Abstract
Despite the crucial role of microbial communities in agroecosystem functioning, a clear picture of how nitrogen shapes rhizosphere microbial complexity and community structure across diverse maize lines remains elusive. To address this gap, we conducted 16S amplicon sequencing of the rhizosphere microbial communities across a diverse range of maize inbred lines (305 genotypes) and their F1 hybrids (196 genotypes) cultivated in both low-nitrogen (unfertilized) and high-nitrogen (fertilized) soils. Our findings reveal that N fertilizer treatment had contrasting effects on the rhizosphere microbial communities of inbreds and hybrids. N fertilization increased alpha diversity but decreased the abundance of Pseudomonas taxa in inbred lines, while the opposite was true for hybrids. The proportion of variance determined by plant host factors was also better explained under low-N, demonstrating that N fertilization reduced the influence of the host over the rhizosphere microbial community. Microbial networks revealed significant differences in the number of nodes and clustering coefficients between the rhizosphere microbial communities of inbred and hybrid maize, with these differences being further differentiated by changes in nitrogen levels. Overall, our study reveals the interplay among rhizosphere microbiomes, abiotic stress induced by low soil nitrogen, and plant host factors facilitating the identification of stable microbial communities in response to environmental stress. These findings contribute to the potential engineering of resilient microbial consortia highlighting the importance of the influence of plant genotype and the environment on the rhizosphere microbiome.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The plant rhizosphere is an essential zone directly adjacent to the plant root system that hosts a diverse and functionally dynamic microbial consortium (Philippot et al. 2013; Shi et al. 2016; Ling et al. 2022; Mukhtar et al. 2023). These microbes play a vital role in nutrient cycling, organic matter decomposition, and plant defense against different stressors and diseases (Haney et al. 2015; Hu et al. 2018; Gu et al. 2020). Therefore, it is important to comprehend the shift in rhizosphere microbial community composition and stability to changing edaphic environments across diverse plant genotypes. Such knowledge is crucial not only to engineer resilient microbial consortia for varying field conditions but also to model the global microbial distribution in agroecosystems.
Nitrogen (N) fertilization is one of the most important factors influencing the soil edaphic environment and, subsequently, soil microbial community diversity and composition (van der Bom et al. 2018; Mukhtar et al. 2023). Previous research has demonstrated varying impacts of N application on belowground microbial communities, yielding contrasting results and conclusions. These range from substantial positive or negative influences (Yang et al. 2022; Mukhtar et al. 2023) to instances where no significant alterations in community diversity and composition were observed (French et al. 2017; Ouyang and Norton 2020; Duff et al. 2022). For example, N fertilization has been reported to enhance nutrient availability and to support a diverse microbial community (van der Bom et al. 2018; Li et al. 2020), while alleviating N stress can foster stable microbial communities (Hernandez et al. 2021). Conversely, excessive N fertilization (and N deposition) can adversely affect soil microbial diversity (Zhang et al. 2018; Hao et al. 2022; Yang et al. 2022; Wang et al. 2023), thereby negatively impacting ecosystem functioning. Excess N fertilization also led to increased carbon deposition that resulted in differences in the labeled carbon uptake by microbial groups in two grass species (Leptin et al. 2022) highlighting the importance of N additions in influencing plant root carbon release and the subsequent impact on the soil microbiome. Similarly, the complexity and modularity of soil microbial networks have also been shown to vary with N application across various field conditions (Li et al. 2021; Liu et al. 2021; Zhang et al. 2021). While the responses of soil microbial communities have been documented, our current understanding of how rhizosphere microbial communities respond to N fertilization (and N stress) remains inconclusive.
Host plant-microbiome interactions introduce an additional layer of complexity and uncertainty in discerning the microbial response to N levels. On the one hand, the genetic makeup of the host plant profoundly influences the physical and chemical characteristics of the rhizosphere (Sasse et al. 2018; Wang et al. 2022; Lopes et al. 2023), which, in turn, filters and recruits different microbial taxa. Notably, variations in primary and secondary metabolites in root exudates play a pivotal role in shaping the soil microbiota, as these compounds serve as an essential source of energy and a specialized environment that enhances growth as well as influences the composition of microbial communities (Sasse et al. 2018; Zhalnina et al. 2018; Jacoby et al. 2021). On the other hand, host plant genotypes exhibit diverse root system architecture, which may directly or indirectly exert influence on the composition and functionality of microbial communities (Szoboszlay et al. 2015; Saleem et al. 2018) by altering various soil biophysical and edaphic properties. These variations in root exudates and root structure orchestrate the changes in the structure and function of soil microbes and gene expression in the rhizosphere, which varies depending on soil properties and developmental stages (Hao et al. 2022; Lopes et al. 2023).
Plants may also adjust their nutrient acquisition strategies in response to N availability—e.g., changes in resource allocation (e.g., enzyme production) under differing nutrient conditions —impacting the composition of root exudates (Zhu et al. 2016; Hao et al. 2022). Changing N conditions also shifts gene expression profiles and morphological characteristics of plant root systems (Chen et al. 2011; Gaudin et al. 2011; Singh et al. 2022). This could influence the proportion of variance in microbial relative abundance associated with plant genotype. Nevertheless, these evolving plant root- microbe interactions are likely to exert their influence on soil biogeochemical cycling (Mukhtar et al. 2023) and have been shown to impact plant phenology and expression of heterosis (O’Brien et al. 2021; Wagner et al. 2021). These complex interactions and feedback make it essential to understand the shift in rhizosphere microbial community composition, and its complexity and stability in response to changing N levels and how these responses differ between diverse plant genotypes.
