Abstract
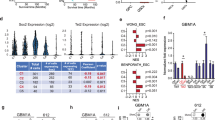
Favorable outcome after chemotherapy of glioblastomas cannot unequivocally be linked to promoter hypermethylation of the O 6-methylguanine-DNA methyltransferase (MGMT) gene encoding a DNA repair enzyme associated with resistance to alkylating agents. This indicates that molecular mechanisms determining MGMT expression have not yet been fully elucidated. We here show that glioblastomas are capable to downregulate MGMT expression independently of promoter methylation by elongation of the 3′-UTR of the mRNA, rendering the alternatively polyadenylated transcript susceptible to miRNA-mediated suppression. While the elongated transcript is poorly expressed in normal brain, its abundance in human glioblastoma specimens is inversely correlated with MGMT mRNA expression. Using a bioinformatically guided experimental approach, we identified miR-181d, miR-767-3p, and miR-648 as significant post-transcriptional regulators of MGMT in glioblastomas; the first two miRNAs induce MGMT mRNA degradation, the latter affects MGMT protein translation. A regression model including the two miRNAs influencing MGMT mRNA expression and the MGMT methylation status reliably predicts The Cancer Genome Atlas MGMT expression data. Responsivity of MGMT expressing T98G glioma cells to temozolomide was significantly enhanced after transfection of miR-181d, miR-767-3p, and miR-648. Taken together, our results uncovered alternative polyadenylation of the MGMT 3′-UTR and miRNA targeting as new mechanisms of MGMT silencing.





Similar content being viewed by others
References
Bady P, Sciuscio D, Diserens AC, Bloch J, van den Bent MJ, Marosi C, Dietrich PY, Weller M, Mariani L, Heppner FL, McDonald DR, Lacombe D, Stupp R, Delorenzi M, Hegi ME (2012) MGMT methylation analysis of glioblastoma on the Infinium methylation BeadChip identifies two distinct CpG regions associated with gene silencing and outcome, yielding a prediction model for comparisons across datasets, tumor grades, and CIMP-status. Acta Neuropathol 124(4):547–560. doi:10.1007/s00401-012-1016-2
McLendon, Friedman A, Bigner D et al (2008) Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455(7216):1061–1068. doi:10.1038/nature07385
Di Giammartino DC, Nishida K, Manley JL (2011) Mechanisms and consequences of alternative polyadenylation. Mol Cell 43(6):853–866. doi:10.1016/j.molcel.2011.08.017
Everhard S, Tost J, El Abdalaoui H, Criniere E, Busato F, Marie Y, Gut IG, Sanson M, Mokhtari K, Laigle-Donadey F, Hoang-Xuan K, Delattre JY, Thillet J (2009) Identification of regions correlating MGMT promoter methylation and gene expression in glioblastomas. Neuro Oncol 11(4):348–356. doi:10.1215/15228517-2009-001
Hegi ME, Diserens AC, Gorlia T, Hamou MF, de Tribolet N, Weller M, Kros JM, Hainfellner JA, Mason W, Mariani L, Bromberg JE, Hau P, Mirimanoff RO, Cairncross JG, Janzer RC, Stupp R (2005) MGMT gene silencing and benefit from temozolomide in glioblastoma. N Engl J Med 352(10):997–1003. doi:10.1056/NEJMoa043331
Helfer S, Schott J, Stoecklin G, Forstemann K (2012) AU-rich element-mediated mRNA decay can occur independently of the miRNA machinery in mouse embryonic fibroblasts and Drosophila S2-cells. PLoS One 7(1):e28907. doi:10.1371/journal.pone.0028907
Hermisson M, Klumpp A, Wick W, Wischhusen J, Nagel G, Roos W, Kaina B, Weller M (2006) O6-methylguanine DNA methyltransferase and p53 status predict temozolomide sensitivity in human malignant glioma cells. J Neurochem 96(3):766–776. doi:10.1111/j.1471-4159.2005.03583.x
