Abstract
The base excision repair machinery protects DNA in cells from the damaging effects of oxidation, alkylation, and deamination; it is specialized to fix single-base damage in the form of small chemical modifications. Base modifications can be mutagenic and/or cytotoxic, depending on how they interfere with the template function of the DNA during replication and transcription. DNA glycosylases play a key role in the elimination of such DNA lesions; they recognize and excise damaged bases, thereby initiating a repair process that restores the regular DNA structure with high accuracy. All glycosylases share a common mode of action for damage recognition; they flip bases out of the DNA helix into a selective active site pocket, the architecture of which permits a sensitive detection of even minor base irregularities. Within the past few years, it has become clear that nature has exploited this ability to read the chemical structure of DNA bases for purposes other than canonical DNA repair. DNA glycosylases have been brought into context with molecular processes relating to innate and adaptive immunity as well as to the control of DNA methylation and epigenetic stability. Here, we summarize the key structural and mechanistic features of DNA glycosylases with a special focus on the mammalian enzymes, and then review the evidence for the newly emerging biological functions beyond the protection of genome integrity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The integrity of genetic information is under constant threat by the tendency of DNA to engage in chemical reactions in its cellular environment. These can damage the DNA in various ways, most frequently by oxidation, alkylation, or deamination of the coding bases (Lindahl and Wood 1999). Damage to DNA bases may affect their base-pairing properties and, therefore, needs to be fixed to maintain the template function of the DNA (Kunz et al. 2009a). Many base lesions are pro-mutagenic, i.e., they give rise to genetic mutations if not repaired. One such example is 7,8-dihydro-8-oxoguanine (8-oxoG), a frequent product of DNA oxidation. 8-oxoG tends to base-pair with adenine, thus giving rise to G•C to T•A transversion mutations. Likewise, hydrolytic deamination of cytosine and 5-methylcytosine (5-meC) gives rise to uracil and thymine mispaired with guanine, respectively, causing C•G → T•A transition mutations if not repaired. Alkylation can generate a variety of DNA base lesions comprising O6-methylguanine (6-meG), N7-methylguanine (7-meG), or N3-methyladenine (3-meA). While 6-meG is pro-mutagenic by its property to pair with thymine, 7-meG and 3-meA block replicative DNA polymerases and are therefore cytotoxic (Lindahl and Wood 1999).
These and many other forms of DNA base damage arise in cells at least 10,000 times every day and only the continuous action of specialized DNA repair systems can prevent a rapid decay of genetic information. Single-base lesions are eliminated by base excision repair (BER), a pathway initiated by DNA glycosylases that recognize and excise damaged bases. Base removal by a DNA glycosylase generates a so-called apurinic/apyrimidinic site (AP-site) in DNA, which is then further processed by specific AP-endonuclease, DNA polymerase, and DNA ligase activities to restore the original DNA sequence (Fig. 1) (Almeida and Sobol 2007). Accordingly, cells lacking DNA glycosylase functions generally show increased levels of base damage in their DNA, elevated mutation rates, and hypersensitivity to specific DNA damaging agents. Surprisingly, however, the phenotype of DNA glycosylase disruptions in mice is usually rather moderate (reviewed in Robertson et al. 2009), the only known exception being the thymine DNA glycosylase (TDG), which was recently reported to be essential for embryonic development in mouse (Cortazar et al. 2011; Cortellino et al. 2011).
The core pathway short-patch BER. The base-excision repair pathway addresses single-base lesions (a). BER is initiated by a DNA glycosylase, e.g., UNG, specifically recognizing and binding a base lesion. Upon encountering a substrate base, e.g., uracil for UNG, the glycosylase flips the base out of the base-stack into its catalytic site pocket where specific contacts examine the substrate base and position it for nucleophilic attack to the N-glycosidic bond (b). Release of the substrate base results in an abasic site (c), which is further processed by the AP-endonuclease, APE1, that cleaves the phosphate backbone 5’ to the abasic site, producing a 3’OH and a 5’deoxyribose-phosphate moiety (5’dRP) (d). Polymerase β (Polβ) hydrolyzes the 5’dRP and fills in the single nucleotide gap, which is subsequently sealed by the DNA ligase III (LigIII), supported by the scaffold protein XRCC1 (e), thus restoring the original base sequence (f). The increase of DNA bending from UNG to Polβ might support the directionality of the handover from one BER factor to the next
In this review, we will focus on the mammalian DNA glycosylases, for which we will briefly summarize the key structure–function concepts and discuss their role in the repair of DNA base lesions. We will further elaborate on their newly emerging functions beyond canonical DNA repair, e.g., in innate and adaptive immunity and in DNA methylation control.
DNA glycosylases—an ancient family of DNA repair proteins
The consideration that cells must possess an ability to remove uracil from DNA, which arises either by misincorporation of deoxyuridine monophosphate (dUMP) during DNA replication or by hydrolytic deamination of cytosine, led to the discovery of an enzyme capable of cleaving uracil–deoxyribose bonds, the uracil N-glycosidase (Ung) in Escherichia coli (Lindahl 1974). This finding was followed by the isolation of many other DNA glycosylases in species from all kingdoms of life. Eleven DNA glycosylases have been identified in mammals and these can be subdivided into four structurally distinct superfamilies; the uracil DNA glycosylases (UDGs), the helix-hairpin-helix (HhH) glycosylases, the 3-methyl-purine glycosylase (MPG), and the endonuclease VIII-like (NEIL) glycosylases (Table 1).
The uracil DNA glycosylases
E. coli Ung turned out to be the founding member of a large superfamily of glycosylases, which now includes six subfamilies, three of which are present in the eukarya. Besides the UNG subfamily, these include the mismatch-specific uracil DNA glycosylases (MUGs) (Gallinari and Jiricny 1996) and the single-strand-specific monofunctional uracil DNA glycosylases (SMUGs) (Haushalter et al. 1999). Despite a considerable amino-acid sequence divergence, all UDGs share a common alpha–beta fold structured catalytic domain (Aravind and Koonin 2000).
Members of the UNG subfamily have been characterized in organisms from bacteria and yeasts to humans and large "eukaryotic" viruses. These are highly conserved both at the amino-acid sequence and gene structure levels; the yeast and human proteins share 40.3% amino-acid sequence similarity and the human, mouse, and fish genes have identical exon–intron boundaries, indicating that the exon–intron organization has not changed for more than 450 million years (Krokan et al. 1997). Alternative splicing as well as transcription from two distinct start sites gives rise to the specific mitochondrial and nuclear isoforms UNG1 and UNG2 in mouse and human cells (Nilsen et al. 1997). UNG is highly specific for processing of uracil in DNA but also excises DNA-incorporated 5-fluorouracil (5-FU), a uracil analog used in cancer therapy (Pettersen et al. 2011). Interactions with PCNA and RPA target the nuclear UNG2 to sites of DNA synthesis, where its main function is to rapidly excise uracil that gets incorporated opposite from adenine (Otterlei et al. 1999). Accordingly, mouse cells deficient in UNG accumulate ~100-fold increased levels of uracil in their DNA but, notably, do not show a significant mutator phenotype (Nilsen et al. 2000). This is unlike human cells where inhibition of UNG appears to elevate the mutation frequency mildly (Radany et al. 2000). However, mice lacking Ung do develop B-cell lymphomas and show disturbances of antibody diversification, implicating a specific function of UNG in processing deamination-induced U•G mismatches at immunoglobulin loci to facilitate somatic hypermutation and class switch recombination (Rada et al. 2002; Nilsen et al. 2003). Consistently, mutations in the human UNG gene have been associated with a subgroup of hyper-IgM syndrome patients, showing impaired class switch recombination (Imai et al. 2003).
The MUG subfamily of UDGs emerged with the identification of TDG, an enzyme capable of excising thymine from G•T mismatches. Nonetheless, the family was named after the E. coli Mug protein (Gallinari and Jiricny 1996), giving credit to the fact that the G•U rather than the G•T mismatch represents the common most efficiently processed substrate for the members of this subfamily. MUG orthologs have been described in Schizosaccharomyces pombe, Drosophila melanogaster (Hardeland et al. 2003), and mammals (Neddermann et al. 1996). Crystal structural analyses revealed a striking resemblance in the overall fold of the glycosylase domains of E. coli Mug and Ung, despite the absence of a notable sequence similarity (Barrett et al. 1998). Unlike UNG, however, the MUG glycosylases have a spacious and rather non-discriminating active site pocket, accommodating a broad range of substrates including pyrimidine derivates like 5-FU, 5-hydroxymethyluracil (5-hmU), and 3,N4-ethenocytosine (Table 1) (reviewed in Cortazar et al. 2007), and they process these substrates with an extremely low turnover rate (Waters and Swann 1998; Hardeland et al. 2000). Compared to E. coli Mug, which consists of the catalytic core only, TDG contains additional N- and C-terminal domains, providing non-specific DNA interaction and regulatory functions (Hardeland et al. 2002; Steinacher and Schär 2005; Baba et al. 2005). Knockout of Tdg in mouse is embryonic lethal, suggesting that, unlike other UDGs, it has a non-redundant essential function in embryonic development (Cortazar et al. 2011; Cortellino et al. 2011). TDG-deficient cells do not show increased sensitivity towards agents that would cause TDG-relevant DNA base lesions, nor do they show increased levels of spontaneous mutations (Cortazar et al. 2011), implicating functions beyond canonical DNA repair, which will be discussed below. An involvement of TDG in DNA repair becomes obvious, however, in the processing of the anti-cancer drug 5-FU when incorporated into the DNA. In this special case, the repair activity of TDG does not provide drug resistance as might be expected; excision of the base analog by TDG results in an accumulation of toxic AP-site intermediates and DNA strand breaks and, thus, mediates the DNA-directed cytotoxic effect of 5-FU (Kunz et al. 2009b).
SMUG-family glycosylases were initially identified as a uracil-excising activity in Xenopus, insect, and human cells (Haushalter et al. 1999). As such, it appears to serve as back-up for UNG in limiting uracil accumulation (U•A) in genomic DNA and in preventing C → T mutation following cytosine deamination (U•G) (Haushalter et al. 1999; An et al. 2005). While Xenopus SMUG has a preference for uracil in single-stranded DNA, hence the name “single-strand-specific monofunctional uracil DNA glycosylase”, the human homolog processes uracil also in double-stranded DNA (Table 1) (Haushalter et al. 1999; Kavli et al. 2002). SMUGs show only limited amino-acid sequence similarity with members of other UDG subfamilies and the conservation seems restricted to catalytic site residues, showing mosaic features of the UNG and MUG enzymes. Crystallographic analysis of SMUG1 identified a pyrimidine binding pocket topologically similar to other UDGs and implicated a water displacement/replacement mechanism to account for the enzyme's preference for uracil over thymine (Wibley et al. 2003). Like TDG, SMUG1 is active on 5-FU but, unlike TDG, appears to protect cells from the cytotoxic effects of the drug as shown in siRNA knockdown experiments (An et al. 2007). Notably, a 5-hmU DNA glycosylase activity originally discovered in calf thymus was later identified as SMUG1 (Cannon-Carlson et al. 1989; Boorstein et al. 2001). So, like TDG, SMUG1 is capable of processing the deamination product of 5-hydroxymethylcytosine (5-hmC), a substrate that has recently gained attention in the context of active DNA demethylation.
Additional subfamilies of UDGs appear to have evolved in archaeal and bacterial organisms thriving under extreme environmental conditions such as high temperature, favoring hydrolytic deamination of cytosine and 5-meC. As these will not be further discussed here, the reader is referred to the excellent classification of the UDG superfamily originally published by Aravind and Koonin (2000).
The helix-hairpin-helix glycosylases
The second superfamily of DNA glycosylases, characterized by a shared helix-hairpin-helix (HhH) domain, comprises a diverse group of enzymes present in organisms throughout all kingdoms of life. Phylogenetic analysis in 94 genomes from bacteria, archaea, and eukaryotes identified six distinct families of HhH DNA glycosylases: Nth (homologs of the E. coli EndoIII protein), OggI (8-oxoG DNA glycosylase I), MutY/Mig (A/G-mismatch-specific adenine glycosylase), AlkA (alkyladenine-DNA glycosylase), MpgII (N-methylpurine-DNA glycosylase II), and OggII (8-oxoG DNA glycosylase II) (Denver et al. 2003). The Nth and MutY/Mig family glycosylases as well as some of the MpgII type proteins contain iron–sulfur [4Fe4S] clusters that are thought to play a structural role in DNA binding and substrate recognition (Cunningham et al. 1989; Kuo et al. 1992; Guan et al. 1998; Porello et al. 1998; Begley et al. 1999).