In this study, by using 305 inbreds and 196 F1 hybrids, we investigated the potential impacts of maize genotype on the rhizosphere microbial community composition, microbial heritability, and co-occurrence patterns under two distinct N schemes: nutrient deficient soils (low-N) and fertilized soils (high-N). Maize was chosen as the focal crop because of its economic importance, widespread global cultivation, the remarkable adaptability of hybrids within agroecosystems and considerable difference in microbial community composition between inbred lines and their hybrid rhizospheres (Wagner et al. 2020). Although, soil edaphic and climatic conditions (Bahram et al. 2018; Mukhtar et al. 2020, 2023), as well as the endosphere (Lopes et al. 2022; Wang et al. 2022), significantly influence belowground microbial community structure, this study primarily focused on changes in soil microbial community dynamics within the rhizosphere of a large population of maize inbreds and hybrids in response to different N treatments. We hypothesized that N input will: 1) differentially influence the microbial diversity of rhizospheres across different plant groups (inbred lines and their hybrids), 2) reduce the proportion to which the variation in microbial relative abundance could be attributed to plant genetic factors with a stronger impact in hybrid maize, and 3) increase the modularity and complexity of microbial networks, especially in rhizosphere of inbreds. The outcome of this study contributes to elucidating the combined effects of diverse plant lineages and contrasting N availability that ultimately shapes the structure of rhizosphere microbial communities in maize production ecosystems.
Material and methods
Experimental design and sample collection
A total of more than 500 maize inbred lines and hybrids were planted (in May, 2022) at a University of Nebraska—Lincoln, facility (40.867695, -96.603074). The area was rain-fed and characterized by a mean annual temperature of approximately 12 °C and a mean annual precipitation of around 760 mm. The soil was slightly acidic, primarily silty clay loam, with an organic matter content varying from 2.3% to 3.6% and a nitrate level from 4.1 to 7.0 ppm prior to fertilization. The inbred lines are doubled haploid (DH) lines released from the USDA GEM (Germplasm Enhancement of Maize) project (Vanous et al. 2018). Hybrids were generated in 2020 and 2021 by crossing of BGEM lines with two tester inbred lines B73 and Mo17. Specifically, BGEM lines derived from PHZ51 as the recurrent parent were crossed with B73, while BGEM lines from PHB47 were crossed with Mo17. The selection of B73 or Mo17 lines was due to their high importance as stiff-stalk and non-stiff-stalk varieties. A description about BGEM lines, parental race, and the origin has been previously described (Vanous et al. 2018).
We further grouped the inbred lines into eight subgroups based on stalk stiffness (stiff or non-stiff), flowering time (early or late), and plant height (short or tall). Hybrid groups were similarly categorized into eight subgroups, corresponding to their respective parental BGEM inbred lines and their crosses with either B73 or Mo17. Each subgroup comprised between 14 to 54 different genotypes, and these genotypes were replicated in four blocks (following an incomplete block design): two blocks for unfertilized soils, referred to as the 'low nitrogen treatment' (low-N), and two blocks for fertilized soils with 168 kg/ha of N as urea, referred to as the 'high nitrogen treatment' (high-N). Each row spanned 4.5 m, and 20 seeds were planted per row.
After eight weeks of planting (July 2022), the plant roots were extracted directly from the field soil at a depth of 20 to 30 cm. The roots were shaken to remove loosely adherent bulk soil. To collect rhizosphere samples in the field, 5 – 8 segments of randomly selected roots, including adherent rhizosphere soil was placed into 50 ml tubes containing 35 ml of phosphate buffer and then vortexed for two minutes to remove most of adherent rhizosphere soil. Roots were then removed, and tubes were placed on ice and brought into the laboratory. The rhizosphere suspensions were then filtered through a 100 µM sterile mesh into a fresh 50 ml tube to remove root debris. Rhizosphere samples were then spun down, followed by removal of the supernatant. Then 2 ml of phosphate buffer was added to the rhizosphere soil which was then resuspended and transferred to a 2 ml tube. This was then spun down again to allow for the removal of the supernatant and the rhizosphere soil was then frozen at –20 °C. For a detailed protocol please see (McPherson et al. 2018).
16S library preparation and Sequencing data analysis
DNA was extracted from the rhizosphere soil using a MagAttract PowerSoil Pro DNA Kit Soil DNA Isolation kit (Qiagen LLC, Germantown, MD). The V4 region of the 16S rRNA gene was then amplified from the rhizosphere DNA using the primers 515F and 806R (Caporaso et al. 2012). To amplify this region PCR was used with initial denaturing at 95 °C for 5 min, 30 cycles of 95 °C for 40 s, 65 °C for 30 s and 72 °C for 1 min, followed by final extension at 72 °C for 10 min. The sequencing libraries were prepared using a dual-index primer system (Kozich et al. 2013). Amplicons were purified using an NGS 96-well normalization kit (Norgen Biotek Corp, ON, Canada) and then normalized based on the DNA concentrations which were quantified using QuantiFluor ONE dsDNA Dye (Promega Corporation, Madison WI) and then pooled in an equimolar mixture, and further purified using the SPRIselect beads (Beckman Coulter, Brea, CA). Sequencing was then performed on an Illumina MiSeq platform using the MiSeq 600 cycles v3 kit. Sequences generated in this study are found on NCBI (accession number: PRJNA1092665).
Raw sequence data of forward and reverse reads (FASTQ) was processed within the QIIME2 environment (Bolyen et al. 2019) (release 2023.5) and denoised sequences within QIIME2 using the DADA2 pipeline. We clustered the remaining sequences into amplicon sequence variants (ASVs) against the SILVA 138.1 database (Quast et al. 2013) using a pre-trained Naïve Bayes feature classifier. Raw ASV reads were subjected to a series of filters to produce a final ASV table with biologically relevant and reproducible 16S sequences. Samples with 1) sequencing depth < 3000, 2) ASVs that were not observed in at least 25 samples were removed. Chloroplast and mitochondrial sequences were eliminated from all samples. A total of 7,007 ASVs—with a total frequency of 41,912,761 and a median frequency per sample of 20,153—were obtained and classified into the 293 genera and 24 known phyla.
Microbial abundance and diversity analyses
The relative abundance (%) of microbial taxa was individually quantified at both the phylum and genus levels for each sample, and subsequently averaged across inbred and hybrid plants, including their subgroups, as well as across various nutrient treatments by using base functions in R. Alpha and beta diversity were assessed using the ‘vegan’ package in R (Oksanen et al. 2022). Alpha diversity was measured using the inverse Simpson index, which provides insights into species richness and evenness within individual samples. ANOVA, followed by a post-hoc test (Tukey's test) that was used to determine whether there were significant differences between groups (inbred and hybrids) and subgroups based on stalk stiffness (stiff or non-stiff), flowering time (early or late), and plant height (short or tall) under low- and high-N treatments (n = 14 – 54 for alpha diversity).
