Abstract
Diversification of agricultural practices, including changes in crop rotation, intercropping or cover cropping, influence the soil microbiome. Here the impact of tillage and crop diversification on the soil microbiome is reported, being one of the few boreal studies. The field experiment consisted of four treatments with four replications all having a short cereal rotation practice namely an oat (Avena sativa) – spring barley (Hordeum vulgare) – wheat (Triticum aestivum) rotation for the past 10 years until spring 2018. During that period two of the treatments were conventionally tilled with moldboard ploughing whereas the other two were no-tillage treatments. From the growing season 2018 until fall 2020 the main crop in all treatments was spring barley. The first conventional tillage treatment was diversified with English ryegrass (Lolium perenne) as an undersown cover crop for the next three growing seasons. The first no-tillage treatment continued with spring barley only. The second conventional tillage and no-tillage treatment had winter rapeseed in rotation in 2019. Bulk soils were sampled in May 2018 before diversification and then in October 2018, 2019, and 2020. The results showed a clear effect of tillage on the beta-diversity of the soil microbiome and an increase in fungal richness. Barley monoculture interrupted with winter rapeseed resulted in a minor change of the fungal and bacterial community composition. Other fungal and bacterial alpha diversity measures did not react to tillage or diversification nor did the gene copy abundances involved in the N cycle. In conclusion tillage had a profound effect on the soil microbiome hindering impact of the diversification.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Diversification of agricultural practices has the goal to increase the resilience of crop systems and improve soil health (Hertel et al. 2021). This can be achieved by diversifying the crop rotation and through intercropping or cover cropping. Cover cropping is especially important to mitigate soil erosion in boreal regions where soil is left bare during the fall and winter season (Lizarazo et al. 2020). Diversifying the plant cover leads to diversification of litter and root exudates both having the possibility to diversify the soil microbiome (Griffiths et al. 2022). A diversified microbiome can lead to improved ecosystem services, like suppression of pathogens (Wang et al. 2022), improved harvest yields (Stefan et al. 2021; Tosi et al. 2022), soil aggregation, water retention, decomposition of plant matter and nutrient cycling (Cappelli et al. 2022) or increased nutrient input through N fixation in the case of legume cultivation (Mbuthia et al. 2015; Ai et al. 2018).
Also improved C sequestration into the soil is linked to a change in the soil microbiome where especially the increase in mass and species diversity of soil fungi seems to be important in this process (Yang et al. 2022). It has been shown that microbial-derived soil organic matter (SOM) is stable (Kallenbach et al. 2016) and for instance soybean (Glycine max) intercropping into sugarcane (Saccharum sp) monocultures was effective in this respect (Chen et al. 2023). The soil microbiome also contributes indirectly to soil C stabilization by enhancing soil aggregate formation (Oades 1993; Rillig et al. 2015). Thus, it is important to document the impacts of different diversification trials on the soil microbiome. As stated by Khmelevtsova et al. (2022) the number of soil microbial taxa is huge, but the “ideal” composition of the community is unknown.
Nitrogen is a critical and essential nutrient for plants ensuring crop growth (Hirsch and Mauchline 2015). Microbes conduct the whole nitrogen cycle including atmospheric N2 fixation, nitrification and finally denitrification where nitrate is reduced to N2 again to complete the cycle, or partially reduced to generate nitrous oxide. Most often, however, mineral N fertilizers have replaced the microbial processes to increase crop yields and to replenish the N removed from the fields at harvest. Excessive use of mineral fertilizers can lead to negative phenomena such as increased N leaching leading to pollution of water bodies. Diversification of crop rotations can be based on the exploitation of N2 fixing symbiotic rhizobia in leguminous plants or by promoting microbial mineralization to release ammonia or nitrate from organic material such as from green manure. The use of crops in diversification that have symbiotic arbuscular mycorrhizal fungi (AMF) can promote the transfer of N from soil organic matter (SOM) (Leigh et al. 2009), nitrate, and ammonia (Fellbaum et al. 2012). Diversification trials having conventional mineral N fertilization produced a positive impact on the soil microbiome by increasing richness, diversity, and promoting a more elaborated network structure (Liu et al. 2023). Changes in soil microbiomes are most likely reflected on the efficiency of N fertilization. Therefore, it is relevant to link the impact of agricultural practices also to microbial processes related to the N cycle.
Agricultural practices performed before and during agricultural diversification can have an impact on the on the reaction of the soil microbiome upon diversification measures. One practice commonly used in agriculture is soil tillage which is to be compared to no-till practices. A meta-analysis has shown that no-tillage had different effects on bacterial and fungal community diversity; increasing soil bacterial diversity, with no significant change to fungal diversity when compared to tilled systems (Li et al. 2020). Therefore, diversification of tilled and no-tilled agricultural systems can have a different outcome on the soil microbiome (Kim et al. 2020) but information concerning the boreal agricultural environment does not exist.
Here we report on a boreal field study where after 10 years of short cereal rotation (oat/barley/wheat) with and without tillage has been diversified over three years by cover cropping or introduction of new crops in the rotation sequence. The soil microbiome, fungi and bacteria, has been analyzed before diversification and thereafter for the following three falls to document a) the impact of tilling and b) the impact of diversification. We hypothesize that 1) tilling has an impact on the soil microbiome, 2) it is stronger than the impact of diversification and 3) diversification changes the soil microbiome and increases bacterial and fungal alpha-diversity. We used amplicon sequencing to study the soil microbiome (fungi and bacteria) and qPCR to measure the abundance of amoA, narG and nirK genes and the potential nitrification to document their activity in the N cycle under the treatments.
Material and methods
Site treatments and sampling
The Kotkanoja long-term field trial locates in Jokioinen southern Finland (N 60° 82’, E 23° 51’). According to the World Reference Base for Soil Resources (WRB) the soil type is Protovertic Luvisol. Sand, silt, and clay contents of the soil were 4.7%, 30.5% and 64.8%, respectively.
The experimental field consists of four treatments (Table 1) each having four study plots of a size of 33 m × 33 m, respectively. Two of the treatments (A and C) were conventionally tilled (CT; moldboard ploughing to a depth of 20 cm) whereas the other two treatments (B and D) were no-till (NT) treatments. All four treatments had the same short cereal monoculture rotation practice namely an oat (Avena sativa) – spring barley (Hordeum vulgare) – wheat (Triticum aestivum) rotation for the preceding 10 years prior to this study, 2008—2018. Thereafter, from the growing season 2018 until fall 2020, the treatments, except B, were diversified, and all started with spring barley as the main crop. Treatment A continued with the CT treatment having spring barley and English ryegrass (Lolium perenne) as an under sown cover (or catch) crop for the next three growing seasons. Treatment B continued as NT with spring barley only, thus being a cereal monoculture. Treatment C continued as CT with a spring barley—winter rapeseed (Brassica napus)—spring barley rotation and treatment D continued as NT with the same rotation as treatment C but received a one-time moldboard ploughing in 2018 to relieve the problems related to phosphorus accumulation in the top layer of the soil. The farming practices, including fertilization, are presented in Table 1.
Soil sampling was conducted in May 2018 before diversification and spring barley sowing and in each October 2018–2020 after harvesting. The May 2018 sampling enabling a comparison of 10 years CT vs NT practice on the measured variables whereas the October sampling represented diversification. The winter rapeseed was sown in August 2018 into tilled soil and harvested in September 2019. According to the instructions of the “Diverfarming” project each study plot bulk soil sample was composed of a minimum of nine subsamples taken approximately from a 1 m2 area and between 5 to 10 cm depth except for the chemical analysis where the 0–10 cm depth was used. In addition, on each sampling occasion, one of the study plots of each treatment was randomly chosen and sampled twice. This bulk soil of pseudo-replication was sampled from the whole study plot area. This was performed due to the “Diverfarming” project handbook guidance to have the treatment replication of n = 5. The samples were cleaned from possible litter, roots, and stones, sieved (2 mm) and mixed well before being stored in -20 °C for DNA extraction or air drying for soil analysis.