The founding member of the Nth family was originally discovered as an endonuclease activity (EndoIII) in E. coli (Radman 1976) but then turned out to be a DNA glycosylase with an associated AP-lyase activity. Nth proteins appear to be the most highly conserved subfamily within the HhH glycosylases (Denver et al. 2003). The mammalian homolog, NTHL1 (endonuclease III-like 1), acts on ring fragmented purines or oxidized pyrimidine residues like thymine glycol (Tg), formamidopyrimidine (FaPy), 5-hydroxycytosine (5-hC), and 5-hydroxyuracil (5-hU), preferentially when placed opposite guanine (Table 1) (Dizdaroglu et al. 1999; Eide et al. 2001). Nth1 knockout mice show no overt abnormalities presumably because the loss of its repair function can be compensated for by NEIL glycosylases (see below) (Ocampo et al. 2002; Takao et al. 2002).
MutY was first identified in E. coli as an enzyme excising adenine from A•G mispairs (Au et al. 1988), while Mig.Mth was discovered in hyperthermophilic archaea by its ability to excise U or T mispaired with G (Horst and Fritz 1996). MutY/Mig homologs are widespread in bacterial genomes, but only about half of the eukaryotes and less than a third of the archaeal species analyzed have glycosylases of this family (Denver et al. 2003). The mammalian homolog of MutY, termed MYH, excises adenine opposite 8-oxoG, guanine, or cytosine (Table 1) (McGoldrick et al. 1995), contributing to a multimodal defense against the mutability of guanine oxidation (van Loon et al. 2010). Facilitating the replacement of A opposite 8-oxoG with a C, MYH produces the preferred substrate for the 8-oxoG directed DNA glycosylase OGG1. Hence, disruption of Myh in mice does not produce a mutator phenotype per se because it is masked by OGG1, which corrects the bulk of oxidized guanines before replicative DNA polymerases get a chance to misinsert adenine opposite the damaged base. Knocking out both Myh and Ogg1, however, results in a synergistic increase in G → T mutations (Russo et al. 2004). In humans, germline mutations in the MYH gene have been associated with a predisposition to colorectal cancer (Al-Tassan et al. 2002; Jones et al. 2002).
The Ogg1 protein family is less well represented across the phylogeny. While present in most eukaryotic genomes, Ogg1 encoding genes seem to be missing in bacteria and archaea (Denver et al. 2003). Ogg1 was originally discovered in yeast and later also identified in mammals (Nash et al. 1996; Lu et al. 1997; van der Kemp et al. 1996; Radicella et al. 1997), where it provides the major activity for the removal of 8-oxoG opposite cytosine (Friedberg et al. 2006). It does, however, also excise other oxidized pyrimidines or ring-fragmented purines like formamidopyrimidine (FaPy) (Table 1) (Dherin et al. 1999; Karahalil et al. 1998). In Saccharomyces cerevisiae, inactivation of OGG1 results in an accumulation of G → T transversion mutations (Thomas et al. 1997). Ogg1 null mice are viable but exhibit a 2-fold increase in chromosomal 8-oxoG and moderately elevated spontaneous mutation frequencies (Klungland et al. 1999). Polymorphisms in the human OGG1 gene impairing the 8-oxoG incision activity were found to be associated with non-small cell lung cancer (Janik et al. 2011) and an increased risk of childhood acute lymphoblastic leukemia (Stanczyk et al. 2011).
Although closely related to the Nth and MutY proteins, MBD4, also known as MED1, is special in two ways; it has a methyl-CpG binding domain (MBD) and is therefore also a member of the MBD protein family, and it functionally interacts with MLH1, a protein of the postreplicative mismatch repair system (Hendrich and Bird 1998; Bellacosa et al. 1999). Like the structurally unrelated TDG, MBD4 is a mismatch-directed DNA glycosylase processing a wide range of G-mispaired base lesions, including thymine, uracil, 5-FU, and 3,N4-ethenocytosine (Table 1) (Petronzelli et al. 2000; Cortellino et al. 2003). Its methyl-CpG binding domain in addition to its activity on the deamination product of 5-meC has made MBD4 a prime candidate for an active DNA demethylase, an epigenetic function likely to be important in embryogenesis. However, Mbd4 knockout mice show no developmental defects, but a mild increase in C → T mutation frequency and a predisposition to gastrointestinal cancer in APC-deficient tumor models (Millar et al. 2002; Wong et al. 2002), consistent with a role of MBD4 in repair of cytosine or 5-meC deamination damage.
E. coli AlkA, the founding member of the AlkA family of HhH glycosylases, acts on alkylated bases, e.g., 3-meA. While homologs are present in many bacterial and eukaryotic genomes (Denver et al. 2003), mammals appear to be devoid of this particular class of enzyme. Instead, they use a structurally unrelated enzyme, MPG, to eliminate specific forms of base alkylation damage.
The 3-methyl-purine glycosylase (MPG)
MPG, also known as AAG or MDG, originally identified in rat (O'Connor and Laval 1990) and later in human, is a DNA glycosylase excising a range of alkylated bases from DNA, including 3-meA, 7-meG, 3-methylguanine (3-meG) as well as ethylated bases in single- and double-stranded DNA (Table 1) (O'Connor 1993; Lee et al. 2009). MPGs form a structurally distinct class of glycosylases; they lack helix-hairpin-helix motifs nor do they have an alpha–beta fold structure characteristic of UDGs. Mice lacking MPG are viable and show a mild increase in the frequency of spontaneous mutation (Engelward et al. 1997; Hang et al. 1997), and they are more prone to develop azoxymethane-induced colon cancer than their wild-type counterparts (Wirtz et al. 2010). Reminiscent of TDG’s role in the DNA-directed cytotoxic effect of 5-FU, MPG drives alkylation-induced retinal degeneration in mice by generating cytotoxic BER intermediates (Meira et al. 2009).
The endonuclease VIII-like glycosylases
Although their substrate spectrum overlaps with that of endonuclease III (Nth), the homologs of E. coli endonuclease VIII, encoded by the nei gene, are structurally related to the formamidopyrimidine-DNA glycosylase Fpg and form a separate family of DNA glycosylases. Nei was discovered in E. coli as a second activity next to Nth acting on thymine glycol (Tg) and urea (Melamede et al. 1994).
The mammalian counterparts are termed Nei-like (NEIL) 1, 2, and 3, and share a conserved helix-two-turn-helix motif with the E. coli Fpg and EndoVIII proteins. The preferred substrates of NEIL1 and NEIL2 are oxidized pyrimidines such as Tg, 5-hC, FaPyA, and FaPyG (Hazra et al. 2002; Morland et al. 2002; Rosenquist et al. 2003), but also 5-hydroxyuracil (5-hU) and 8-oxoG in DNA bubble structures (Table 1) (Dou et al. 2003). NEIL3, on the other hand, excises FaPy but is inactive on 8-oxoG (Liu et al. 2010). Mice with a targeted inactivation of the Neil1 gene exhibit a phenotype reminiscent of the metabolic syndrome, as well as increased levels of DNA damage in mitochondrial DNA (Vartanian et al. 2006). Neil3 knockout mice are viable and fertile, and the protein appears to be preferentially expressed in hematopoietic tissues (Torisu et al. 2005), implicating a possible function in hematopoiesis or the immune system.
Structure function aspects
DNA glycosylases evolved to counter the many different forms of chemical damage occurring to DNA bases. They are highly specialized enzymes with distinct structures and substrate specificities, but they all share a common principle of action. DNA glycosylases recognize their cognate substrates by rotating bases out of the DNA helix into a specifically fitting pocket that harbors the active site. Substrate selectivity is mostly achieved by steric exclusion from the binding pocket of bases that are not to be processed, and to some extent also by the catalytic efficiency of the active site configuration. If base fitting is successful, cleavage of the N-glycosidic bond will occur, resulting in the release of a free base and the generation of a base-less sugar, an AP-site in the DNA.
Detection and verification of base damage
Detecting a single damaged base in a vast excess of regular bases in the genome amounts to the proverbial task of searching a needle in a haystack, and this is not helped by the fact that the lesions addressed by BER do not usually cause notable distortions to the DNA helix. A human cell suffers about 104 base lesions per day, translating into roughly one lesion every 10 s within a genome of about 14 billion nucleotides that must be spotted and repaired. How DNA glycosylases manage to efficiently search for and recognize these lesions is not clear but biochemical and structural work has provided some insight into possible mechanisms.
Scanning the genome for damaged bases
Little is known about how DNA glycosylases find damaged bases in the genome. One idea, proposed on the basis of biochemical evidence and theoretical considerations, postulates the association of the glycosylase with undamaged DNA by non-specific interactions, facilitating sliding along the DNA duplex for a certain distance and scanning the sequence for irregular bases (Berg et al. 1981). Considering the structural and functional diversity of DNA glycosylases, however, there are likely to be different translocation mechanisms, variably involving features of tracking, diffusion, and hopping on the DNA (Blainey et al. 2009; Steinacher and Schär 2005; Friedman and Stivers 2010). Recently, an appealing DNA scanning concept was proposed for DNA glycosylases harboring an [4Fe4S] cluster. The underlying observation was that E. coli MutY and Nth change the oxidation state of their iron–sulfur cluster from [4Fe4S]2+ to [4Fe4S]3+ upon contact with DNA, which stabilizes the interaction. Thus, if such DNA glycosylases bind in the vicinity of each other, they might act as electron donors and acceptors for each other, making use of the DNA for charge transfer. This may facilitate the dissociation of one glycosylase upon binding of another by reduction of its [4Fe4S] cluster. If the electron transfer involved is perturbed by a base lesion between the two glycosylases, however, both will stay bound to the DNA, increasing the likelihood of damage detection (Boal et al. 2009). This way, [4Fe4S] clusters may support the search for base damage without a need of scanning the entire DNA sequence.
Detection of DNA base damage by DNA glycosylases ultimately requires a full examination of the chemical surface of single bases. To minimize the effort, DNA glycosylases employ strategies of damage pre-selection. OGG1 and UNG, for instance, were proposed to pre-select substrates by establishing superficial base contacts through conserved residues in close proximity to the mouth of their catalytic pocket. This allows potential substrates to be identified without fully inserting every base into the active site pocket (Fig. 2) (Banerjee et al. 2005; Parker et al. 2007). Consistently, NMR studies on human UNG showed the glycosylase to undergo a conformational change upon DNA binding, allowing for oscillation between an open form, loosely interacting with the DNA in an unspecific manner, and a closed form engaged in base examination without fully rotating the base out of the helix (Fig. 2) (Friedman et al. 2009). Base pre-scanning is likely to be facilitated by DNA breathing, considering that many base lesions affect base pairing dynamics to some extent and are therefore likely to enhance local DNA melting.
