Abstract
Pathogenic variants in RAC3 cause a neurodevelopmental disorder with brain malformations and craniofacial dysmorphism, called NEDBAF. This gene encodes a small GTPase, which plays a critical role in neurogenesis and neuronal migration. We report a 31 weeks of gestation fetus with triventricular dilatation, and temporal and perisylvian polymicrogyria, without cerebellar, brainstem, or callosal anomalies. Trio whole exome sequencing identified a RAC3 (NM_005052.3, GRCh38) probably pathogenic de novo variant c.276 T>A p.(Asn92Lys). Eighteen patients harboring 13 different and essentially de novo missense RAC3 variants were previously reported. All the patients presented with corpus callosum malformations. Gyration disorders, ventriculomegaly (VM), and brainstem and cerebellar malformations have frequently been described. The only previous prenatal case associated with RAC3 variant presented with complex brain malformations, mainly consisting of midline and posterior fossa anomalies. We report the second prenatal case of NEDBAF presenting an undescribed pattern of cerebral anomalies, including VM and polymicrogyria, without callosal, cerebellar, or brainstem malformations. All neuroimaging data were reviewed to clarify the spectrum of cerebral malformations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
De novo pathogenic variants of RAC3 cause neurodevelopmental disorder with structural brain anomalies and dysmorphic facies (NEDBAF) (OMIM #618577) [1], whereas some de novo missense RAC1 variants with dominant-negative effect have been associated with cerebral malformations and highly variable head circumference (OMIM #617751) [2].
RAC3 is expressed in the central nervous system, whereas RAC1 is expressed ubiquitously. RAC3 belongs, like RAC1, to the Rho family of guanosine triphosphatases (GTPases), which are key regulators of cytoskeletal dynamics and intracellular signalling. RAC3 is involved in neuronal migration, and both RAC1 and RAC3 are involved in neurogenesis. These proteins share a highly conserved G domain that mediates GDP/GTP binding and GTP hydrolysis. This region includes two regions, Switch-I and Switch-II, where most RAC1 and RAC3 pathogenic variants are located [3]. Hinde et al. demonstrated that dimeric RAC1 is inactive and selectively exported to the cytoplasm, whereas monomeric RAC1 is active and retained in the nucleus [4].
Here, we report a fetus from an unrelated and healthy Caucasian couple. Ultrasonography performed at 28 weeks of gestation (WG) demonstrated triventricular dilatation with square frontal horns, frontal lobes hypoplasia, insufficient Sylvian operculization and bilateral clubfeet (Fig. 1A).
A Sonographic and MRI examinations performed respectively at 28 and 31 WG showed ventriculomegaly and square shape of the frontal horns with enlarged pericerebral spaces and hypoplastic frontal lobes (arrow) (1,5), abnormal Sylvian operculization (2,6: arrows), the normal corpus callosum (3 between crosses, 7) and the normal aspect of the brainstem and cerebellum (4,7,8 arrows). B The 3D structures of wild-type (left side) and variant (right side) RAC3 proteins were predicted with the PyMOL application (2C2H Crystal structure of RAC3 in complex with GDP from the Protein Data Bank: https://www.rcsb.org/). The wild-type (WT) and variant amino acids are represented in green and red respectively. In silico protein modeling indicates that the variant could interfere with RAC3 dimer formation, because of its position at the interface between both RAC3 proteins. Cytoplasmic accumulation of active monomeric forms of RAC3 could be suggested, as previously described with RAC1 variants [4]. C Schematic overview of the canonical RAC3 transcript. Coding exons of RAC3 are illustrated by light blue boxes (ENST00000306897.8), and introns by black lines. Residues 1–192 were indicated within the corresponding exons. The G boxes are represented by the green boxes. Switch I region (red) extends from residues 25–39 and Switch II region (purple) from residues 57–75. The variant in RAC3 identified in this study is indicated by the green line, and the eighteen different variants in RAC3 reported in previous publications are indicated using dark blue lines. Exon lengths, variants/Switch domains/G boxes positions are represented approximately to scale, but intron lengths are not
White matter thinning, expanded subarachnoid spaces, and temporal and perisylvian polymicrogyria were identified on MRI performed at 31 WG (Fig. 1A).