Soil analysis
The C (TOC) and N contents were measured with a LECO TruMac CN Carbon/Nitrogen Determinator (LECO corporation, USA) as described by the manufacturer and the results were used to calculate the C/N ratio. Sum of base cations (Bs), cation exchange capacity (CEC), electrical conductivity (EC) and pH were calculated using a standard soil fertility analysis (SFS EN ISO/IEC 17025:2005 FINAS T096) provided by a commercial service (Eurofins agro testing Finland ltd, Finland) according to accredited standard methodology. The soil dry mass (dm) was analyzed with thermogravimetric analyzer (TGA701, LECO corporation, USA).
Potential nitrification (PN) was measured using a microscale method as described by Hoffmann et al. (2007; ISO 15685 (2012)). Nitrite production was measured by reading the absorbance at 540 nm with a CLARIOstar Plus plate reader (BMG LABTECH, Germany). According to test runs the optimum concentration of sodium chlorate in the analysis was 15 mmol L−1.
DNA extraction
At each of the four sampling times soil DNA was extracted according to DNeasy PowerSoil Kit (Qiagen, Germany) with some modifications to the standard protocol. Modifications for the experienced user protocol steps were: (1) 0.5 g soil sample and initial mixing in horizontally placed tubes for 30 min. (5) the tubes were centrifuged for 2 min at 16,100 × g. (7 and 10) the tubes were incubated for 15 min at 4 °C. (8 and 11) the tubes were centrifuged for 2 min at 16,100 × g. (16) the tubes we centrifuged for 5 min at 16,100 × g. (20) the tubes were centrifuged for 3 min at 16,100 × g. The quality of extracted DNA was analyzed by calculating A260/A280 and A260/A230 ratios with the NanoDrop One (ThermoFrisher Scientific, USA). Minimum quality values of DNA accepted for downstream processes were 1.8 and 1.9, respectively. The concentration of DNA was measured with a Qubit (ThermoFisher Scientific, USA) fluorometer.
Quantitative PCR (qPCR)
qPCR was used to quantify the gene abundancy of ammonia mono-oxygenase (bacterial amoA), nitrate reductase (narG), and nitrite reductase (nirK) using primers described in Rotthauwe et al. (1997), Bru et al. (2007), and Henry et al. (2004), respectively. Thermal cycling conditions were as reported in the original papers except for the following modifications. Initial denaturation step was 95 °C for 2 min for all genes. The thermal conditions for a total of 40 cycles in amoA were as follows: denaturation 95 °C for 15 s, annealing at 58 °C for 30 s and extension 72 °C for 45 s. Fluorescence for analysis was collected at the end of each extension step. Genes were amplified in Rotor-Gene 6000 (Corbett Research ltd., Australia) qPCR machine in 20 µL reactions containing 10 ng of total DNA (5 µL), 1 × QuantiNova SYBR Green (Qiagen, Germany), and 0.375 µM of forward and reverse primers. Standards for absolute quantification (approx. 107 to 102 copies/reaction) were cloned from PCR products into topGEM®-T Easy vector system as described by the manufacturer (Promega Corp., USA). Cloned plasmids were purified with QIAprep Spin Miniprep Kit (Qiagen, Germany) as described by the manufacturer. One empty cloning-vector was purified to be used as spiked exogenous DNA for inhibition tests for each soil sample DNA (10 ng). Inhibition tests were performed similarly as qPCR-reactions for amoA described above with following modifications: forward primer (SP6; ATTTAGGTGACACTATAG), reverse primer (T7; TAATACGACTCACTATAGGG), amplification template 104 copies per reaction of empty cloning vector, annealing at 50 °C for 30 s and extension 72 °C for 60 s. The control reaction in an inhibition test was a spiked sample without soil DNA. All qPCR-reactions were run at least in duplicate. The equations for the qPCR standard curves and reactions efficiencies were y = -3.54x + 35.881 (efficiency = 0.92; R2 = 0.99923), y = -3.898x + 40.732 (efficiency = 0.81; R2 0.99964), and y = -3.418x + 33.5 (efficiency = 0.91; R2 = 0.99837), respectively, for amoA, narG, and nirK genes. All standard curves showed a linear response from approximately 102 to 107 copies per reaction. No inhibition (not shown) was observed in samples spiked with empty cloning vector and subsequent amplification with SP6 and T7 primers compared control reactions without soil DNA.
PCR and sequencing
The bacterial community was determined through the next-generation-sequencing of bacterial 16S hypervariable regions using the Ion Torrent™ Personal Genome Machine™ (PGM, London, UK) System. Bacterial 16S regions were amplified using an Ion 16S™ Metagenomics Kit (Thermo Fisher Scientific, Waltham, MA, USA) with two different degenerate primer sets to amplify regions V2–4–8 and V3–6, V7–9. The amplified 16S amplicons were then processed using an Ion Xpress™ Plus Fragment Library Kit in combination with an Ion Xpress™ Barcode Adapter 1–96 Kit (Thermo Fisher Scientific, Waltham, MA, USA).
All the purification processes between incubation and the amplification reactions of library preparation were processed using DynaMag™-2 magnetic racks (Thermo Fisher Scientific, Waltham, MA, USA) and an AMPure XP Purification Kit (Beckman Coulter, Brea, CA, USA). Library preparation and barcoding were followed by the determination of the size and concentration of the final libraries using an Agilent 2100 Bioanalyzer system and the Agilent High Sensitivity DNA kit (Agilent, Santa Clara, CA). Sequencing templates were prepared using an Ion One Touch 2 System and an Ion PGM™ Hi-Q™ View OT2 Kit (Thermo Fisher Scientific, Waltham, MA, USA). The sequencing reaction was performed using Ion Torrent PGM (Thermo Fisher Scientific, Waltham, MA, USA) with an Ion PGMTM Hi-QTM View Sequencing Kit (Thermo Fisher Scientific, Waltham, MA, USA).
Sequencing libraries for fungal ITS1 were prepared using a modified version of the method proposed by Smith and Peay (2014). Sequencing libraries were generated by PCR amplification using locus specific primers (ITS1f-ITS2) tailed with the Illumina adapters. The reverse primers were barcoded to allow multiplexing using the 12-base Golay barcodes (Caporaso et al. 2012). PCR amplification was performed in a final volume of 30 µl containing 3 µl of buffer 10X, 0.7 µl of each primer (10 mM), 0.9 µl of 50 mM MgSO4, 0.6 µl of 10 mM dNTP and 0.12 µl of Invitrogen Platinum Taq DNA polymerase High Fidelity (Cat N° 11,304–011). The thermal cycling conditions used for the PCR were: initial denaturation at 95 °C for 3 min followed by 35 cycles of denaturation at 95 °C for 45 s, annealing at 50 °C for 1 min and extension at 72 °C for 1 min terminating with a final extension at 72 °C for 10 min. Before sequencing, amplicon libraries were mixed with 10% PhiX control library to increase the sequences diversity. Fungal libraries were sequenced on the Illumina MiSeq platform generating 300 bp paired end reads.
Sequencing and data processing
For bacterial raw sequences, barcodes, adaptors, and primers were trimmed by BaseCaller application. First, the raw sequences were denoised with ACACIA (Bragg et al. 2012), and imported to Quantitative Insights into Microbial Ecology 2 (QIIME2) v2019.1.0 (Bolyen et al. 2019). Then, imported sequences were denoised using the DADA2 algorithm with the sequences truncated with a Q > 25 (Callahan et al. 2016) on average. The sequences were clustering into Operational Taxonomic Units (OTUs) based on 97% similarity with the SILVA reference database using VSEARCH (Rognes et al. 2016). Low-confident OTUs were removed, and the sequences were classified using “classify-consensus-vsearch” command with SILVA 132 database. The average number of bacterial 16S rRNA raw reads in the sample library (n = 80) was 56 004 (SD = 31 556). After trimming and assembly filtering, removal of singletons (n = 1) and discarding of OTUs without bacterial matches there were a total of 3 499 156 sequences left (43 739 in average, SD = 23 993) with 8542 bacterial OTUs for analyses.