Mechanism of base removal by monofunctional and bifunctional glycosylases. While all DNA glycosylases share a common principle of action, they differ in details of damage search, recognition, and excision. Among the monofunctional DNA glycosylases, UNG searches for uracil in DNA by oscillating between an open conformation, making unspecific contacts with the DNA, and a closed conformation for base-interrogation (a), involving conserved structures at the mouth of the catalytic site pocket (b). Pre-selected bases are flipped out of the DNA duplex and inserted into the catalytic pocket, where specific hydrogen bonds align the base for nucleophilic attack by an activated water molecule strategically positioned by a conserved aspartic acid residue (red asterisk, c). TDG differs from UNG by its non-specific interaction with DNA through a clamp formed by the N-terminal domain (a), its larger catalytic pocket that accommodates a broader range of substrates, and its ability to involve the opposing base in lesion recognition (b). The catalytic residue in TDG is an asparagine (black asterisk) that positions, but does not activate, a water molecule which can then act as a weak nucleophile (c). TDG stays firmly bound to the abasic site upon base release (d) until SUMOylation induces dissociation by neutralizing the N-terminal DNA binding activity (e). Similar to UNG, the bifunctional OGG1 pre-selects bases for flipping by interactions with conserved amino-acid residues at the mouth of the catalytic cavity (a, b), and similar to TDG, it also takes the opposing base into account for substrate selection (c). Bifunctional glycosylases use a conserved amino-acid residue (blue asterisk) for nucleophilic attack, which results in a covalent intermediate between the glycosylase and the DNA substrate (d). Resolution of this intermediate produces a DNA nick that is further processed by APE1 (e)
Formation of a mature enzyme substrate complex
For final damage verification, the base needs to be flipped out of the DNA helix and accommodated in the active site cavity of the glycosylase. This increases the surface for molecular interactions, providing for a sensitive discrimination of even minor base alterations and, ultimately, catalysis of the base release. One of the best-studied glycosylases with regard to damage recognition is UNG. On its surface, UNG forms a positively charged groove that accommodates the minor groove of the DNA and contains the active site cavity. Upon encountering a uracil in DNA, UNG uses a hydrophobic side chain of a conserved leucine to push the uracil out of the base stack and insert it into the catalytic pocket (Fig. 3). The same residue stabilizes the double helix by occupying the vacated space (Mol et al. 1995; Savva et al. 1995). At the same time, UNG pinches and compresses the double-stranded DNA backbone, thereby inducing DNA bending by about 45° at the site of the damage (Fig. 1) (Parikh et al. 1998). Selectivity for uracil is mediated by several structural features: purine bases are sterically excluded from the active site by the overall narrow geometry of the binding pocket, while the entry of thymine and other 5-methylated pyrimidines is blocked by a side chain of a conserved tyrosine residue. Accordingly, UNG can be converted into a thymine-processing enzyme by an amino acid substitution that eliminates this sterical block (Kavli et al. 1996). Cytosine can enter the active site but, due to unfavorable hydrogen bonding at the bottom of the cavity, cannot be positioned correctly for catalysis (Kavli et al. 1996; Slupphaug et al. 1996).
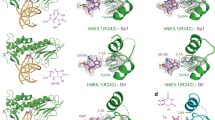
The base-flipping intermediate captured in a crystal structure of substrate bound human UNG. The UNG double mutant L272R/D145N, stabilizing the glycosylase–substrate complex, was co-crystallized with an oligonucleotide bearing a U•G mismatch. Uracil (red) is flipped into an extrahelical position (a) and inserted deeply into the tight fitting active site pocket of UNG (b), where it is positioned by specific molecular interactions for the nucleophilic attack on the N-glycosidic bond (Slupphaug et al. 1996)
While UNG is highly specialized to the excision of DNA uracil, TDG is a good example of a broad spectrum UDG showing mismatch dependency. Unlike UNG, the bacterial and human TDG orthologs have rather spacious pyrimidine-binding pockets, accommodating a large variety of base derivatives (Barrett et al. 1998, 1999; Baba et al. 2005; Maiti et al. 2008), although the preferred substrates are G-mismatched deaminated pyrimidines (Waters and Swann 1998; Hardeland et al. 2003). Because TDG acts on thymine, it needs to be able to discriminate between a regular thymine in DNA (A•T) and one that resulted from deamination of 5-meC (G•T). Substrate selection thus has to take into account not only the base structure itself but also the opposite base. Structural and biochemical studies of E. coli Mug and human TDG (Barrett et al. 1998; Maiti et al. 2008, 2009) have shed light on how this might be achieved. In contact with DNA, TDG undergoes a conformational change in its N-terminal domain, forming a clamp-like structure that permits TDG to track along the DNA (Fig. 2) (Steinacher and Schär 2005). Upon encountering a G-mismatched substrate (G•T), the substrate base is pushed out of the DNA helix by an insertion loop wedging into the DNA helix. This same wedge stabilizes the base stack and forms specific hydrogen bonds with the widowed guanine, mimicking Watson–Crick base pairing (Barrett et al. 1998, 1999; Maiti et al. 2008). These interactions then cooperate with the non-specific DNA binding activity of the N terminus to stabilize the TDG–substrate complex for efficient base excision (Hardeland et al. 2000; Steinacher and Schär 2005).
Another DNA glycosylase well studied with regard to damage recognition is OGG1. Following successful pre-selection of a potential 8-oxoG substrate (Fig. 2) (Banerjee et al. 2005), the oxidized base is flipped into the active site cavity of OGG1, inducing DNA bending of about ~70° due to a tyrosine residue that wedges between the opposing C and its 5′ neighbor. Unstacking of the widowed C facilitates enzyme–DNA contacts that maintain opposite base selectivity. The void generated by 8-oxoG rotation is occupied by a conserved asparagine residue, which contributes to hydrogen bonding with the opposing cytosine. While A, C, and T are excluded from the active site pocket by several strategically positioned amino-acids, a conserved glycine is the only residue discriminating between G and 8-oxoG (Bruner et al. 2000).
Catalysis of base removal
The catalytic mechanism subdivides DNA glycosylases into monofunctional and bifunctional enzymes (Table 1). Monofunctional glycosylases perform base excision only, using an activated water molecule for nucleophilic attack on the N-glycosidic bond, while bifunctional glycosylases use an amino group of a lysine side chain for the same purpose, forming a Schiff-base intermediate, and subsequently cleave the DNA backbone 3′ to the lesion.
Monofunctional DNA glycosylases
To illustrate the monofunctional mode of action, we will focus on UNG and TDG as two well-studied UDGs showing contrasting catalytic features. Upon recognition, uracil is tightly fitted into the active site pocket of UNG (Fig. 3). Interactions between the uracil and amino-acid residues at the bottom of the pocket position the N-glycosidic bond for hydrolysis (Mol et al. 1995; Savva et al. 1995; Slupphaug et al. 1996). Structure and mutational analyses have established a catalytic mechanism involving the polarization of the N-glycosidic bond by a conserved histidine to make it susceptible for nucleophilic attack, and the positioning and deprotonation of a water molecule by a conserved catalytic aspartate, which then attacks the C1 of the deoxyribose (Fig. 2) (Mol et al. 1995; Slupphaug et al. 1996). N-glycosidic bond cleavage is completed by addition of the water nucleophile to uracil, resulting in a free base and an AP-site.
The mismatch-specific uracil glycosylases, e.g., Mug and TDG, interact less specifically with the substrate base within the catalytic pocket and use a less potent mechanism of catalysis, as first revealed in the crystal structure of E. coli Mug (Barrett et al. 1998, 1999). In place of the catalytic aspartate in UNG, Mug/TDG enzymes have an asparagine. This asparagine positions a water molecule but, unlike the aspartate in UNG, is unable to protonate it for an efficient nucleophilic attack on the N-glycosidic bond (Fig. 2). Also, an appropriate residue for polarization of the N-glycosidic bond appears to be missing in Mug/TDG, altogether explaining the comparably low catalytic efficiency of the MUG enzymes (Hardeland et al. 2000; Maiti et al. 2009). These differences illustrate that the mode of catalysis can vary considerably even within one DNA glycosylase superfamily, most likely reflecting the requirement to fine tune substrate spectrum and catalytic efficiency in the evolution of subfamilies with distinct biological functions.
Bifunctional DNA glycosylases
DNA glycosylases that use an amino group of a lysine side chain as a nucleophile for base cleavage form a covalent Schiff-base intermediate with the substrate. The resolution of this reaction intermediate incises the DNA 3′ to the product AP-site, generating a strand break with 3′ phosphate and 5′ OH ends. Thus, these enzymes couple base excision with an AP-lyase step, as best illustrated by the bifunctional mechanism proposed for OGG1 (Sun et al. 1995; Nash et al. 1997). Once an 8-oxoG is stably fitted in the active site cavity of OGG1, the side chain amino group of a suitably positioned catalytic lysine is activated as a nucleophile to attack the C1 of the deoxyribose. The resulting rearrangement to a covalent Schiff-base intermediate releases the damaged base from the DNA, which is then held in position to further participate in the catalysis of the β-lyase reaction, cleaving the DNA strand at the 3′ phosphate. Notably, all steps of the OGG1 catalyzed base release involve the transfer of protons and this is promoted by the excised base itself (Bruner et al. 2000; Fromme et al. 2003). An interesting variation on the bifunctional mode of action is displayed by the mammalian NEIL proteins and their E. coli counterpart Nei. These enzymes couple base excision to beta and delta elimination, incising the DNA strand both 3′ and 5′ to the AP-site (Takao et al. 2002). Moreover, unlike OGG1, E. coli Nei ejects the excised base from the catalytic site immediately, i.e., is capable to perform the AP-lyase reaction without contribution of the damaged base (Zharkov et al. 2002).
AP-site dissociation and turnover of glycosylases
Upon base release, DNA glycosylases tend to stay bound to the product of their action, the AP-site. In fact, many glycosylases display a higher affinity to AP-sites than to their actual substrate base (Parikh et al. 1998; Waters et al. 1999; Hardeland et al. 2000; Hill et al. 2001; Pope et al. 2002). The release of the glycosylase from the AP-site is thus rate limiting in the BER process (Fig. 1). Since AP-sites are chemically unstable and lack base coding potential, the binding of the glycosylase might serve to protect cells against their cytotoxic and mutagenic effects. This considered, it appears reasonable that the release of the AP-site is coordinated with the recruitment and assembly of the downstream acting BER factors. Consistently, the AP-endonuclease APE1 was reported to stimulate the turnover of several DNA glycosylases including UNG2, TDG, and OGG1 (Parikh et al. 1998; Waters et al. 1999; Hill et al. 2001), and a similar effect was observed for the XPC protein on TDG and SMUG1 (Shimizu et al. 2003, 2010). It remains to be clarified, though, to what extent these stimulatory effects reflect active processes or simply a passive competition for a common DNA substrate.
Another, perhaps more sophisticated, way to regulate AP-site interaction of the glycosylase is by posttranslational modification. In this direction, UNG2 was reported to be cell cycle specifically phosphorylated at serine 23, which markedly increases its association with replicating chromatin but also its turnover rate, apparently to facilitate efficient correction of misincorporated uracil during ongoing DNA replication (Hagen et al. 2008). In the case of TDG, which binds AP-sites very rigidly, dissociation is regulated by modification of a C-terminal lysine residue with small ubiquitin-like modifiers (SUMOs). This induces a conformational change that neutralizes the non-specific DNA-binding capacity of its N-terminal domain, thereby facilitating AP-site dissociation (Fig. 2) (Hardeland et al. 2002; Steinacher and Schär 2005). SUMO modification might be triggered by the presence of downstream acting BER factors, providing for a controlled handover of the AP-site intermediate in the repair process (R. Steinacher and P. Schär, unpublished data).
It appears to be a general feature of DNA glycosylases to bend DNA upon establishing a mature enzyme–substrate complex; UNG for example induces a bend of ~45° (Parikh et al. 1998), OGG1 of even ~70° (Bruner et al. 2000). On the basis of these and similar observations with other BER factors, it was proposed that the DNA bending might serve as a structural determinant to orchestrate the handover from one step in the repair process to the next (Fig. 1) (Parikh et al. 1999; Wilson and Kunkel 2000).
Functions of DNA glycosylases beyond DNA repair
Although DNA glycosylases are optimally suited for the repair of damaged DNA bases, their structural and biochemical properties would support a much wider spectrum of genetic functions. The ability to recognize and excise chemically modified bases can be used to edit the DNA at specifically marked sites. The UDG superfamily in particular appears to comprise enzymes with specialized functions, e.g., in innate immunity and antibody diversification, as well as in the regulation of gene expression and epigenetic maintenance.
DNA glycosylases in immunity
UDGs haven been functionally associated with mechanisms providing innate immunity against viral infection as well as antibody diversity in the adaptive immune system. All these processes are triggered by enzymatic deamination of cytosine by members of the apolipoprotein B mRNA editing catalytic polypeptide (APOBEC) family of proteins, including the activation-induced cytidine deaminase (AID) (Conticello et al. 2005).
DNA glycosylases providing innate immunity
Proteins of the human APOBEC3 subfamily were shown to inhibit replication of a variety of retroviruses, including human immunodeficiency viruses (HIV) (reviewed in Holmes et al. 2007). APOBEC3G was discovered by its ability to restrict replication of a mutant HIV-1 lacking the viral infectivity factor (Vif) (Sheehy et al. 2003). Cells infected with such a virus package APOBEC3G into HIV-1 virions. When these infect new cells, APOBEC3G will deaminate multiple cytosines in the viral cDNA during reverse transcription, which inactivates the provirus (reviewed in Holmes et al. 2007). While APOBEC3G appears to intervene with the viral life cycle at several steps, the antiviral activity mediated by uracilation of the viral genome is coupled to the action of UNG2 and APE1. UNG2 is thought to introduce AP-sites into the deaminated viral cDNA, triggering the cleavage and thus degradation by APE1 (Harris et al. 2003; Yang et al. 2007). Consistently, inhibition and/or downregulation of either UNG2 or APE1 was shown to decrease the antiviral potency of APOBEC3G (Yang et al. 2007). Given that HIV-1 evolved Vpr, a small protein specifically targeting UNG2 and SMUG1 for degradation by the ubiquitin–proteasome system, supports a critical role of uracil base excision in antiviral defense (Schrofelbauer et al. 2005, 2007).
