Abstract
Isothermal titration calorimetry (ITC) is a widely used tool to experimentally probe the heat signal of the formation of the protein corona around macromolecules or nanoparticles. If an appropriate binding model is applied to the ITC data, the heat of binding and the binding stoichiometry as well as the binding affinity per protein can be quantified and interpreted. However, the binding of the protein to the macromolecule is governed by complex microscopic interactions. In particular, due to the steric and electrostatic protein–protein interactions within the corona as well as cooperative, charge renormalization effects of the total complex, the application of standard (e.g., Langmuir) binding models is questionable and the development of more appropriate binding models is very challenging. Here, we discuss recent developments in the interpretation of the Langmuir model applied to ITC data of protein corona formation, exemplified for the well-defined case of lysozyme coating highly charged dendritic polyglycerol sulfate (dPGS), and demonstrate that meaningful data can be extracted from the fits if properly analyzed. As we show, this is particular useful for the interpretation of ITC data by molecular computer simulations where binding affinities can be calculated but it is often not clear how to consistently compare them with the ITC data. Moreover, we discuss the connection of Langmuir models to continuum binding models (where no discrete binding sites have to be assumed) and their possible extensions toward the inclusion of leading order cooperative electrostatic effects.

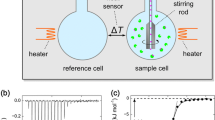
Schematic plot of the ITC experiment and the binding cooperativity of protein corona formation revealed from computer simulations.
Similar content being viewed by others
Introduction
The rational design of macromolecular polymeric drugs and nanocarriers has become a central task in medicine and pharmacy in the recent years [1,2,3]. Proteins typically bind strongly to the macromolecular or nanoparticle surface and thereby form a protein “corona,” a dense shell of proteins that can entirely coat the macromolecule [3,4,5,6,7,8,9,10,11,12]. As a consequence, the solution environment, be it in vivo or in vitro, does not see the macromolecule anymore but only the protein corona, which has important implications for the biological immune response to the macromolecule, its metabolic fate, and thus the function of such a complex in biomedical or biotechnological applications [3,4,5,6,7,8,9,10,11,12]. Many fundamental questions about the properties of the corona have kept the scientific community busy in the last years, for example, what is the protein composition of such a corona [4], how does it change dynamically in time [13, 14], and what are the underlying microscopic mechanisms and interactions that control the formation, evolution, and stability of such a corona [15]?
Calorimetry, in particular isothermal titration calorimetry (ITC), has become an important tool to characterize the protein corona [4,5,6, 9, 10, 17,18,19]. ITC provides the calorimetric heat, and, if a suitable binding model for data analysis is applied and correctly interpreted, it delivers important quantities such as the stoichiometry of binding and the binding affinity (binding constant) Kb of the proteins, as well as the heat of binding [19, 20]. However, the full quantification of the macromolecule-protein complexation based on ITC is often out of reach because the molecular and mechanistic processes are not well understood, and the application of standard binding models questionable. Naturally, the underlying interactions are governed by a complex interplay between electrostatic, solvation, and steric effects. Theoretical and simulation concepts are in need that allow a quantitative assessment of these forces [21,22,23,24]. Moreover, for the protein corona, cooperative effects must be discussed because the proteins interact with each other and also change some properties of the macromolecule; in particular, electrostatic interactions play a decisive role [6, 10, 13, 19]. Similar challenges arise also in the sorption of proteins to charged nano- and microgels [25]. The interpretation of ITC data and the definition and application of appropriate binding models are therefore very important but challenging tasks.
Hence, it is desirable to identify well-defined model experimental systems where the protein corona is sufficiently simple and accessible for interpretation. “Simple” should mean that the binding partners are structurally well characterized and the corona has only one protein component in a fully binding equilibrium. For such a simple system, also computer simulations and theoretical, physical binding models are easier to devise. Recently, Xu et al. presented a study of such a simple system by combining coarse-grained (CG) molecular computer simulations and ITC data of the corona formation of only one type of protein on the dendritic polyglycerol macromolecule terminated with sulfate (dPGS) [16, 26, 27]. dPGS has received much attention in the last decade because of its high potential in drug design: it exhibits significant anti-inflammatory action during disordered immune response [28,29,30,31] and has high efficacy and functionality in many other biomedical problems [32,33,34,35,36]. The dPGS is well characterized by now theoretically [37, 38] and experimentally [39]. In their work, Xu et al. analyzed ITC data of dPGS-protein complexation for various generations (sizes) of dPGS in particular using lysozyme as a well-defined model protein. Due to its almost complete sulfate termination, dPGS is highly negatively charged at relevant pH values, while the lysozyme proteins carry a positive net charge. Lysozyme complexes strongly with dPGS and forms a well-developed protein corona: Fig. 1 illustrates the lysozyme corona by snapshots taken from the CG computer simulation model [16].
Illustrating the protein corona for a lysozyme-dPGS (dendritic polyglycerol sulfate) system. Panel (a): Top of the panel: Coarse-grained representation of the highly charged dPGS macromolecule (charged surface groups in red with a shell of counterions in yellow). Bottom of the panel: Coarse-grained representation of the protein lysozyme. Positively charged beads are in green, negative charges in red. Panel (b): The lysozyme proteins adsorb strongly on the dPGS surface and form the protein corona. Snapshot taken from coarse-grained computer simulations [16]. Panel (c): Simplified two-dimensional sketch of the protein corona around dPGS. Proteins (green spheres) of radius Rp and charge Qp adsorb in a shell of width δ on the dPGS macromolecule (red sphere of radius Rm and charge \({Q_{m}^{0}}\)) to form a dense shell (corona)
In the CG simulation model, the solvent acts only as implicit background, while salt and dPGS monomers, as well as protein amino acids, are modeled as coarse-grained beads. Importantly, the original shape and charge structure are conserved in this model. In particular, the salt ions, sulfate groups, and charged acidic and basic amino acids carry monovalent integer charges of appropriate sign. Hence, while most hydration effects (i.e., those beyond simple dielectric screening) are neglected, all steric and electrostatic interactions are still well resolved and accounted for. The simulations showed strong cooperative effects in protein binding due to (excluded-volume) packing and electrostatic interactions in the dense protein corona. Consequently, binding affinities were found to depend on the density (coverage) of the proteins on the macromolecular surface. Despite this complexity, Xu et al. showed that still meaningful comparisons to ITC data are possible if the fits by the applied binding models, such as the standard Langmuir binding model, are properly analyzed and interpreted [16].