Raw fungal reads were filtered using Trimmomatic (Bolger et al. 2014) to remove adapters and low-quality reads using a quality cutoff of 20 in a 20 bp sliding windows. Paired end reads were merged using Pear (Zhang et al. 2014). Sequences were generated using the QIIME2 DADA2 plugin (Callahan et al. 2016), where sequences were clustered into OTUs based on a 97% sequence similarity using QIIME2 vsearch cluster-features-open-reference tool (Rognes et al. 2016). Taxonomy was assigned using the UNITE database version 8.2, dynamic (Abarenkov et al. 2020). The average amount of fungal ITS1 raw reads in the sample library (n = 74) was 122 896 (SD = 51 312). After the trimming and assembly filtering, removal of singletons (n = 1) and discarding of OTUs without fungal matches there were a total of 6 220 525 sequences (84 061 in average, SD = 39 407) left with 1797 fungal OTUs for analyses. With FUNGuild the major functional groups, symbiotic (SYM), saprotrophic (SAP) and pathogenic (PAT) fungi were obtained from the OTU data using the FUNGuild.py script (Nguyen et al. 2016) and investigated further.
Raw sequence data is deposited to the European Nucleotide Archive (ENA) database under the project name “CS12 Diverfarming project” with the accession number PRJEB60421 for 16S rRNA and ITS1 data.
Community and statistical analyses
All results except OTUs are reported per soil dry mass (dm) either in g or kg. The R-studio version 1.4.1106 (RStudio Team 2022) including the R version 4.2.1 (R Core Team 2022) were used for visualization and analyses of OTU data. Alpha-diversity measures as three Hill’s numbers; Q0 for species richness, Q1 the exponential of Shannon’s entropy index, and Q2 for the inverse of Simpson’s concentration index, were obtained from the bacterial and fungal OTU data with package ‘hillR’ (Chao et al. 2014). For beta-diversity analyses both bacterial and fungal OTU data were normalized using the geometric mean of pairwise ratios (GMPR) method (Chen et al. 2018) to diminish the effect of variable sample library sizes and to increase the resolution for zero-inflated data. Beta-diversity was investigated by using nonmetric multidimensional scaling (NMDS) with stable solution from random starts, axis scaling and species scores with function metaMDS and fitted environmental variables (Table S1) with function envfit from ‘vegan’ package 2.6–4 (Oksanen et al. 2022) by using Bray–Curtis dissimilarity index. A permutational multivariate analysis of variance (PERMANOVA) using Bray–Curtis distance matrices with function adonis2 from ‘vegan’ package was executed to test the effect of treatment and sampling time and their interaction on the fungal and bacterial OTU composition.
To identify OTUs that were significantly differentially abundant between different treatments, we conducted a differential abundance analysis (DAA) using the ‘ancombc’ function of the ANCOM-BC R package (Lin and Peddada 2020). The analysis was conducted at the OTU level using the default settings except for the following: zero_cut = 0.75, lib_cut = 1000, struc_zero = TRUE, conserve = TRUE. Results are investigated as paired comparisons between treatments and treatment groupings. Significance is determined at the P < 0.05 level for fungi and at the P < 0.0001 level for bacteria. The tillage effect was tested by combining the May 2018 treatment A and C data which was then compared to the combined treatment B and D OTU data. The impact of winter rapeseed was evaluated by comparing the October 2019 data between treatment grouping A and B against the treatment grouping C and D. Also, within treatment October 2018 vs October 2020 comparisons were made to identify individual proliferation of each diversification. The proportions (%) of shared and unique OTUs between treatments from different data sets were visualized with Venn diagrammes by applying script ps_venn.R (Russel 2021) from the phyloseq object and package eulerr (Micallef and Rodgers 2014; Wilkinson 2012).
Generalized linear mixed models (GLMM) were used to analyze the microbial diversity (Hill’s numbers), fungal guilds, gene abundance (qPCR) and potential nitrification data by utilizing the GLIMMIX procedure of the SAS Enterprise Guide 7.15 (SAS Institute Inc., Cary, NC, USA). Also, the physicochemical data of Table S1 was analyzed with the same model structure. The categorical variables of treatment (A-D) and sampling time (May 2018, Oct 2018, Oct 2019, and Oct 2020), and their interaction was used as fixed effects. Due to skewed distributions, the assumption of gamma (with a log link) and lognormal (with an identity link) distributions were used. Each row of the field design was used as a block, and thus both managements, CT and NT, had two plots per row. The effect of management (CT and NT) was not included in the model because the treatments (A-D) did not occur in both management plots, but the comparison of managements was made using the estimated means of the treatments in CT and NT plots (AC vs. BD).
All models were estimated by using the residual pseudo-likelihood (REPL) estimation method, for the models with the assumptions of gamma distribution, and the residual maximum likelihood (REML) estimation method for others. Degrees of freedom were calculated using the Kenward–Roger method. Pairwise comparisons of the estimated means were analyzed using the Tukey’s method with a significance level of α = 0.05. The comparison of managements was coded separately by using the method of Westfall (Westfall 1997). Finally, the normality of the model residuals was confirmed through using several residual plots.
Results
Soil physicochemical data
Table S1 presents the analyzed physicochemical soil data. From the measured variables the only which was significantly (p = 0.034) different due to tillage was the electrical conductivity (EC) which showed lower estimated means for the CT (A, C) than for the NT (B, D) treatments (statistical result data not shown) in May 2018. The modelling showed casual significant (p < 0.05) differences between treatments for pH, EC, C/N and Bs at different sampling times but no interpretable trend could be identified due to diversification (data not shown).
Effect of tillage and crop rotation on microbial community
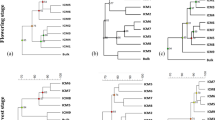
NMDS (Fig. 1) and PERMANOVA (Table 2) revealed that tillage practices affected the fungal community composition after 10 years of short cereal rotation. Samplings performed in May 2018 showed differences in fungal community between CT (A and C treatments) and NT (B and D treatments). This difference between CT and NT was persistent over the whole study period including the diversification trials. Within the CT treatment the different diversification treatments between treatments A and C induced a difference in the fungal community in October 2019 but this reversed by October 2020. Within the NT treatments the fungal community was similar in May 2018 and October 2020. In October 2018 and 2019 the community differed reflecting the one-time moldboard ploughing and the diversification performed to NT treatment D. PERMANOVA indicated significant differences between the treatments having the different tillage and diversification treatments and sampling times but no interaction between these two factors (Table 2). The treatment explained 16% and the sampling time 9.5% of the variation. Like the fungi, the bacterial community also reacted to the tillage (Fig. 2). In the NMDS analysis the CT grouped separately from the NT in May 2018. The one-time moldboard ploughing in the D treatment in summer 2018 induced a change in bacterial community composition which led to the D treatment community resembling more the A and C treatment CT communities in the NMDS analysis in October 2018. From October 2019 onwards the D treatment bacterial community overlapped with the C treatment community. Still the general separation between CT and NT treatments stayed but was not as strong as in May 2018. PERMANOVA (Table 2) indicated significant differences between the treatments having the different tillage and diversification treatments and sampling times but no interaction between these two factors. The treatment explained 9.9% and the sampling time 19% of the variation. The soil physico-chemical data of Table S1 fitting the fungal and bacterial community composition were different for the two domains and changed with the sampling occasion (Figs. 1 and 2). The diversification effect of winter rapeseed on the fungal community is correlated with an increase in soil electric conductivity (Fig. 1c). The tillage effect on the bacterial community is correlated with pH and Bs (Fig. 2a).
