Abstract
Background
Gestational diabetes mellitus (GDM) poses significant health risks for both mothers and children, contributing to long-term complications such as type 2 diabetes and cardiovascular disease. This study explores the potential of microRNAs (miRNAs) as biomarkers for GDM by analyzing peripheral blood samples from GDM patients.
Method
Ten samples, including peripheral blood from 5 GDM patients and 5 controls, were collected to perform the RNA sequencing analysis. Differentially expressed miRNAs were further validated by quantitative real-time polymerase chain reaction.
Results
A total of 2287 miRNAs were identified, 229 of which showed differential expression. Validation by qRT-PCR confirmed significant up-regulation of miR-5193, miR-5003-3p, miR-3127-5p, novel-miR-96, miR-6734-5p, and miR-122-5p, while miR-10395-3p was down-regulated. Bioinformatics analyses revealed the involvement of these miRNAs in pathways associated with herpes simplex virus 1 infection.
Conclusion
This study provides insights into the differential expression of miRNAs in GDM patients and their potential roles in disease pathogenesis. It suggests that the differentially expressed miRNAs could serve as potential biomarkers for GDM, shedding light on the complex molecular mechanisms involved.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
This study identifies specific miRNAs that are differentially expressed in gestational diabetes mellitus (GDM) patients, suggesting their potential as biomarkers for GDM diagnosis and highlighting their roles in the disease pathogenesis, particularly in pathways associated with herpes simplex virus 1 infection. |
Introduction
Gestational diabetes mellitus (GDM) is a common complication of pregnancy, referring to diabetes diagnosed for the first time in the mid or late stages of pregnancy, which is neither pre-existing type 1 nor type 2 diabetes [1, 2]. Risk factors for GDM include overweight/obesity, Westernized diet and micronutrient deficiencies, advanced maternal age, insulin resistance, and/or family history of diabetes [3,4,5]. Although most GDM patients can return to normal after delivery, poor glycemic control in some cases can lead to serious risks and long-term damage [6]. These damages include the development of type 2 diabetes (T2DM) and cardiovascular disease (CVD), the increased risk of preeclampsia and the increased need for cesarean section in mothers, as well as the complications in newborns, such as preterm birth, polyhydramnios, macrosomia, shoulder dystocia, admission to the neonatal intensive care unit, neonatal respiratory distress syndrome, neonatal hypoglycemia, and hyperbilirubinemia. In addition, there is an increased risk of future obesity, CVD, T2DM, and/or GDM in children [1, 3, 4, 7]. With the increasing trend toward obesity, the incidence of GDM is also increasing significantly. According to 2017 data from the International Diabetes Federation, approximately 14% of pregnant women worldwide (about 18 million people) are affected by GDM each year [8]. The prevalence of GDM is even higher in Southeast Asia, reaching 24.2%. Although 70–85% of GDM patients can be treated through appropriate physical activity, diet, and lifestyle changes, 15–30% of patients still require medication [9]. These medications include insulin and oral glucose-lowering drugs [10]. Although newer oral antidiabetic drugs such as glipizide and metformin have emerged, there are still concerns about their long-term safety for both mother and child [3, 4]. Currently, there is no effective cure or prevention strategy for GDM. One reason is that the molecular mechanisms of GDM are still unclear [1].
Non-coding RNAs (ncRNAs) account for about 60% of the transcriptional products in the human genome and play a central regulatory role in many physiological and pathological processes, including cell proliferation, differentiation, apoptosis, and disease occurrence and development [11]. ncRNAs are divided into two major classes: small ncRNAs, such as microRNAs, which are less than 200 nucleotides in length, and long ncRNAs (lncRNAs), which are longer than 200 nucleotides [12, 13]. MiRNAs play a leading role in RNA silencing. MiRNA function by pairing with target mRNA sequences, which can be located in coding regions or intronic regions of non-coding transcripts, and even in exonic regions [14]. According to the latest statistics from 2018, 2654 mature microRNA sequences have been discovered in humans [15]. The functions of many miRNAs are still being defined. Existing research data has shown that a large number of miRNAs play a key role in gene regulation. Their dysregulation is associated with various diseases, from cancer to metabolic diseases [16, 17]. Multiple studies have shown that obesity, T2DM, and cardiovascular disease are associated with miRNA dysregulation or dysfunction, potentially playing a role in regulating beta-cell function and quality, as well as metabolic processes [18, 19]. According to the whole genome analysis, there are more than 600 miRNAs present in the placenta, which may play an important role in pregnancy and GDM [18, 20, 21]. Cao et al. found that the expression of miR-98 in placental tissue of pregnant women with GDM at 37–40 weeks was significantly higher than that of normal pregnant women (n = 202). MiR-98 can directly regulate the transcription factor methyl-CpG binding protein 2, interfere with glucose uptake in GDM by controlling the activity of transient receptor potential cation channel subfamily C member 3 [22]. Nair et al. found that compared to normal controls, there were 13 up-regulated miRNAs and 14 down-regulated miRNAs in GDM. It should be noted that the predicted target genes of these miRNAs are largely involved in glucose metabolism, including the phosphatidylinositol 3-kinase (PI3K)/protein kinase B (AKT) signaling pathway [23, 24]. Li et al. showed that miR-96 was down-regulated in GDM placental tissue and may alter beta-cell function by targeting p21 (RAC1) activated kinase 1 [25, 26].