In this contribution, we systematically describe the challenges and new developments in the interpretation of ITC data probing the protein corona. We start by recalling what is actually measured by ITC and what information is usually obtainable. We then introduce and derive the concept of a coverage-dependent binding affinity in the Langmuir model that serves for a better interpretation of the ITC data as well as for extensions of standard binding models. With these prerequisites, we show how binding affinities from simulations, namely the free energy of binding per protein extracted from a potential of mean force calculation, can be compared with ITC data fitted by the standard Langmuir model. Moreover, we discuss the relation of Langmuir models to continuum binding models (where no discrete binding sites have to be assumed) as well as simple extensions of binding models toward the inclusion of leading order cooperative electrostatic effects. Our perspectives and extensions serve as starting points for the development of more elaborate binding models that may be directly applicable to fit ITC data in future studies.
Analysis of ITC data
The adsorption of a protein onto a macromolecule is accompanied by the release of heat, ΔHITC, as measured by ITC. In general, we have to assume for cooperative adsorption that this heat per adsorbing protein depends on the number of already bound proteins, i.e., it is a function of coverage 𝜃, usually defined as the ratio of bound proteins \({N_{p}^{b}}\) to the total number of binding sites N, i.e., \(\theta = {N_{p}^{b}}/N\). We note that typically linked equilibria can contribute to the measured heat by ITC, not directly related to the binding event [19]. ΔHITC is thus described often as marker enthalpy and is not necessarily the same as the binding enthalpy ΔHb (see, e.g., discussions and references in [19, 25]). The total in ITC released heat can then be written for a total of \({N_{p}^{b}}\) bound proteins as:
where ΔHITC(i) is the measured heat by ITC for the titration of the i th protein, cm the concentration of macromolecules, and V is the total solution volume. In the ITC experiments, Q is measured vs. the total protein concentration \(c_{p}^{tot}\). Introducing the molar ratio \(x = c_{p}^{tot}/c_{m}\) and going to the continuum limit we can write formally:
where \(H_{\text {ITC}}({N_{p}^{b}})\) is the total heat generated per macromolecule after the adsorption of \({N_{p}^{b}}\) proteins, and \({N_{p}^{b}}(x)\) represents the binding isotherm, i.e., the number of bound proteins versus molar ratio at fixed temperature.
Instead of the total heat, the incremental heat \(Q^{\prime }(x)=dQ/d x\) is typically employed for fitting to better track the changes in the heat during titration. The quantity of interest is the incremental heat per increment of protein, given by:
Due to the rather complex molecular interactions governing the protein adsorption process, the function \(H_{\text {ITC}}({N_{p}^{b}})\) is typically unknown and it is virtually always assumed that ΔHITC is constant and independent of coverage. In that case, it is:
Let us discuss in the following the behavior of this relation for the typical case of a standard Langmuir isotherm. The latter is based on identifying the simple association reaction equilibrium A+B→AB, with a binding constant Kb = [AB]/[A][B], where the square brackets denote concentrations. The binding constant is associated with a binding energy ΔGb through \(K_{b} = v_{0} \exp (-\upbeta {\Delta } G_{b})\), where v0 = l/mol is the standard volume [20, 21, 40]. Strictly speaking, ΔGb is a free energy, but we will call it in the following simply binding energy or Langmuir energy to distinguish from other free energies we will approach in our disucssions. With the assumption of N discrete binding sites, the standard Langmuir model yields then [20, 41,42,43]:
where \(c_{p}(x) = c_{m}[x - {N_{p}^{b}}(x)]\) is the free (bulk) protein concentration. The latter depends on \({N_{p}^{b}}\) in the experiments with a fixed number of proteins in the sample (i.e., a canonical thermodynamic ensemble). In the standard Langmuir model, the binding constant Kb is independent of protein concentration, and solving (5) for \({N_{p}^{b}}(x)\) yields:
with ξ = 1 + x/N + 1/(NKbcm). Using Eq. 6 in Eq. 1 with constant ΔHITC gives the total heat as:
The incremental heat is then [44]:
The fitting of \(Q^{\prime }(x)\) to the experimental data then yields the unknown constants Kb, N, and ΔHITC. Equation 8 describes a sigmoidal curve typically measured by ITC. In particular, deeper inspection of Eq. 6 shows that for large binding affinities, Kb ≫ 1/(Ncm), and very small x ≪ N, it is \({N_{p}^{b}}(x) = x\) (independent of Kb), and Eq. 8 exhibits a plateau at the beginning of the ITC titration of height ΔHITC, independent of the binding constant. This is indeed a typical ITC signature in strongly associating systems [44]. In the limit of very large molar ratios \(x\rightarrow \infty \), all of the titrated proteins will adsorb for a non-vanishing Kb and \({N_{p}^{b}}(x)\) saturates to a constant value, the maximum coverage. Then \(Q^{\prime }(x)=0\) and the ITC curve carries not much information anymore. For large Kb, the inflection point between the two saturating limits is at \(x\simeq N\), becoming exactly equal for \(K_{b}\rightarrow \infty \). For low affinities \(K_{b}\lesssim 1/(Nc_{m})\), the curve behaves differently, but these “low-signal” cases are often not well accessible by ITC [44]. The just explained behavior is illustrated in Fig. 2 for various values of the so-called Wiseman parameter c = cmKb [44].
Further analysis of Eq. 8 shows that for large Kb the binding affinity is determined essentially by the slope of the inflection point of \(Q^{\prime }(x=N)\), where we find \(d^{2}Q/dx^{2}|_{x=N} \propto \sqrt {K_{b}}\) (see Appendix). For larger slopes, automatically, the transition becomes also sharper, cf. Fig. 2. Hence, as an important conclusion, we find that fitting the standard Langmuir model to any sigmoidal function with sharp transitions probes essentially only the very vicinity of the inflection point x = N of the \(Q^{\prime }(x)\) curves. In other words, fitting to Langmuir isotherms is thus most sensitive close to \(x\simeq N\), where the molar ratio equals the number of available binding sites. Importantly, the extracted apparent binding constant Kb thus describes only the binding affinity right at x = N. This finding comes in useful when we have situations, like here for the protein corona, where the binding constant is actually not a constant but depends on the coverage (and thus on the molar ratio x).
In Fig. 3a, we show the experimental ITC data for \(Q^{\prime }(x)\) (symbols) for the specific case of lysozyme adsorbing onto dPGS of various generations G2, G4, G4.5, and G5.5 (with fractional values due to incomplete synthesis [16]) at a relatively low salt concentration of cs = 10 mM. Indeed we see that the systems exhibit wide plateau regions and very sharp transitions, indicating very large binding affinities. The standard Langmuir model fits relatively well and extracts thus the binding constant right at the inflection point x = N. The binding coordination, i.e., the maximum number of proteins constituting the protein corona, increases with dPGS generation (size) from about N = 3 for G2 to about N = 13 for G5.5. For increasing salt concentrations cs, we see in Fig. 3b for generation G2 how the curves shift and the transition becomes softer, as demonstrated also in Fig. 2, pointing to decreasing binding affinities with increasing salt concentration.