DNA glycosylases providing adaptive immunity
The cooperation of cytidine deaminase and UDG activities is also central to the genetic transactions associated with antibody diversification in the adaptive immune system, i.e., somatic hypermutation (SHM) and class switch recombination (CSR) (Fig. 4) (Muramatsu et al. 2000; Rada et al. 2002).
Uracil DNA glycosylase in antibody diversification. UNG plays a central role in somatic hypermutation (SHM) and class switch recombination (CSR). UNG2 is targeted to immunoglobulin loci in activated B-cells by AID converting cytosine to uracil in single-stranded DNA during transcription. While replication across unprocessed uracil itself will generate C → T mutations, uracil excision by UNG provides for a wider range of mutations both at the C•G and nearby A•T base pairs; (1) by generating non-instructive AP-sites which, upon DNA replication give rise to transition or transversion mutations, (2) by initiating long-patch BER or, (3) following cleavage by APE1, by providing a DNA strand nick for activation of MMR. Error-prone synthesis associated with long-patch BER and MMR would then produce mutations at A•T base pairs. Similarly, DNA strand breaks occurring through BER of deaminated cytosines in switch regions of immunoglobulin heavy chain loci may induce CSR (blue panel)
SHM introduces point mutations in the light chain variable (V) region of immunoglobulin loci. Mutagenesis is initiated by AID, which is targeted to these loci to deaminate multiple cytosines in single-stranded DNA occurring during transcription (reviewed in Pavri and Nussenzweig 2011). Mutations can arise from these U•G mismatches in several ways (Fig. 4): (1) in the absence of uracil excision, replication across the U•G mismatch would generate a C → T mutation in one of the daughter strands; (2) uracil excision by a UDG and subsequent replication across the resulting AP-site can potentially give rise to any type of base substitution at the site of deamination, although replicative DNA polymerases preferentially insert A opposite an AP-site; (3) long-patch BER, initiated by a UDG, or MMR coupled to error-prone DNA synthesis would cause mutations in proximity to the deaminated cytosine, allowing for mutations to occur also at A•T base pairs.
An involvement of UNG2 in antibody diversification was first implicated by a general perturbance of both SHM and CSR in an UNG2 inhibited chicken B-cell line as well as in UNG-deficient mice (Di Noia and Neuberger 2002; Rada et al. 2002). While the loss of UNG mainly affected SHM at G•C base pairs, inactivation of the mismatch repair system (MMR) was found to diminish hypermutation at A•T pairs (Rada et al. 1998; Wiesendanger et al. 2000). This suggested that error prone MMR contributes to mutagenesis at sites away from the deaminated cytosine (Wilson et al. 2005). Later work then implicated that MMR can indeed operate at an AID induced G•U mismatch if assisted by UNG2, providing a nick at a nearby G•U for initiation of strand excision (Frieder et al. 2009; Schanz et al. 2009).
AID, UNG2, and APE1 were also shown to play a crucial role in the initiation of CSR, a specialized recombination process switching the Ig isotype of an antibody without affecting its antigen specificity. CSR rearranges the antibody constant region of heavy chain genes (CH) by strand breakage in and joining of two selected switch (S) regions located upstream of every CH-coding segment. The DNA strand breaks required for initiation of CSR were shown to arise through cytosine deamination, subsequent uracil excision, and AP-site cleavage by AID, UNG2, and an AP-endonuclease, respectively (Fig. 4) (Imai et al. 2003). Consistently, murine B-cells lacking UNG2 show a severe reduction of CSR and in humans, recessive mutations in the UNG gene have been associated with the hyper-IgM syndrome caused by a deficiency in CSR (Rada et al. 2002; Imai et al. 2003). It is also reported, however, that the catalytic activity of UNG2 is dispensable for efficient CSR, while an N-terminal sequence motif appears to be important, suggesting that UNG2 is not responsible for the DNA cleavage step of CSR (Begum et al. 2007, 2009). It is possible that UNG2 simply marks AID-induced G•U mismatches for further processing by other factors, whereby the N terminus is required to mediate specific protein–protein interactions.
Residual CSR and SHM in UNG2-deficient mice hint at a possible contribution of other UDGs. Indeed, overexpression of SMUG1 in MSH2 UNG2 double-deficient cells could partially restore SHM and CSR. A biological role of SMUG1 in antibody diversification, however, is questionable as it is downregulated upon B-cell activation (Di Noia et al. 2006). Likewise, the U•G mismatch-directed glycosylase MBD4 is unlikely to play a major role in either SHM or CSR as a knockout of the gene in mouse showed no effect on either processes (Bardwell et al. 2003). The situation is less clear for TDG. While the impact of a TDG-deficiency on antibody diversification remains to be investigated, its upregulation in activated mouse B-cells hints at a specific function in B-cell maturation (Christophe Kunz and Primo Schär, unpublished data). In this context, TDG might be simply required to prevent mutations at non-Ig genes arising from mistargeted AID. It might, however, also directly contribute to SHM and CSR. Considering its tight interaction with AP-sites, TDG would be optimally suited to delay processing of these repair intermediates, thereby favoring error-prone translesion synthesis and recombination repair.
DNA glycosylases in DNA methylation control
Distinct patterns of DNA methylation and histone modifications are established during cell lineage restriction to determine and maintain cell-type-specific gene expression programs. In mammals, DNA methylation occurs mainly in the form of 5-meC in CpG dinucleotide sequences and is controlled by a methylation machinery consisting of both methylating and demethylating components. While the de novo establishment and the maintenance of CpG methylation can be rationalized by the biochemical features of the DNA methyltransferase (DNMT) activities involved (Hermann et al. 2004), the reverse process of demethylation has remained elusive. In principle, however, DNA demethylation can be achieved through active and/or passive mechanisms. Whereas passive demethylation occurs upon DNA replication with downregulation or inhibition of the maintenance DNA methyltransferase, active demethylation describes the enzymatic removal of 5-meC and replacement with C. Active mechanisms have been implicated in several important biological processes, including the demethylation of the paternal pronucleus in the murine zygote (Mayer et al. 2000; Oswald et al. 2000), the genome-wide methylation erasure and reset of parental imprinting during gametogenesis (Monk et al. 1987; Kafri et al. 1992), and the reprogramming of methylation patterns observed after transfer of somatic cell nuclei to enucleated oocytes (Dean et al. 2001; Simonsson and Gurdon 2004). Active demethylation has also been reported to be targeted to select gene regulatory regions during neurogenesis, memory formation, and immune response (Bruniquel and Schwartz 2003; Miller and Sweatt 2007; Ma et al. 2009).
Mechanistically, the conversion of 5-meC to C in DNA can occur in different ways: (1) by a direct removal of the methyl group, (2) the replacement of 5-meC with C by excision repair, or (3) the deamination or oxidation of 5-meC followed by replacement of the nucleotide by BER. Consistent with an excision repair scenario, an increasing number of observations point at an engagement of DNA glycosylases in active demethylation. In plants, the removal of 5-meC by at least four bifunctional DNA glycosylases is well established; ROS1, DEMETER (DME), and the DME-like (DML) 2 and 3 all process 5-meC in CpG and non-CpG sequence contexts (reviewed in Zhu 2009). Mutations in these glycosylases affect cytosine methylation at specific loci but not in the overall genome, suggesting that they act in a targeted rather than a global manner (Penterman et al. 2007; Lister et al. 2008). In vertebrates, the situation is less clear; MBD4 and TDG have been implicated in DNA demethylation but they do not seem to be potent enough to excise 5-meC directly.
Concepts of DNA glycosylase-mediated demethylation in vertebrates
First evidence for an involvement of DNA glycosylases in active demethylation in vertebrates came with the discovery of a 5-meC DNA glycosylase activity in extracts of chicken embryos (Jost et al. 1995). The responsible enzyme later revealed itself as a homolog of the human TDG (Zhu et al. 2000b). 5-meC DNA glycosylase activity was then also reported for the human TDG and MBD4 proteins by the same group (Zhu et al. 2000a, 2001). Yet, as several attempts to reproduce these finding were unsuccessful, the idea of TDG and MBD4 acting as demethylating glycosylases did not achieve broad acceptance. Recent observations, however, indicate that the difficulty in reproduction might lie in the necessity of unknown co-factors that either boost the catalytic efficiency of these glycosylases or convert 5-meC into a more favorable substrate.
MBD4 was recently reported to control CpG methylation in the context of parathyroid (PTH) hormone-induced gene activation. This was shown for the CYP27B1 promoter, which undergoes active demethylation upon hormone stimulation (Kim et al. 2009). Both promoter activation and cytosine demethylation coincided with and depended on the physical association of MBD4 and downstream BER factors, but not of the functionally related TDG. Remarkably, this study further showed that phosphorylation of MBD4 by PKC may potentiate its activity to process 5-meC, suggesting that posttranslational modification might be required to unleash a potentially harmful but dormant 5-meC glycosylase activity in certain DNA glycosylases for targeted demethylation under specific conditions (Fig. 5a). This might apply as well to TDG, given its propensity to posttranslational modification by SUMOylation (Hardeland et al. 2002), ubiquitylation (Hardeland et al. 2007), phosphorylation (Um et al. 1998), and acetylation (Tini et al. 2002). It will thus be necessary to revisit TDG's activity as a 5-meC glycosylase under conditions that support the formation of these posttranslational modifications.
Possible pathways of active DNA demethylation involving BER. Enzymatic removal of 5-meC has been suggested to be accomplished by different DNA glycosylase mediated mechanisms. Direct excision of 5-meC by mammalian DNA glycosylases has been tested with contradicting results, suggesting the possibility that posttranslational modification might induce a shift in the substrate spectrum, allowing for the removal of 5-meC. (a) Another possibility is the enzymatic conversion of 5-meC to substrates more favorable to DNA glycosylases, such as deamination to thymine (b), hydroxylation to 5-hydroxymethylcytosine (c) and/or further oxidation or deamination of 5-hmC to 5-formylcytosine (5-fC) and 5-carboxylcytosine (5-caC) (d) or 5-hydroxymethyluracil (e), respectively. All these processes could be accomplished by a cooperation of 5-meC deaminases like AID, hydroxylases like the TET proteins, and DNA glycosylases like TDG (G•T, G•5hmU, G•fC, G•caC), MBD4 (G•T, ?) and possibly SMUG1 (G•T, G•5hmU, ?)
Other lines of investigation support demethylation scenarios that involve the conversion of 5-meC to more favorable substrates for DNA glycosylases. Deamination of 5-meC by a cytidine deaminase is one possibility. This would generate a G•T mismatch that can be acted on by MBD4 or TDG (Fig. 5b). Along these lines, it was shown that the estrogen responsive pS2/TFF1 promoter undergoes cyclic DNA methylation and demethylation during transcriptional activation, involving 5-meC deamination by the methyltransferase Dnmt3a/b itself and subsequent TDG-dependent BER (Metivier et al. 2008; Kangaspeska et al. 2008). The demethylation complex also includes the RNA helicase p68, implicating an RNA component in either targeting and/or stabilization of the complex, consistent with previously reported observations on the chicken 5-methylcytosine DNA glycosylase (Jost et al. 1997; Schwarz et al. 2000). Additional support for a deamination-coupled demethylation pathway came from a study in zebrafish embryos, implicating AID in the deamination of 5-meC and MBD4 in the excision of the resulting G•T mismatch, as well as Gadd45 as an auxiliary factor (Rai et al. 2008). The same concept was adopted more recently to explain TDG-mediated 5-meC demethylation as it may occur during somatic differentiation of the developing mouse embryo (Cortellino et al. 2011). This, however, was mainly inferred from the co-immunoprecipitation of overexpressed AID and Gadd45 with TDG and therefore needs to be corroborated by more direct functional evidence. Finally, an AID-dependent and, thus, deamination-mediated mechanism has been associated with global DNA demethylation occurring in primordial germ cells as well as during the reprogramming of somatic cell nuclei towards pluripotency (Bhutani et al. 2010; Popp et al. 2010). Obviously, genome-wide 5-meC deamination would generate massive amounts of G•T mismatches, and all of these would have to be repaired by the G•T-directed DNA glycosylases MBD4 or TDG. A strong prediction of a deamination-based demethylation model is therefore that a failure of G•T repair would result in genome-wide C → T mutagenesis. This has not been observed so far in MBD4 and/or TDG-deficient cells (Cortazar et al. 2011) and will have to be tested more carefully.