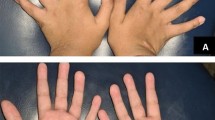
No cerebellar, brainstem, or callosal anomalies were observed. Parents asked for termination of the pregnancy at 32 WG. Post mortem external examination showed craniofacial dysmorphism with a large forehead, hypertelorism, downslanting palpebral fissures, bilateral epicanthus, short nose with anteverted nostrils, small mouth, thin upper lip, micro retrognathia, and short neck.
Molecular karyotype on DNA extracted from amniotic cells and blood samples from the parents was normal. Trio whole exome sequencing (WES) identified a novel de novo heterozygous variant c.276 T>A p.(Asn92Lys) in RAC3 (NM_005052.3, GRCh38), absent from control databases (gnomAD 3.1), involving a highly conserved residue and predicted to be deleterious by most in silico tools (CADD score 20.5). Sequence alignment to the human reference genome, variants calling and annotation were performed using an in‐house bioinformatic pipeline. The variant is located outside the Switch domains, but in silico protein modeling suggests that it could interfere with the RAC3 dimer formation (Fig. 1B). Considering the strong genotype/phenotype correlation, this variant was classified as probably pathogenic (Class 4).
Eighteen patients, including 16 unrelated patients and 2 half-sibs, were previously reported, harboring 13 different and essentially de novo missense RAC3 variants [1, 5,6,7,8,9] (Table 1; Fig. 1C).
Only one prenatal case has been previously reported [5]. The hallmark malformation concerned the corpus callosum (CC): thick genu, splenium thinning, and partial or total agenesis. Here, we report the first case without callosal abnormalities, suggesting that an apparently normal CC does not exclude the involvement of RAC3. Gyration disorders, including polymicrogyria at various locations, are the second most commonly reported neuroimaging characteristic. Cerebral parenchymal thinning with subsequent enlarged subarachnoid spaces is also often described, in contrast to a normal head circumference. Ventriculomegaly (VM) was present in more than half of cases. Until now, square frontal horns have not been reported to be associated with RAC3-related neurodevelopmental disorders. However, we noticed square frontal horns in previously reported cases after reviewing the existing neuroimaging data (Table 1). Structural defects of the brainstem and cerebellum are also part of the NEDBAF features with midbrain hypoplasia, elongated, flattened and/or bifid pons, abnormal cerebellar foliation, and vermis defects. Midline malformations, gray matter heterotopia, and Arnold-Chiari malformation type I have also been described.
In vitro functional studies performed on 12 variants [7, 8] have demonstrated abnormal neuronal morphology in transfected cells in comparison to the control. A gain-of-function effect with an overall increase in GDP/GTP exchange activity and reduction or suppression of GTPase hydrolytic activity. In vivo functional studies performed on five variants [7, 8] have shown severe anomalies in neuronal migration as well as defects in axon extension to the contralateral hemisphere during corticogenesis. Only two RAC3 pathogenic variants have been previously reported outside the Switch domains [8]. There seems to be no difference in the phenotype, whether the variant is within or outside the domain.
We report here the second prenatal case of NEDBAF syndrome. The previous fetal case presented with complex brain malformations mainly consisting of midline and posterior fossa anomalies [5], whereas we identified mostly cortical malformations. Both fetuses had a fetal akinesia sequence and similar craniofacial dysmorphic features. The variation between both fetuses could be explained, at least in part, by the different localization of the variants.