Exemplification of the behavior of the incremental heat \(Q^{\prime }\) versus rescaled molar ratio x/N for the standard Langmuir description (8) for various Wiseman parameters c = cmKb [44] (that is, rescaled binding constant Kb = c/cm). Only for large Kb ≫ 1/(Ncm) the curve develops a plateau for small x/N and the inflection point is at \(x\simeq N\)
a Differential heat from ITC of lysozyme-dPGS complexation for dPGS generations G2 to G5.5 at a temperature of 310 K and cs = 10 mM salt concentration [16]. The solid lines correspond to the fits by the standard Langmuir model, Eq. 8. b ITC isotherms of lysozyme-G2 complexation at different salt concentrations cs (see legend) and fitted by the Langmuir model. The inset displays the salt dependence of the binding constant Kb on a log-log scale. According to limiting laws [19], \(-\mathrm {d}\ln K_{b}/\mathrm {d} \ln c_{s} = N_{\text {CR}}=3.1\pm 0.1\) counterions are released upon binding. c CG simulation results of the PMF, Gi(r), as a function of the center-of-mass distance r between G5 and lysozyme for the successive binding of i = 1 to 15 proteins in 10 mM salt concentration [16], color-coded according to the scale. d The simulation binding free energy ΔGsim(𝜃) (solid symbols; units kBT = 1/β) plotted versus coverage 𝜃 for G2, G4, and G5, respectively, reads off from the global minimum of the PMFs, as such in (c). The large open circle, triangle, and square symbols indicate the simulation-referenced Langmuir binding free energy \({\Delta } G^{\text {ITC}}_{\text {sim}}(\theta ^{*})\), Eq. 25, for G2, G4, and G5.5 at their respective coverage \(\theta ^{*} \simeq 0.95 \)
The Langmuir model with coverage-dependent binding constant
If cooperativity effects are at play, we need to consider that the binding affinity of a protein depends on the number of already bound proteins. One way of dealing with cooperativity is to introduce binding polynomials [43, 45, 46], where every binding step A+B→AB, A+AB→2AB, A + 2AB→3AB, etc., has its individual binding constant and equilibrium coverages are calculated by averaging over all states. Here, we choose a different but, as we will see, related route: we assume a Langmuir binding energy that continuously changes with successive binding and thus with coverage 𝜃, i.e., ΔGb = ΔGb(x) = ΔGb(𝜃). As we show in the following, this can be readily incorporated into the derivation of the Langmuir model in the canonical ensemble and leads to a proper definition of a coverage-dependent binding constant Kb(𝜃).
For this, consider a finite region with volume V in which a much smaller subspace (macromolecule on which the proteins bind) with N binding sites available. The binding sites can be all of the same kind. The cooperativity is assumed to enter through protein–protein interactions as well as the change of global properties (like the total charge of the macromolecule–protein complex). We assume a canonical ensemble with in total \(N_{p}^{tot}\) proteins in the cell volume V which now contains only one macromolecule. We define the fraction of bound protein particles by \(\theta ={N_{p}^{b}}/N\). The number of available and independent binding states is then [42, 47, 48]
from the combinatorial possibilities of distributing \({N_{p}^{b}}\) indistinguishable proteins on N sites, and ζ is the microscopic partition sum of a single protein particle in the bound state [21, 49, 50]. Different to the standard Langmuir model, we now introduce a coverage-dependent binding energy \({\Delta } G_{b}({N_{p}^{b}})\) associated with the binding of a protein from a reference state to one Langmuir site, given that \({N_{p}^{b}}-1\) proteins are already adsorbed. This leads to the partition sum:
and we can define the free energy of the macromolecule-protein complex as
Using the Stirling approximation \(\ln (m!) = m\ln (m)-m\) leads (within an unimportant constant) to the free energy of the complex normalized per binding site:
where we defined \(\zeta ^{{N_{p}^{b}}}=(v_{0}/{\Lambda }^{3})^{{N_{p}^{b}}}\) in terms of an effective configurational volume v0 divided by the cubed thermal (de Broglie) wavelength Λ3. “Effective” means that also restrictions on internal vibrational and orientational degrees of freedom upon binding as expressed by the full microscopic partition sum [21, 49, 50] are adsorbed in the volume v0.
The total Helmholtz free energy F of the canonical system (including the finite bath of free protein surrounding the macromolecule) is thus:
where we introduced the ideal gas free energy \(\upbeta F_{id}=(N_{p}^{tot}-{N_{p}^{b}})[\ln ((N_{p}^{tot}-{N_{p}^{b}}){\Lambda }^{3}/V)-1]\) of the canonical reservoir. Then, \(c_{p} = N_{p}/V = (N_{p}^{tot}-{N_{p}^{b}})/V\) is the density of unbound (i.e., free bulk) proteins in V. The total free energy \(\tilde f=\upbeta F/N\) per binding site is then:
The minimization of the free energy with respect to the coverage of bound proteins:
yields then the final relation between the binding energy and the fraction of bound proteins in dependence of the free protein concentration cp:
where \({\Delta } G_{b}^{\prime }(\theta ) = \partial {\Delta } G_{b}/\partial \theta \). The left-hand side of the equation defines a coverage-dependent binding constant Kb(𝜃). As we see, we cannot simply define Kb(𝜃) solely by ΔGb(𝜃) but have to consider also the changes (derivative) of the latter with 𝜃. The physical interpretation of this term is that a newly binding protein changes the coverage and thus the binding energy (and thus binding equilibrium) for all other bound proteins. If \({\Delta } G_{b}^{\prime } =0\) for all 𝜃, then ΔGb = const., and we recover the standard Langmuir model with \(K_{b}=v_{0}\exp (-\upbeta {\Delta } G_{b}) =\) const. In some cases, like in the application discussed in this work, it is \(\theta {\Delta } G_{b}^{\prime } \ll {\Delta } G_{b}\), and the second term in the exponent can be neglected. Note also, that if we rewrite (17) in 𝜃 = cpKb(𝜃)/(1 + cpKb(𝜃)), it is relatively simple to see that we can generate the conventional binding polynomials [43, 45, 46] by a Taylor expansion of Kb(𝜃(x)) with respect to molar ratio x, i.e., \(K_{b}(\theta )\simeq K_{0} + K_{1}x + K_{2}x^{2} + ...\), which may serve for interesting interpretations of the constant coefficients Ki in future work.
It is for analysis of some problems reasonable to assume that the binding energy ΔGb splits into an intrinsic process, ΔGb,int which is independent of any cooperative effects, and an excess contribution, ΔGb,exc(𝜃), that accounts for the cooperativity. With this and neglecting the \({\Delta } G_{b}^{\prime }\) term in Eq. 17, we can write:
Such a splitting was applied, e.g., in the case of charged proteins binding to hydrogels [51], or in the splitting of electrostatic contributions from intrinsic ones for small molecular ligands binding to membranes [52,53,54]. Note again that Eq. 18 is only accurate strictly if \(\theta {\Delta } G_{b}^{\prime } \ll {\Delta } G_{b}\), that is, the explicit variation of the binding energy with coverage is much smaller than the absolute binding energy.