Non-metric multidimensional scaling (NMDS) plot showing the composition the fungal ITS derived OTUs obtained from the different treatments (A, B, C, D) at different sampling times a) May 2018, b) October 2018, c) October 2019 and d) October 2020. Ellipses show the confidence intervals of the mean according to separations by treatments. Arrows show environmental variables from Table S1 that fit the x–y data of the NMDS significantly (P < 0.05). TOC, total organic C; Nt, total N; Bs, base cations; pH, soil pH; EC, electric conductivity
Non-metric multidimensional scaling (NMDS) plot showing the composition of the bacterial 16S rRNA derived OTUs obtained from the different treatments (A, B, C, D) at different sampling times a) May 2018, b) October 2018, c) October 2019 and d) October 2020. Ellipses show the confidence intervals of the mean according to separations by treatments. Arrows show environmental variables from Table S1 that fit the x–y data of the NMDS significantly (P < 0.05). TOC, total organic C; Nt, total N; Bs, base cations; pH, soil pH; EC, electric conductivity; CN; ratio of C and N
We identified fungal and bacterial OTUs which were differently abundant between two groups. The first group was formed from the May 2018 data to compare CT (A, C treatments) and NT (B, D treatments) and the second group was formed of the October 2019 data combining AB and CD treatments to elucidate the winter rapeseed effect (Table S2). The DAA on the fungal Phylum level for the CT and NT treatments highlighted 7 OTUs (3 Ascomycota, 2 unknown, 1 Basidiomycota and 1 Mortierellomycota) and 4 OTUs (3 Ascomycota and 1 Glomeromycota) which were differentially abundant in the CT and NT treatments, respectively. On the genus level, many of these OTUs remained unidentified but from those which could be identified the CT treatment had differentially abundant OTUs that were representatives of one Mortierella (SAP), and one Geminibasidium (SAP). Identified differentially abundant OTUs for the NT treatment at the genus level included only 1 Peziza (SAP-SYM). Additionally, there was 1 SYM OTU belonging to the Glomeraceae Family. The AB group in October 2019 was characterized by 4 OTUs including 2 Ascomycota, 1 Basidiomycota and 1 unknown Phylum. None of them were identified to the genus level or to any functional guild. The CD grouping was characterized by 10 fungal OTUs including on the Phylum level 4 Ascomycota, 3 Basidiomycota, and 1 of each Glomeromycota (SYM), Olpidiomycota and Blastocladiomycota. To the genus level we could identify Pseudoheletium, Olphidium (PAT), Naganishia (PAT) and Amaurodon (SYM). 1 OTU belonging to the Serendipitaceae Family belonged to the fungal guild of SYM.
For bacteria (Table S3) the observed differentially abundant OTUs were identified on the Phylum level. 3 (1 Acidobacteria, 1 Actinobacteria and 1 Bacteroidetes) and 9 OTUs (4 Proteobacteria, 2 Chloroflexi, 1 Acidobacteria, 1 Actinobacteria and 1 Rokubacteria (NC-10)) characterized the tillage (CT) versus no-till (NT) practice in the May 2018 soils, respectively. Nocardioides and Flavisolibacter were the OTUs identified to the genus level in the CT group. In the NT group Candidatus Genus Koribacter, Aquicella and Geobacter were the ones identified to the genus level. There were no differentially abundant bacterial OTUs characterizing the AB vs CD grouping in October 2019.
The Venn analysis of the May 2018 data showed that the CT and NT treatments shared 40% of their fungal OTUs but both groups had a large percentage, ca 30% of unique OTUs only found in the CT or NT treatments, respectively (Fig. 3). A similar partitioning of shared and non-shared OTUs for the CT and NT treatment was also seen for the bacterial OTUs (Fig. 4). The Venn analysis performed to the individual diversification treatments was elucidated by comparing the October samplings picturing the succession within the treatments over three seasons. This showed an increase in the non-shared fungal OTUs following diversification. Treatment A (barley/ryegrass diversification) and B (barley monoculture continued under NT) had the highest values of non-shared fungal OTUs in Oct 2020 and treatment C (barley winter rapeseed barley rotation) and D (barley winter rapeseed barley rotation with one-time tillage in summer 2018) in Oct 2019 (Fig. 3). The bacterial Venn analysis (Fig. 4) also showed an increasing trend of non-shared OTUs being at the beginning in October 2018 between 8–15% and at the end in October 2020 between 31–38% for the treatments A – D. The Venn analysis of the October 2019 data combining AB and CD treatments to elucidate the winter rapeseed effect showed 45 and 46% of shared fungal and bacterial OTUs, respectively whereas the winter rapeseed diversification had 33 and 28% unique fungal and bacterial OTUs, respectively (Figs. 3 and 4).
Effect of tillage and diversification on microbial diversity
According to the GLMMs (Table 3; Tables S4 and S5; Table S6 for CT vs NT results) the 10-year tillage managements did not influence the fungal or bacterial alpha-diversity measures HillQ0, HillQ1 and HillQ2 (Figs. S1 and S2) except for the ITS HillQ0 measure (Table S6). The diversification treatments as the main factor had no significant effect on the fungal and bacterial diversity measures whereas time was always significant showing an in- or decrease of the measures at the different sampling times. Also, the treatment x time interaction was significant for all ITS HillQs and for instance in October 2018 treatment D was different from treatment A and treatment B for HillQ0 (Table S4; Fig. S1). HillQ1 and Q2 diversity measures showed different behavior with time but mostly the interaction term was significant due to different CT or NT treatment result differences at different sampling times. These are not under research since the tillage effect was elucidated from the May 2018 data.
Effect of tillage and diversification on fungal functionality
The three functional fungal guilds, saprotrophs (SAP), symbiotrophs (SYM) and pathotrophs (PAT) constituted 3%, 2% and 0.1% of all reads, respectively. GLMM identified that the relative proportion of SYM (consisting of 84% of arbuscular mycorrhizal fungi, AMF, reads), SAP and PAT fungi did not react to tillage (Table S6) and the proportion of SYM and SAP fungi did not react to the diversification treatments, but due to interaction with time this was investigated more closely (Tables 3 and S5). The proportion of SYM fungi increased over time with the highest modelled values for the treatments having winter rapeseed in the rotation and treatment C being different from treatment B in October 2019 and 2020 (p < 0.08). The proportion of SAP fungi did not react to time but had significant time x treatment interactions and the D treatment differed from the A (p < 0.06) and B (p < 0.05) treatment in October 2018. The proportion of PAT fungi reacted significantly to the diversification being higher in the treatments C and D in October 2019 when these plots were under winter rapeseed (Tables 3 and S5). The modelled values decreased in October 2020, but treatment D was still significantly different (p < 0.05) from treatment B like in October 2018.
Effect of tillage and diversification on the N cycle
The qPCR abuncances of amoA (involved in ammonia oxidation), narG (involved in denitrification), and nirK (involved in denitrification) did not react to the 10 years of tillage treatments (Table S6). Also, the diversification treatment had no influence on the results (Tables 3 and S5). However, differences were found between sampling times and for gene levels within sampling times and for nirK a nearly significant (p = 0.072) diversification effect was noted. The treatments C and D having the winter rapeseed diversification differed from the A and B treatments in October 2019 and D also in 2020 which was also seen in the significant time and treatment and time interaction (Table 3). The winter rapeseed diversification treatments (C and D) showed higher nirK qPCR abundances in October 2020 when all were cultivated with barley but lower values in October 2019 when the plots were cultivated with winter rapeseed (Table S5). The D treatment had higher amoA values in October 2018 and 2019 but decreased towards the mean of the other treatments in October 2020 (Table S5).
Tillage had no effect on the potential nitrification (PN) rate whereas (Table S6) diversification with winter rapeseed increased the rate significantly (Tables 3 and S5). In treatment D the rate increased over the three sampling times between October 2018 and 2020 irrespective of the one-time tillage performed and in treatment C the rate increased between October 2019 and 2020. Also, the diversification with ryegrass (treatment A) increased the potential nitrification with time from October 2019 onwards. With the cereal monoculture, (treatment B), the potential nitrification first increased until October 2019 and then decreased. Treatment and time interaction was significant and in October 2019 and 2020 the D treatment differed (in 2019 D vs B p < 0.09; the rest p < 0.05) from all the others.
Discussion
The NMDS performed to the fungal OTU data shows that the tillage treatment is stronger than the diversification treatment (Fig. 1). This becomes apparent within the CT treatments where the A treatment was diversified with barley together with a cover crop, ryegrass, for three years and the C treatment with a spring barley—winter rapeseed—spring barley rotation thus interrupting in both cases the short rotation based on cereals carried out for the past 10 years. In the October 2019 sampling the A and C treatments had different fungal communities. This was probably induced by the winter rapeseed of the rotation of the C treatment, since when both were under barley (Oct 2018 and 2020) they clustered together even though one of them, the A treatment, had the ryegrass diversification treatment. Still the CT treatments were separated from the NT treatments, and the diversification could not overcome this. The argument, that tillage induces a more distinct change to the fungal community than diversification is also undermined with the NMDS data from the NT treatment where the D treatment, which had a single moldboard ploughing in summer 2018, separated from the B treatment in October 2018 when both were under barley (Fig. 1). This difference remained in October 2019 when the D treatment was under winter rapeseed diversification but merged in October 2020. Thus, within both CT and NT treatments the winter rapeseed diversification induced a community change. This diversification treatment did not overcome the tillage treatment shown by the fact that the C and D treatments having the same rotation did never cluster together. Therefore, the first hypothesis (Hypothesis 1) is supported i.e., that tillage has a profound effect on the soil microbiome as also the second one (Hypothesis 2) that tillage has a stronger impact on the soil microbiome than our diversification since this trend is also seen with the bacterial OTU data (Fig. 2). Also, the Venn analysis (Figs. 3 and 4) underlined that the tillage had a profound effect on the fungal and bacterial community since only 40% of the OTUs were shared between the till and no-till treatments, respectively.
