402
Ma Y, Greider CW (2009) Kinase-independent functions of TEL1 in telomere maintenance. Mol Cell Biol 29:5193–5202
Makovets S, Blackburn EH (2009) DNA damage signalling prevents deleterious telomere addition at DNA breaks. Nat Cell Biol 11:1383–1386
Mallory JC, Petes TD (2000) Protein kinase activity of Tel1p and Mec1p, two Saccharomyces cerevisiae proteins related to the human ATM protein kinase. Proc Natl Acad Sci U S A 97:13749–13754
Maringele L, Lydall D (2002) EXO1-dependent single-stranded DNA at telomeres activates subsets of DNA damage and spindle checkpoint pathways in budding yeast yku70Delta mutants. Genes Dev 16:1919–1933
McEachern MJ, Haber JE (2006) Break-induced replication and recombinational telomere elongation in yeast. Annu Rev Biochem 75:111–135
McGee JS, Phillips JA, Chan A, Sabourin M, Paeschke K, Zakian VA (2010) Reduced Rif2 and lack of Mec1 target short telomeres for elongation rather than double-strand break repair. Nat Struct Mol Biol 17:1438–1445
Michelson RJ, Rosenstein S, Weinert T (2005) A telomeric repeat sequence adjacent to a DNA double-stranded break produces an anticheckpoint. Genes Dev 19:2546–2559
Morrow DM, Tagle DA, Shiloh Y, Collins FS, Hieter P (1995) TEL1, an S. cerevisiae homolog of the human gene mutated in ataxia telangiectasia, is functionally related to the yeast checkpoint gene MEC1. Cell 82:831–840
Nakada D, Matsumoto K, Sugimoto K (2003) ATM-related Tel1 associates with double-strand breaks through an Xrs2-dependent mechanism. Genes Dev 17:1957–1962
Nakada D, Hirano Y, Sugimoto K (2004) Requirement of the Mre11 complex and exonuclease 1 for activation of the Mec1 signaling pathway. Mol Cell Biol 24:10016–10025
Nakada D, Hirano Y, Tanaka Y, Sugimoto K (2005) Role of the C terminus of Mec1 checkpoint kinase in its localization to sites of DNA damage. Mol Biol Cell 16:5227–5235
Negrini S, Ribaud V, Bianchi A, Shore D (2007) DNA breaks are masked by multiple Rap1 binding in yeast: implications for telomere capping and telomerase regulation. Genes Dev 21:292–302
Pardo B, Marcand S (2005) Rap1 prevents telomere fusions by nonhomologous end joining. EMBO J 24:3117–3127
Porter SE, Greenwell PW, Ritchie KB, Petes TD (1996) The DNA-binding protein Hdf1p (a putative Ku homologue) is required for maintaining normal telomere length in Saccharomyces cerevisiae. Nucleic Acids Res 24:582–585
Ray A, Runge KW (1999a) Varying the number of telomere-bound proteins does not alter telomere length in tel1Δ cells. Proc Natl Acad Sci U S A 96:15044–15049
Ray A, Runge KW (1999b) The yeast telomere length counting machinery is sensitive to sequences at the telomere–nontelomere junction. Mol Cell Biol 19:31–45
Ray A, Runge KW (2001) Yeast telomerase appears to frequently copy the entire template in vivo. Nucleic Acids Res 29:2382–2394
Ritchie K, Mallory J, Petes TD (1999) Interactions of TLC1 (which encodes the RNA subunit of telomerase), TEL1 and MEC1 in regulating telomere length in the yeast Saccharomyces cerevisiae. Mol Cell Biol 19:6065–6075
Rose MD, Winston F and Hieter P (1990) Methods in yeast genetics: a laboratory course manual. Cold Spring Harbor Laboratory
Rouse J, Jackson SP (2002a) Interfaces between the detection, signaling, and repair of DNA damage. Science 297:547–551
Rouse J, Jackson SP (2002b) Lcd1p recruits Mec1p to DNA lesions in vitro and in vivo. Mol Cell 9:857–869
Runge KW (2006) Telomeres and aging in the yeast model system. In: Conn PM (ed) Handbook of models for human aging. Elsevier Academic Press, New York, pp 191–206
Sabourin M, Tuzon CT, Zakian VA (2007) Telomerase and Tel1p preferentially associate with short telomeres in S. cerevisiae. Mol Cell 27:550–561
Smogorzewska A, de Lange T (2004) Regulation of telomerase by telomeric proteins. Annu Rev Biochem 73:177–208
Spellman PT, Sherlock G, Zhang MQ, Iyer VR, Anders K, Eisen MB, Brown PO, Botstein D, Futcher B (1998) Comprehensive identification of cell cycle-regulated genes of the yeast Saccharomyces cerevisiae by microarray hybridization. Mol Biol Cell 9:3273–3297
Strahl-Bolsinger S, Hecht A, Luo K, Grunstein M (1997) SIR2 and SIR4 interactions differ in core and extended telomeric heterochromatin in yeast. Genes Dev 11:83–93
Teng SC, Zakian VA (1999) Telomere–telomere recombination is an efficient bypass pathway for telomere maintenance in Saccharomyces cerevisiae. Mol Cell Biol 19:8083–8093
Teng SC, Chang J, McCowan B, Zakian VA (2000) Telomerase-independent lengthening of yeast telomeres occurs by an abrupt Rad50p-dependent, Rif-inhibited recombinational process. Mol Cell 6:947–952
Tran PT, Erdeniz N, Dudley S, Liskay RM (2002) Characterization of nuclease-dependent functions of Exo1p in Saccharomyces cerevisiae. DNA Repair (Amst) 1:895–912
Tsubouchi H, Ogawa H (2000) Exo1 roles for repair of DNA double-strand breaks and meiotic crossing over in Saccharomyces cerevisiae. Mol Biol Cell 11:2221–2233
Usui T, Ogawa H, Petrini JH (2001) A DNA damage response pathway controlled by Tel1 and the Mre11 complex. Mol Cell 7:1255–1266
Wellinger RJ, Ethier K, Labrecque P, Zakian VA (1996) Evidence for a new step in telomere maintenance. Cell 85:423–433
Zakian VA (1996) Structure, function and replication of Saccharomyces cerevisiae telomeres. Annu Rev Genet 30:141–172
Zhang W, Durocher D (2010) De novo telomere formation is suppressed by the Mec1-dependent inhibition of Cdc13 accumulation at DNA breaks. Genes Dev 24:502–515
Acknowledgements
We would like to thank Drs. Tom Petes and Victoria Lundblad for the yeast strains and plasmids, Steven Sanders, Jinyu Wang, and Brian Yard for the comments on the manuscript, and Anna Yakubenko, Bianca Calderon, Matthew Shaughnessy, and Thomas Klein for the technical support during the course of this work. This work was supported by the US National Institutes of Health grants R01 GM050752 to KWR. REH was supported by F32 AG022743 and T32 HD007104.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Supplemental Material
Below is the link to the electronic supplementary material.
412_2011_359_MOESM1_ESM.doc
The Supplementary Material containing three figures and two tables can be found with this article on-line at http://xxxx. (DOC 958 kb)
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Hector, R.E., Ray, A., Chen, BR. et al. Mec1p associates with functionally compromised telomeres. Chromosoma 121, 277–290 (2012). https://doi.org/10.1007/s00412-011-0359-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-011-0359-0