It is important to note that the volume v0 depends on the exact nature of the bound state and is typically not known. Per convention in experiments, the standard volume v0 = l/mol \(\simeq 1.6\) nm3, corresponding to the standard concentration co = 1 M (1 mol/l) is employed. In that case, the determined ΔGb = ΔGo is then called the standard free energy of binding [20]. Hence, in experiments, the discussion of origins or values of v0 and ΔGb individually without knowing microscopic details is in principle not feasible. However, if we consider computer simulations (see section “Comparing Langmuir to computer simulation binding free energies”) where those quantities can (or must) be calculated independently, one has to discuss the origins and microscopic definition of v0 much more thoroughly. Since, in general, v0 will depend on the specific microscopic processes, one has to take care how to convert a theoretical or simulation derived free energy to the standard energy of binding ΔGo, as already discussed in a comprehensive way for other associating systems [21, 40].
Connection to continuum binding models
It is helpful for interpretation of ITC data to also consider continuum binding models, i.e., to dismiss the assumption of discrete binding sites. In particular, for proteins binding to a macromolecule, it is often not clear what the presumed discrete binding sites actually are. For a relatively homogeneous macromolecular/nanoparticle surface, a continuum picture, where adsorbed proteins may freely move and diffuse on the macromolecule surface, could be much more appropriate. Inspired from our previous work on protein–hydrogel interactions [51], such a continuum binding model for the current case of a protein corona could be sketched as in the following. We make the following Boltzmann ansatz for the number of adsorbed proteins via:
where we split (for reasons becoming clear below) the total binding free energy in the Boltzmann exponent into a macromolecule–protein, ΔGmp(x), and a protein–protein, ΔGpp(x), contribution. (We intentionally avoided the subscript “b” here to make this free energy of binding in the continuum model distinct from the Langmuir binding energy discussed above.) Furthermore, Vb = Amδ is the effective binding volume (say a spherical shell of thickness δ, cf. Fig. 1) on the macromolecular surface with area Am = 4π(Rm + Rp)2, where Rm and Rp are the macromolecule and protein radii, respectively. A surface density τ(x) (number per area) can now be defined by \(\tau (x) = {N_{p}^{b}}(x)/A_{m}\), and accordingly a surface packing fraction Apτ < 1, where \(A_{p}\simeq \pi {R_{p}^{2}}\) is the area taken effectively by a protein.
We stay in line with the coverage-dependent Langmuir model Eq. 17 introduced in the previous section and assume in Eq. 19 that the macromolecule–protein binding free energy ΔGmp(𝜃) is in general coverage dependent, thus also a function of x, ΔGmp(x). The free energy ΔGpp(x) considers the change of free energy of inserting one protein into the quasi-two-dimensional liquid of already adsorbed proteins on the surface. It is a contribution purely coming from direct protein–protein interactions. It thus vanishes, in contrast to ΔGmp(x), in the limit \(\tau (x) \rightarrow 0\). For proteins interacting only with hard cores, it could be modeled by the excess free energy of hard discs where the equation of state is well known [55]. Disregarding any specific protein–protein interactions right now, we can always make a virial expansion in orders of the density τ. Considering only the first leading order, \(\upbeta {\Delta } G_{pp}(\tau ) \simeq 2B_{2}\tau \), where B2 is the (two-dimensional) second virial coefficient, expanding the exponent in Eq. 19 for small coverages via \(\exp [-\upbeta {\Delta } G_{pp}(\tau )] \simeq (1-2B_{2}\tau )\), and rearranging to solve for \({N_{p}^{b}}(x)\), we obtain a closed form for the binding isotherm through:
If we now make the substitution N = Am/(2B2) (for reasons becoming clearer below), express \({N_{p}^{b}}\) by coverage \(\theta = {N_{p}^{b}}/N\), and define a binding constant \(\tilde K_{b}(x)=\tilde v_{0}\exp [-\upbeta {\Delta } G_{mp}(x)]\) with \(\tilde v_{0} = 2V_{b}B_{2}/A_{m} = V_{b}/N\), we find exactly the Langmuir type of form:
The key steps for such a mapping of the continuum model to the Langmuir model is on one hand the two-body approximation in the protein–protein correlations, and on the other hand, the definition of the effective binding volume \(\tilde v_{0} = V_{b}/N\). With that we recognize that in the limit of not too large coverages (where only 2-body interactions dominate), the continuum model and the discrete Langmuir model constitute essentially (mathematically) the same binding models, albeit with different interpretations. N = Am/(2B2) in the continuum picture is not the number of binding sites but defines the maximal number of binding spaces limited by the (2-body) protein–protein interactions. This becomes immediately clear if we identify B2 with the excluded interaction area, e.g., for the case for hard-disk like particles of radius R where \(B_{2}^{HD}=2\pi R^{2}\). Then, not more than N adsorbed particles will fit on the surface simply by packing constraints.
Note that if B2 < 0 or higher order terms contribute, that is, protein–protein interactions are attractive or strongly correlated, respectively, such a simple interpretation of N and relation to Langmuir would not be so easily possible anymore. Hence, while a mapping of the more general continuum picture on the specific Langmuir model is in principle possible in some limits, details are subtle, in particular, how to define which interaction contributions actually enter the binding constant Kb and which enter in the effective binding volume. A direct connection of the continuum free energy of binding to the coverage-dependent model Eq. 17 is thus generally not possible—it may be feasible only for certain specific physical binding processes. As we will see in the following, the continuum view is important for the comparison of ITC fits to computer simulation data.
Comparing Langmuir to computer simulation binding free energies
Now we come to one of the key questions, how can we connect and compare the free energy of binding calculated from a computer simulation to the one obtained from a binding model fitted to ITC data. In particular, we want to address the non-trivial question of what can be done if the (typically complex) binding mechanisms and cooperative effects are not known, and an appropriate binding model is missing.
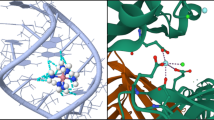
Let us briefly summarize what we typically do in a computer simulation. In a canonical ensemble simulation setup (that is, a fixed box volume V and fixed number of proteins), we can access the equilibrium free energy of binding of a protein i to a binding region by calculating the so-called potential of mean force (PMF) Gi(r) [49, 50]. This is done along the reaction coordinate r, which is typically the center-of-mass distance between a protein and the macromolecule/protein complex. Such a simulation setup is illustrated in Fig. 4 for a coarse-grained, implicit-solvent simulation of lysozyme associating with dPGS [16]. Since the degrees of freedom are massively reduced in such a coarse-grained representation, the equilibrium PMF, as shown in [16], can be well sampled by simple statistical averages for every constraint of the distance reaction coordinate r, in contrast to explicit-water all-atom simulations which are usually harder to converge. The calculations can be done for a single protein i also if already i − 1 proteins are adsorbed. We can thus evaluate the PMF for every successive binding event in the protein corona. PMFs calculated from coarse-grained simulations for the specific example of lysozyme to dPGS of generation G5 are shown in Fig. 3c. For large distances r, Gi(r ≫ r0) = 0 is set to 0 to define the free unbound reference system. For small \(r\simeq \) 3–4 nm, we see a deep global minimum which defines the bound state at distance r0. The simulation binding free energy \({\Delta } G_{\text {sim}}({N_{p}^{b}})\) is then calculated from the PMF differences between the global minimum at the bound state at r0 and the free bulk state for large r ≫ r0, that is \({\Delta } G_{\text {sim}}({N_{p}^{b}}) = G_{i={N_{p}^{b}}}(r=r_{0})\), if \({N_{p}^{b}}-1\) proteins are already present in the corona.