Many of the brain malformations described in the NEDBAF condition are prenatal cerebral features suggestive of tubulinopathies. Cabet et al. [10] described two distinct prenatal imaging patterns related to tubulinopathies: a severe form characterized by enlarged germinal matrices, microlissencephaly, and kinked brainstem, and a mild form associated with asymmetric brainstem, corpus callosal dysgenesis, lack of Sylvian fissure operculization, and distortion of the anterior part of the interhemispheric fissure, in the absence of additional extracerebral anomalies. In the largest series of 19 fetuses with prenatal diagnosis of tubulinopathy, Brar et al. [11] reported VM, cerebellar hypoplasia, brainstem kinking, abnormalities of the CC, flat appearing Sylvian fissure, microlissencephaly, microcephaly and absence of cavum septum pellucidum. Square-shaped anterior horns described in some cases of NEDBAF are also compatible with tubulinopathies [10]. Cerebral malformations are highly variable in both conditions, and can affect the supratentorial and infratentorial parts of the brain.
In this study, we report a second prenatal case of NEDBAF presenting an undescribed pattern of cerebral anomalies, including VM with square-shaped anterior horns, and temporal and perisylvian polymicrogyria, without callosal, cerebellar, or brainstem malformations. NEDBAF should be considered as a differential diagnosis of tubulinopathies because of the similarity of cerebral malformations. Enlarged germinal matrices, microlissencephaly or microcephaly should lead to suspicion of tubulinopathy in the first place. On the other hand, extracerebral features, such as arthrogryposis, should point towards RAC3-fetopathy. Furthermore, we outline that various brain malformations can lead to the same diagnosis, highlighting the benefit of fast whole exome sequencing in prenatal cases.
Data availability
No datasets were generated or analysed during the current study.
References
Costain G, Callewaert B, Gabriel H, Tan TY, Walker S, Christodoulou J, Lazar T, Menten B, Orkin J, Sadedin S, Snell M, Vanlander A, Vergult S, White SM, Scherer SW, Hayeems RZ, Blaser S, Wodak SJ, Chitayat D, Marshall CR, Meyn MS (2019) De novo missense variants in RAC3 cause a novel neurodevelopmental syndrome. Genet Med Off J Am Coll Med Genet 21:1021–1026. https://doi.org/10.1038/s41436-018-0323-y
Banka S, Bennington A, Baker MJ, Rijckmans E, Clemente GD, Ansor NM, Sito H, Prasad P, Anyane-Yeboa K, Badalato L, Dimitrov B, Fitzpatrick D, Hurst ACE, Jansen AC, Kelly MA, Krantz I, Rieubland C, Ross M, Rudy NL, Sanz J, Stouffs K, Xu ZL, Malliri A, Kazanietz MG, Millard TH (2022) Activating RAC1 variants in the switch II region cause a developmental syndrome and alter neuronal morphology. Brain 145:4232–4245. https://doi.org/10.1093/brain/awac049
Scala M, Nishikawa M, Nagata K, Striano P (2021) Pathophysiological mechanisms in neurodevelopmental disorders caused by Rac GTPases dysregulation: what’s behind neuro-RACopathies. Cells 10:3395. https://doi.org/10.3390/cells10123395
Hinde E, Yokomori K, Gaus K, Hahn KM, Gratton E (2014) Fluctuation-based imaging of nuclear RAC1 activation by protein oligomerisation. Sci Rep 4:4219. https://doi.org/10.1038/srep04219
Cabet S, Vasiljevic A, Putoux A, Labalme A, Sanlaville D, Chatron N, Lesca G, Guibaud L (2022) Prenatal imaging features related to RAC3 pathogenic variant and differential diagnoses. Prenat Diagn 42:478–481. https://doi.org/10.1002/pd.6106
Hiraide T, Kaba Yasui H, Kato M, Nakashima M, Saitsu H (2019) A de novo variant in RAC3 causes severe global developmental delay and a middle interhemispheric variant of holoprosencephaly. J Hum Genet 64:1127–1132. https://doi.org/10.1038/s10038-019-0656-7