Given that a single miRNA can target hundreds of mRNAs, and a specific target mRNA is typically controlled by several different miRNAs, to study such complex network control mechanism of miRNAs in GDM is a time-consuming and inefficient choice using traditional biomedical research methods. This study used RNA sequencing transcriptomics analysis, which has the ability to study large amounts of data using computer technology, to compare and analyze the variations of plasma miRNAs in GDM patients and normal pregnant women. It provides scientific data and evidence for further research and explanation of the complex pathological mechanisms of GDM. These abnormally expressed miRNAs in GDM may also become early diagnostic biomarkers or therapeutic targets for GDM in the future.
Methods
Ethics and subjects
The GDM patients were all recruited from the outpatient clinic of The Frist Hospital of PuTian City. Normal controls exhibited normal glucose tolerance. GDM diagnosis was based on the 75 g oral glucose tolerance test (OGTT), with the diagnosis criteria fasting plasma glucose ≥5.1 mmol/L, 1 h plasma glucose ≥10.0 mmol/L or 2 h plasma glucose ≥8.5 mmol/L following the 75 g OGTT. The patients were excluded with the following criteria: known pre-existing diabetes mellitus before pregnancy; history of chronic renal disease or hepatic dysfunction; any significant medical condition that could interfere with glucose metabolism; multiple pregnancies (e.g., twins or triplets); incomplete or missing data from the oral glucose tolerance test (OGTT). The study protocol received approval from the institutional ethics committee, and informed consent was obtained from all participants. The confidentiality and privacy of participant information were strictly maintained throughout the study.
Peripheral blood collection for miRNA sequencing
Ten samples were collected, including peripheral blood from five GDM patients and five controls. Blood was stored in an anticoagulant tube containing EDTA, immediately frozen in liquid nitrogen, and stored at −80 ℃ for subsequent tests.
RNA extraction and miRNA sequencing
Total RNA was extracted from peripheral blood using the miRNeasy Mini Kit (Cat No. 217004, Qiagen, CA, USA). Total RNA quantity and purity were analyzed using RNA Nano 6000 Assay Kit of the Agilent Bioanalyzer 2100 System (Agilent Technologies, CA, USA) with RIN number ≥ 8. Then, approximately 1.5 μg of total RNA was used to build a small RNA library using the Illumina VAHTSTM Small RNA Library Prep Kit (Illumina Inc., CA, USA). Subsequently, unique sequences with length of 18–30 nt were mapped to specific species precursors with miRbase v22 by BLAST search. All procedures were performed by Biomarker Technologies.
qRT-PCR for further verification
Total RNA was extracted from peripheral blood using the miRNeasy Mini Kit (Cat No. 217004, Qiagen, CA, USA) and U6 was used as an internal reference to normalize miRNA expression. Total RNA from each sample was reverse-transcribed using M-MuLV Reverse Transcriptase (Cat No. P7040L, Enzymatics, USA), according to the manufacture’s protocol. qRT-PCR was performed with a 2× PCR master mix (Cat No. AS-MR-006-5, Arraystar, MD, USA) to quantify miRNA expression. After pre-denaturation at 95 ℃ for 10 min, 40 PCR cycles were performed (95 ℃ for 10 s, and 60 ℃ for 60 s). The expression of target genes was analyzed using the 2−ΔΔCt method [27]. The primer information for U6 and miRNAs is shown in Table S1.
Statistical analyses
Data were analyzed using SPSS 22.0 software (IBM Corporation, Armonk, NY, USA) and GraphPad Prism version 8.0 (GraphPad Software Inc., San Diego, CA, USA). Variables were expressed as the mean ± standard deviation. Differences between groups were analyzed using Student’s t test. P < 0.05 was considered statistically significant.
