Abstract
Objective
Cervical cancer (CC) is one of the most common types of malignant female cancer, and its incidence and mortality are not optimistic. Protein panels can be a powerful prognostic factor for many types of cancer. The purpose of our study was to investigate a proteomic panel to predict the survival of patients with common CC.
Methods and results
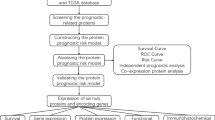
The protein expression and clinicopathological data of CC were downloaded from The Cancer Proteome Atlas and The Cancer Genome Atlas database, respectively. We selected the prognosis-related proteins (PRPs) by univariate Cox regression analysis and found that the results of functional enrichment analysis were mainly related to apoptosis. We used Kaplan–Meier analysis and multivariable Cox regression analysis further to screen PRPs to establish a prognostic model, including BCL2, SMAD3, and 4EBP1-pT70. The signature was verified to be independent predictors of OS by Cox regression analysis and the area under curves. Nomogram and subgroup classification were established based on the signature to verify its clinical application. Furthermore, we looked for the co-expressed proteins of three-protein panel as potential prognostic proteins.
Conclusion
A proteomic signature independently predicted OS of CC patients, and the predictive ability was better than the clinicopathological characteristics. This signature can help improve prediction for clinical outcome and provides new targets for CC treatment.










Similar content being viewed by others
Availability of data and materials
We obtained the data sets from TCPA (https://www.tcpaportal.org/) and TCGA (http://cancergenome.nih.gov/).
References
Siegel RL, Miller KD, Fuchs HE, Jemal A (2022) Cancer statistics, 2022. CA A Cancer J Clin 72:7–33
Nogueira-Rodrigues A, Ferreira CG, Bergmann A, de Aguiar SS, Thuler LC (2014) Comparison of adenocarcinoma (ACA) and squamous cell carcinoma (SCC) of the uterine cervix in a sub-optimally screened cohort: a population-based epidemiologic study of 51,842 women in Brazil. Gynecol Oncol 135:292–296
Bhatla N, Aoki D, Sharma DN, Sankaranarayanan R (2018) Cancer of the cervix uteri. Int J Gynaecol Obstet 143(Suppl 2):22–36
Dasari S, Wudayagiri R, Valluru L (2015) Cervical cancer: Biomarkers for diagnosis and treatment. Clin Chim Acta 445:7–11
Shukla HD (2017) Comprehensive analysis of cancer-proteogenome to identify biomarkers for the early diagnosis and prognosis of cancer. Proteomes 5:28
Hirose S, Murakami N, Takahashi K et al (2020) Genomic alterations in STK11 can predict clinical outcomes in cervical cancer patients. Gynecol Oncol 156:203–210
Hu X, Schwarz JK, Lewis JS Jr et al (2010) A microRNA expression signature for cervical cancer prognosis. Cancer Res 70:1441–1448
Mao X, Qin X, Li L et al (2018) A 15-long non-coding RNA signature to improve prognosis prediction of cervical squamous cell carcinoma. Gynecol Oncol 149:181–187
Cai S, Yu X, Gu Z et al (2020) A 10-gene prognostic methylation signature for stage I-III cervical cancer. Arch Gynecol Obstet 301:1275–1287
Choi CH, Chung JY, Kang JH et al (2020) Chemoradiotherapy response prediction model by proteomic expressional profiling in patients with locally advanced cervical cancer. Gynecol Oncol 157(2):437–443
Masuda M, Yamada T (2019) Utility of reverse-phase protein array for refining precision oncology. Adv Exp Med Biol 1188:239–249
Shan Z, Luo D, Liu Q et al (2021) Proteomic profiling reveals a signature for optimizing prognostic prediction in Colon Cancer. J Cancer 12:2199–2205
Chu G, Xu T, Zhu G, Liu S, Niu H, Zhang M (2021) Identification of a novel protein-based signature to improve prognosis prediction in renal clear cell carcinoma. Front Mol Biosci 8:623120
Chen MM, Li J, Wang Y et al (2019) TCPA v3.0: an integrative platform to explore the pan-cancer analysis of functional proteomic data. Mol Cell Proteomics 18:S15-s25