As has previously been shown, conventional tillage changes the microbial community composition (Sharma-Poudyal et al. 2017; Rincon-Florez et al. 2020; Wipf et al. 2021; Hartmann and Six 2023; Khmelevtsova et al. 2022; Mackay et al. 2023). The differences are mainly attributed to the fact that tillage introduces fresh organic matter into the deeper soil which impacts the community (Naylor et al. 2022). In the case of Fungi and Actinobacteria also the tillage induced hyphal dis-rupture affects the community composition. But also, other tillage induced soil impacts are attributed to affect the microbial community composition like disruption of soil aggregates, compactness of soil, and reduction in spaces between soil particles, which lead to alteration in movements of water and gas into the soil, and thus, eventually affect the soil as a habitat for living organisms (Mangalassery et al. 2015). As such our 10 years lasting experiment comparing CT and NT effects on the microbiome under cereal crop rotation confirms existing knowledge (Gupta et al. 2022). The reaction of the soil microbiome to tillage seems to be quite sensitive since even single tillage events have been reported to affect the soil bacterial community (Kraut-Cohen et al. 2020). This was also seen in our study.
We accept with caution Hypothesis 3—diversification changes the soil microbiome and increases bacterial and fungal alpha-diversity. None of the fungal and bacterial alpha-diversity measures reacted to the model main factors tillage or diversification except that in May 2018 the 10-year tillage had significantly increased ITS HillQ0 values (Table S6). But a time x treatment interaction, being related only to the diversification treatment at different times, exists for the fungal species richness (HillQ0) but is not very interpretive showing an effect of winter rapeseed cultivation (only treatment D) being higher in October 2018. As treatment C did not show the same response and the timeframe was not 2019 where cultivation with winter rapeseed could theoretically have an effect, we do not consider this a diversification phenomenon. But the winter rapeseed diversification shows changes in the fungal and bacterial community composition which leads to accepting the hypothesis. One can argue if the microbiome change in October 2019 of the D treatment is due to the one-time tillage performed in 2018 or due to the diversification. A one-time tillage effect on the soil microbiome has been reported to last for one (Liu et al. 2016) or one to three years following tillage (Wortmann et al. 2008). This effect is seen in our NMDS analysis in October 2018. But as also the C treatment microbiome reacted to the same diversification; we argue that the changes in 2019 are winter rapeseed induced. Wooliver et al. (2022) discusses the role of microbial diversity and community composition in relation to diversification and concludes that the effect is more to be seen on the community composition as also our results show but the intensity of changes seems to be connected to the site of diversification performance (Castellano-Hinojosa et al. 2023). Venn analysis of the fungal and bacterial OTUs gives the impression that diversification increases the percentage of non-shared OTUs. The fungal OTUs steadily increased with the cover crop diversification, and it increased with the crop rotation treatments so that it was highest in October 2019 when winter rapeseed interrupted the cereal monoculture especially so under treatment C. But as treatment B had also a steady increase of non-shared OTUs over the three seasons and as no diversification was performed to it, it points to the conclusion that other factors than diversification control the partitioning of shared and non-shared fungal OTUs, and the bacterial Venn analysis shows the same trends. Similarly, the Venn analysis performed on the October 2019 data when AB treatment grouping was compared to the CD grouping elucidated the winter rapeseed effect. Both fungi and bacteria shared the same amount of OTUs between the two groups being around 45% (Figs. 3 and 4). The winter rapeseed treatments (CD) had a little higher unique fungal OTU proportion being 33%, whereas the bacterial portion was 28% leaving respectively 23 and 26% unique OTUs for the AB group which is more influenced by barley.
The data obtained from differential abundance analysis did not provide conclusive results. We could not identify a clear picture when each treatment was compared using the October 2018 vs October 2020 sampling data. The idea was to see the proliferation of the soil microbiome due to diversification. But as treatment B, like the other treatments, had > 30 and > 50 differently abundant bacterial OTUs characterizing the October 2018 and 2020 samplings (data not shown) even though no diversification was performed, we are unsure what the identified OTUs are really standing for except for yearly variation. As tillage influenced the soil microbiome, we decided to have a closer look at differential abundance analysis results comparing the CT and NT treatments of the May 2018 OTU data. The saprotrophic soil fungi Geminibasidium which is reported as heat-resistant and xerotolerant (Nguyen et al. 2013) was more abundant in CT. Geminibasidium is suggested to be one of the keystone saprotrophic taxa under continuous cropping by playing an essential role in increasing the content of available soil nutrients (Wang et al. 2021). In turn, two separate taxa of the Order Pezizales (Genus Peziza and Family Pyrenomataceae) were more abundant in NT and are to be found as saprotrophs in soil, dung, or rotten wood, and some are mycorrhizal or hypogeous, and others associated with fire sites (Cannon and Kirk 2007). Pezizales could be a good indicator for NT since they declined after a single tillage event in a long-term NT farming system (Fang et al. 2022). In addition, also AMF taxa of the Glomeraceae Family were more abundant in NT. The result is in parallel to the previous finding that higher contribution of Glomeraceae in NT systems is suggested to be related partially to the lack of disruption of the hyphal network and the composition of the soil propagules in this system (Schalamuk and Cabello 2010).
Due to winter rapeseed influencing the fungal community we also investigated differentially abundant OTUs using the October 2019 AB treatment vs CD treatment grouping (Table S2). Diversification by rapeseed did show an increase the AMF belonging to the Paraglomerales Order, as well as the Serendipitaceae Family (serendipitoids) having both endophytic and mycorrhizal properties. This is surprising since rapeseed is a non-mycorrhizal plant. Rapeseed also increased the presence of the rapeseed pathogen Olphidium. However, Floc’h et al. (2022) showed that AMF fungi are common under winter rapeseed monocultures. Paraglomaceae has been shown to be common in soils under organic management (Gosling et al. 2014) and Serendipitoids are reported to enhance the growth and stress resistance of barley (Sepehri et al. 2021). They are suggested to be able to decompose SOM and contribute to efficient SOM turnover and thus prevent losses of C and nutrients (Craven and Ray 2019). Thus, diversification with rapeseed has the potential to increase the amounts of beneficial fungi promoting plant growth and soil health.
Smith et al. (2016) showed that Actinobacteria, Firmicutes, and Planctomycetes were more abundant in NT fields while Bacteroidetes and Proteobacteria were more abundant in CT. A similar pattern could not be detected with our differential abundance analysis using the bacterial OTU CT vs NT data from May 2018 (Table S3). However, the same authors concluded that it is difficult to generalize at the bacterial phyla level. The Candidate Rokubacteria NC10 Phylum was among the differentially abundant taxa in NT. NC10 can couple methane oxidation to the reduction of nitrite to dinitrogen (Ettwig et al. 2010). High relative abundance of Rokubacteria have been reported under NT treatments (Sui et al. 2022). NT also favored two representatives of oligotrophic Chloroflexi supporting earlier observations (Pathan et al. 2021). When examined at lower taxonomic levels, representatives of Candidatus Koribacter (Acidobacteria) and Geobacter (Proteobactria) genera were also among NT characterizing taxa. Candidatus Koribacter are capable of oxidation of carbon monooxide and degradation of complex plant polymers (Ward et al. 2009). There is also evidence that Koribacter plays a role in the global N cycle by the reduction of nitrate, nitrite, and possible nitric oxide (Ward et al. 2009). Furthermore, nifH phylotypes related to Geobacter were one of the key microbes of dinitrogen fixation in soils associated with NT management (Calderoli et al. 2017). Geobacter has also been suggested to have an important role in regulating emissions and biogeochemical cycling of soil-derived greenhouse gases (Li and Qixing 2020). The actinobacterial Nocardioides was more abundant in the CT soil. These are common endophytes and have potent biocontrol activities against cereal pathogens (Coombs et al. 2004). Abundance of any specific bacterial taxa seems not to be largely affected by the diversification with winter rapeseed since no differential abundant bacterial OTUs were observed when October 2019 AB vs CD treatment grouping data was compared. However, at a less strict p-value (< 0.001) there was one OTU (lfc value 3.5) belonging to the Nocardioides that was observed as differentially abundant after winter rapeseed (data not shown). Some Nocardioides are also capable of dinitrogen fixation (Nafis et al. 2019) or reducing nitrate (Toth et al. 2008; Dastager et al. 2010; Wang et al. 2016). Rapeseed might benefit from bacterial derived N because of its low N use efficiency (Bouchet et al. 2016).