Beta diversity was evaluated using the Bray–Curtis dissimilarity index, which quantified the variation in species composition between different samples. The data were rarefied for alpha diversity, ensuring that all samples had the same number of sequences, while non-rarefied data were used for beta diversity and network analysis, preserving the original sequencing depth for a comprehensive assessment of community composition and interactions. Additionally, statistical tests such as Analysis of Similarities (ANOSIM, method = Bray) and Multivariate Analysis of Variance Using Distance Matrices (Adonis) were applied to determine the significance of dissimilarity between sample groups (n = 14 – 54), allowing for the comprehensive analysis of diversity patterns in the dataset.
Plant genotype driven microbial variation
We estimated the proportion to which the variation in microbial relative abundance could be attributed to plant genetic factors (hereafter referred to as VPM) for microbial groups from the rhizosphere of all genotypes in two replicated blocks at the genus and phylum levels. The VPM was estimated across the experiment including both inbred and hybrid rhizospheres, as well as separately for each group under low-N and high-N treatments. We characterized both higher and lower taxonomic levels to obtain a more comprehensive understanding of the plant genotype-associated microbial taxa and to investigate whether, instead of the entire community, only a subset of the microbiome is responsive to host variation. VPM was estimated by using proportion of variance explained by the genotype (Vg) relative to the total variance, which includes both (Vg) and error (Ve) and N is the number of replicates (2) (Eq-1).
Taxa that appeared in fewer than 200 samples, representing roughly 10% of the total samples, were omitted, and the remaining data were log-transformed before conducting the VPM analysis. The 'lmer' function from the R package 'lme4' was used to construct linear mixed-effects models with all random effects. Finally, a permutation test with 5000 iterations was conducted to assess whether the observed VPM is attributable to random variability or was statistically significant. Subsequently, rhizosphere microbial taxa that were significantly associated with plant genetic factors (p < 0.05) are referred to as plant genotype-associated microbial taxa (HT), while microbial taxa that were not significant were removed from analysis.
Network analysis
Co-occurrence networks were constructed to investigate microbial interactions within subgroups of both inbreds and hybrids under low-N and high-N conditions. We created distinct networks for each of the eight inbred subgroups, as opposed to forming two large networks encompassing both inbred and hybrid individuals by utilizing all genotypes within the subgroups and their two replications. This approach allowed for a more nuanced analysis of network structures while avoiding the potential confounding effects of amalgamating all inbreds into a single network. We constructed co-occurrence networks using function sparcc (iter = 100, and other parameters with default values) from R package “SpiecEasi” with ASVs present in > 10% samples. To enhance network clarity and reduce the occurrence of false positives, we applied a correlation cutoff of 0.3 (|correlation| \(\ge\) 0.3), accompanied by a significance threshold of 0.05 (p < 0.05). Significance of correlations was calculated using 5000 bootstraps. Group used for the network statistical analysis contained between n = 14 – 54 genotypes.
The co-occurrence network graphs were generated using the 'igraph' package in R (Csardi and Nepusz 2006). Comprehensive insights into network properties were obtained through the computation of key parameters, including node count, edge count, modularity (M) (from absolute weight), and clustering coefficients (CC, type = ’global’). All statistical analyses were conducted using the R programming software, unless stated otherwise.
Results
Rhizosphere microbial diversity and community structure
We first assessed the relative abundance of microbial groups in the rhizosphere at differing taxonomic levels between maize inbreds and hybrids under low- and high-N conditions. A total of 152 and 142 out of 293 microbial groups (at the genus level) differed between inbred and hybrid under low-N and high-N treatments, respectively. Pseudomonas was the dominant bacterial taxon at the genus level and was significantly more abundant in the rhizosphere of hybrids (ANOVA, p < 0.05, Supplementary Table 1). The mean relative abundance of Pseudomonas in hybrids was 28.3% for low-N and 41.5% for high-N, compared to inbreds with 22.4% for low-N and 20.1% for high-N. The second most abundant taxon at the genus level in the rhizosphere of hybrids was an unclassified taxon of the phylum Acidobacteriota whose mean relative abundance ranged from 6.9% in low-N to 5.6% for high-N (Fig. 1a). In contrast Ralstonia was the second most abundant in the rhizosphere of inbreds ranging from 4.8% in low-N and 7.6% in high-N. Moreover, the BCP (Burkholderia-Caballeronia-Paraburkholderia), commonly found in bulk soil and the rhizosphere soil was in lower abundance (p < 0.05) in the rhizosphere of hybrids compared to inbred lines. The relative abundance of BCP in the rhizosphere of inbreds was on average 2.9% in low-N and 4.2% in high-N conditions, while its abundance was less than 1% in the hybrid samples under low-N conditions. At the phylum level, a total of 15 phyla under low-N and 12 under high-N conditions (out of 24) significantly differed (p < 0.05) in relative abundance between inbreds and hybrids. Among them, Proteobacteria, Acidobacteriota, and Actinobacteriota were the dominant bacterial taxa, with mean relative abundances of 49.6%, 11.5%, and 8.3%, respectively, across samples.