Bioinformatics analyses
The differential expression analysis of two groups was performed using the DESeq2 R package (1.10.1). miRNAs with |log2(FC)| ≥ 0.58, and P < 0.05 were assigned as differentially expressed. The target genes of the differentially expressed miRNAs were predicted with selectively predicted algorithms in two independent online software programs: miRanda v3.3a and TargetScanS v 5.0 software. Novel miRNA prediction was carried out using miRDeep2 [28]. Gene Ontology (GO) enrichment analysis of the differentially expressed genes was implemented by the ClusterProfiler R packages based on Wallenius non-central hyper-geometric distribution. Kyoto Encyclopedia of Gene and Genomes (KEGG) pathway analysis was carried out to investigate the potential significant pathways (http://www.genome.jp/kegg/) through the KOBAS software.
Results
Clinical characteristics
A total of ten women were included in the study, five with normal healthy pregnancies and five with GDM. Maternal demographics are detailed in Table 1. Although no change in HbA1c between groups was observed in the first trimester of pregnancy ruling out pre-existing diabetes mellitus, the 1 h OGTT test revealed higher blood glucose concentrations in GDM versus normal pregnancies (p = 0.003). There were no significant differences in BMI, age, weight, or height between the two groups.
Identification of differentially expressed miRNAs
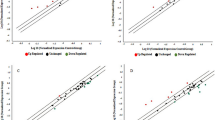
A total of 2287 miRNAs were identified, including 1420 known miRNAs in miRbase and 867 novel miRNAs predicted by miRDeep2. In total, 229 differentially expressed miRNAs were determined to have a |log2(FC)| ≥ 0.58, and P < 0.05 by DESeq2 R package (1.10.1); 114 miRNAs were up-regulated and 115 miRNAs were down-regulated (Table S2). Volcano plot (Fig. 1) represented the significant differences in miRNA expression between the control (CTL) and GDM groups.
Validation of differentially expressed miRNA by qRT-PCR
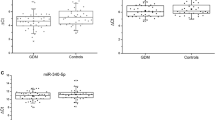
Among the differentially expressed miRNAs mentioned above, 13 significant differentially expressed miRNAs (7 up-regulated miRNAs and 6 down-regulated miRNAs) with |log2(FC)| > 1.4 were selected for validation by qRT-PCR (Table 2). As shown in Fig. 2, the expression levels of miR-5193, miR-5003-3p, miR-3127-5p, novel-miR-96, miR-6734-5p, and miR-122-5p in peripheral blood of GDM patient were significantly up-regulated (P < 0.05), and the expression levels of miR-10395-3p were significantly down-regulated (P < 0.05). However, the qRT-PCR results of other selected miRNAs were not consistent with the next-generation sequencing results.
The function of the target genes of miRNAs
Target genes of differentially expressed miRNAs were predicted by the miRanda and TargetScanS software. A total of 11,046 target genes were predicted. Gene Ontology (GO) and Kyoto Encyclopedia of Gene and Genomes (KEGG) pathway analyses were conducted to explore the main functions of target genes (Fig. 3). GO analysis of target genes showed that the most potential target genes are involved in the regulation of transcription, biosynthetic process, integral component membrane, and DNA binding (Fig. 3A–C). Regulation of transcription and biosynthetic process were in biological processes, integral component membrane was encompassed in cellular component, and DNA binding was involved in molecular function. KEGG pathways for these genes were mainly enriched in the herpes simplex virus 1 infection (Fig. 3D).
Function analysis of target genes of miRNAs. A, B and C The most enriched GO terms of target genes of miRNA. X-axis represents the gene ratio, and Y-axis represents the GO terms. D Enrichment analysis of target gene KEGG pathway. X-axis represents the enriched factor and Y-axis represents the KEGG pathways. The color of the dot indicates the different q-values. The size of the dot indicates the gene number
Discussion
GDM is the most common pregnancy complication that imposes a serious short- and long-term health risk to mother and child. While most GDM patients can recover afterward, poor glycemic control in some cases may result in the development of type 2 diabetes and cardiovascular diseases [6]. An effective therapy that can reduce the incidence of GDM is a major research priority for public health. Many molecular biomarkers for GDM have been investigated, including single-nucleotide polymorphisms (SNPs), metabolites, miRNAs, and proteins.
With the development of molecular biotechnology, non-coding RNA have received increasing attention in recent years. miRNAs are a subset of non-coding RNAs that act as negative regulators in gene expression. Many studies have shown that miRNA is widely involved in cell growth and development, metabolism, and apoptosis, and is strongly associated with human diseases. Over the years, evidence has increasingly shown that miRNA dysregulation has been linked to diabetes. It was reported that miRNA play an important role in type 1 and type 2 diabetes, including in beta-cell biology, insulin resistance, and diabetes complications [29]. In recent years, several miRNAs up-regulated in GDM patients have been identified. miR-29a, its serum expression was significantly down-regulated in pregnant women with GDM [30]. miR-657 affects macrophage-mediated immunity and inflammation in GDM [31]. miR-770-5p and miR-96 play protective roles in GDM by contributing to β-cell proliferation [26, 32]. Serum aberrant expression of miR-132 may exert a protective role against GDM by reducing the inhibition of high glucose in trophoblast cell proliferation [33]. These findings clearly establish the importance of serum miRNA in GDM patients. Moreover, miRNA can be collected from peripheral blood. Therefore, detecting the expression of miRNA in peripheral blood is a novel and simple method for GDM.