The rotation effect of winter rapeseed and barley on the soil microbiome has not been an object of study so far and thus other cereal winter rapeseed rotations must serve for a comparison. Hansen et al. (2019) observed that the spring wheat—winter rapeseed rotation had no effect on the microbial community when compared to rapeseed only. This was true for the 0–5 cm soil layer but not when the 5–10 cm layers were compared. In that depth the rotation had a different community indicating that introduction of rapeseed into the rotation can be used to impact the soil microbiome. Lay et al. (2018) compared a wheat- rapeseed rotation to a rapeseed monoculture and concluded that the rapeseed rhizosphere has a core microbiome which is also reflected into the surrounding soil. The same result on the soil fungal community was verified by Sommermann et al. (2018) in a different rotation setup where rapeseed or maize was rotated with winter weed under two different tillage treatments. Having winter rapeseed in the rotation can for instance influence the soil fungal community since they do not have AMF whereas spring barley and wheat are AMF plants. Roy et al. (2021) demonstrated that the pre-crop mycorrhizal status affected the richness and composition of AMF available to the succeeding crop. Therefore, crop rotation schemes that involve a breaking crop, i.e., a non-AMF host from the Brassicaceae family like in this experiment are used to decrease the load of AMF to the next AMF host crop. This has been shown to benefit the following crop in terms of yield (Angus et al. 2015) and influences the soil and rhizosphere fungal community. Our modelling results show that the soil of the C treatment had a higher portion of symbiotic (mainly AMF) fungi compared to treatment B in 2019 and 2020 (Tables 3 and S5). Floc’h et al. (2022) showed that AMF fungi survive under winter rapeseed monocultures for even 10 years due to forming interactions to cohort soil fungi and bacteria and this observation might hold also for our study. Maybe this interaction with microbial cohorts explains also why under rapeseed cultivation the modelled proportion of pathogenic fungi increased in October 2019 (Table 3 and S5). As discussed above the differential abundance analyses identified AMF belonging to the Paraglomerales Order, as well as the Serendipitaceae Family to characterize the winter rapeseed diversification.
Verifying the impact of the cover crop ryegrass within the CT treatment on the soil fungal community seems to be not possible. The only time the A treatment had a distinguished fungal community compared to the C treatment was in October 2019 when the C treatment was under winter rapeseed. In October 2018 and 2020, when both treatments were under barley, they clustered together irrespective of one having the cover crop. Still cover crops can influence the soil microbiome. Aiyer et al. (2022) showed that the choice of cover crop, including ryegrass, in combination with the cash crop barley changed the soil fungal and bacterial community but Lucadamo et al. (2022) argued the cover crop influence to be weak. According to a meta-analysis, plants generally used as cover crops increased parameters of soil microbial abundance, activity, and diversity by 27%, 22%, and 2.5% respectively, compared to those of bare fallow (Kim et al. 2020).
Surprisingly the qPCR abuncances of the amoA, nirK and narG involved in the N turnover did not react to tillage or diversification directly (Tables 3, S5 and S6). This might be an indication of functional redundancy often seen with microbial community changes (Chen et al. 2022). Crop diversification does not directly affect the abundance of the soil bacteria and archaea carrying the amoA but in the long term, if diversification increases soil organic matter, then also amoA can increase, especially the one carried by bacteria (Hao et al. 2022). The same meta-analysis (Hao et al. 2022) reports about the increase of the nirK and narG abundances with diversification in a 10-year long term perspective but no responses are to be expected in short-term studies. The duration of diversification, ecosystem type, planting pattern, crop species, as well as soil layer and texture are important factors regulating the gene abundances of N-cycle (Hao et al. 2022). Diversified cash crop rotation can increase the potential nitrification rate (Raglin et al. 2022). In our study the potential nitrification did not follow the qPCR result of the amoA, but as the outcome of nitrification is nitrate the nirK, involved in denitrification, could correlate to the potential nitrification increase over time. As this was not apparent in our study, we conclude that our laboratory conducted nitrification potential measurement is not correlated to the N-cycle genes retrieved from field conditions. This can be due to using DNA based measures instead of RNA which would more truly reflect the field activity or gross rates nitrification measurements (Elrys et al. 2021) that could have been applied to get a better view of the nitrification rate.
Conclusion
Diversification of agricultural cultivation can be performed in many ways and here, reflecting boreal circumstances, interruption of tilled and non-tilled cereal monocultural rotation was performed with an oil plant, winter rapeseed, or cultivation of a cover crop, ryegrass, extending the length of the field being vegetated. Our results show that tillage has a profound effect on the soil microbiome which is stronger than that of the diversification trials. Interruption of the barley monoculture with winter rapeseed was briefly reflected in the soil microbiome as a change of the fungal and bacterial community composition. It probably needs years using winter rapeseed diversification before the microbial community composition shows permanent differences to a non-diversified control. Future research needs to connect changes in microbial community composition to agricultural field ecosystem services like yield, food quality or pathogen resistances and to soil C sequestration.