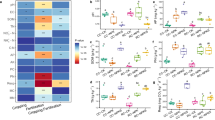
Dominant microbial taxa and alpha-diversity in the rhizosphere across maize inbreds and hybrids under different N treatments. (a) The relative abundance of the dominant microbial taxa (top 10 dominant taxa) across inbred lines and hybrids at the genus level under low-N (LN) and high-N (HN). Microbial alpha-diversity (measured as the Inverse Simpson index) (b) across inbred and hybrid lines and (c) across different subgroups of inbreds—classified based on stalk stiffness, flowering time, and plant height (SE, TE, SL and TL)—and corresponding hybrids (H1 to H4) under LN and HN treatments. S: Stiff stalk, N: Non-stiff stalk. SE: Short-early, TE: Tall-early SL: Short-late, and TL: Tall-late. WD2101 (soil type). * p < 0.05 and ** p < 0.01
The alpha diversity was significantly influenced by both plant groups (inbreds and hybrids) and N treatments (low- and high-N) (ANOVA, F: 160.0 and 8.9, respectively, p < 0.01) (Fig. 1b). The alpha diversity in the rhizosphere of inbreds was significantly higher than their hybrid counterparts, particularly under high-N conditions (Fig. 1b). Moreover, the alpha diversity for the rhizosphere microbial communities of the hybrids under low-N (ranging from 1.5 to 41.5, mean: 13.4) was, on average, more than two-fold higher than those values observed under high-N treatments (ranging from 1.3 to 35.4, mean: 7.7). However, the N treatment significantly increased (p < 0.05) the alpha diversity in inbred rhizosphere (Fig. 1b) suggesting a contrasting influence of N treatment on rhizosphere microbial alpha diversity across different plant groups.
We also assessed the change in alpha diversity between subgroups of inbred lines and their corresponding hybrids. The subgroups SE (short-early), SL (short-late), TE (tall-early), and TL (tall-late) represent plant height and flowering time, with S and N indicating stiff stalk and non-stiff stalk, respectively. The corresponding hybrids are represented by H1, H2, H3, and H4, respectively (Fig. 1c). Within inbred lines, the highest alpha diversity was observed for TL (20.3 for S and 22.4 for N), followed by SE (18.2 for S stalks) under high-N conditions. The rhizosphere communities of the maize hybrids, on the other hand, exhibited a significant reduction in the alpha diversity for almost all groups compared to their parental inbreds (BGEM). The most significant difference in alpha diversity between inbred and hybrid rhizosphere microbial communities was observed between inbred subgroup TL (20.3 for S stalk and 22.4 for N stalk) and their corresponding hybrid subgroup H4 (8.6 for S stalk and 5.9 for N stalk) under high-N treatments. Hybrid subgroups (H1 to H4) were statistically indistinguishable (p > 0.05) from each other under high-N treatments.
The plant groups (inbred and hybrid) and N treatments (low- and high-N) also significantly influenced the microbial community composition across the maize rhizosphere (Fig. 2). Adonis (permutational multivariate analysis of variance using distance matrices) showed that microbial community composition differed significantly across both plant groups (F = 105.6, p < 0.01) and N treatments (F = 41.5, p < 0.01), as well as the interaction between plant groups and N treatments (F = 23.6, p < 0.01). We further conducted ANOSIM (Analysis of Similarities, distance matrix: Bray–Curtis) to assess whether dissimilarity within the plant groups was different (i.e., smaller) than those between groups. The bacterial and archaeal communities in the rhizosphere of hybrids were distinct from those observed in the rhizosphere of inbreds under high-N treatments (R = 0.1, p < 0.001), while no significant difference was observed under low-N treatments (Fig. 2a).
Microbial beta-diversity and clustering across maize inbred and hybrid rhizosphere communities under different N conditions. Changes in rhizosphere microbial community composition (Bray–Curtis) between maize inbreds and hybrids under (a) low-N and (b) high-N treatments. The ellipses are drawn at the 95% confidence level
Plant genotype-driven rhizosphere microbial traits
We investigated the proportion of variation in microbial relative abundance associated with plant genotype (abbreviated as VPM) at both the genus and phylum levels under different N treatments. A high VPM value would suggest that maize genetic diversity plays a significant role in shaping the microbial communities, while a low value would indicate that environmental factors may have a stronger influence. When measuring VPM across the samples including both inbreds and hybrids, a total of 38 microbial taxa under low-N and 21 under high-N, at the genus level, were detected as significantly plant genotype-associated microbial taxa (HT) with a mean VPM of 0.21 and 0.25 (p < 0.05) (Fig. 3). Among HT, Ralstonia exhibited the highest proportion of variance associated with plant genotype (VPM = 0.41, p < 0.05), while the BCP was the second most plant genotype-associated taxon (VPM: 0.34, p < 0.05) and Niastella was the third most plant genotype-associated taxon (VPM = 0.33, p < 0.05) under low-N treatments (Fig. 3a). In contrast, Amycolatopsis showed the highest VPM (0.39, p < 0.05), followed by Sphingobium in second place and Ralstonia in third (VPM: 0.36, p < 0.05) under high-N treatments. Moreover, Pseudomonas, the most dominant taxon in our samples, displayed a moderate VPM (0.26, p < 0.05) under high-N. We also noted that VPM was positively correlated with microbial relative abundance under low-N treatments. The correlation between microbial relative abundance and VPM was r = 0.48 (Pearson’s correlation test, p = 0.002) for low-N and r = 0.26 (Pearson’s correlation test, p = 0.25) for high-N.
The proportion of variance in microbial relative abundance associated with plant genotype (VPM) in the rhizosphere at the genus level. a the top 10 plant genotype-associated microbial taxa (HT) (ranked based on VPM values) along with their relative abundance under both low-N (LN) and high-N (HN) conditions (* p < 0.05). b the magnitude of microbial-plant genotype associations (HT × VPM), and (c) VPM for rhizobiome traits under low-N and high-N treatments for both inbred and hybrid lines. The colors indicate whether VPM are significant (p < 0.05) among either inbred (pink), hybrid (pink), or both (green), while gray represents taxa with p > 0.05. NA* Unclassified taxa
We further quantified the VPM for taxa in rhizospheres of inbreds and hybrids separately and identified those that exhibited significant VPM in either inbred or hybrid samples, or in both groups. When measuring VPM across the rhizosphere of inbreds, the number of HT taxa was 31 in low-N treatments and 17 in high-N treatments, each with a mean VPM > 0.2 (p < 0.05). However, the HT was reduced in the rhizospheres of hybrids, particularly under high-N treatment, with only 7 microbial taxa with a mean VPM of 0.25 (Fig. 3b). Moreover, Pseudomonas, the most dominant taxon in our samples, displayed a nonsignificant association with both the rhizosphere of inbreds and hybrids (p > 0.05) under both low-N and high-N (Fig. 3a). When identifying plant genotype-associated microbial taxa present in both inbred and hybrid rhizospheres, a total of 12 microbial groups (9 under low-N and 3 under high-N) were found to be in common (Fig. 3c). Among them, all plant genotype-associated microbial groups under high-N conditions and 8/9 microbial groups under low-N were associated with Proteobacteria, while one microbial group (under low-N) was associated with Actinobacteriota.