In this study, we collect peripheral blood of GDM patients to identify potential miRNA biomarker. A total of 2287 miRNAs were identified, including 229 differentially expressed genes. We performed qRT-PCR on 13 differentially expressed miRNAs based on our predicted sequences. The results showed that the expression of partial selected miRNAs was in line with the expression in sequencing. The expression levels of miR-5193, miR-5003-3p, miR-3127-5p, novel-miR-96, miR-6734-5p, and miR-122-5p in peripheral blood of GDM patients were significantly increased (P < 0.05), and the expression levels of miR-10395-3p were significantly decreased (P < 0.05). It has been shown that miR-5193 plays a significant role in prostate cancer and ovarian cancer [34, 35]. EIF4A3-regulated circ_0087429 can reverse EMT and inhibit cervical cancer progression via miR-5003-3p-dependent up-regulation of OGN expression [36]. The involvement of miR-3127-5p in preeclampsia has been documented, attributed to the fact that overexpression of its target gene HOXA7 can reverse the effects of miR-3127-5p in trophoblast cells [37]. The aberrant functionality of trophoblast cells is a crucial factor in the occurrence of preeclampsia. A previous study suggested that miR-6734-5p could be a notable miRNA associated with high-grade serous ovarian cancer [38]. miR-10395-3p was poorly studied, and currently, its dysregulation has only been supposed to play a significant role in lung cancer [39]. Unfortunately, in even GDM diabetes models, these miRNAs have not been reported to have relevant effects. This study firstly shows that these miRNAs may have essential roles in GDM. miRNA showing dysregulated expression in GDM might represent potential therapeutic targets for novel interventions. Further research is needed to validate the diagnostic, prognostic, and therapeutic utility of the identified miRNA biomarkers in larger patient cohorts and diverse populations.
Previously, miR-122-5p showed 2.55-fold increase in the T2D-liver cancer [40]. miRNA profiles in extracellular vesicles from GDM showed that miR-122-5p showed significantly higher levels in GDM cases than in controls [41]. In the insulin receptor signaling pathway, increased expression of miR-122-5p is predicted to inhibit insulin binding to the insulin receptor protein [41]. In this study, miR-122-5p was significantly up-regulated in GDM cases (P < 0.05), which is consistent with previous report. It indicates that they may play an important role in the pathogenesis of GDM and are expected to become new molecular biomarkers of GDM.
In recent years, extensive studies have shown that several key pathways involved in the development of GDM are associated with the pathophysiology of T2DM. The nuclear factor-kappa light chain enhancer of activated B cells (NF-κB) signaling pathway has critical role in gene expression for immune and inflammatory responses [42]. The importance of the NF-κB pathway extends to GDM and has been reported in the GDM placenta with increased NF-κB mRNA [43]. It is also worth noting that increased levels of chorionic gonadotrophin (CG) in GDM can impair insulin signaling in adipocytes through the NF-κB pathway [44]. Toll-like receptors (TLRs) are key surface molecules with and an essential role in triggering and inflammatory innate immune response [45]. Tangeras et al. showed that most TLRs are functionally active in human placenta. GDM is associated with increased expression of MyD88 and TLR4 mRNA in the placenta [43, 46]. The PI3K/mTOR pathway is required for survival in environments with variable nutrient availability. Compared with the expression of mTOR pathway in placentas of normal term babies and GDM babies, the increased expression of the ribosomal protein p-p70S6K, a downstream component of the mTOR signaling network, indicates that mTOR plays a role in the observed pathology in the placenta of GDM births [47]. Glycogen synthase kinase 3 (GSK3), a serine/threonine protein kinase, was originally found to have a role in the storage of glucose in glycogen [48]. Women with GDM showed significantly reduced GSK3β serine phosphorylation in their skeletal muscle and omental adipose tissue [49]. Similarly, AMPK activity was also significantly reduced in skeletal muscle and adipose tissues of GDM patients [50, 51]. In pregnant adipose tissue, the inflammasome processes IL-1β secretion in TLRs and pro-inflammatory cytokine signaling pathways [52]. Recently, it also reported that placental VEGF and CD31 expression in pregnancies complicated by GDM show influence on pregestational BMI and gestational weight gain in women with GDM [53].