References
Abarenkov K, Zirk A, Piirmann T, Pöhönen R, Ivanov F, Nilsson RH, Kõljalg U (2020) UNITE QIIME release for Fungi 2. Version 04.02.2020. UNITE Community. https://doi.org/10.15156/BIO/786387
Ai C, Zhang S, Zhang X, Guo D, Zhou W, Huang S (2018) Distinct responses of soil bacterial and fungal communities to changes in fertilization regime and crop rotation. Geoderma 319:156–166. https://doi.org/10.1016/j.geoderma.2018.01.010
Aiyer H, Fofana B, Fraser T, Caldwell C, McKenzie-Gopsill A, Mills A, Foster A (2022) Choice of cover crop influences soil fungal and bacterial communities in Prince Edward Island, Canada. Can J Microbiol 68:465–482. https://doi.org/10.1139/cjm-2021-0360
Angus JF, Kirkegaard JA, Hunt JR, Ryan MH, Ohlander L, Peoples MB (2015) Break crops and rotations for wheat. Crop Pasture Sci 66:523–552. https://doi.org/10.1071/CP14252
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, Alexander H, Alm EJ, Arumugam M, Asnicar F, Bai Y, Bisanz JE, Bittinger K, Brejnrod A, Brislawn CJ, Brown CT, Callahan BJ, Caraballo-Rodríguez AM, Chase J, Cope EK, Da Silva R, Diener C, Dorrestein PC, Douglas GM, Durall DM, Duvallet C, Edwardson CF, Ernst M, Estaki M, Fouquier J, Gauglitz JM, Gibbons SM, Gibson DL, Gonzalez A, Gorlick K, Guo J, Hillmann B, Holmes S, Holste H, Huttenhower C, Huttley GA, Janssen S, Jarmusch AK, Jiang L, Kaehler BD, Kang KB, Keefe CR, Keim P, Kelley ST, Knights D, Koester I, Kosciolek T, Kreps J, Langille MGI, Lee J, Ley R, Liu Y-X, Loftfield E, Lozupone C, Maher M, Marotz C, Martin BD, McDonald D, McIver LJ, Melnik AV, Metcalf JL, Morgan SC, Morton JT, Naimey AT, Navas-Molina JA, Nothias LF, Orchanian SB, Pearson T, Peoples SL, Petras D, Preuss ML, Pruesse E, Rasmussen LB, Rivers A, Robeson MS II, Rosenthal P, Segata N, Shaffer M, Shiffer A, Sinha R, Song SJ, Spear JR, Swafford AD, Thompson LR, Torres PJ, Trinh P, Tripathi A, Turnbaugh PJ, Ul-Hasan S, van der Hooft JJJ, Vargas F, Vázquez-Baeza Y, Vogtmann E, von Hippel M, Walters W, Wan Y, Wang M, Warren J, Weber KC, Williamson CHD, Willis AD, Xu ZZ, Zaneveld JR, Zhang Y, Zhu Q, Knight R, Caporaso JG (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Bouchet AS, Laperche A, Bissuel-Belaygue C, Snowdon R, Nesi N, Stahl A (2016) Nitrogen use efficiency in rapeseed. A Review Agron Sustain Dev 36:38. https://doi.org/10.1007/s13593-016-0371-0
Bragg L, Stone G, Imelfort M, Hugenholtz P, Tyson GW (2012) Fast, accurate error-correction of amplicon pyrosequences using Acacia. Nat Methods 9:425–426. https://doi.org/10.1038/nmeth.1990
Bru D, Sarr A, Philippot L (2007) Relative abundances of proteobacterial membrane-bound and periplasmic nitrate reductases in selected environments. Appl Environ Microbiol 73:18. https://doi.org/10.1128/AEM.00643-07
Calderoli PA, Collavino MM, Kraemer FB, Morrás HJM, Aguilar OM (2017) Analysis of nifH-RNA reveals phylotypes related to Geobacter and Cyanobacteria as important functional components of the N2 -fixing community depending on depth and agricultural use of soil. MicrobiologyOpen 6:e00502. https://doi.org/10.1002/mbo3.502
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583. https://doi.org/10.1038/nmeth.3869
Cannon PF, Kirk PM (2007) Fungal families of the world. CAB International, Cambridge, UK. ISBN: 978-0851998275
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Huntley J, Fierer N, Owens SM, Betley J, Fraser L, Bauer M, Gormley N, Gilbert JA, Smith G, Knight R (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6:1621–1624. https://doi.org/10.1038/ismej.2012.8
Cappelli SL, Domeignoz-Horta LA, Loaiza V, Laine A-L (2022) Plant biodiversity promotes sustainable agriculture directly and via belowground effects. Trends Plant Sci 27:674–687. https://doi.org/10.1016/j.tplants.2022.02.003
Castellano-Hinojosa A, Kanissery R, Strauss SL (2023) Cover crops in citrus orchards impact soil nutrient cycling and the soil microbiome after three years but effects are site-specifc. Biol Fertil Soils 59:659–678. https://doi.org/10.1007/s00374-023-01729-1
Chao A, Chiu C-H, Jost L (2014) Unifying species diversity, phylogenetic diversity, functional diversity, and related similarity/differentiation measures through Hill numbers. Annu Ecol Evol Syst 45:297–324. https://doi.org/10.1146/annurev-ecolsys-120213-091540
Chen L, Reeve J, Zhang L, Huang S, Wang X, Chen J (2018) GMPR: a robust normalization method for zero-inflated count data with application to microbiome sequencing data. Peer J 6:e4600. https://doi.org/10.7717/peerj.4600
Chen H, Ma K, Lu C, Fu Q, Qiu Y, Zhao J, Huang Y, Yang Y, Schadt CW, Chen H (2022) Functional redundancy in soil microbial community based on metagenomics across the globe. Front Microbiol 13:878978. https://doi.org/10.3389/fmicb.2022.878978
Chen D, Wang C, Ma X, Chen K, Wang Z, Wang Q, Zhang J, Zhou Q, Shen W (2023) Dynamic changes in soil fungal communities and functional groups in response to sugarcane/soybean intercropping with reduced nitrogen fertilizer application. Biol Fertil Soils 59:363–378. https://doi.org/10.1007/s00374-023-01709-5
Coombs JT, Michelsen PP, Franco CMM (2004) Evaluation of endophytic actinobacteria as antagonists of Gaeumannomyces graminis var. tritici in wheat. Biol Control 29:359–366. https://doi.org/10.1016/j.biocontrol.2003.08.001
Craven KD, Ray R (2019) More than serendipity: The potential to manage soil carbon and emissions while promoting low-input agriculture with serendipitoid mycorrhizae. Phytobiomes J 3:161–164. https://doi.org/10.1094/PBIOMES-12-18-0058-P
Dastager SG, Lee JC, Pandey A, Kim CJ (2010) Nocardioides mesophilus sp. nov., isolated from soil. Int J Syst Evol Micr 60:2288–2292. https://doi.org/10.1099/ijs.0.019059-0
Elrys AS, Wang J, Metwally MAS, Cheng Y, Zhang J-B, Cai Z-C, Chang SX, Müller C (2021) Global gross nitrification rates are dominantly driven by soil carbon-to-nitrogen stoichiometry and total nitrogen. Glob Change Biol 27:6512–6524. https://doi.org/10.1111/gcb.15883
Ettwig KF, Butler MK, Le Paslier D, Pelletier E, Mangenot S, Kuypers MMM, Schreiber F, Dutilh BE, Zedelius J, de Beer D, Gloerich J, Wessels HJCT, van Alen T, Luesken F, Wu ML, van de Pas-Schoonen KT, Op den Camp HJM, Janssen-Megens EM, Francoijs K-J, Stunnenberg H, Weissenbach J, Jetten MSM, Strous M (2010) Nitrite-driven anaerobic methane oxidation by oxygenic bacteria. Nature 464:543–548. https://doi.org/10.1038/nature08883
Fang Y, Van Zwieten L, Rose MT, Vasileiadis S, Donner E, Vancov T, Rigg JL, Weng ZH, Lombi E, Drigo B, Conyers M, Tavakkoli E (2022) Unraveling microbiomes and functions associated with strategic tillage, stubble, and fertilizer management. Agric Ecosyst Environ 323:107686. https://doi.org/10.1016/j.agee.2021.107686
Fellbaum CR, Gachomo EW, Beesetty Y, Choudhari S, Strahan GD, Pfeffer PE, Toby Kier E, Bücking H (2012) Carbon availability triggers fungal nitrogen uptake and transport in arbuscular mycorrhizal symbiosis. PNAS 109:2666–2671. https://doi.org/10.1073/pnas.1118650109
Floc’h J-B, Hamel C, Laterrière M, Tidemann B, St-Arnaud M, Hijri M (2022) Long-term persistence of arbuscular mycorrhizal fungi in the rhizosphere and bulk soils of non-host Brassica napus and their networks of co-occurring microbes. Front Plant Sci 13:828145. https://doi.org/10.3389/fpls.2022.828145
Gosling P, Proctor M, Jones J, Bending GD (2014) Distribution and diversity of Paraglomus spp. in tilled agricultural soils. Mycorrhiza 24:1–11. https://doi.org/10.1007/s00572-013-0505-z
Griffiths M, Delory BM, Jawahir V, Wong KM, Bagnal GC, Dowd TG, Nusinow DA, Miller AJ, Topp CN (2022) Optimisation of root traits to provide enhanced ecosystem services in agricultural systems: A focus on cover crops. Plant Cell Environ 45:751–770. https://doi.org/10.1111/2Fpce.14247