The cooperativity effects in the protein binding can be clearly observed in the simulation results for \({\Delta } G_{\text {sim}}({N_{p}^{b}})\), summarized in Fig. 3d: the attraction per protein decreases with successive binding, indicating negative cooperativity, i.e., the more proteins are loaded into the corona the less favorable the binding becomes. From equilibrium simulations [16], it was found that for dGPS G5 the maximum protein occupancy was N = 13. The PMF for this in Fig. 3c is the third (reddish color) from the top. The two top ones still exhibit a global binding minimum but also a significant barrier for crossing at \(r\simeq 6\) nm, kinetically hindering the binding of the corresponding proteins. (Note that for radial crossing of barriers in 3D space, the effective free energy landscape has to be corrected by a distance-dependent entropic factor [56, 57].) We see also from Fig. 3d that the assumption \(\theta {\Delta } G_{b}^{\prime } \ll {\Delta } G_{b}\) made in section “The Langmuir model with coverage-dependent binding constant” for the considered system holds well, that is, the changes \(\theta [{\Delta } G_{\text {sim}}({N_{p}^{b}})\)-\({\Delta } G_{\text {sim}}({N_{p}^{b}}+1)]\) are typically much smaller than \({\Delta } G_{\text {sim}}({N_{p}^{b}})\). However, the simulation binding free energy cannot be related directly to the binding energy derived from ITC, as we will argue in the following paragraphs.
We now show how to connect the simulation free energy to the Langmuir results. In the simulations, with the constraint of having \({N_{p}^{b}}-1\) proteins bound in the simulation, we are probing the equilibrium \({N_{b}^{p}}-1 \rightleftharpoons {N_{p}^{b}}\) binding in a domain (say spherical shell) of volume Vb ≪ V. This translates into the Boltzmann equilibrium:
where in this case cp is the concentration of the free (unbound) proteins for which the equilibrium of \({N_{p}^{b}}\) bound proteins is established. The binding volume Vb is a quantity that can be calculated in the simulations [16]. Equation 22 is similar to the continuum binding model introduced in the previous section in Eq. 19, but all the binding contributions are clumped together in one single binding free energy ΔGsim.
Illustration of a PMF (potential of mean force) calculation in a coarse-grained computer simulation [16] to obtain the equilibrium binding free energy of a protein binding into an incomplete corona. A single protein (right) is continuously moved along the center-to-center distance r between the protein and the complex (left; dPGS in red), where already a few proteins are bound. The PMF, Gi(r) (blue curve), obtained from sampling and integrating the equilibrium mean force along r, exhibits a deep global binding minimum at a close distance r0. With Gi(r ≫ r0) = 0, the value of Gi(r0) defines the simulation binding free energy ΔGsim
We emphasize again the nature of \({\Delta } G_{\text {sim}}({N_{p}^{b}})\) in the simulations: it includes all entropic (including steric) and energetic interactions of the \({N_{p}^{b}}\)th protein with the macromolecule and already bound proteins. In comparison, in the Langmuir model with coverage-dependent binding constant Kb(𝜃), Eq. 17, the binding equilibrium for the protein coverage 𝜃 at concentration cp is:
Comparing Eqs. 22 and 23 shows that there is the following relation between the simulation free energy ΔGsim and the Langmuir binding energy ΔGb:
ΔGsim includes all contributions to the transfer free energy from bulk to the binding volume Vb. The Langmuir energy ΔGb in contrast considers only the energy of binding into one of the Langmuir binding boxes with effective volume v0. Thus, to link to the simulation free energy we have to add the entropic correction terms (the \(\ln \)-terms) which take care of the confinement and configurational arrangements in the N Langmuir binding boxes of volume v0. Hence, the right side of the equation represents the total Langmuir free energy, with entropy in reference to simulation binding volume. This allows a direct comparison between the ITC evaluation and the simulation PMFs on the full free energy level. As a conclusion, ΔGsim and ΔGb (or the standard energy ΔG0 ) cannot be compared directly, unless we are in the low coverage limit 𝜃 ≪ 1 and the simulation binding volume is for some reason exactly given by the standard binding volume.
This mapping is directly related to section “Connection to continuum binding models,” Eq. 21, where we showed that the Langmuir picture can be translated to a continuum picture if v0 = Vb/N and v0 can be interpreted as an effective configurational volume constrained by the binding shell and pair interactions. Contributions from ΔGsim are thus split into ΔGb, \(k_{B}T\ln (1-\theta )\), and \(k_{B}T\ln (v_{0}N/V_{b})\) terms in rather non-trivial ways. Then, Eq. 24 can also be interpreted such that the Langmuir binding free energy ΔGb includes all transfer free energy contributions, apart from 1- and 2-body confinement and protein–protein interaction contributions, implicitly included in the definition of v0 and the Langmuir assumption of discrete binding boxes. Hence, the ITC/Langmuir derived ΔGb includes apart from the energetic macromolecule–protein contributions also many-body (larger than 2-body) packing correlations, which makes it difficult to compare specific interaction contributions between simulations and the Langmuir fit.
We can now use Eq. 24 to make a one-to-one comparison between the simulation results reported in Fig. 3c and the Langmuir fits to ITC in Fig. 3a. Recall from our discussion in section “Analysis of ITC data” (cf. Fig. 2) that the ITC fit provides information about the binding affinity Kb(𝜃) only for a certain coverage 𝜃∗ := 𝜃(x = N). Only there Eq. 24 can be evaluated and we formally rewrite:
where \({\Delta } G_{\text {sim}}^{\text {ITC}}(\theta ^{*})\) defines the total binding free energy of ITC that can be compared with simulation results. Thus, we can refer to it as the “simulation-referenced” Langmuir free energy [16].