The recent discovery of DNA dioxygenases acting on 5-meC in DNA introduced another conceptual framework for active demethylation. The principle of oxidative demethylation of DNA bases was first described for the bacterial DNA repair protein AlkB. AlkB belongs to a large superfamily of Fe(II)/2-oxoglutarate (2-OG)-dependent hydroxylases and catalyzes the hydroxylation of N-methylated bases like 1-methyladenine (1-meA) and 3-methylcytosine (3-meC), ultimately resulting in demethylation upon release of the hydroxymethyl moiety as formaldehyde (Falnes et al. 2002; Trewick et al. 2002). The superfamily of Fe(II)/2-OG hydroxylases also contains the kinetoplastid base J binding proteins (JBP). Base J stands for β-d-glucosyl(hydroxymethyl)uracil, an abundant base in the genome of kinetoplastida, synthesized through a 5-hmU intermediate generated by enzymatic hydroxylation of thymine by JBP1/2 (reviewed in Borst and Sabatini 2008). Computational analyses identified the mammalian oncogenic TET proteins as close relatives of the JBPs (Iyer et al. 2009). The ultimate finding that TETs comprise a catalytic domain capable of catalyzing the oxidation of 5-meC to 5-hmC uncovered a functional link between these proteins and DNA methylation, possibly demethylation (Tahiliani et al. 2009; Ito et al. 2010). Indeed, 5-hmC was suggested to trigger passive demethylation through inhibition of the maintenance methyltransferase DNMT1 (Valinluck and Sowers 2007). Besides that, 5-hmC might represent an intermediate of active DNA demethylation through stepwise oxidation of 5-meC coupled to either excision repair or decarboxylation (Fig. 5d).
Direct excision of 5-hmC by a DNA glycosylase would seem a plausible scenario (Fig. 5c). A 5-hmC DNA glycosylase was reported to be active in calf thymus extracts (Cannon et al. 1988). The responsible protein, however, has never been purified, nor has 5-hmC glycosylases activity been associated with any known mammalian DNA glycosylase. Given their activity on 5-hmU, TDG and SMUG1 would seem good candidates for 5-hmC processing, but recent evidence shows that at least TDG fails to do so at an appreciable rate (He et al. 2011; Maiti and Drohat 2011). Thus, while direct 5-hmC excision by DNA glycosylases may occur and contribute to DNA demethylation, the underlying enzymatic pathway remains to be clarified.
Another possible route of demethylation would be the further conversion of 5-hmC to an intermediate for DNA glycosylase mediated excision. The deamination of 5-hmC by a specific deaminase (e.g., AID), for example, would give rise to 5-hmU mismatched with G (Rusmintratip and Sowers 2000), which is a substrate for SMUG1 and TDG (Fig. 5e) (Boorstein et al. 2001; Hardeland et al. 2003; Cortellino et al. 2011). As discussed above for 5-meC deamination, however, the concept of demethylation through a pro-mutagenic 5-hmU intermediate is debatable as this would require an immensely efficient and accurate coupling of the deamination and repair processes, particularly in densely methylated DNA sequences, if genome integrity is to be maintained.
Less problematic in this regard seems a more recently implicated mode of TET-mediated demethylation. The key discovery was that TET proteins do not only generate 5-hmC but can oxidize this intermediate further to 5-formylcytosine (5-fC), and 5-carboxylcytosine (5-caC) (Ito et al. 2011), which are good substrates for excision by TDG (He et al. 2011; Maiti and Drohat 2011) (Fig. 5d). Given this, a DNA glycosylase-mediated conversion of 5-meC to C may thus occur without a need to deaminate, i.e., to generate a mutagenic intermediate. However, such a pathway would still trigger massive DNA incision activity in the context of genome-wide active demethylation, which seems a genetically risky and energetically wasteful way to erase an epigenetic mark. A more straightforward way to eliminate 5-caC under these conditions would be to couple the 5-meC oxidation cascade with a decarboxylation step to generate C and CO2 as final products. A 5-caC decarboxylase is thus an activity to watch out for.
G•T-directed DNA glycosylases and epigenetic stability
Active demethylation of 5-meC in mammalian cells occurs under different circumstances for different purposes, such as genome-wide epigenetic reprogramming, activation of tissue-specific genes during embryogenesis, and the maintenance of active and bivalent chromatin states during and after cell-lineage commitment. These circumstances generate specific demands for a demethylation mechanism (e.g., targeting, catalysis, processivity, efficiency) which are most likely addressed by distinct pathways.
Any of the above considered routes of DNA repair-mediated active demethylation of 5-meC requires an enzyme capable of recognizing and excising a cytosine derivative (5-meC, 5-hmC, 5-caC, T, 5-hmU) in a base (mis)pairing configuration with guanine. On the basis of their substrate spectra, the two DNA glycosylases MBD4 and TDG appear to be most suitable for this purpose, and both have been implicated in one way or another in DNA demethylation as discussed. The biological functions associated with these activities, however, still need to be clarified. Considering, for instance, the reported involvement of MBD4 in hormone-induced promoter demethylation, and the requirement of such demethylation for derepression of select genes during embryogenesis, it is surprising that Mbd4 knockout mice develop normally and display no apparent epigenetic abnormality (Millar et al. 2002; Wong et al. 2002). So, either promoter demethylation during embryogenesis is not developmentally important, or MBD4 is not or only redundantly involved in these processes. The phenotype of Tdg knockout mice, however, has provided more direct evidence for an epigenetic function.
Disruption of TDG in mouse causes embryonic lethality (Cortazar et al. 2011; Cortellino et al. 2011), most likely because TDG-deficient cells fail to properly establish and/or maintain cell-type-specific gene expression programs during cell lineage commitment (Cortazar et al. 2011). This phenotype coincides with the occurrence of aberrant chromatin modifications at promoters of misregulated genes: a loss of active histone marks (H3K4me2), a gain of repressive histone marks (H3K9me3, H3K27me3) and, nota bene, a gradual accumulation of CpG methylation (Cortazar et al. 2011). Together with evidence for a TDG-dependent engagement of BER at affected gene promoters (Cortazar et al. 2011; Cortellino et al. 2011), this suggests that TDG protects CpG-rich promoters from aberrant hypermethylation by active demethylation of erroneously methylated cytosines.
TDG may also contribute to active demethylation of originally methylated sequences. Tissues of TDG deficient embryos showed hypermethylation at the Alb1 enhancer and the Tat glucocorticoid-responsive unit, both undergoing demethylation in the process of tissue specific gene activation (Cortellino et al. 2011). These results can, however, be interpreted in two ways; while the hypermethylation measured in the absence of TDG can indeed be explained by inefficient active CpG demethylation during tissue differentiation, it can equally well be accounted for by a lack of TDG-dependent maintenance of the unmethylated state following successful active demethylation through a different pathway. Hence, while the concept of TDG-mediated active demethylation in the maintenance of hypomethylated states at CpG-rich gene promoters is well supported, its potential contribution to demethylation of methylated sequences requires further validation.
The functions of MBD4 and TDG in mediating DNA demethylation are clearly distinct but there might be a partial overlap; TDG might compensate for the loss of MBD4 in knockout mice but obviously not vice versa. A plausible scenario would be that MBD4 is primarily involved in the demethylation of methylated sequences in the context of developmental gene activation, whereas TDG’s function is to protect unmethylated promoter sequences across the genome from aberrant de novo methylation. This would be supported by the ability of MBD4 to bind to methylated CpGs, and the preferential association of TDG with unmethylated gene promoters.
Conclusion
Given their ability to interrogate the surface of DNA bases by flipping them out of the helix into a selective active site pocket, DNA glycosylases represent efficient tools to specifically recognize unduly modified bases and eliminate them from the DNA, thereby enforcing genetic integrity. Yet, while being perfectly equipped to function in DNA repair, their structure and mode of action could provide for more. This is supported by accumulating evidence for non-canonical functions of these DNA-probing enzymes, the most recent and most intensely discussed being an involvement in DNA demethylation and epigenetic control. Further studies will shed light on the mechanism surrounding the action of DNA glycosylases in such a context, from the signals required for the temporal and spatial regulation of their action to the co-factors necessary for efficient base-flipping and excision.
References
Almeida KH, Sobol RW (2007) A unified view of base excision repair: lesion-dependent protein complexes regulated by post-translational modification. DNA Repair (Amst) 6:695–711
Al-Tassan N, Chmiel NH, Maynard J, Fleming N, Livingston AL, Williams GT, Hodges AK, Davies DR, David SS, Sampson JR, Cheadle JP (2002) Inherited variants of MYH associated with somatic G:C– > T:A mutations in colorectal tumors. Nat Genet 30:227–232
An Q, Robins P, Lindahl T, Barnes DE (2005) C – > T mutagenesis and gamma-radiation sensitivity due to deficiency in the Smug1 and Ung DNA glycosylases. EMBO J 24:2205–2213
An Q, Robins P, Lindahl T, Barnes DE (2007) 5-Fluorouracil incorporated into DNA is excised by the Smug1 DNA glycosylase to reduce drug cytotoxicity. Cancer Res 67:940–945
Aravind L, Koonin EV (2000) The alpha/beta fold uracil DNA glycosylases: a common origin with diverse fates. Genome Biol 1, RESEARCH0007
Au KG, Cabrera M, Miller JH, Modrich P (1988) Escherichia coli mutY gene product is required for specific A-G––C.G mismatch correction. Proc Natl Acad Sci USA 85:9163–9166
Baba D, Maita N, Jee JG, Uchimura Y, Saitoh H, Sugasawa K, Hanaoka F, Tochio H, Hiroaki H, Shirakawa M (2005) Crystal structure of thymine DNA glycosylase conjugated to SUMO-1. Nature 435:979–982
Banerjee A, Yang W, Karplus M, Verdine GL (2005) Structure of a repair enzyme interrogating undamaged DNA elucidates recognition of damaged DNA. Nature 434:612–618
Bardwell PD, Martin A, Wong E, Li Z, Edelmann W, Scharff MD (2003) Cutting edge: the G-U mismatch glycosylase methyl-CpG binding domain 4 is dispensable for somatic hypermutation and class switch recombination. J Immunol 170:1620–1624