At the phylum level, Desulfobacterota showed significant VPM (p < 0.05) under high-N treatments, while Bacteroidota, Actinobacteriota, and Crenarchaeota taxa showed significant VPM (p < 0.05) under low-N treatments (Fig. 4) when measuring VPM across the rhizosphere samples of both inbreds and hybrids. On the other hand, in the rhizosphere of inbreds, there were four microbial taxa at the phylum level in low-N treatments with a mean VPM > 0.15 (p < 0.05). There was one taxon under low-N and one taxon under high-N treatment with a mean VPM > 0.15 (p < 0.05) in the rhizosphere of hybrids.
The proportion of variance in microbial relative abundance associated with plant genotype (VPM) in the rhizosphere at the phylum level. (a) the magnitude of microbial-plant genotype associations (no of genotype-associated microbial taxa (HT) × VPM), (b) the plant genotype-associated microbial taxa (HT) (ranked based on VPM values across samples) under both low-N (LN) and high-N (HN) conditions (* p < 0.05) and (c) VPM for rhizobiome traits under low-N and high-N treatments for both inbred and hybrid lines. The colors indicate whether VPM are significant (p < 0.05) among either inbred (pink), hybrid (pink), or both (green), while gray represents taxa with p > 0.05. UC: Unclassified taxa
Microbial network topology and complexity
Co-occurrence networks were established to better understand change in interaction between microbial taxa at the genus level across inbred and hybrid rhizospheres in response to N treatment. Microbial networks were constructed for each subgroup of maize inbred lines (classified based on stalk stiffness, plant height, and flowering time) and their hybrids (with Mo17 and B73 inbred lines, H1-H4) under both low- and high-N treatments (Fig. 5 and Supplementary material Fig. S1). The differences between networks were measured by using different parameters including the network size (total number of nodes, N), network connectivity (total number of links, E), modularity (M), and clustering coefficient (the extent to which nodes are clustered, CC).
Changes in microbial network topological properties across plant groups and N treatments. Boxplots illustrate topological properties of microbial networks associated with subgroups of non-stiff stalk (NSS) maize inbred lines and their hybrid rhizospheres under low- and high-N treatments. Network topological properties include (a) no of nodes (N), (b) network connectivity (total number of links, E), (c) modularity (M), and (d) clustering coefficient (the extent to which nodes are clustered, CC). Note: Microbial networks representing weak and nonsignificant interactions with a correlation < 0.3 and p > 0.05 were excluded from the analysis. * p < 0.05 and ** p < 0.01
Microbes in the rhizosphere of non-stiff stalk inbreds had a significantly lower number of nodes (N: 173 and 159) compared to their hybrids (N: 206.5 and 196.5) under both low-N and high-N conditions (p < 0.05). Moreover, stiff stalk inbreds under high-N conditions had significantly fewer nodes than their hybrids under low-N conditions (p < 0.05). We also found that hybrids had a relatively higher number (p = 0.05) of edges (E: 1361.5) compared to inbreds under low-N conditions (E: 811.75).
Among other network parameters, the clustering coefficient was significantly higher for inbreds compared to hybrids under high-N conditions, a trend observed in both stiff and non-stiff stalk maize (Fig. 5 and Figure S1). Additionally, stiff stalk inbreds had a significantly higher clustering coefficient (CC: 0.40) under high-N conditions than hybrids under low-N conditions (CC: 0.29). Finally, different genotype groups (inbred and hybrid) and N treatments had minimal impact on network modularity for both stiff and non-stiff stalk maize.
Discussion
By leveraging large-scale maize field experiments, we measured the root rhizosphere microbial community responses to N treatment and assessed how the responses of these communities vary depending on the host which differed between inbred and their hybrids. The focus of this study was on the rhizosphere because it is a hotspot for diverse microbial activity and biogeochemical cycling (Zhang et al. 2019; Meier et al. 2022; Chai et al. 2023). We found a strong influence of both plant genetic factors and environmental filtering in shaping rhizosphere microbial community structure, variation in microbial relative abundance associated with the host, and network interactions. Our results contribute new insights into how host plant-microbial interactions are shaped across the rhizosphere of diverse maize lines in nutrient-deficient and fertilized soils.
The N input significantly increased the diversity of the rhizosphere microbial communities in inbred but decreased it in hybrid maize. This observation was true for all the different subgroups of inbreds lines (classified based on stalk stiffness, plant height and flowering time) and their hybrids. There is evidence suggesting contrasting influence of N fertilization on soil microbial diversity across varying field conditions (Zhang et al. 2018; Li et al. 2020; Mukhtar et al. 2023) including cropping systems (Kong et al. 2023). Our results support these findings and further suggest that the magnitude of N impact on microbial diversity differs between inbreds and their hybrid rhizospheres in agroecosystems. In this study, N treatment increased the average alpha diversity by 1.2 times in the rhizosphere of inbreds while decreasing alpha diversity by more than twofold in hybrids (Fig. 1). This underscore the significant role of host genotype (Chai et al. 2023) resulting in maize heterosis and overall plant vigor that impacts the composition and dynamics of the microbial community (Picard and Bosco 2005; Brisson et al. 2019; Wagner et al. 2020). As shown previously, hybrids exhibit differences in root traits and exudate production compared to inbred lines, and generally support more auxin and antibiotic producing rhizobacteria and genetically diverse Pseudomonas populations (Picard and Bosco 2005; Picard et al. 2008; Schmidt et al. 2016). Along with differences in genotype between plant groups, N application may also alter the maize root exudation production (Zhu et al. 2016), further influencing the rhizosphere microbial diversity. Consequently, soil microbes in the rhizosphere of hybrid lines under high-N treatment may undergo enhanced environmental filtering (compared to inbred lines), supporting higher relative abundance of a specific microbial taxa, such as Pseudomonas (Fig. 1). Taken together, the rhizosphere of hybrids appears to be exhibit heightened selectivity for microbial taxa compared to inbreds especially in N fertilized soils accentuating the compound impact of host plant genetic factors and edaphic conditions in shaping rhizosphere microbial community structure.