Nishikawa M, Scala M, Umair M, Ito H, Waqas A, Striano P, Zara F, Costain G, Capra V, Nagata K-I (2023) Gain-of-function p. F28S variant in RAC3 disrupts neuronal differentiation, migration and axonogenesis during cortical development, leading to neurodevelopmental disorder. J Med Genet 60:223–232. https://doi.org/10.1136/jmedgenet-2022-108483
Scala M, Nishikawa M, Ito H, Tabata H, Khan T, Accogli A, Davids L, Ruiz A, Chiurazzi P, Cericola G, Schulte B, Monaghan KG, Begtrup A, Torella A, Pinelli M, Denommé-Pichon AS, Vitobello A, Racine C, Mancardi MM, Kiss C, Guerin A, Wu W, Gabau Vila E, Mak BC, Martinez-Agosto JA, Gorin MB, Duz B, Bayram Y, Carvalho CMB, Vengoechea JE, Chitayat D, Tan TY, Callewaert B, Kruse B, Bird LM, Faivre L, Zollino M, Biskup S, Undiagnosed diseases network, telethon undiagnosed diseases program, Striano P, Nigro V, Severino M, Capra V, Costain G, Nagata KI (2022) Variant-specific changes in RAC3 function disrupt corticogenesis in neurodevelopmental phenotypes. Brain J Neurol 145:3308–3327. https://doi.org/10.1093/brain/awac106
White JJ, Mazzeu JF, Coban-Akdemir Z, Bayram Y, Bahrambeigi V, Hoischen A, van Bon BWM, Gezdirici A, Gulec EY, Ramond F, Touraine R, Thevenon J, Shinawi M, Beaver E, Heeley J, Hoover-Fong J, Durmaz CD, Karabulut HG, Marzioglu-Ozdemir E, Cayir A, Duz MB, Seven M, Price S, Ferreira BM, Vianna-Morgante AM, Ellard S, Parrish A, Stals K, Flores-Daboub J, Jhangiani SN, Gibbs RA, Center B-H, for Mendelian Genomics, Brunner HG, Sutton VR, Lupski JR, Carvalho CMB (2018) WNT Signaling Perturbations Underlie the Genetic Heterogeneity of Robinow Syndrome. Am J Hum Genet 102:27–43. https://doi.org/10.1016/j.ajhg.2017.10.002
Cabet S, Karl K, Garel C, Delius M, Hartung J, Lesca G, Chaoui R, Guibaud L (2021) Two different prenatal imaging cerebral patterns of tubulinopathy. Ultrasound Obstet Gynecol 57:493–497. https://doi.org/10.1002/uog.22010
Brar BK, Thompson MG, Vora NL, Gilmore K, Blakemore K, Miller KA, Giordano J, Dufke A, Wong B, Stover S, Lianoglou B, Van den Veyver I, Dempsey E, Rosner M, Chong K, Chitayat D, Sparks TN, Norton ME, Wapner R, Baranano K, Jelin AC, Fetal sequencing consortium (2022) prenatal phenotyping of fetal tubulinopathies: a multicenter retrospective case series. Prenat Diagn 42:1686–1693. https://doi.org/10.1002/pd.6269
Acknowledgements
The authors thank the patients for their participation in this study. We are also grateful to Dr. Scala and Dr. Costain for helping us complete the neuroimaging data for their previously published patients.
Author information
Authors and Affiliations
Contributions
C.M. performed genetic consultations, wrote the manuscript, and created the Table and Fig. 1a and c. M.C. performed sonographic and MRI examinations, reviewed neuroimaging data and commented Fig. 1a. K.K. referred the family. N.S. created the bioinformatic tool for molecular study. O.M. performed molecular study and created the in silico protein modeling. A.T. wrote the manuscript and supervised the study. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
All genetic testing were performed by clinical service under clinical consent forms. Informed consent was obtained from legal guardians. Termination of pregnancy was performed according to Belgian law.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Meunier, C., Cassart, M., Kostyla, K. et al. An unusual presentation of de novo RAC3 variation in prenatal diagnosis. Childs Nerv Syst 40, 1597–1602 (2024). https://doi.org/10.1007/s00381-024-06285-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00381-024-06285-z