How do we evaluate the coverage 𝜃∗? Given the sigmoidal differential heat as in Fig. 3a, \(Q^{\prime }(x)\), the inflection point directly delivers the binding stoichiometry N. The protein coverage 𝜃 follows from the normalized integrated incremental heat as:
With that, one can define the protein coverage at which the binding affinity is evaluated by 𝜃∗ = 𝜃(x = N) as well as the mean coordination number \(N_{p}^{b*} = N \theta ^{*}\) at the respective binding equilibrium. In Fig. 5, we exemplify the calculation of 𝜃∗ for dPGS generations G2 and G5.5-dPGS, respectively, directly from the ITC measurements. The coverage 𝜃∗ is found as 𝜃∗ = 0.92 for G2 and 𝜃∗ = 0.97 for G5.5, which corresponds to the mean coordination numbers \(N_{p}^{b*} = 2.2\) for G2 and \(N_{p}^{b*} = 12.2\) for G5.5, respectively.
Top panels: Protein coverage 𝜃(x) as a function of molar ratio x for the generations (a) G2 and (b) G5.5. In the lower panel, the corresponding ITC differential heat \(Q^{\prime }(x)\) is displayed. 𝜃 is obtained by integration of \(Q^{\prime }(x)\), see Eq. 26. The dashed line indicates the binding stoichiometry N defined by the inflection point. The slope of \(Q^{\prime }(x)\) at this point yields the binding affinity K(𝜃∗) from the Langmuir fits (see section “Analysis of ITC data” and the Appendix). The coverage at this point is \(\theta ^{*}(x=N) = N_{p}^{b*}/N\)
Thus, as an important finding, the coverages at which the binding affinity is probed by ITC are already very close to saturation. The binding affinity is therefore related to those few, 1-2 proteins which finally complete the protein corona at high protein concentration. As we see from the simulations in Fig. 3d, the proteins that start forming the corona have a very different, about 10kBT more attractive, binding free energy than the finally binding ones. In Fig. 3d, we also plot the results for the simulation-referenced Langmuir free energy (25), for generations G = 2,4, and 5.5. The ITC-based results are all very similar for the various generations at around \({\Delta } G^{\text {ITC}}_{\text {sim}}\simeq 14\) to 15 kBT and depicted by large open symbols at 𝜃∗ in Fig. 2d. They match very well the simulation free energies at coverages of \(\theta \simeq 0.95\), consistently right at the 𝜃∗ values where the ITC binding affinity was determined. Hence, our comparison on the total free energy level shows good quantitative agreement between ITC and the computer simulations. Some more physical interpretation of the data based on the dominant electrostatic interactions in this system follows in the next section.
Electrostatic excess contributions and cooperativity
When many charged proteins bind to a charged region cooperativity effects come into play, in leading order simply due to the change of the global electrostatic properties during adsorption. This has been formulated, for instance, in the Guoy-Chapman-Stern theory for the binding of charged ligands to charged surfaces [52,53,54] or the binding of net-charged proteins to microgels [51], where the successive binding incrementally changes the overall surface or Donnan electrostatic potential, respectively. Consequently, the total binding constant Kb = Kb(x) = Kb(𝜃) has to be defined more generally and split up into an intrinsic part Kb,int and an excess part Kb,exc, as discussed before in Eq. 18. For the purpose here, we can equate the excess contribution with a cooperative electrostatic contribution, via:
i.e., ΔGb(x) = ΔGb,int + ΔGb,elec(x). Per definition, the intrinsic, x-independent binding constant Kb,int only contains contributions from local and specific interaction between the protein and the macromolecule, and the nonspecific, global, and cooperative electrostatic effect has been separated out. To be more precise with the word “global,” we have in mind a leading order multipole expansion (monopole, dipole, etc.) of the electrostatic potential of the whole complex, whose electrostatic properties are changing during binding. Specific local interactions may include local solvation effects, e.g., hydrophobic, or possibly highly localized interactions, such as H-bonds or salt bridges.
For proteins interacting with the highly charged dPGS macromolecule, it was found that the intrinsic part is dominated by a highly localized electrostatic effect, the counterion-release (CR) contribution: For the highly charged dPGS macromolecule, strong charge-renormalization was observed by a massive uptake of counterions [37]. A few of those counterions “condensed” on the dPGS surface layer are liberated when the protein binds, whereupon an oppositely charged protein patch becomes a multivalent counterion for the polyelectrolyte [58,59,60,61,62,63,64]. The resulting favorable (purely entropic) free energy in dependence of the salt concentration cs can be formulated as:
where cci (typically ≫ cs) is the local concentration of condensed counterions, cs the bulk salt concentration, and NCR denotes the number of those counterions released after binding. Equation 28 follows in some limits from the pioneering considerations of Record, Anderson, and Lohman [65] in the realm of DNA-protein complexation that culminated in the leading-order expression for the binding constant purely from counterion release, \(d\ln K_{b}/d \ln c_{s} = -N_{\text {CR}}\). More detailed discussions on the derivation and consequences of these processes and the interpretation of NCR can be found in the original work [65] and partially in more recent references [19, 66].
Apart from some extreme scenarios, e.g., total charge reversal of the complex, or large bulk salt concentration changes with titration, it can be safely assumed that the counterion-release part is an intrinsic contribution to the macromolecule-protein binding term (cf. Eq. 19) whose magnitude does not depend on protein coverage. The CR mechanism should play a role for every protein that carries a significant positive patch (even net-neutral or net-negatively charged ones [16, 26, 27]) and according to Eq. 28 it can be assumed that the number of released ions, NCR, scales with patch size, that is, this interaction is protein specific. We see in the inset of Fig. 3b that indeed the CR signature is found very well expressed in the dPGS-lysozyme system, where \(\ln K_{b}\) is a linear function of \(\ln c_{s}\). The slope delivers the number of released counterions for the dPGS G2 of about \(N_{\text {CR}} \simeq 3\), which well matches the computer simulation results in which ions can be explicitly counted [16]. CR is a very large driving force because the local concentration of condensed ions, cci, is typically in the molar range compared with only millimolar salt bulk concentrations cs. As a consequence, Eq. 28 predicts that only a single released ion can contribute as much as 3–4 kBT (entropic) free energy to the binding. More details on computational characterization of the CR effects in dPGS-lysozyme complexation can be found in the previous joint simulation/ITC works [16, 26, 27, 66].
For the non-specific electrostatic part, one could envision a multipole expansion for the charged binding partners. In first order, the proteins are simply charged spheres of size Rp carrying a net charge Qp = Zpe. Analogously in first order the macromolecule/protein complex is a sphere of size Rm + Rp and a coverage-dependent charge
depending on the number of bound proteins, because they modify the net charge of the complex. This is arguably the simplest realization of electrostatic cooperativity, in which the screened monopole electrostatic potential between the complex (including macromolecule and corona) and an approaching protein depends on the coverage which renormalizes the charge \({Q_{m}^{0}}\). This effect can be positively or negatively cooperative depending on the signs of the involved charges. On a simple Debye-Hückel level, we can write for \({\Delta } G_{b,\text {elec}}({N_{p}^{b}})\):
where λB = e2/(4π𝜖0𝜖kBT) is the Bjerrum length and \(\kappa = \sqrt {4\pi \lambda _{B} c_{s}}\) the inverse (Debye) screening length for a monovalent salt at concentration cs. Note that Eq. 30 with the simple addition of macromolecule and protein charges as defined in Eq. 29 involves both macromolecule–protein and protein–protein interactions which would enter in principle in both binding energies in the continuum ansatz Eq. 19, respectively.