Tan GS, Lim KH, Tan HT et al (2014) Novel proteomic biomarker panel for prediction of aggressive metastatic hepatocellular carcinoma relapse in surgically resectable patients. J Proteome Res 13:4833–4846
Liu X, Wu Y, Liu P, Zhang X (2022) Developing a validated nomogram for predicting ovarian metastasis in endometrial cancer patients: a retrospective research. Arch Gynecol Obstet 305:719–729
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A (2015) Global cancer statistics, 2012. CA Cancer J Clin 65:87–108
Matsuo K, Chang EJ, Matsuzaki S, Shimada M, Ciccone MA, Roman LD (2021) Recent changes in demographics and outcomes of cervical cancer in the United States. Arch Gynecol Obstet 304:1–3
Shi X, Li Y, Sun Y et al (2020) Genome-wide analysis of lncRNAs, miRNAs, and mRNAs forming a prognostic scoring system in esophageal squamous cell carcinoma. PeerJ 8:e8368
Hou Z, Yang J, Wang H, Liu D, Zhang H (2019) A potential prognostic gene signature for predicting survival for glioblastoma patients. BioMed Research Int 2019:9506461
Fulda S (2009) Tumor resistance to apoptosis. Int J Cancer 124:511–515
Ngoi NYL, Choong C, Lee J et al (2020) Targeting mitochondrial apoptosis to overcome treatment resistance in cancer. Cancers (Basel) 12:574
Leverson JD, Cojocari D (2018) Hematologic tumor cell resistance to the BCL-2 Inhibitor venetoclax: a product of its microenvironment? Front Oncol 8:458
Kalkavan H, Green DR (2018) MOMP, cell suicide as a BCL-2 family business. Cell Death Differ 25:46–55
Woo SM, Kwon TK (2019) E3 ubiquitin ligases and deubiquitinases as modulators of TRAIL-mediated extrinsic apoptotic signaling pathway. BMB Rep 52:119–126
Ni T, Li W, Zou F (2005) The ubiquitin ligase ability of IAPs regulates apoptosis. IUBMB Life 57:779–785
Millet C, Zhang YE (2007) Roles of Smad3 in TGF-beta signaling during carcinogenesis. Crit Rev Eukaryot Gene Expr 17:281–293
Padua D, Massague J (2009) Roles of TGFbeta in metastasis. Cell Res 19:89–102
Yang S, Sun Y, Jiang D et al (2021) MiR-362 suppresses cervical cancer progression via directly targeting BAP31 and activating TGFβ/Smad pathway. Cancer Med 10:305–316
Chang QQ, Chen CY, Chen Z, Chang S (2019) LncRNA PVT1 promotes proliferation and invasion through enhancing Smad3 expression by sponging miR-140-5p in cervical cancer. Radiol Oncol 53:443–452
Fan Q, Qiu MT, Zhu Z et al (2015) Twist induces epithelial-mesenchymal transition in cervical carcinogenesis by regulating the TGF-beta/Smad3 signaling pathway. Oncol Rep 34:1787–1794
Xia Q, Li C, Bian P, Wang J, Dong S (2014) Targeting SMAD3 for inhibiting prostate cancer metastasis. Tumour Biol 35:8537–8541
Chen N, Balasenthil S, Reuther J et al (2013) DEAR1 is a chromosome 1p35 tumor suppressor and master regulator of TGF-beta-driven epithelial-mesenchymal transition. Cancer Discov 3:1172–1189
Karlsson E, Perez-Tenorio G, Amin R et al (2013) The mTOR effectors 4EBP1 and S6K2 are frequently coexpressed, and associated with a poor prognosis and endocrine resistance in breast cancer: a retrospective study including patients from the randomised Stockholm tamoxifen trials. Breast Cancer Res 15:R96
Zhu J, Wang M, Hu D (2020) Development of an autophagy-related gene prognostic signature in lung adenocarcinoma and lung squamous cell carcinoma. PeerJ 8:e8288
Rho SB, Byun HJ, Kim BR, Lee CH (2021) Knockdown of LKB1 sensitizes endometrial cancer cells via AMPK activation. Biomol Ther (Seoul) 29:650–657
Yu G, Chen L, Hu Y, Yuan Z, Luo Y, Xiong Y (2021) Antitumor effects of baicalein and its mechanism via TGFβ pathway in cervical cancer HeLa cells. Evid Based Complement Alternat Med 2021:5527190
Wang S, Pang T, Gao M et al (2013) HPV E6 induces eIF4E transcription to promote the proliferation and migration of cervical cancer. FEBS Lett 587:690–697