We performed GO and KEGG pathway analyses to explore the biological functions and potential pathways in genes of differentially expressed miRNAs. In the GO enrichment analysis, the most significant biological processes are regulation of the transcription and biosynthetic process, the predominant cellular component is the integral membrane component, and the principal molecular functions are transcription factor activity and sequences-specific DNA binding. KEGG pathway analysis showed that the most significantly enriched pathway is the herpes simplex virus 1 infection pathway. Type 1 herpes simplex virus (HSV-1) is one of the most common human antigens, infecting billions of people. Each year, between 250,000 and 500,000 of every million virus-infected individuals experience severe symptoms, leading to herpes simplex encephalitis, and most of these unfortunate cases occur in children under the age of 3 years [54]. Through RIG-I affinity purification and RNA sequencing of cells infected with the herpes simplex virus, it was revealed that small non-coding RNAs, especially RNA5SP141, constitute a class of intracellular ligands for RIG-I [55]. Gene mutations in nucleic acid sensing components such as TLR3 or its downstream signaling molecules (including UNC93B1, TLR3, TRIF, TRAF3, TBK1, IRF3, etc.) are one of the causes of herpes simplex encephalitis. It was reported that TLR3 ligand significantly increased the expression of a number of inflammatory markers in the skeletal muscle of pregnant woman [56]. Additionally, treatment of skeletal muscle of pregnant women with poly(I:C) could result in a decrease in glucose uptake [57]. All these findings strongly indicate that differentially expressed miRNAs in GDMs, along with their target genes, play a crucial role in the pathogenesis of the disease.
Conclusion
In conclusion, this study provides insights into the differential expression of miRNAs in GDM patients and their potential roles in disease pathogenesis. Identified miRNAs, especially miR-5193, miR-5003-3p, miR-3127-5p, novel-miR-96, miR-6734-5p, miR-122-5p, and miR-10395-3p, may serve as novel biomarkers for GDM. More research is warranted to elucidate the specific functions and mechanisms of these miRNAs in GDM, paving the way for improved diagnosis and therapeutic interventions.
Availability of data and materials
The data sets used and analyzed during this study are available and under the domain of the corresponding author (plar000@163.com).
References
Plows JF, Stanley JL, Baker PN, Reynolds CM, Vickers MH (2018) The pathophysiology of gestational diabetes mellitus. Int J Mol Sci 19. https://doi.org/10.3390/ijms19113342
American Diabetes A (2018) 2. Classification and diagnosis of diabetes: standards of medical care in diabetes-2018. Diabetes Care 41:S13–S27. https://doi.org/10.2337/dc18-S002
Camelo Castillo W, Boggess K, Sturmer T, Brookhart MA, Benjamin DK Jr, Jonsson Funk M (2015) Association of adverse pregnancy outcomes with glyburide vs insulin in women with gestational diabetes. JAMA Pediatr 169:452–458. https://doi.org/10.1001/jamapediatrics.2015.74
Feig DS, Moses RG (2011) Metformin therapy during pregnancy: good for the goose and good for the gosling too? Diabetes Care 34:2329–2330. https://doi.org/10.2337/dc11-1153
Dincgez B, Ercan I, Sahin I, Erturk NK (2024) The risk of developing gestational diabetes mellitus in maternal subclinical hypothyroidism: a systematic review and meta-analysis. Arch Gynecol Obstet 309:765–774. https://doi.org/10.1007/s00404-023-07137-y
Yogev Y, Xenakis EM, Langer O (2004) The association between preeclampsia and the severity of gestational diabetes: the impact of glycemic control. Am J Obstet Gynecol 191:1655–1660. https://doi.org/10.1016/j.ajog.2004.03.074
Group HSCR, Metzger BE, Lowe LP, Dyer AR, Trimble ER, Chaovarindr U, Coustan DR, Hadden DR, McCance DR, Hod M, McIntyre HD, Oats JJ, Persson B, Rogers MS, Sacks DA (2008) Hyperglycemia and adverse pregnancy outcomes. N Engl J Med 358:1991–2002. https://doi.org/10.1056/NEJMoa0707943