Together with Eq. 28, the non-specific electrostatic contribution Eq. 30 in the coverage-dependent Langmuir model Eq. 27 would be a first attempt of a binding model that includes electrostatic cooperativity effects. If we compare back to Eq. 17, however, we see that we would need to include the first derivative of Eq. 30 with respect to coverage (the \({\Delta } G_{b}^{\prime }\) term in Eq. 19) to include the cooperative effect more completely to consistently describe the total binding equilibrium. Similar models, additionally including Born self-energy terms and dipolar contributions, albeit neglecting the \({\Delta } G_{b}^{\prime }\) term, were devised for proteins binding to hydrogels in equilibrium [51, 67] as well as for protein sorption kinetics to core-shell hydrogels [68] and molecular cargo to hollow hydrogels [69].
We finally note that some of the findings in the ITC and simulation data for the dPGS-lysozyme complexation can be already well interpreted by such a simple electrostatic extension of the binding models. For instance, we observe in Fig. 3d that the binding affinity does not depend much on the dPGS generation. The reason is that charge renormalization of the dPGS leads to effective macromolecular charges that change only little with generations (although the bare charge changes substantially) [37]. As a consequence, the global electrostatics in leading order is very similar for the various generations. For high coverages 𝜃∗ near saturation, where we evaluated the ITC data in section “Comparing Langmuir to computer simulation binding free energies,” the dPGS is almost completely neutralized by proteins and the binding affinity reflects mostly the intrinsic contribution, that is, the CR contribution, Eq. 28. Comparing the binding free energy at 𝜃∗ to Eq. 28 with independent estimates of cci and NCR indeed confirmed that CR is the dominant intrinsic mechanism to binding [16]. Recall that the CR process is well captured in the (implicit water) coarse-grained simulations due to the explicit resolution of the ions. The good agreement of simulations and experiments in Fig. 3d indicates that water hardly contributes to the total free energy of binding. A deeper discussion of water effects on the (temperature-dependent) binding enthalpy ΔHb and the corresponding entropy of binding can be found in a recent publication [66].
Summary and concluding remarks
Clearly, the complexation of proteins with a macromolecule or a nanoparticle involves complex processes. ITC can probe these processes but only implicitly by measuring the incremental heat released in the complexation equilibrium. Binding models are thus in need which either take into account the microscopic interaction details, or, if not available, standard binding models need to be properly interpreted. Nowadays, with the great help of molecular computer simulations of increasing quality and efficiency we are in a position to learn a lot about the complexation process and devise, modify, and interpret binding models more systematically. As a start, in this work, we introduced the concept of a coverage-dependent binding affinity in the Langmuir model that serves for better interpretation of ITC data as well as for extensions of standard binding models. We showed how binding affinities calculated from computer simulations, namely the free energy of binding per protein extracted from a potential of mean force calculation, can be consistently compared with experimental ITC data fitted by the standard Langmuir model. Moreover, we discussed the relation of Langmuir models to continuum binding models as well as simple extensions of binding models toward the inclusion of leading order cooperative electrostatic effects. Our perspectives will hopefully serve as starting points for the development of more elaborate binding models that may be directly applicable to fit ITC data in future studies of the protein corona or similar complex and cooperative problems.
Change history
04 September 2021
A Correction to this paper has been published: https://doi.org/10.1007/s00396-021-04861-2
References
Lee CC, MacKay JA, Frechet JMJ, Szoka FC (2005) . Nat Biotechnol 23:1517
Svenson S, Tomalia DA (2005) . Drug Adv Deliv Rev 57:2106
Owens DE, Peppas NA (2006) . Intern J Pharamceutics 307:93
Cedervall T, Lynch I, Lindman S, Berggård T, Thulin E, Nilsson H, Dawson KA, Linse S (2007) . Proc Natl Acad Sci USA 104(7):2050
Lindman S, Lynch I, Thulin E, Nilsson H, Dawson KA, Linse S (2007) . Nano Lett 7(4):914
Baier G, Costa C, Zeller A, Baumann D, Sayer C, Araujo PHH, Mailánder V, Musyanovych A, Landfester K (2011) . Macromol Biosci 11:628
Monopoli MP, Åberg C, Salvati A, Dawson KA (2012) . Nat Nanotechnol 7(12):779
Wang F, Yu L, Monopoli MP, Sandin P, Mahon E, Salvati A, Dawson KA (2013) . Nanomed: Nanotechnol Biol Med 9(8):1159
Winzen S, Schoettler S, Baier G, Rosenauer C, Mailaender V, Landfester K, Mohr K (2015) . Nanoscale 7:2992
Schöttler S, Landfester K, Mailänder V (2016) . Angew Chem (Intern Ed) 55:8806
Giudice MCL, Herda LM, Polo E, Dawson KA (2016) . Nat Commun 7:13475
Boselli L, Polo E, Castagnola V, Dawson KA (2017) . Angew Chem Int Ed 56(15):4215
Casals E, Pfaller T, Duschl A, Oostingh GJ, Puntes V (2010) . ACS Nano 4:3623
Angioletti-Uberti S, Ballauff M, Dzubiella J (2018) . Mol Phys 116:3154
Wei Q, Becherer T, Angioletti-Uberti S, Dzubiella J, Wischke C, Neffe AT, Lendlein A, Ballauff M, Haag R (2014) . Angew Chem (Intern Ed) 53:8004
Xu X, Ran Q, Dey P, Nikam R, Haag R, Ballauff M, Dzubiella J (2018) . Biomacromolecules 19:409
Jelesarov I, Bosshard HR (1999) . J Mol Recognit 12(1):3
Chaires JB (2008) . Annu Rev Biophys 37:135
Xu X, Ballauff M (2019) . J Phys Chem B 123(39):8222
Atkins P, de Paula J (2010) Physical chemistry. W. H Freeman and Company
Gilson MK, Zhou HX (2007) . Annu Rev Biophys 36:21
Durrant JD, McCammon JA (2011) . BMC Biol 9:71