Barrett TE, Savva R, Panayotou G, Barlow T, Brown T, Jiricny J, Pearl LH (1998) Crystal structure of a G:T/U mismatch-specific DNA glycosylase: mismatch recognition by complementary-strand interactions. Cell 92:117–129
Barrett TE, Schärer OD, Savva R, Brown T, Jiricny J, Verdine GL, Pearl LH (1999) Crystal structure of a thwarted mismatch glycosylase DNA repair complex. EMBO J 18:6599–6609
Begley TJ, Haas BJ, Noel J, Shekhtman A, Williams WA, Cunningham RP (1999) A new member of the endonuclease III family of DNA repair enzymes that removes methylated purines from DNA. Curr Biol 9:653–656
Begum NA, Izumi N, Nishikori M, Nagaoka H, Shinkura R, Honjo T (2007) Requirement of non-canonical activity of uracil DNA glycosylase for class switch recombination. J Biol Chem 282:731–742
Begum NA, Stanlie A, Doi T, Sasaki Y, Jin HW, Kim YS, Nagaoka H, Honjo T (2009) Further evidence for involvement of a noncanonical function of uracil DNA glycosylase in class switch recombination. Proc Natl Acad Sci USA 106:2752–2757
Bellacosa A, Cicchillitti L, Schepis F, Riccio A, Yeung AT, Matsumoto Y, Golemis EA, Genuardi M, Neri G (1999) MED1, a novel human methyl-CpG-binding endonuclease, interacts with DNA mismatch repair protein MLH1. Proc Natl Acad Sci USA 96:3969–3974
Berg OG, Winter RB, von Hippel PH (1981) Diffusion-driven mechanisms of protein translocation on nucleic acids. 1. Models and theory. Biochemistry 20:6929–6948
Bhutani N, Brady JJ, Damian M, Sacco A, Corbel SY, Blau HM (2010) Reprogramming towards pluripotency requires AID-dependent DNA demethylation. Nature 463:1042–1047
Blainey PC, Luo G, Kou SC, Mangel WF, Verdine GL, Bagchi B, Xie XS (2009) Nonspecifically bound proteins spin while diffusing along DNA. Nat Struct Mol Biol 16:1224–1229
Boal AK, Genereux JC, Sontz PA, Gralnick JA, Newman DK, Barton JK (2009) Redox signaling between DNA repair proteins for efficient lesion detection. Proc Natl Acad Sci USA 106:15237–15242
Boorstein RJ, Cummings A Jr, Marenstein DR, Chan MK, Ma Y, Neubert TA, Brown SM, Teebor GW (2001) Definitive identification of mammalian 5-hydroxymethyluracil DNA N-glycosylase activity as SMUG1. J Biol Chem 276:41991–41997
Borst P, Sabatini R (2008) Base J: discovery, biosynthesis, and possible functions. Annu Rev Microbiol 62:235–251
Bruner SD, Norman DP, Verdine GL (2000) Structural basis for recognition and repair of the endogenous mutagen 8-oxoguanine in DNA. Nature 403:859–866
Bruniquel D, Schwartz RH (2003) Selective, stable demethylation of the interleukin-2 gene enhances transcription by an active process. Nat Immunol 4:235–240
Cannon SV, Cummings A, Teebor GW (1988) 5-Hydroxymethylcytosine DNA glycosylase activity in mammalian tissue. Biochem Biophys Res Commun 151:1173–1179
Cannon-Carlson SV, Gokhale H, Teebor GW (1989) Purification and characterization of 5-hydroxymethyluracil-DNA glycosylase from calf thymus. Its possible role in the maintenance of methylated cytosine residues. J Biol Chem 264:13306–13312
Conticello SG, Thomas CJ, Petersen-Mahrt SK, Neuberger MS (2005) Evolution of the AID/APOBEC family of polynucleotide (deoxy)cytidine deaminases. Mol Biol Evol 22:367–377
Cortazar D, Kunz C, Saito Y, Steinacher R, Schär P (2007) The enigmatic thymine DNA glycosylase. DNA Repair (Amst) 6:489–504
Cortazar D, Kunz C, Selfridge J, Lettieri T, Saito Y, MacDougall E, Wirz A, Schuermann D, Jacobs AL, Siegrist F, Steinacher R, Jiricny J, Bird A, Schär P (2011) Embryonic lethal phenotype reveals a function of TDG in maintaining epigenetic stability. Nature 470:419–423
Cortellino S, Turner D, Masciullo V, Schepis F, Albino D, Daniel R, Skalka AM, Meropol NJ, Alberti C, Larue L, Bellacosa A (2003) The base excision repair enzyme MED1 mediates DNA damage response to antitumor drugs and is associated with mismatch repair system integrity. Proc Natl Acad Sci USA 100:15071–15076
Cortellino S, Xu J, Sannai M, Moore R, Caretti E, Cigliano A, Le Coz M, Devarajan K, Wessels A, Soprano D, Abramowitz LK, Bartolomei MS, Rambow F, Bassi MR, Bruno T, Fanciulli M, Renner C, Klein-Szanto AJ, Matsumoto Y, Kobi D, Davidson I, Alberti C, Larue L, Bellacosa A (2011) Thymine DNA glycosylase is essential for active DNA demethylation by linked deamination–base excision repair. Cell 146:67–79
Cunningham RP, Asahara H, Bank JF, Scholes CP, Salerno JC, Surerus K, Munck E, McCracken J, Peisach J, Emptage MH (1989) Endonuclease III is an iron–sulfur protein. Biochemistry 28:4450–4455
Dean W, Santos F, Stojkovic M, Zakhartchenko V, Walter J, Wolf E, Reik W (2001) Conservation of methylation reprogramming in mammalian development: aberrant reprogramming in cloned embryos. Proc Natl Acad Sci USA 98:13734–13738
Denver DR, Swenson SL, Lynch M (2003) An evolutionary analysis of the helix-hairpin-helix superfamily of DNA repair glycosylases. Mol Biol Evol 20:1603–1611
Dherin C, Radicella JP, Dizdaroglu M, Boiteux S (1999) Excision of oxidatively damaged DNA bases by the human alpha-hOgg1 protein and the polymorphic alpha-hOgg1(Ser326Cys) protein which is frequently found in human populations. Nucleic Acids Res 27:4001–4007
Di Noia J, Neuberger MS (2002) Altering the pathway of immunoglobulin hypermutation by inhibiting uracil-DNA glycosylase. Nature 419:43–48
Di Noia JM, Rada C, Neuberger MS (2006) SMUG1 is able to excise uracil from immunoglobulin genes: insight into mutation versus repair. EMBO J 25:585–595
Dizdaroglu M, Karahalil B, Senturker S, Buckley TJ, Roldan-Arjona T (1999) Excision of products of oxidative DNA base damage by human NTH1 protein. Biochemistry 38:243–246
Dou H, Mitra S, Hazra TK (2003) Repair of oxidized bases in DNA bubble structures by human DNA glycosylases NEIL1 and NEIL2. J Biol Chem 278:49679–49684
Eide L, Luna L, Gustad EC, Henderson PT, Essigmann JM, Demple B, Seeberg E (2001) Human endonuclease III acts preferentially on DNA damage opposite guanine residues in DNA. Biochemistry 40:6653–6659
Engelward BP, Weeda G, Wyatt MD, Broekhof JL, de Wit J, Donker I, Allan JM, Gold B, Hoeijmakers JH, Samson LD (1997) Base excision repair deficient mice lacking the Aag alkyladenine DNA glycosylase. Proc Natl Acad Sci USA 94:13087–13092
Falnes PO, Johansen RF, Seeberg E (2002) AlkB-mediated oxidative demethylation reverses DNA damage in Escherichia coli. Nature 419:178–182
Friedberg EC, Walker GC, Siede W, Wood RD, Schultz RA, Ellenberger T (2006) DNA repair and mutagenesis, 2nd edn. ASM, Washington
Frieder D, Larijani M, Collins C, Shulman M, Martin A (2009) The concerted action of Msh2 and UNG stimulates somatic hypermutation at A. T base pairs. Mol Cell Biol 29:5148–5157
Friedman JI, Stivers JT (2010) Detection of damaged DNA bases by DNA glycosylase enzymes. Biochemistry 49:4957–4967
Friedman JI, Majumdar A, Stivers JT (2009) Nontarget DNA binding shapes the dynamic landscape for enzymatic recognition of DNA damage. Nucleic Acids Res 37:3493–3500
Fromme JC, Bruner SD, Yang W, Karplus M, Verdine GL (2003) Product-assisted catalysis in base-excision DNA repair. Nat Struct Biol 10:204–211
Gallinari P, Jiricny J (1996) A new class of uracil-DNA glycosylases related to human thymine-DNA glycosylase. Nature 383:735–738
Guan Y, Manuel RC, Arvai AS, Parikh SS, Mol CD, Miller JH, Lloyd S, Tainer JA (1998) MutY catalytic core, mutant and bound adenine structures define specificity for DNA repair enzyme superfamily. Nat Struct Biol 5:1058–1064
Hagen L, Kavli B, Sousa MM, Torseth K, Liabakk NB, Sundheim O, Pena-Diaz J, Otterlei M, Horning O, Jensen ON, Krokan HE, Slupphaug G (2008) Cell cycle-specific UNG2 phosphorylations regulate protein turnover, activity and association with RPA. EMBO J 27:51–61
Hang B, Singer B, Margison GP, Elder RH (1997) Targeted deletion of alkylpurine-DNA-N-glycosylase in mice eliminates repair of 1, N6-ethenoadenine and hypoxanthine but not of 3, N4-ethenocytosine or 8-oxoguanine. Proc Natl Acad Sci U S A 94:12869–12874
Hardeland U, Bentele M, Jiricny J, Schär P (2000) Separating substrate recognition from base hydrolysis in human thymine DNA glycosylase by mutational analysis. J Biol Chem 275:33449–33456
Hardeland U, Steinacher R, Jiricny J, Schär P (2002) Modification of the human thymine-DNA glycosylase by ubiquitin-like proteins facilitates enzymatic turnover. EMBO J 21:1456–1464
Hardeland U, Bentele M, Jiricny J, Schär P (2003) The versatile thymine DNA-glycosylase: a comparative characterization of the human, Drosophila and fission yeast orthologs. Nucleic Acids Res 31:2261–2271
Hardeland U, Kunz C, Focke F, Szadkowski M, Schär P (2007) Cell cycle regulation as a mechanism for functional separation of the apparently redundant uracil DNA glycosylases TDG and UNG2. Nucleic Acids Res 35:3859–3867
Harris RS, Bishop KN, Sheehy AM, Craig HM, Petersen-Mahrt SK, Watt IN, Neuberger MS, Malim MH (2003) DNA deamination mediates innate immunity to retroviral infection. Cell 113:803–809
Haushalter KA, Todd Stukenberg MW, Kirschner MW, Verdine GL (1999) Identification of a new uracil-DNA glycosylase family by expression cloning using synthetic inhibitors. Curr Biol 9:174–185
Hazra TK, Izumi T, Boldogh I, Imhoff B, Kow YW, Jaruga P, Dizdaroglu M, Mitra S (2002) Identification and characterization of a human DNA glycosylase for repair of modified bases in oxidatively damaged DNA. Proc Natl Acad Sci USA 99:3523–3528
He YF, Li BZ, Li Z, Liu P, Wang Y, Tang Q, Ding J, Jia Y, Chen Z, Li L, Sun Y, Li X, Dai Q, Song CX, Zhang K, He C, Xu GL (2011) Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science [Epub ahead of print]
Hendrich B, Bird A (1998) Identification and characterization of a family of mammalian methyl-CpG binding proteins. Mol Cell Biol 18:6538–6547
Hermann A, Gowher H, Jeltsch A (2004) Biochemistry and biology of mammalian DNA methyltransferases. Cell Mol Life Sci 61:2571–2587
Hill JW, Hazra TK, Izumi T, Mitra S (2001) Stimulation of human 8-oxoguanine-DNA glycosylase by AP-endonuclease: potential coordination of the initial steps in base excision repair. Nucleic Acids Res 29:430–438
Holmes RK, Malim MH, Bishop KN (2007) APOBEC-mediated viral restriction: not simply editing? Trends Biochem Sci 32:118–128
Horst JP, Fritz HJ (1996) Counteracting the mutagenic effect of hydrolytic deamination of DNA 5-methylcytosine residues at high temperature: DNA mismatch N-glycosylase Mig. Mth of the thermophilic archaeon Methanobacterium thermoautotrophicum THF. EMBO J 15:5459–5469
Imai K, Slupphaug G, Lee WI, Revy P, Nonoyama S, Catalan N, Yel L, Forveille M, Kavli B, Krokan HE, Ochs HD, Fischer A, Durandy A (2003) Human uracil-DNA glycosylase deficiency associated with profoundly impaired immunoglobulin class-switch recombination. Nat Immunol 4:1023–1028
Ito S, D'Alessio AC, Taranova OV, Hong K, Sowers LC, Zhang Y (2010) Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466:1129–1133