Federation IDJIDF (2017) IDF diabetes atlas 8th ed. pp. 905–911
Johns EC, Denison FC, Norman JE, Reynolds RM (2018) Gestational diabetes mellitus: mechanisms, treatment, and complications. Trends Endocrinol Metab 29:743–754. https://doi.org/10.1016/j.tem.2018.09.004
Lende M, Rijhsinghani A (2020) Gestational diabetes: overview with emphasis on medical management. Int J Environ Res Public Health 17. https://doi.org/10.3390/ijerph17249573
Filardi T, Catanzaro G, Mardente S, Zicari A, Santangelo C, Lenzi A, Morano S, Ferretti E (2020) Non-coding RNA: role in gestational diabetes pathophysiology and complications. Int J Mol Sci 21. https://doi.org/10.3390/ijms21114020
Anfossi S, Babayan A, Pantel K, Calin GA (2018) Clinical utility of circulating non-coding RNAs—an update. Nat Rev Clin Oncol 15:541–563. https://doi.org/10.1038/s41571-018-0035-x
Chen B, Huang S (2018) Circular RNA: an emerging non-coding RNA as a regulator and biomarker in cancer. Cancer Lett 418:41–50. https://doi.org/10.1016/j.canlet.2018.01.011
Ha M, Kim VN (2014) Regulation of microRNA biogenesis. Nat Rev Mol Cell Biol 15:509–524. https://doi.org/10.1038/nrm3838
Kozomara A, Birgaoanu M, Griffiths-Jones S (2019) miRBase: from microRNA sequences to function. Nucleic Acids Res 47:D155–D162. https://doi.org/10.1093/nar/gky1141
Peng Y, Croce CM (2016) The role of microRNAs in human cancer. Signal Transduct Target Ther 1:15004. https://doi.org/10.1038/sigtrans.2015.4
Soifer HS, Rossi JJ, Saetrom P (2007) MicroRNAs in disease and potential therapeutic applications. Mol Ther 15:2070–2079. https://doi.org/10.1038/sj.mt.6300311
Guay C, Regazzi R (2016) New emerging tasks for microRNAs in the control of beta-cell activities. Biochim Biophys Acta 1861:2121–2129. https://doi.org/10.1016/j.bbalip.2016.05.003
Liu ZN, Jiang Y, Liu XQ, Yang MM, Chen C, Zhao BH, Huang HF, Luo Q (2021) MiRNAs in gestational diabetes mellitus: potential mechanisms and clinical applications. J Diabetes Res 2021:4632745. https://doi.org/10.1155/2021/4632745
Chen DB, Wang W (2013) Human placental microRNAs and preeclampsia. Biol Reprod 88:130. https://doi.org/10.1095/biolreprod.113.107805
Iljas JD, Guanzon D, Elfeky O, Rice GE, Salomon C (2017) Review: bio-compartmentalization of microRNAs in exosomes during gestational diabetes mellitus. Placenta 54:76–82. https://doi.org/10.1016/j.placenta.2016.12.002
Cao JL, Zhang L, Li J, Tian S, Lv XD, Wang XQ, Su X, Li Y, Hu Y, Ma X, Xia HF (2016) Up-regulation of miR-98 and unraveling regulatory mechanisms in gestational diabetes mellitus. Sci Rep 6:32268. https://doi.org/10.1038/srep32268
Nair S, Jayabalan N, Guanzon D, Palma C, Scholz-Romero K, Elfeky O, Zuniga F, Ormazabal V, Diaz E, Rice GE, Duncombe G, Jansson T, McIntyre HD, Lappas M, Salomon C (2018) Human placental exosomes in gestational diabetes mellitus carry a specific set of miRNAs associated with skeletal muscle insulin sensitivity. Clin Sci (Lond) 132:2451–2467. https://doi.org/10.1042/CS20180487
Nguyen-Ngo C, Jayabalan N, Salomon C, Lappas M (2019) Molecular pathways disrupted by gestational diabetes mellitus. J Mol Endocrinol 63:R51–R72. https://doi.org/10.1530/JME-18-0274
Ahn M, Yoder SM, Wang Z, Oh E, Ramalingam L, Tunduguru R, Thurmond DC (2016) The p21-activated kinase (PAK1) is involved in diet-induced beta cell mass expansion and survival in mice and human islets. Diabetologia 59:2145–2155. https://doi.org/10.1007/s00125-016-4042-0
Li L, Wang S, Li H, Wan J, Zhou Q, Zhou Y, Zhang C (2018) microRNA-96 protects pancreatic beta-cell function by targeting PAK1 in gestational diabetes mellitus. BioFactors 44:539–547. https://doi.org/10.1002/biof.1461
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Friedlander MR, Mackowiak SD, Li N, Chen W, Rajewsky N (2012) miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res 40:37–52. https://doi.org/10.1093/nar/gkr688
Fernandez-Valverde SL, Taft RJ, Mattick JS (2011) MicroRNAs in beta-cell biology, insulin resistance, diabetes and its complications. Diabetes 60:1825–1831. https://doi.org/10.2337/db11-0171