The number of plant genotype-associated microbial taxa (HT) under low-N was significantly higher than under high-N treatments, with inbred rhizospheres supporting a greater number of HT than hybrid rhizospheres. In fact, HT (across the samples) under low-N were almost double (HT = 38) than those observed under high-N (HT = 21), supporting the hypothesis that key microbial taxa selected by the host may be a benefit to plant adaptation under nutrient-limited conditions (Zhang et al. 2019; Meier et al. 2022; Chai et al. 2023). Moreover, the proportion of variance in microbial relative abundance associated with plant genotype (refer as VPM) in this study (0.14 to 0.41) were within the same range as heritability values determined in other studies on rhizosphere communities of maize regardless of different environmental conditions (background variations) and the relative abundance of different microbial groups (Walters et al. 2018; Deng et al. 2021; Meier et al. 2022). In our study, the maize rhizosphere of inbred and hybrid was dominated by Pseudomonas which could be linked to the timing of our sampling, conducted at 8 weeks after planting. For instance, a bloom of Pseudomonas was observed in the maize rhizosphere between weeks 7 and 8 of development (Walters et al. 2018), which may not always occur in similar development stages (i.e., seven weeks after planting) (Wagner et al. 2020). Interestingly, the relative abundance of Pseudomonas, on average, was 1.3 (under low-N) to 2.1 times (under high-N) higher in the rhizosphere of hybrid as compared to inbred lines (Fig. 1). This may explain the fewer heritable taxa in hybrids compared to inbreds under both low- and high-N treatments. Despite the significant differences in the relative abundance of more than half of the microbial groups at the genus level between low-N and high-N, we observed minimal difference in VPM for the rhizosphere microbial communities from hybrids at phylum level across different N treatments. This implies that a restricted subset of microbial groups at lower taxonomic levels are responsive to plant genetic factors underscoring the importance of taxonomic resolution when analyzing microbial heritability.
The rhizosphere microbial networks showed a significantly higher number of nodes in hybrids under low-N compared to inbreds under high-N. This is likely due to the fact that: 1) hybrids may experience intermediate or lower stress under different N levels, which is not surprising given their higher overall vigor and greater adaptability than inbreds (Ruiz et al. 2019), and 2) the high selectivity for specific microbial taxa in the rhizosphere of hybrids (e.g., Pseudomonas) (Fig. 1). On the other hand, the clustering coefficient was higher in the microbial communities in the rhizosphere of inbreds as compared to hybrids only under high-N conditions suggesting changes in the cooperative interactions in the rhizosphere microbial communities in response to changing N availability. Taken together, microbial network topology across inbred and hybrid rhizospheres highlights a fundamental caveat in their functioning which depends on the variation in species response to soil N conditions. These intraspecies differences in maize should be considered when developing effective plant-specific soil inoculants and biofertilizers, especially under nutrient-deficient soils for low-input agriculture.
Conclusions
This study focused on the rhizosphere microbial communities, a region of the soil in agroecosystems that is a hotspot for taxonomical and functional diverse microbes, influenced by both soil conditions and plant root functional characteristics. We showed that the response of the microbial community structure and network topology to N application were based on plant genetic architecture. This may have also been influenced by sampling time with a proliferation of Pseudomonas taxa in the rhizosphere. Therefore, ignoring the influence of plant genotype on soil rhizosphere microbiota may result in either underestimation or overestimation of microbial responses to N availability and thereby agroecosystem functioning. However, it remains to be confirmed whether microbial responses to N levels in the rhizosphere of maize inbreds and hybrids will exhibit similar patterns under different climate and soil conditions, and how these patterns vary across different sampling times.
Data Availability
Sequences generated in this study can be found on NCBI (accession number: PRJNA1092665).
References
Bahram M, Hildebrand F, Forslund SK, Anderson JL, Soudzilovskaia NA, Bodegom PM, Bengtsson-Palme J, Anslan S, Coelho LP, Harend H (2018) Structure and function of the global topsoil microbiome. Nature 560:233
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, Alexander H, Alm EJ, Arumugam M, Asnicar F (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857
Brisson VL, Schmidt JE, Northen TR, Vogel JP, Gaudin ACM (2019) Impacts of maize domestication and breeding on rhizosphere microbial community recruitment from a nutrient depleted agricultural soil. Sci Rep 9:15611
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Huntley J, Fierer N, Owens SM, Betley J, Fraser L, Bauer M (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6:1621–1624
Chai YN, Qi Y, Goren E, Chiniquy D, Sheflin AM, Tringe SG, Prenni JE, Liu P, Schachtman DP (2023) Root-associated bacterial communities and root metabolite composition are linked to nitrogen use efficiency in sorghum. Msystems 9(1):e01190-23
Chen R, Tian M, Wu X, Huang Y (2011) Differential global gene expression changes in response to low nitrogen stress in two maize inbred lines with contrasting low nitrogen tolerance. Genes Genomics 33:491–497. https://doi.org/10.1007/s13258-010-0163-x