Ito S, Shen L, Dai Q, Wu SC, Collins LB, Swenberg JA, He C, Zhang Y (2011) Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science [Epub ahead of print]
Iyer LM, Tahiliani M, Rao A, Aravind L (2009) Prediction of novel families of enzymes involved in oxidative and other complex modifications of bases in nucleic acids. Cell Cycle 8:1698–1710
Janik J, Swoboda M, Janowska B, Ciesla JM, Gackowski D, Kowalewski J, Olinski R, Tudek B, Speina E (2011) 8-Oxoguanine incision activity is impaired in lung tissues of NSCLC patients with the polymorphism of OGG1 and XRCC1 genes. Mutat Res 709–710:21–31
Jones S, Emmerson P, Maynard J, Best JM, Jordan S, Williams GT, Sampson JR, Cheadle JP (2002) Biallelic germline mutations in MYH predispose to multiple colorectal adenoma and somatic G:C– > T:A mutations. Hum Mol Genet 11:2961–2967
Jost JP, Siegmann M, Sun L, Leung R (1995) Mechanisms of DNA demethylation in chicken embryos. Purification and properties of a 5-methylcytosine-DNA glycosylase. J Biol Chem 270:9734–9739
Jost JP, Fremont M, Siegmann M, Hofsteenge J (1997) The RNA moiety of chick embryo 5-methylcytosine–DNA glycosylase targets DNA demethylation. Nucleic Acids Res 25:4545–4550
Kafri T, Ariel M, Brandeis M, Shemer R, Urven L, McCarrey J, Cedar H, Razin A (1992) Developmental pattern of gene-specific DNA methylation in the mouse embryo and germ line. Genes Dev 6:705–714
Kangaspeska S, Stride B, Metivier R, Polycarpou-Schwarz M, Ibberson D, Carmouche RP, Benes V, Gannon F, Reid G (2008) Transient cyclical methylation of promoter DNA. Nature 452:112–115
Karahalil B, Girard PM, Boiteux S, Dizdaroglu M (1998) Substrate specificity of the Ogg1 protein of Saccharomyces cerevisiae: excision of guanine lesions produced in DNA by ionizing radiation- or hydrogen peroxide/metal ion-generated free radicals. Nucleic Acids Res 26:1228–1233
Kavli B, Slupphaug G, Mol CD, Arvai AS, Peterson SB, Tainer JA, Krokan HE (1996) Excision of cytosine and thymine from DNA by mutants of human uracil-DNA glycosylase. EMBO J 15:3442–3447
Kavli B, Sundheim O, Akbari M, Otterlei M, Nilsen H, Skorpen F, Aas PA, Hagen L, Krokan HE, Slupphaug G (2002) hUNG2 is the major repair enzyme for removal of uracil from U:A matches, U:G mismatches, and U in single-stranded DNA, with hSMUG1 as a broad specificity backup. J Biol Chem 277:39926–39936
Kim MS, Kondo T, Takada I, Youn MY, Yamamoto Y, Takahashi S, Matsumoto T, Fujiyama S, Shirode Y, Yamaoka I, Kitagawa H, Takeyama K, Shibuya H, Ohtake F, Kato S (2009) DNA demethylation in hormone-induced transcriptional derepression. Nature 461:1007–1012
Klungland A, Rosewell I, Hollenbach S, Larsen E, Daly G, Epe B, Seeberg E, Lindahl T, Barnes DE (1999) Accumulation of premutagenic DNA lesions in mice defective in removal of oxidative base damage. Proc Natl Acad Sci USA 96:13300–13305
Krokan HE, Standal R, Slupphaug G (1997) DNA glycosylases in the base excision repair of DNA. Biochem J 325:1–16
Kunz C, Saito Y, Schär P (2009a) DNA repair in mammalian cells: mismatched repair: variations on a theme. Cell Mol Life Sci 66:1021–1038
Kunz C, Focke F, Saito Y, Schuermann D, Lettieri T, Selfridge J, Schär P (2009b) Base excision by thymine DNA glycosylase mediates DNA-directed cytotoxicity of 5-fluorouracil. PLoS Biol 7:e91
Kuo CF, McRee DE, Fisher CL, O'Handley SF, Cunningham RP, Tainer JA (1992) Atomic structure of the DNA repair [4Fe-4S] enzyme endonuclease III. Science 258:434–440
Lee CY, Delaney JC, Kartalou M, Lingaraju GM, Maor-Shoshani A, Essigmann JM, Samson LD (2009) Recognition and processing of a new repertoire of DNA substrates by human 3-methyladenine DNA glycosylase (AAG). Biochemistry 48(9):1850–1861
Lindahl T (1974) An N-glycosidase from Escherichia coli that releases free uracil from DNA containing deaminated cytosine residues. Proc Natl Acad Sci USA 71:3649–3653
Lindahl T, Wood RD (1999) Quality control by DNA repair. Science 286:1897–1905
Lister R, O'Malley RC, Tonti-Filippini J, Gregory BD, Berry CC, Millar AH, Ecker JR (2008) Highly integrated single-base resolution maps of the epigenome in Arabidopsis. Cell 133:523–536
Liu M, Bandaru V, Bond JP, Jaruga P, Zhao X, Christov PP, Burrows CJ, Rizzo CJ, Dizdaroglu M, Wallace SS (2010) The mouse ortholog of NEIL3 is a functional DNA glycosylase in vitro and in vivo. Proc Natl Acad Sci USA 107:4925–4930
Lu R, Nash HM, Verdine GL (1997) A mammalian DNA repair enzyme that excises oxidatively damaged guanines maps to a locus frequently lost in lung cancer. Curr Biol 7:397–407
Ma DK, Jang MH, Guo JU, Kitabatake Y, Chang ML, Pow-Anpongkul N, Flavell RA, Lu B, Ming GL, Song H (2009) Neuronal activity-induced Gadd45b promotes epigenetic DNA demethylation and adult neurogenesis. Science 323:1074–1077
Maiti A, Drohat AC (2011) Thymine DNA glycosylase can rapidly excise 5-formylcytosine and 5-carboxylcytosine: potential implications for active demethylation of CpG sites. J Biol Chem [Epub ahead of print]
Maiti A, Morgan MT, Pozharski E, Drohat AC (2008) Crystal structure of human thymine DNA glycosylase bound to DNA elucidates sequence-specific mismatch recognition. Proc Natl Acad Sci USA 105:8890–8895
Maiti A, Morgan MT, Drohat AC (2009) Role of two strictly conserved residues in nucleotide flipping and N-glycosylic bond cleavage by human thymine DNA glycosylase. J Biol Chem 284:36680–36688
Mayer W, Niveleau A, Walter J, Fundele R, Haaf T (2000) Demethylation of the zygotic paternal genome. Nature 403:501–502
McGoldrick JP, Yeh YC, Solomon M, Essigmann JM, Lu AL (1995) Characterization of a mammalian homolog of the Escherichia coli MutY mismatch repair protein. Mol Cell Biol 15:989–996
Meira LB, Moroski-Erkul CA, Green SL, Calvo JA, Bronson RT, Shah D, Samson LD (2009) Aag-initiated base excision repair drives alkylation-induced retinal degeneration in mice. Proc Natl Acad Sci USA 106:888–893
Melamede RJ, Hatahet Z, Kow YW, Ide H, Wallace SS (1994) Isolation and characterization of endonuclease VIII from Escherichia coli. Biochemistry 33:1255–1264
Metivier R, Gallais R, Tiffoche C, Le Peron C, Jurkowska RZ, Carmouche RP, Ibberson D, Barath P, Demay F, Reid G, Benes V, Jeltsch A, Gannon F, Salbert G (2008) Cyclical DNA methylation of a transcriptionally active promoter. Nature 452:45–50
Millar CB, Guy J, Sansom OJ, Selfridge J, MacDougall E, Hendrich B, Keightley PD, Bishop SM, Clarke AR, Bird A (2002) Enhanced CpG mutability and tumorigenesis in MBD4-deficient mice. Science 297:403–405
Miller CA, Sweatt JD (2007) Covalent modification of DNA regulates memory formation. Neuron 53:857–869
Mol CD, Arvai AS, Slupphaug G, Kavli B, Alseth I, Krokan HE, Tainer JA (1995) Crystal structure and mutational analysis of human uracil-DNA glycosylase: structural basis for specificity and catalysis. Cell 80:869–878
Monk M, Boubelik M, Lehnert S (1987) Temporal and regional changes in DNA methylation in the embryonic, extraembryonic and germ cell lineages during mouse embryo development. Development 99:371–382
Morland I, Rolseth V, Luna L, Rognes T, Bjoras M, Seeberg E (2002) Human DNA glycosylases of the bacterial Fpg/MutM superfamily: an alternative pathway for the repair of 8-oxoguanine and other oxidation products in DNA. Nucleic Acids Res 30:4926–4936
Muramatsu M, Kinoshita K, Fagarasan S, Yamada S, Shinkai Y, Honjo T (2000) Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell 102:553–563
Nash HM, Bruner SD, Schärer OD, Kawate T, Addona TA, Spooner E, Lane WS, Verdine GL (1996) Cloning of a yeast 8-oxoguanine DNA glycosylase reveals the existence of a base-excision DNA-repair protein superfamily. Curr Biol 6:968–980
Nash HM, Lu R, Lane WS, Verdine GL (1997) The critical active-site amine of the human 8-oxoguanine DNA glycosylase, hOgg1: direct identification, ablation and chemical reconstitution. Chem Biol 4:693–702
Neddermann P, Gallinari P, Lettieri T, Schmid D, Truong O, Hsuan JJ, Wiebauer K, Jiricny J (1996) Cloning and expression of human G/T mismatch-specific thymine-DNA glycosylase. J Biol Chem 271:12767–12774
Nilsen H, Otterlei M, Haug T, Solum K, Nagelhus TA, Skorpen F, Krokan HE (1997) Nuclear and mitochondrial uracil-DNA glycosylases are generated by alternative splicing and transcription from different positions in the UNG gene. Nucleic Acids Res 25:750–755
Nilsen H, Rosewell I, Robins P, Skjelbred CF, Andersen S, Slupphaug G, Daly G, Krokan HE, Lindahl T, Barnes DE (2000) Uracil-DNA glycosylase (UNG)-deficient mice reveal a primary role of the enzyme during DNA replication. Mol Cell 5:1059–1065
Nilsen H, Stamp G, Andersen S, Hrivnak G, Krokan HE, Lindahl T, Barnes DE (2003) Gene-targeted mice lacking the Ung uracil-DNA glycosylase develop B-cell lymphomas. Oncogene 22:5381–5386
Ocampo MT, Chaung W, Marenstein DR, Chan MK, Altamirano A, Basu AK, Boorstein RJ, Cunningham RP, Teebor GW (2002) Targeted deletion of mNth1 reveals a novel DNA repair enzyme activity. Mol Cell Biol 22:6111–6121
O'Connor TR (1993) Purification and characterization of human 3-methyladenine-DNA glycosylase. Nucleic Acids Res 21:5561–5569
O'Connor TR, Laval F (1990) Isolation and structure of a cDNA expressing a mammalian 3-methyladenine-DNA glycosylase. EMBO J 9:3337–3342
Oswald J, Engemann S, Lane N, Mayer W, Olek A, Fundele R, Dean W, Reik W, Walter J (2000) Active demethylation of the paternal genome in the mouse zygote. Curr Biol 10:475–478
Otterlei M, Warbrick E, Nagelhus TA, Haug T, Slupphaug G, Akbari M, Aas PA, Steinsbekk K, Bakke O, Krokan HE (1999) Post-replicative base excision repair in replication foci. EMBO J 18:3834–3844
Parikh SS, Mol CD, Slupphaug G, Bharati S, Krokan HE, Tainer JA (1998) Base excision repair initiation revealed by crystal structures and binding kinetics of human uracil-DNA glycosylase with DNA. EMBO J 17:5214–5226
Parikh SS, Mol CD, Hosfield DJ, Tainer JA (1999) Envisioning the molecular choreography of DNA base excision repair. Curr Opin Struct Biol 9:37–47
Parker JB, Bianchet MA, Krosky DJ, Friedman JI, Amzel LM, Stivers JT (2007) Enzymatic capture of an extrahelical thymine in the search for uracil in DNA. Nature 449:433–437
Pavri R, Nussenzweig MC (2011) AID targeting in antibody diversity. Adv Immunol 110:1–26
Penterman J, Zilberman D, Huh JH, Ballinger T, Henikoff S, Fischer RL (2007) DNA demethylation in the Arabidopsis genome. Proc Natl Acad Sci USA 104:6752–6757
Petronzelli F, Riccio A, Markham GD, Seeholzer SH, Genuardi M, Karbowski M, Yeung AT, Matsumoto Y, Bellacosa A (2000) Investigation of the substrate spectrum of the human mismatch-specific DNA N-glycosylase MED1 (MBD4): fundamental role of the catalytic domain. J Cell Physiol 185:473–480
Pettersen HS, Visnes T, Vagbo CB, Svaasand EK, Doseth B, Slupphaug G, Kavli B, Krokan HE (2011) UNG-initiated base excision repair is the major repair route for 5-fluorouracil in DNA, but 5-fluorouracil cytotoxicity depends mainly on RNA incorporation. Nucleic Acids Res. doi:gkr563 [pii]