Zhao C, Dong J, Jiang T, Shi Z, Yu B, Zhu Y, Chen D, Xu J, Huo R, Dai J, Xia Y, Pan S, Hu Z, Sha J (2011) Early second-trimester serum miRNA profiling predicts gestational diabetes mellitus. PLoS ONE 6:e23925. https://doi.org/10.1371/journal.pone.0023925
Wang P, Wang Z, Liu G, Jin C, Zhang Q, Man S, Wang Z (2019) miR-657 promotes macrophage polarization toward M1 by targeting FAM46C in gestational diabetes mellitus. Mediators Inflamm 2019:4851214. https://doi.org/10.1155/2019/4851214
Zhang YL, Chen XQ (2020) Dysregulation of microRNA-770-5p influences pancreatic-beta-cell function by targeting TP53 regulated inhibitor of apoptosis 1 in gestational diabetes mellitus. Eur Rev Med Pharmacol Sci 24:793–801. https://doi.org/10.26355/eurrev_202001_20062
Zhou X, Xiang C, Zheng X (2019) miR-132 serves as a diagnostic biomarker in gestational diabetes mellitus and its regulatory effect on trophoblast cell viability. Diagn Pathol 14:119. https://doi.org/10.1186/s13000-019-0899-9
Song Z, Guo Q, Wang H, Gao L, Wang S, Liu D, Liu J, Qi Y, Lin B (2020) miR-5193, regulated by FUT1, suppresses proliferation and migration of ovarian cancer cells by targeting TRIM11. Pathol Res Pract 216:153148. https://doi.org/10.1016/j.prp.2020.153148
Pan Y, Zhang R, Chen H, Chen W, Wu K, Lv J (2019) Expression of tripartite motif-containing proteactiin 11 (TRIM11) is associated with the progression of human prostate cancer and is downregulated by microRNA-5193. Med Sci Monit 25:98–106. https://doi.org/10.12659/MSM.911818
Yang M, Hu H, Wu S, Ding J, Yin B, Huang B, Li F, Guo X, Han L (2022) EIF4A3-regulated circ_0087429 can reverse EMT and inhibit the progression of cervical cancer via miR-5003-3p-dependent upregulation of OGN expression. J Exp Clin Cancer Res 41:165. https://doi.org/10.1186/s13046-022-02368-4
Li J, Han J, Zhao A, Zhang G (2022) CircPAPPA regulates the proliferation, migration, invasion, apoptosis, and cell cycle of trophoblast cells through the miR-3127-5p/HOXA7 axis. Reprod Sci 29:1215–1225. https://doi.org/10.1007/s43032-021-00802-0
Wang R, Du X, Zhi Y (2020) Screening of critical genes involved in metastasis and prognosis of high-grade serous ovarian cancer by gene expression profile data. J Comput Biol 27:1104–1114. https://doi.org/10.1089/cmb.2019.0235
Zhang D, Yang Y, Kang Y, Xie D, Zhang X, Hao J (2023) Dysregulated expression of microRNA involved in resistance to osimertinib in EGFR mutant non-small cell lung cancer cells. J Thorac Dis 15:1978–1993. https://doi.org/10.21037/jtd-23-401
Lee HM, Wong WKK, Fan B, Lau ES, Hou Y, O CK, Luk AOY, Chow EYK, Ma RCW, Chan JCN, Kong APS (2021) Detection of increased serum miR-122-5p and miR-455-3p levels before the clinical diagnosis of liver cancer in people with type 2 diabetes. Sci Rep 11:23756https://doi.org/10.1038/s41598-021-03222-x
Gillet V, Ouellet A, Stepanov Y, Rodosthenous RS, Croft EK, Brennan K, Abdelouahab N, Baccarelli A, Takser L (2019) miRNA profiles in extracellular vesicles from serum early in pregnancies complicated by gestational diabetes mellitus. J Clin Endocrinol Metab 104:5157–5169. https://doi.org/10.1210/jc.2018-02693
Baldwin AS Jr (1996) The NF-kappa B and I kappa B proteins: new discoveries and insights. Annu Rev Immunol 14:649–683. https://doi.org/10.1146/annurev.immunol.14.1.649
Feng H, Su R, Song Y, Wang C, Lin L, Ma J, Yang H (2016) Positive correlation between enhanced expression of TLR4/MyD88/NF-kappaB with insulin resistance in placentae of gestational diabetes mellitus. PLoS ONE 11:e0157185. https://doi.org/10.1371/journal.pone.0157185
Ma Q, Fan J, Wang J, Yang S, Cong Q, Wang R, Lv Q, Liu R, Ning G (2015) High levels of chorionic gonadotrophin attenuate insulin sensitivity and promote inflammation in adipocytes. J Mol Endocrinol 54:161–170. https://doi.org/10.1530/JME-14-0284
Vasselon T, Detmers PA (2002) Toll receptors: a central element in innate immune responses. Infect Immun 70:1033–1041. https://doi.org/10.1128/IAI.70.3.1033-1041.2002
Mrizak I, Grissa O, Henault B, Fekih M, Bouslema A, Boumaiza I, Zaouali M, Tabka Z, Khan NA (2014) Placental infiltration of inflammatory markers in gestational diabetic women. Gen Physiol Biophys 33:169–176. https://doi.org/10.4149/gpb_2013075