Csardi G, Nepusz T (2006) The igraph software package for complex network research. InterJ Complex Syst 1695:1–9
Deng S, Caddell DF, Xu G, Dahlen L, Washington L, Yang J, Coleman-Derr D (2021) Genome wide association study reveals plant loci controlling heritability of the rhizosphere microbiome. ISME J 15:3181–3194
Duff AM, Forrestal P, Ikoyi I, Brennan F (2022) Assessing the long-term impact of urease and nitrification inhibitor use on microbial community composition, diversity and function in grassland soil. Soil Biol Biochem 170:108709
French KE, Tkacz A, Turnbull LA (2017) Conversion of grassland to arable decreases microbial diversity and alters community composition. Appl Soil Ecol 110:43–52
Gaudin ACM, Mcclymont SA, Holmes BM, Lyons E, Raizada MN (2011) Novel temporal, fine-scale and growth variation phenotypes in roots of adult-stage maize (Zea mays L.) in response to low nitrogen stress. Plant Cell Environ 34:2122–2137. https://doi.org/10.1111/j.1365-3040.2011.02409.x
Gu S, Wei Z, Shao Z, Friman V-P, Cao K, Yang T, Kramer J, Wang X, Li M, Mei X (2020) Competition for iron drives phytopathogen control by natural rhizosphere microbiomes. Nat Microbiol 5:1002–1010
Haney CH, Samuel BS, Bush J, Ausubel FM (2015) Associations with rhizosphere bacteria can confer an adaptive advantage to plants. Nat Plants 1:1–9
Hao C, Dungait JAJ, Wei X, Ge T, Kuzyakov Y, Cui Z, Tian J, Zhang F (2022) Maize root exudate composition alters rhizosphere bacterial community to control hotspots of hydrolase activity in response to nitrogen supply. Soil Biol Biochem 170:108717
Hernandez DJ, David AS, Menges ES, Searcy CA, Afkhami ME (2021) Environmental stress destabilizes microbial networks. ISME J 15:1722–1734
Hu L, Robert CAM, Cadot S, Zhang XI, Ye M, Li B, Manzo D, Chervet N, Steinger T, Van Der Heijden MGA (2018) Root exudate metabolites drive plant-soil feedbacks on growth and defense by shaping the rhizosphere microbiota. Nat Commun 9:2738
Jacoby RP, Koprivova A, Kopriva S (2021) Pinpointing secondary metabolites that shape the composition and function of the plant microbiome. J Exp Bot 72:57–69. https://doi.org/10.1093/jxb/eraa424
Kong W, Qiu L, Ishii S, Jia X, Su F, Song Y, Hao M, Shao M, Wei X (2023) Contrasting response of soil microbiomes to long-term fertilization in various highland cropping systems. ISME Commun 3:81
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD (2013) Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microbiol 79:5112–5120
Leptin A, Whitehead D, Orwin KH, McNally SR, Hunt JE, Cameron KC, Lehto NJ (2022) High additions of nitrogen affect plant species-specific differences in the composition of main microbial groups and the uptake of rhizodeposited carbon in a grassland soil. Biol Fertil Soils 58:149–165
Li Y, Tremblay J, Bainard LD, Cade-Menun B, Hamel C (2020) Long-term effects of nitrogen and phosphorus fertilization on soil microbial community structure and function under continuous wheat production. Environ Microbiol 22:1066–1088
Li B-B, Roley SS, Duncan DS, Guo J, Quensen JF, Yu H-Q, Tiedje JM (2021) Long-term excess nitrogen fertilizer increases sensitivity of soil microbial community to seasonal change revealed by ecological netwAork and metagenome analyses. Soil Biol Biochem 160:108349
Ling N, Wang T, Kuzyakov Y (2022) Rhizosphere bacteriome structure and functions. Nat Commun 13:836
Liu J, Li S, Yue S, Tian J, Chen H, Jiang H, Siddique KHM, Zhan A, Fang Q, Yu Q (2021) Soil microbial community and network changes after long-term use of plastic mulch and nitrogen fertilization on semiarid farmland. Geoderma 396:115086
Lopes LD, Wang P, Futrell SL, Schachtman DP (2022) Sugars and jasmonic acid concentration in root exudates affect maize rhizosphere bacterial communities. Appl Environ Microbiol 88:e00971-e1022
Lopes LD, Futrell SL, Bergmeyer E, Hao J, Schachtman DP (2023) Root exudate concentrations of indole-3-acetic acid (IAA) and abscisic acid (ABA) affect maize rhizobacterial communities at specific developmental stages. FEMS Microbiol Ecol 99:fiad019
McPherson MR, Wang P, Marsh EL, Mitchell RB, Schachtman DP (2018) Isolation and analysis of microbial communities in soil, rhizosphere, and roots in perennial grass experiments. J Vis Exp 137:e57932
Meier MA, Xu G, Lopez-Guerrero MG, Li G, Smith C, Sigmon B, Herr JR, Alfano JR, Ge Y, Schnable JC (2022) Association analyses of host genetics, root-colonizing microbes, and plant phenotypes under different nitrogen conditions in maize. Elife 11:e75790
Mukhtar H, Lin C-M, Wunderlich RF, Cheng L-C, Ko M-C, Lin Y-P (2020) Climate and land cover shape the fungal community structure in topsoil. Sci Total Environ 751:141721
Mukhtar H, Wunderlich RF, Muzaffar A, Ansari A, Shipin OV, Cao TN-D, Lin Y-P (2023) Soil microbiome feedback to climate change and options for mitigation. Sci Total Environ 882:163412
O’Brien AM, Ginnan NA, Rebolleda-Gómez M, Wagner MR (2021) Microbial effects on plant phenology and fitness. Am J Bot 108:1824–1837
Oksanen J, Simpson G, Blanchet F, Kindt R, Legendre P, Minchin P, O'Hara R, Solymos P, Stevens M, Szoecs E, Wagner H, Barbour M, Bedward M, Bolker B, Borcard D, Carvalho G, Chirico M, De Caceres M, Durand S, Evangelista H, FitzJohn R, Friendly M, Furneaux B, Hannigan G, Hill M, Lahti L, McGlinn D, Ouellette M, Ribeiro Cunha E, Smith T, Stier A, Ter Braak C, Weedon J (2022) _vegan: Community Ecology Package_. R package version 2.6–4. https://CRAN.R-project.org/package=vegan
Ouyang Y, Norton JM (2020) Short-term nitrogen fertilization affects microbial community composition and nitrogen mineralization functions in an agricultural soil. Appl Environ Microbiol 86:e02278-e2319