Hu J, Lutz CS, Wilusz J, Tian B (2005) Bioinformatic identification of candidate cis-regulatory elements involved in human mRNA polyadenylation. RNA 11(10):1485–1493. doi:10.1261/rna.2107305
Ji Z, Lee JY, Pan Z, Jiang B, Tian B (2009) Progressive lengthening of 3′ untranslated regions of mRNAs by alternative polyadenylation during mouse embryonic development. Proc Natl Acad Sci USA 106(17):7028–7033. doi:10.1073/pnas.0900028106
Ji Z, Tian B (2009) Reprogramming of 3′ untranslated regions of mRNAs by alternative polyadenylation in generation of pluripotent stem cells from different cell types. PLoS One 4(12):e8419. doi:10.1371/journal.pone.0008419
John B, Enright AJ, Aravin A, Tuschl T, Sander C, Marks DS (2004) Human MicroRNA targets. PLoS Biol 2(11):e363. doi:10.1371/journal.pbio.0020363
Kaina B, Christmann M, Naumann S, Roos WP (2007) MGMT: key node in the battle against genotoxicity, carcinogenicity and apoptosis induced by alkylating agents. DNA Repair (Amst) 6(8):1079–1099. doi:10.1016/j.dnarep.2007.03.008
Kitange GJ, Carlson BL, Schroeder MA, Grogan PT, Lamont JD, Decker PA, Wu W, James CD, Sarkaria JN (2009) Induction of MGMT expression is associated with temozolomide resistance in glioblastoma xenografts. Neuro Oncol 11(3):281–291. doi:10.1215/15228517-2008-090
Kitange GJ, Mladek AC, Carlson BL, Schroeder MA, Pokorny JL, Cen L, Decker PA, Wu W, Lomberk GA, Gupta SK, Urrutia RA, Sarkaria JN (2012) Inhibition of histone deacetylation potentiates the evolution of acquired temozolomide resistance linked to MGMT upregulation in glioblastoma xenografts. Clin Cancer Res 18(15):4070–4079. doi:10.1158/1078-0432.CCR-12-0560
Kreth S, Heyn J, Grau S, Kretzschmar HA, Egensperger R, Kreth FW (2010) Identification of valid endogenous control genes for determining gene expression in human glioma. Neuro Oncol 12(6):570–579. doi:10.1093/neuonc/nop072
Kreth S, Thon N, Eigenbrod S, Lutz J, Ledderose C, Egensperger R, Tonn JC, Kretzschmar HA, Hinske LC, Kreth FW (2011) O-methylguanine-DNA methyltransferase (MGMT) mRNA expression predicts outcome in malignant glioma independent of MGMT promoter methylation. PLoS One 6(2):e17156. doi:10.1371/journal.pone.0017156
Kreth S, Thon N, Kreth FW (2012) Epigenetics in human gliomas. Cancer Lett. doi:10.1016/j.canlet.2012.04.008
Lavon I, Fuchs D, Zrihan D, Efroni G, Zelikovitch B, Fellig Y, Siegal T (2007) Novel mechanism whereby nuclear factor kappaB mediates DNA damage repair through regulation of O(6)-methylguanine-DNA-methyltransferase. Cancer Res 67(18):8952–8959. doi:10.1158/0008-5472.CAN-06-3820
Lewis BP, Burge CB, Bartel DP (2005) Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120(1):15–20. doi:10.1016/j.cell.2004.12.035
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, Sweet-Cordero A, Ebert BL, Mak RH, Ferrando AA, Downing JR, Jacks T, Horvitz HR, Golub TR (2005) MicroRNA expression profiles classify human cancers. Nature 435(7043):834–838. doi:10.1038/nature03702
Malley DS, Hamoudi RA, Kocialkowski S, Pearson DM, Collins VP, Ichimura K (2011) A distinct region of the MGMT CpG island critical for transcriptional regulation is preferentially methylated in glioblastoma cells and xenografts. Acta Neuropathol 121(5):651–661. doi:10.1007/s00401-011-0803-5
Mayr C, Bartel DP (2009) Widespread shortening of 3′UTRs by alternative cleavage and polyadenylation activates oncogenes in cancer cells. Cell 138(4):673–684. doi:10.1016/j.cell.2009.06.016