Wang RX, Lai LH, Wang SM (2002) . J Comput-Aided Molec Des 16:11
Perez A, Morrone JA, Simmerling C, Dill KA (2016) . Curr Opin Struct Biol 36:25
Welsch N, Becker AL, Dzubiella J, Ballauff M (2012) . Soft Matter 8:1428
Ran Q, Xu X, Dzubiella J, Haag R, M Ballauff (2018) (3), 9086
Ran Q, Xu X, Dey P, Yu S, Dzubiella J, Haag R, Ballauff M (2018) . J Chem Phys 149:163324
Türk H, Haag R, Alban S (2004) . Bioconjug Chem 15:162
Dernedde J, Rausch A, Weinhart M, Enders S, Tauber R, Licha K, Schirner M, Zügel U, von Bonin A, Haag R (2010) . Proc Natl Acad Sci USA 107(46):19679
Calderón M, Quadir MA, Sharma SK, Haag R (2010) . Adv Mater 22:190
Reimann S, Gröger D, Kühne C, Riese SB, Dernedde J, Haag R (2015) . Adv Healthcare Mater 4:2154
Maysinger D, Gröger D, Lake A, Licha K, Weinhart M, Chang PKY, Mulvey R, Haag R, McKinney RA (2015) . Biomacromolecules 16:3073
Reimann S, Schneider T, Welker P, Neumann F, Licha K, Schulze-Tanzil G, Wagermaier W, Fratzl P, Haag R (2017) . J Mater Chem B 5:4754
Zhong Y, Dimde M, Stöbener D, Meng F, Deng C, Zhong Z, Haag R (2016) . ACS Appl Mater Interfaces 8:27530
Sousa-Herves A, Würfel P, Wegner N, Khandare J, Licha K, Haag R, Welker P, Calderón M (2015) . Nanoscale 7:3923
Kurniasih IN, Keilitz J, Haag R (2015) . Chem Soc Rev 44:4145
Xu X, Ran Q, Dey P, Nikam R, Haag R, Ballauff M, Dzubiella J (2017) . Macromolecules 50(12):4759
Nikam R, Xu X, Ballauff M, Kanduc M, Dzubiella J (2018) . Soft Matter 14:4300
Weinhart M, Gröger D, Enders S, Riese SB, Dernedde J, Kainthan RK, Brooks DE, Haag R (2011) . Macromol Biosci 11:1088
General IJ (2010) . J Chem Theory Comput 6:2520–2524
Langmuir I (1918) . J Am Chem Soc 40(9):1361
McQuarrie DA (2000) Statistical Mechanics. University Science Books
Dill KA, Bromberg S (2003) Statistical thermodynamics in chemistry and biology. Garland Science, New York
Turnbull WB, Daranas AH (2003) . J Am Chem Soc 125:14859
Wyman J Jr (1964) . Adv Protein Chem 19:223
Schellman JA (1975) . Biopolymers 14:999
Volmer MA, Mahnert P (1925) . Z Phys Chem 115:253
Masel R (1996) Principles of adsorption and reaction on solid surfaces. Wiley Interscience
Deng Y, Roux B (2009) . J Phys Chem B 113:2234–2246
Gumbart JC, Roux B, Chipot C (2013) . J Chem Theo Comp 9:794
Yigit C, Welsch N, Ballauff M, Dzubiella J (2012) . Langmuir 28:14373
Aveyard R, Haydon R (1973) Introduction to the principles of surface chemistry. Cambridge University Press, Cambridge
Seelig J (2004) . Biochimica et Biophysica Acta-Biomembranes 1666:40
McLaughlin S (1989) . Annu Rev Biophys Biophys Chem 18:113
Santos A, Lãspez de Haro M, Bravo Yuste S (1995) . J Chem Phys 103(11):4622
Zaccone A, Terentjev EM (2012) . Phys Rev Lett 108:038302
Ness C, Zaccone A (2017) . Ind Eng Chem Res 56:3726
Wittemann A, Ballauff M (2006) . Phys Chem Chem Phys 8:5269
Henzler K, Haupt B, Lauterbach K, Wittemann A, Borisov O, Ballauff M (2010) . J Am Chem Soc 132(9):3159
Becker AL, Henzler K, Welsch N, Ballauff M, Borisov O (2012) . Curr Opin Coll Interf Sci 17:90
Yu S, Xu X, Yigit C, van der Giet M, Zidek W, Jankowski J, Dzubiella J, Ballauff M (2015) . Soft Matter 11(23):4630
Yigit C, Heyda J, Ballauff M, Dzubiella J (2015) . J Chem Phys 143(1):064905
Yigit C, Kanduc M, Ballauff M, Dzubiella J (2016) . Langmuir 33:417
Xu X, Kanduc M, Wu J, Dzubiella J (2016) . J Chem Phys 145(3):034901
Record MT, Anderson CF, Lohman TM (1978) . Q Rev Biophys 11(02):103
Xu X, Angioletti-Uberti S, Lu Y, Ballauff M, Dzubiella J (2019) . Langmuir 35:5373
Adroher-Benítez I, Moncho-Jorda A, Dzubiella J (2017) . Langmuir 33:4567
Angioletti-Uberti S, Ballauff M, Dzubiella J (2014) . Soft Matter 10:7932
Moncho-Jorda A, German-Bellod A, Angioletti-Uberti S, Adroher-Benitez I, Dzubiella J (2019) . ACS Nano 13:1603
Acknowledgments
The authors are grateful to Rohit Nikam for a critical reading of the manuscript. Xiao Xu thanks the Chinese Scholar Council and the National Science Foundation of China (21903045) for financial support.
Funding
Open Access funding enabled and organized by Projekt DEAL. Xiao Xu received financial support from the Chinese Scholar Council and the National Science Foundation of China (21903045).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised due to a retrospective Open Access order.
Appendix
Appendix
The equation to fit a ITC sigmoidal curve according to the Langmuir isotherm reads:
with ξ = 1 + x/N + 1/(NKbcm). Now, we write \(x^{\prime } = x/N\) as the fraction between the molar ratio and the number of the binding sites, \(f= \frac {1}{Vc_{m} {\Delta } H_{\text {ITC}}}Q^{\prime }(x)\) denoting the normalized sigmoidal curve, and c = Kbcm as the Wiseman parameter of the ITC experiment [44]. With that, the derivative df/dx with respect to the molar ratio x is:
On that basis, the second-order derivative d2f/d2x follows:
The inflection point of the sigmoidal curve with d2f/d2x = 0 is then determined as \(x^{\prime } = 1-1/c\), which approaches 1 in the limit of \(c\rightarrow \infty \). In that case, the corresponding slope of the titration isotherm at the inflection point is:
which is found proportional to the square root of the binding constant Kb. Thereby, the binding constant is provided by the slope at the inflection point of the sigmoidal curve in the framework of the Langmuir isotherm. Given a coverage-dependent Kb(𝜃((x)), the Langmuir fit then returns the binding constant Kb(x = N) associated with the inflection point at \(x^{\prime }= x/N = 1\) in the limit of \(c \rightarrow \infty \).
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Xu, X., Dzubiella, J. Probing the protein corona around charged macromolecules: interpretation of isothermal titration calorimetry by binding models and computer simulations. Colloid Polym Sci 298, 747–759 (2020). https://doi.org/10.1007/s00396-020-04648-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00396-020-04648-x