Sati L, Soygur B, Celik-Ozenci C (2016) Expression of mammalian target of rapamycin and downstream targets in normal and gestational diabetic human term placenta. Reprod Sci 23:324–332. https://doi.org/10.1177/1933719115602765
Woodgett JR (1990) Molecular cloning and expression of glycogen synthase kinase-3/factor A. EMBO J 9:2431–2438. https://doi.org/10.1002/j.1460-2075.1990.tb07419.x
Lappas M (2014) GSK3beta is increased in adipose tissue and skeletal muscle from women with gestational diabetes where it regulates the inflammatory response. PLoS ONE 9:e115854. https://doi.org/10.1371/journal.pone.0115854
Liong S, Lappas M (2015) Activation of AMPK improves inflammation and insulin resistance in adipose tissue and skeletal muscle from pregnant women. J Physiol Biochem 71:703–717. https://doi.org/10.1007/s13105-015-0435-7
Boyle KE, Hwang H, Janssen RC, DeVente JM, Barbour LA, Hernandez TL, Mandarino LJ, Lappas M, Friedman JE (2014) Gestational diabetes is characterized by reduced mitochondrial protein expression and altered calcium signaling proteins in skeletal muscle. PLoS ONE 9:e106872. https://doi.org/10.1371/journal.pone.0106872
Lappas M (2014) Activation of inflammasomes in adipose tissue of women with gestational diabetes. Mol Cell Endocrinol 382:74–83. https://doi.org/10.1016/j.mce.2013.09.011
Sirico A, Rossi ED, Degennaro VA, Arena V, Rizzi A, Tartaglione L, Di Leo M, Pitocco D, Lanzone A (2023) Placental diabesity: placental VEGF and CD31 expression according to pregestational BMI and gestational weight gain in women with gestational diabetes. Arch Gynecol Obstet 307:1823–1831. https://doi.org/10.1007/s00404-022-06673-3
Whitley RJ, Kimberlin DW (2005) Herpes simplex encephalitis: children and adolescents. Semin Pediatr Infect Dis 16:17–23. https://doi.org/10.1053/j.spid.2004.09.007
Chiu YH, Macmillan JB, Chen ZJ (2009) RNA polymerase III detects cytosolic DNA and induces type I interferons through the RIG-I pathway. Cell 138:576–591. https://doi.org/10.1016/j.cell.2009.06.015
Lappas M (2015) Double stranded viral RNA induces inflammation and insulin resistance in skeletal muscle from pregnant women in vitro. Metabolism 64:642–653. https://doi.org/10.1016/j.metabol.2015.02.002
Kirwan JP, Hauguel-De Mouzon S, Lepercq J, Challier JC, Huston-Presley L, Friedman JE, Kalhan SC, Catalano PM (2002) TNF-alpha is a predictor of insulin resistance in human pregnancy. Diabetes 51:2207–2213. https://doi.org/10.2337/diabetes.51.7.2207
Funding
This work was supported by the Startup Fund for Advanced Talents of Putian University (No. 2020009) and Research Projects of Putian University (No. 2021056).
Author information
Authors and Affiliations
Contributions
HL, XC, and TZ conceived and designed the study. XC and XL collected the blood from patients and controls. SP extracted the total RNA and conducted further validation assay. HL and HC drafted the paper or substantially revised it. The manuscript was written through contributions of all authors. TZ and SP analyzed the data, wrote the main manuscript text, and prepared the paper. All authors reviewed the manuscript and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval and consent to participate
Permission to conduct the study was obtained from the Ethics Committee of the First Hospital of Putian City. All methods were performed in accordance with the relevant guidelines and regulations. The informed consent was obtained from all participants. Confidentiality and privacy of participant information were strictly maintained throughout the study.
Consent to publish
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lin, H., Chen, X., Wang, L. et al. Unraveling the role of microRNAs: potential biomarkers for gestational diabetes mellitus revealed through RNA sequencing analysis. Arch Gynecol Obstet 310, 1255–1264 (2024). https://doi.org/10.1007/s00404-024-07518-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00404-024-07518-x