Pang T, Wang S, Gao M et al (2015) HPV18 E7 induces the over-transcription of eIF4E gene in cervical cancer. Iran J Basic Med Sci 18:684–690
Zhang YF, Zhang ZH, Li MY et al (2021) Britannin stabilizes T cell activity and inhibits proliferation and angiogenesis by targeting PD-L1 via abrogation of the crosstalk between Myc and HIF-1α in cancer. Phytomedicine 81:153425
Wang L, Guo C, Li X et al (2019) Design, synthesis and biological evaluation of bromophenol-thiazolylhydrazone hybrids inhibiting the interaction of translation initiation factors eIF4E/eIF4G as multifunctional agents for cancer treatment. Eur J Med Chem 177:153–170
Xiao D, Lu Z, Wang Z et al (2020) Synthesis, biological evaluation and anti-proliferative mechanism of fluorine-containing proguanil derivatives. Bioorg Med Chem 28:115258
Linette GP, Hess JL, Sentman CL, Korsmeyer SJ (1995) Peripheral T-cell lymphoma in lckpr-bcl-2 transgenic mice. Blood 86:1255–1260
Xie M, Park D, Sica GL, Deng X (2020) Bcl2-induced DNA replication stress promotes lung carcinogenesis in response to space radiation. Carcinogenesis 41:1565–1575
Yang F, Guo L, Cao Y, Li S, Li J, Liu M (2018) MicroRNA-7-5p promotes cisplatin resistance of cervical cancer cells and modulation of cellular energy homeostasis by regulating the expression of the PARP-1 and BCL2 genes. Med Sci Monit 24:6506–6516
Saito T, Takehara M, Tanaka R et al (2004) Correlation between responsiveness of neoadjuvant chemotherapy and apoptosis-associated proteins for cervical adenocarcinoma. Gynecol Oncol 92:284–292
Hu J, Fang Y, Cao Y, Qin R, Chen Q (2014) miR-449a Regulates proliferation and chemosensitivity to cisplatin by targeting cyclin D1 and BCL2 in SGC7901 cells. Dig Dis Sci 59:336–345
Yu L, Zhou GQ, Li DC (2018) MiR-136 triggers apoptosis in human gastric cancer cells by targeting AEG-1 and BCL2. Eur Rev Med Pharmacol Sci 22:7251–7256
Zhu Y, Wen X, Zhao P (2018) MicroRNA-365 inhibits cell growth and promotes apoptosis in melanoma by targeting BCL2 and cyclin D1 (CCND1). Med Sci Monit 24:3679–3692
Yumol J, Gabrielli B, Tayyar Y, McMillan NA, Idris A (2020) Smart drug combinations for cervical cancer: dual targeting of Bcl-2 family of proteins and aurora kinases. Am J Cancer Res 10:3406–3414
Lin Y, Li Z, Liu M, Ye H, He J, Chen J (2021) CD34 and Bcl-2 as predictors for the efficacy of neoadjuvant chemotherapy in cervical cancer. Arch Gynecol Obstet 304:495–501
Acknowledgements
We acknowledge the contributions of the TCPA Database and the TCGA Database gratefully.
Funding
This work was supported by the Shandong Province Nature Science Foundation (No. ZR2016HM08).
Author information
Authors and Affiliations
Contributions
XJ: Conceptualization, Writing-original draft, Writing-review & editing. GC: Writing-original draft, Methodology, Software. YC: Visualization, Investigation and Writing-review & editing. JJ: Software, Validation. TL: Methodology, Visualization. QY: Writing-Review & Editing, Project administration. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare that are relevant to the content of this article.
Consent for publication
All authors have seen the manuscript and approved to submit it to your journal. Neither the entire paper nor any part of its content has been published or has been accepted elsewhere.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
404_2022_6642_MOESM4_ESM.xlsx
Supplementary Table 4. Co-expressed proteins in a three-protein model. The absolute value of correlation coefficient were greater than 0.3 (XLSX 11 KB)
Rights and permissions
About this article
Cite this article
Ji, X., Chu, G., Chen, Y. et al. Comprehensive analysis of novel prognosis-related proteomic signature effectively improve risk stratification and precision treatment for patients with cervical cancer. Arch Gynecol Obstet 307, 903–917 (2023). https://doi.org/10.1007/s00404-022-06642-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00404-022-06642-w