Pope MA, Porello SL, David SS (2002) Escherichia coli apurinic-apyrimidinic endonucleases enhance the turnover of the adenine glycosylase MutY with G:A substrates. J Biol Chem 277:22605–22615
Popp C, Dean W, Feng S, Cokus SJ, Andrews S, Pellegrini M, Jacobsen SE, Reik W (2010) Genome-wide erasure of DNA methylation in mouse primordial germ cells is affected by AID deficiency. Nature 463:1101–1105
Porello SL, Cannon MJ, David SS (1998) A substrate recognition role for the [4Fe-4S]2+ cluster of the DNA repair glycosylase MutY. Biochemistry 37:6465–6475
Rada C, Ehrenstein MR, Neuberger MS, Milstein C (1998) Hot spot focusing of somatic hypermutation in MSH2-deficient mice suggests two stages of mutational targeting. Immunity 9:135–141
Rada C, Williams GT, Nilsen H, Barnes DE, Lindahl T, Neuberger MS (2002) Immunoglobulin isotype switching is inhibited and somatic hypermutation perturbed in UNG-deficient mice. Curr Biol 12:1748–1755
Radany EH, Dornfeld KJ, Sanderson RJ, Savage MK, Majumdar A, Seidman MM, Mosbaugh DW (2000) Increased spontaneous mutation frequency in human cells expressing the phage PBS2-encoded inhibitor of uracil-DNA glycosylase. Mutat Res 461(1):41–58
Radicella JP, Dherin C, Desmaze C, Fox MS, Boiteux S (1997) Cloning and characterization of hOGG1, a human homolog of the OGG1 gene of Saccharomyces cerevisiae. Proc Natl Acad Sci USA 94:8010–8015
Radman M (1976) An endonuclease from Escherichia coli that introduces single polynucleotide chain scissions in ultraviolet-irradiated DNA. J Biol Chem 251:1438–1445
Rai K, Huggins IJ, James SR, Karpf AR, Jones DA, Cairns BR (2008) DNA demethylation in zebrafish involves the coupling of a deaminase, a glycosylase, and gadd45. Cell 135:1201–1212
Robertson AB, Klungland A, Rognes T, Leiros I (2009) DNA repair in mammalian cells: base excision repair: the long and short of it. Cell Mol Life Sci 66:981–993
Rosenquist TA, Zaika E, Fernandes AS, Zharkov DO, Miller H, Grollman AP (2003) The novel DNA glycosylase, NEIL1, protects mammalian cells from radiation-mediated cell death. DNA Repair (Amst) 2:581–591
Rusmintratip V, Sowers LC (2000) An unexpectedly high excision capacity for mispaired 5-hydroxymethyluracil in human cell extracts. Proc Natl Acad Sci USA 97:14183–14187
Russo MT, De Luca G, Degan P, Parlanti E, Dogliotti E, Barnes DE, Lindahl T, Yang H, Miller JH, Bignami M (2004) Accumulation of the oxidative base lesion 8-hydroxyguanine in DNA of tumor-prone mice defective in both the Myh and Ogg1 DNA glycosylases. Cancer Res 64:4411–4414
Savva R, McAuley-Hecht K, Brown T, Pearl L (1995) The structural basis of specific base-excision repair by uracil-DNA glycosylase. Nature 373:487–493
Schanz S, Castor D, Fischer F, Jiricny J (2009) Interference of mismatch and base excision repair during the processing of adjacent U/G mispairs may play a key role in somatic hypermutation. Proc Natl Acad Sci USA 106:5593–5598
Schrofelbauer B, Yu Q, Zeitlin SG, Landau NR (2005) Human immunodeficiency virus type 1 Vpr induces the degradation of the UNG and SMUG uracil-DNA glycosylases. J Virol 79:10978–10987
Schrofelbauer B, Hakata Y, Landau NR (2007) HIV-1 Vpr function is mediated by interaction with the damage-specific DNA-binding protein DDB1. Proc Natl Acad Sci USA 104:4130–4135
Schwarz S, Bourgeois C, Soussaline F, Homsy C, Podesta A, Jost JP (2000) A CpG-rich RNA and an RNA helicase tightly associated with the DNA demethylation complex are present mainly in dividing chick embryo cells. Eur J Cell Biol 79:488–494
Sheehy AM, Gaddis NC, Malim MH (2003) The antiretroviral enzyme APOBEC3G is degraded by the proteasome in response to HIV-1 Vif. Nat Med 9:1404–1407
Shimizu Y, Iwai S, Hanaoka F, Sugasawa K (2003) Xeroderma pigmentosum group C protein interacts physically and functionally with thymine DNA glycosylase. EMBO J 22:164–173
Shimizu Y, Uchimura Y, Dohmae N, Saitoh H, Hanaoka F, Sugasawa K (2010) Stimulation of DNA glycosylase activities by XPC protein complex: roles of protein–protein interactions. J Nucleic Acids 2010
Simonsson S, Gurdon J (2004) DNA demethylation is necessary for the epigenetic reprogramming of somatic cell nuclei. Nat Cell Biol 6:984–990
Slupphaug G, Mol CD, Kavli B, Arvai AS, Krokan HE, Tainer JA (1996) A nucleotide-flipping mechanism from the structure of human uracil-DNA glycosylase bound to DNA. Nature 384:87–92
Stanczyk M, Sliwinski T, Cuchra M, Zubowska M, Bielecka-Kowalska A, Kowalski M, Szemraj J, Mlynarski W, Majsterek I (2011) The association of polymorphisms in DNA base excision repair genes XRCC1, OGG1 and MUTYH with the risk of childhood acute lymphoblastic leukemia. Mol Biol Rep 38:445–451
Steinacher R, Schär P (2005) Functionality of human thymine DNA glycosylase requires SUMO-regulated changes in protein conformation. Curr Biol 15:616–623
Sun B, Latham KA, Dodson ML, Lloyd RS (1995) Studies on the catalytic mechanism of five DNA glycosylases. Probing for enzyme-DNA imino intermediates. J Biol Chem 270:19501–19508
Tahiliani M, Koh KP, Shen Y, Pastor WA, Bandukwala H, Brudno Y, Agarwal S, Iyer LM, Liu DR, Aravind L, Rao A (2009) Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324:930–935
Takao M, Kanno S, Kobayashi K, Zhang QM, Yonei S, van der Horst GT, Yasui A (2002) A back-up glycosylase in Nth1 knock-out mice is a functional Nei (endonuclease VIII) homologue. J Biol Chem 277:42205–42213
Thomas D, Scot AD, Barbey R, Padula M, Boiteux S (1997) Inactivation of OGG1 increases the incidence of G. C– > T. A transversions in Saccharomyces cerevisiae: evidence for endogenous oxidative damage to DNA in eukaryotic cells. Mol Gen Genet 254:171–178
Tini M, Benecke A, Um SJ, Torchia J, Evans RM, Chambon P (2002) Association of CBP/p300 acetylase and thymine DNA glycosylase links DNA repair and transcription. Mol Cell 9:265–277
Torisu K, Tsuchimoto D, Ohnishi Y, Nakabeppu Y (2005) Hematopoietic tissue-specific expression of mouse Neil3 for endonuclease VIII-like protein. J Biochem 138:763–772
Trewick SC, Henshaw TF, Hausinger RP, Lindahl T, Sedgwick B (2002) Oxidative demethylation by Escherichia coli AlkB directly reverts DNA base damage. Nature 419:174–178
Um S, Harbers M, Benecke A, Pierrat B, Losson R, Chambon P (1998) Retinoic acid receptors interact physically and functionally with the T:G mismatch-specific thymine-DNA glycosylase. J Biol Chem 273:20728–20736
Valinluck V, Sowers LC (2007) Endogenous cytosine damage products alter the site selectivity of human DNA maintenance methyltransferase DNMT1. Cancer Res 67:946–950
van der Kemp PA, Thomas D, Barbey R, de Oliveira R, Boiteux S (1996) Cloning and expression in Escherichia coli of the OGG1 gene of Saccharomyces cerevisiae, which codes for a DNA glycosylase that excises 7,8-dihydro-8-oxoguanine and 2,6-diamino-4-hydroxy-5-N-methylformamidopyrimidine. Proc Natl Acad Sci USA 93:5197–5202
van Loon B, Markkanen E, Hubscher U (2010) Oxygen as a friend and enemy: how to combat the mutational potential of 8-oxo-guanine. DNA Repair (Amst) 9:604–616
Vartanian V, Lowell B, Minko IG, Wood TG, Ceci JD, George S, Ballinger SW, Corless CL, McCullough AK, Lloyd RS (2006) The metabolic syndrome resulting from a knockout of the NEIL1 DNA glycosylase. Proc Natl Acad Sci USA 103:1864–1869
Waters TR, Swann PF (1998) Kinetics of the action of thymine DNA glycosylase. J Biol Chem 273:20007–20014
Waters TR, Gallinari P, Jiricny J, Swann PF (1999) Human thymine DNA glycosylase binds to apurinic sites in DNA but is displaced by human apurinic endonuclease 1. J Biol Chem 274:67–74
Wibley JE, Waters TR, Haushalter K, Verdine GL, Pearl LH (2003) Structure and specificity of the vertebrate anti-mutator uracil-DNA glycosylase SMUG1. Mol Cell 11:1647–1659
Wiesendanger M, Kneitz B, Edelmann W, Scharff MD (2000) Somatic hypermutation in MutS homologue (MSH)3-, MSH6-, and MSH3/MSH6-deficient mice reveals a role for the MSH2-MSH6 heterodimer in modulating the base substitution pattern. J Exp Med 191:579–584
Wilson SH, Kunkel TA (2000) Passing the baton in base excision repair. Nat Struct Biol 7:176–178
Wilson TM, Vaisman A, Martomo SA, Sullivan P, Lan L, Hanaoka F, Yasui A, Woodgate R, Gearhart PJ (2005) MSH2-MSH6 stimulates DNA polymerase eta, suggesting a role for A:T mutations in antibody genes. J Exp Med 201:637–645
Wirtz S, Nagel G, Eshkind L, Neurath MF, Samson LD, Kaina B (2010) Both base excision repair and O6-methylguanine-DNA methyltransferase protect against methylation-induced colon carcinogenesis. Carcinogenesis 31:2111–2117
Wong E, Yang K, Kuraguchi M, Werling U, Avdievich E, Fan K, Fazzari M, Jin B, Brown AM, Lipkin M, Edelmann W (2002) Mbd4 inactivation increases C → T transition mutations and promotes gastrointestinal tumor formation. Proc Natl Acad Sci USA 99:14937–14942
Yang B, Chen K, Zhang C, Huang S, Zhang H (2007) Virion-associated uracil DNA glycosylase-2 and apurinic/apyrimidinic endonuclease are involved in the degradation of APOBEC3G-edited nascent HIV-1 DNA. J Biol Chem 282:11667–11675
Zharkov DO, Golan G, Gilboa R, Fernandes AS, Gerchman SE, Kycia JH, Rieger RA, Grollman AP, Shoham G (2002) Structural analysis of an Escherichia coli endonuclease VIII covalent reaction intermediate. EMBO J 21:789–800
Zhu JK (2009) Active DNA demethylation mediated by DNA glycosylases. Annu Rev Genet 43:143–166
Zhu B, Zheng Y, Angliker H, Schwarz S, Thiry S, Siegmann M, Jost JP (2000a) 5-Methylcytosine DNA glycosylase activity is also present in the human MBD4 (G/T mismatch glycosylase) and in a related avian sequence. Nucleic Acids Res 28:4157–4165
Zhu B, Zheng Y, Hess D, Angliker H, Schwarz S, Siegmann M, Thiry S, Jost JP (2000b) 5-methylcytosine-DNA glycosylase activity is present in a cloned G/T mismatch DNA glycosylase associated with the chicken embryo DNA demethylation complex. Proc Natl Acad Sci USA 97:5135–5139
Zhu B, Benjamin D, Zheng Y, Angliker H, Thiry S, Siegmann M, Jost JP (2001) Overexpression of 5-methylcytosine DNA glycosylase in human embryonic kidney cells EcR293 demethylates the promoter of a hormone-regulated reporter gene. Proc Natl Acad Sci USA 98:5031–5036
Acknowledgments
We wish to thank Christophe Kunz and David Schuermann for helpful discussion and critical reading of the manuscript. We also like apologize to all colleagues whose work we could not reference due to limited space. This work was supported by a grant form the Swiss National Science Foundation (31003A-122574).
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by: E. Nigg
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Jacobs, A.L., Schär, P. DNA glycosylases: in DNA repair and beyond. Chromosoma 121, 1–20 (2012). https://doi.org/10.1007/s00412-011-0347-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-011-0347-4