Philippot L, Raaijmakers JM, Lemanceau P, Van Der Putten WH (2013) Going back to the roots: the microbial ecology of the rhizosphere. Nat Rev Microbiol 11:789–799
Picard C, Bosco M (2005) Maize heterosis affects the structure and dynamics of indigenous rhizospheric auxins-producing Pseudomonas populations. FEMS Microbiol Ecol 53:349–357
Picard C, Baruffa E, Bosco M (2008) Enrichment and diversity of plant-probiotic microorganisms in the rhizosphere of hybrid maize during four growth cycles. Soil Biol Biochem 40:106–115
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596
Ruiz MB, D’Andrea KE, Otegui ME (2019) Phenotypic plasticity of maize grain yield and related secondary traits: Differences between inbreds and hybrids in response to contrasting water and nitrogen regimes. F Crop Res 239:19–29
Saleem M, Law AD, Sahib MR, Pervaiz ZH, Zhang Q (2018) Impact of root system architecture on rhizosphere and root microbiome. Rhizosphere 6:47–51
Sasse J, Martinoia E, Northen T (2018) Feed Your Friends: Do Plant Exudates Shape the Root Microbiome? Trends Plant Sci 23:25–41. https://doi.org/10.1016/j.tplants.2017.09.003
Schmidt JE, Bowles TM, Gaudin ACM (2016) Using ancient traits to convert soil health into crop yield: impact of selection on maize root and rhizosphere function. Front Plant Sci 7:373
Shi S, Nuccio EE, Shi ZJ, He Z, Zhou J, Firestone MK (2016) The interconnected rhizosphere: high network complexity dominates rhizosphere assemblages. Ecol Lett 19:926–936
Singh P, Kumar K, Jha AK, Yadava P, Pal M, Rakshit S, Singh I (2022) Global gene expression profiling under nitrogen stress identifies key genes involved in nitrogen stress adaptation in maize (Zea mays L.). Sci Rep 12:4211
Szoboszlay M, Lambers J, Chappell J, Kupper JV, Moe LA, McNear DH Jr (2015) Comparison of root system architecture and rhizosphere microbial communities of Balsas teosinte and domesticated corn cultivars. Soil Biol Biochem 80:34–44
van der Bom F, Nunes I, Raymond NS, Hansen V, Bonnichsen L, Magid J, Nybroe O, Jensen LS (2018) Long-term fertilisation form, level and duration affect the diversity, structure and functioning of soil microbial communities in the field. Soil Biol Biochem 122:91–103
Vanous A, Gardner C, Blanco M, Martin-Schwarze A, Lipka AE, Flint-Garcia S, Bohn M, Edwards J, Lübberstedt T (2018) Association mapping of flowering and height traits in germplasm enhancement of maize doubled haploid (GEM-DH) lines. Plant Genome 11:170083
Wagner MR, Roberts JH, Balint-Kurti P, Holland JB (2020) Heterosis of leaf and rhizosphere microbiomes in field-grown maize. New Phytol 228:1055–1069
Wagner MR, Tang C, Salvato F, Clouse KM, Bartlett A, Vintila S, Phillips L, Sermons S, Hoffmann M, Balint-Kurti PJ (2021) Microbe-dependent heterosis in maize. Proc Natl Acad Sci 118:e2021965118
Walters WA, Jin Z, Youngblut N, Wallace JG, Sutter J, Zhang W, González-Peña A, Peiffer J, Koren O, Shi Q (2018) Large-scale replicated field study of maize rhizosphere identifies heritable microbes. Proc Natl Acad Sci 115:7368–7373
Wang P, Lopes LD, Lopez-Guerrero MG, Van Dijk K, Alvarez S, Riethoven J-J, Schachtman DP (2022) Natural variation in root exudation of GABA and DIMBOA impacts the maize root endosphere and rhizosphere microbiomes. J Exp Bot 73:5052–5066
Wang X, Feng J, Ao G, Qin W, Han M, Shen Y, Liu M, Chen Y, Zhu B (2023) Globally nitrogen addition alters soil microbial community structure, but has minor effects on soil microbial diversity and richness. Soil Biol Biochem 179:108982
Yang Y, Chen X, Liu L, Li T, Dou Y, Qiao J, Wang Y, An S, Chang SX (2022) Nitrogen fertilization weakens the linkage between soil carbon and microbial diversity: a global meta-analysis. Glob Chang Biol 28:6446–6461
Zhalnina K, Louie KB, Hao Z, Mansoori N, Da Rocha UN, Shi S, Cho H, Karaoz U, Loqué D, Bowen BP (2018) Dynamic root exudate chemistry and microbial substrate preferences drive patterns in rhizosphere microbial community assembly. Nat Microbiol 3:470–480
Zhang T, Chen HYH, Ruan H (2018) Global negative effects of nitrogen deposition on soil microbes. ISME J 12:1817–1825
Zhang J, Liu Y-X, Zhang N, Hu B, Jin T, Xu H, Qin Y, Yan P, Zhang X, Guo X (2019) NRT1. 1B is associated with root microbiota composition and nitrogen use in field-grown rice. Nat Biotechnol 37:676–684
Zhang M, Zhang X, Zhang L, Zeng L, Liu Y, Wang X, He P, Li S, Liang G, Zhou W (2021) The stronger impact of inorganic nitrogen fertilization on soil bacterial community than organic fertilization in short-term condition. Geoderma 382:114752
Zhu S, Vivanco JM, Manter DK (2016) Nitrogen fertilizer rate affects root exudation, the rhizosphere microbiome and nitrogen-use-efficiency of maize. Appl Soil Ecol 107:324–333
Acknowledgements
Thanks to Lexie Foster for help with sampling and sample preparation and Nathan Wiest for computer assistance. The work was funded by a grant to DPS and JY—USDA-NIFA 2022-67013-36560. The sequencing was done at the UNMC Genomics Core Facility (NIGMS) INBRE—P20GM103427-19. The experimental design was developed by JY and GX. Field sampling was done by Yang and Schachtman Labs. DNA extractions, library preparation, data analysis, manuscript drafting were done by HM, JH, EB and DPS.
Data generated by this study can be found at accession number: PRJNA1092665
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors have no competing financial or non-financial interests that would conflict with the work presented in this manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mukhtar, H., Hao, J., Xu, G. et al. Nitrogen input differentially shapes the rhizosphere microbiome diversity and composition across diverse maize lines. Biol Fertil Soils (2024). https://doi.org/10.1007/s00374-024-01863-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00374-024-01863-4