Miranda KC, Huynh T, Tay Y, Ang YS, Tam WL, Thomson AM, Lim B, Rigoutsos I (2006) A pattern-based method for the identification of MicroRNA binding sites and their corresponding heteroduplexes. Cell 126(6):1203–1217. doi:10.1016/j.cell.2006.07.031
Ramakrishnan V, Kushwaha D, Koay DC, Reddy H, Mao Y, Zhou L, Ng K, Zinn P, Carter B, Chen CC (2011) Post-transcriptional regulation of O(6)-methylguanine-DNA methyltransferase MGMT in glioblastomas. Cancer Biomark 10(3–4):185–193. doi:10.3233/CBM-2012-0245
Sandberg R, Neilson JR, Sarma A, Sharp PA, Burge CB (2008) Proliferating cells express mRNAs with shortened 3′ untranslated regions and fewer microRNA target sites. Science 320(5883):1643–1647. doi:10.1126/science.1155390
Singh P, Alley TL, Wright SM, Kamdar S, Schott W, Wilpan RY, Mills KD, Graber JH (2009) Global changes in processing of mRNA 3′ untranslated regions characterize clinically distinct cancer subtypes. Cancer Res 69(24):9422–9430. doi:10.1158/0008-5472.CAN-09-2236
Stark A, Brennecke J, Bushati N, Russell RB, Cohen SM (2005) Animal MicroRNAs confer robustness to gene expression and have a significant impact on 3′UTR evolution. Cell 123(6):1133–1146. doi:10.1016/j.cell.2005.11.023
Stupp R, Hegi ME, Mason WP, van den Bent MJ, Taphoorn MJ, Janzer RC, Ludwin SK, Allgeier A, Fisher B, Belanger K, Hau P, Brandes AA, Gijtenbeek J, Marosi C, Vecht CJ, Mokhtari K, Wesseling P, Villa S, Eisenhauer E, Gorlia T, Weller M, Lacombe D, Cairncross JG, Mirimanoff RO (2009) Effects of radiotherapy with concomitant and adjuvant temozolomide versus radiotherapy alone on survival in glioblastoma in a randomised phase III study: 5-year analysis of the EORTC-NCIC trial. Lancet Oncol 10(5):459–466. doi:10.1016/S1470-2045(09)70025-7
Szeliga M, Zgrzywa A, Obara-Michlewska M, Albrecht J (2012) Transfection of a human glioblastoma cell line with liver-type glutaminase (LGA) down-regulates the expression of DNA-repair gene MGMT and sensitizes the cells to alkylating agents. J Neurochem 123(3):428–436. doi:10.1111/j.1471-4159.2012.07917.x
von Deimling A, Korshunov A, Hartmann C (2011) The next generation of glioma biomarkers: MGMT methylation, BRAF fusions and IDH1 mutations. Brain Pathol 21(1):74–87. doi:10.1111/j.1750-3639.2010.00454.x
Wang X, El Naqa IM (2008) Prediction of both conserved and nonconserved microRNA targets in animals. Bioinformatics 24(3):325–332. doi:10.1093/bioinformatics/btm595
Wen PY, Kesari S (2008) Malignant gliomas in adults. N Engl J Med 359(5):492–507. doi:10.1056/NEJMra0708126
Zhang W, Zhang J, Hoadley K, Kushwaha D, Ramakrishnan V, Li S, Kang C, You Y, Jiang C, Song SW, Jiang T, Chen CC (2012) miR-181d: a predictive glioblastoma biomarker that downregulates MGMT expression. Neuro Oncol 14(6):712–719. doi:10.1093/neuonc/nos089
Zhao W, Soejima H, Higashimoto K, Nakagawachi T, Urano T, Kudo S, Matsukura S, Matsuo S, Joh K, Mukai T (2005) The essential role of histone H3 Lys9 di-methylation and MeCP2 binding in MGMT silencing with poor DNA methylation of the promoter CpG island. J Biochem 137(3):431–440. doi:10.1093/jb/mvi048
Acknowledgments
We would like to thank J. Rink and Gaby Groeger for expert technical assistance.
Conflict of interest
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
S. Kreth and E. Limbeck contributed equally.
Rights and permissions
About this article
Cite this article
Kreth, S., Limbeck, E., Hinske, L.C. et al. In human glioblastomas transcript elongation by alternative polyadenylation and miRNA targeting is a potent mechanism of MGMT silencing. Acta Neuropathol 125, 671–681 (2013). https://doi.org/10.1007/s00401-013-1081-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-013-1081-1