Gupta A, Singh UB, Sahu PK, Paul S, Kumar A, Malviya D, Singh S, Kuppusamy P, Singh P, Paul D, Rai JP, Singh HV, Manna MC, Crusberg TC, Kumar A, Saxena AK (2022) Linking soil microbial diversity to modern agriculture practices: A Review. Int J Env Res Pub He 19:3141. https://doi.org/10.3390/2Fijerph19053141
Hansen JC, Schillinger WF, Sullivan TS, Paulitz TC (2019) Soil microbial biomass and fungi reduced with Canola introduced into long-term monoculture wheat rotations. Front Microbiol 10:1488. https://doi.org/10.3389/fmicb.2019.01488
Hao J, Feng Y, Wang X, Yu Q, Zhang F, Yang G, Ren G, Han X, Wang X, Ren C (2022) Soil microbial nitrogen-cycling gene abundances in response to crop diversification: A meta-analysis. Sci Tot Environ 838:156621. https://doi.org/10.1016/j.scitotenv.2022.156621
Hartmann M, Six J (2023) Soil structure and microbiome functions in agroecosystems. Nat Rev Earth Environ 4:4–18. https://doi.org/10.1038/s43017-022-00366-w
Henry S, Baudoin E, López-Gutiérrez JC, Martin-Laurent F, Brauman A, Philippot L (2004) Quantification of denitrifying bacteria in soils by nirK gene targeted real-time PCR. J Microbiol Methods 59:327–335. https://doi.org/10.1016/j.mimet.2004.07.002
Hertel T, Elouafi I, Tanticharoen M, Ewert F (2021) Diversification for enhanced food systems resilience. Nat Food 2:832–834. https://doi.org/10.1038/s43016-021-00403-9
Hirsch PR, Mauchline TH (2015) Chapter Two - The importance of the microbial N cycle in soil for crop plant nutrition. In: Sariaslani S, Gadd GM (eds) Advances in applied microbiology, vol 93. Academic Press, London, pp 45–71
Hoffmann H, Schloter M, Wilke B-M (2007) Microscale-scale measurement of potential nitrification rates of soil aggregates. Biol Fertil Soils 44:411–413. https://doi.org/10.1007/s00374-007-0227-5
ISO 15685 (2012) Soil quality—Determination of potential nitrification and inhibition of nitrification—Rapid test by ammonium oxidation. https://standards.iteh.ai/catalog/standards/sist/ff242fb7-eb9d-4990-9ef1-a0bd570372dc/iso-15685-2012
Kallenbach CM, Frey SD, Grandy AS (2016) Direct evidence for microbial-derived soil organic matter formation and its ecophysiological controls. Nat Commun 7:13630. https://doi.org/10.1038/ncomms13630
Khmelevtsova LE, Sazykin IS, Azhogina TN, Sazykina MA (2022) Influence of agricultural practices on bacterial community of cultivated soils. Agriculture 12:371. https://doi.org/10.3390/agriculture12030371
Kim N, Zabaloy MC, Guan K, Villamil MB (2020) Do cover crops benefit soil microbiome? A meta-analysis of current research. Soil Biol Biochem 142:107701. https://doi.org/10.1016/j.soilbio.2019.107701
Kraut-Cohen J, Zolti A, Shaltiel-Harpaz L, Argaman E, Rabinovich R, Green SJ, Minz D (2020) Effects of tillage practices on soil microbiome and agricultural parameters. Sci Tot Environ 705:135791. https://doi.org/10.1016/j.scitotenv.2019.135791
Lay C-Y, Bell TH, Hamel C, Harker KN, Mohr R, Greer CW, Yergeau É, St-Arnaud M (2018) Canola root–associated microbiomes in the Canadian prairies. Front Microbiol 9:1188. https://doi.org/10.3389/fmicb.2018.01188
Leigh J, Hodge A, Fitter AH (2009) Arbuscular mycorrhizal fungi can transfer substantial amounts of nitrogen to their host plant from organic material. New Phytol 181:199–207. https://doi.org/10.1111/j.1469-8137.2008.02630.x
Li T, Qixing Z (2020) The key role of Geobacter in regulating emissions and biogeochemical cycling of soil-derived greenhouse gases. Env Poll 266:115135. https://doi.org/10.1016/j.envpol.2020.115135
Li Y, Song D, Liang S, Dang P, Qin X, Liao Y, Siddique KHM (2020) Effect of no-tillage on soil bacterial and fungal community diversity: A meta-analysis. Soil Tillage Res 204:104721. https://doi.org/10.1016/j.still.2020.104721
Lin H, Peddada SD (2020) Analysis of compositions of microbiomes with bias correction. Nat Com 11:3514. https://doi.org/10.1038/s41467-020-17041-7
Liu H, Carvalhais LC, Rincon-Florez V, Crawford M, Dang YP, Dennis PG, Schenk PM (2016) One-time strategic tillage does not cause major impacts on soil microbial properties in a no-till Calcisol. Soil Tillage Res 158:91–99. https://doi.org/10.1016/j.still.2015.12.007
Liu B, Schroeder J, Ahnemann H, Poeplau C, Tebbe CC (2023) Crop diversification improves the diversity and network structure of the prokaryotic soil microbiome at conventional nitrogen fertilization. Plant Soil 489:259–276. https://doi.org/10.1007/s11104-023-06011-9
Lizarazo CI, Tuulos A, Jokela V, Mäkelä PSA (2020) Sustainable nixed cropping systems for the Boreal-Nemoral region. Front Sustain Food Syst 4:103. https://doi.org/10.3389/fsufs.2020.00103
Lucadamo EE, Holmes AA, Wortman SE, Yannarell AC (2022) Post-termination effects of cover crop monocultures and mixtures on soil inorganic nitrogen and microbial communities on two organic farms in Illinois. Front Soil Sci 2:824087. https://doi.org/10.3389/fsoil.2022.824087
Mackay JE, Bernhardt LT, Smith RG, Ernakovich JG (2023) Tillage and pesticide seed treatments have distinct effects on soil microbial diversity and function. Soil Biol Biochem 176:108860. https://doi.org/10.1016/j.soilbio.2022.108860
Mangalassery S, Mooney SJ, Sparkes DL, Fraser WT, Sjögersten S (2015) Impacts of zero tillage on soil enzyme activities, microbial characteristics and organic matter functional chemistry in temperate soils. European J Soil Biol 68:9–17. https://doi.org/10.1016/j.ejsobi.2015.03.001
Mbuthia LW, Acosta-Martínez V, DeBryun J, Schaeffer S, Tyler D, Odoi E, Mpheshea M, Walker F, Eash N (2015) Long term tillage, cover crop, and fertilization effects on microbial community structure, activity: Implications for soil quality. Soil Biol Biochem 89:24–34. https://doi.org/10.1016/j.soilbio.2015.06.016
Micallef L, Rodgers P (2014) eulerAPE: Drawing Area-Proportional 3-Venn Diagrams Using Ellipses. PLoS ONE 9:e101717. https://doi.org/10.1371/journal.pone.0101717
Nafis A, Raklami A, Bechtaoui N, El Khalloufi F, El Alaoui A, Glick BR, Hafidi M, Kouisni L, Ouhdouch Y, Hassani L (2019) Actinobacteria from extreme niches in Morocco and their plant growth-promoting potentials. Diversity 11:139. https://doi.org/10.3390/d11080139
Naylor D, McClure R, Jansson J (2022) Trends in microbial community composition and function by soil depth. Microorganisms 10:540. https://doi.org/10.3390/microorganisms10030540
Nguyen HDT, Nickerson NL, Seifert KA (2013) Basidioascus and Geminibasidium: a new lineage of heat-resistant and xerotolerant basidiomycetes. Mycologia 105:1231–1250. https://doi.org/10.3852/12-351
Nguyen NH, Song Z, Bates ST, Branco S, Tedersoo L, Menke J, Schilling JS, Kennedy PG (2016) FUNGuild: An open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol 20:241–248. https://doi.org/10.1016/j.funeco.2015.06.006
Oades JM (1993) The role of biology in the formation, stabilization and degradation of soil structure. Geoderma 56:377–400. https://doi.org/10.1016/0016-7061(93)90123-3
Oksanen J, Simpson GL, Blanchet FG et al (2022) Vegan: community ecology package. R package version 2.6–4. https://CRAN.R-project.org/package=vegan
Pathan SI, Roccotelli A, Petrovičovà B, Romeo M, Badagliacca G, Monti M, Gelsomino A (2021) Temporal dynamics of total and active prokaryotic communities in two Mediterranean orchard soils treated with solid anaerobic digestate or managed under no-tillage. Biol FertilSoils 57:837–861. https://doi.org/10.1007/s00374-021-01569-x
R Core Team (2022) R: a language and environment for statistical computing, R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/
R Studio Team (2022) RStudio: integrated development environment for R. RStudio, PBC, Boston, MA, USA. http://www.rstudio.com/
Raglin SS, Soman C, Ma Y, Kent AD (2022) Long term influence of fertility and rotation on soil nitrification potential and nitrifier communities. Front Soil Sci 2:838497. https://doi.org/10.3389/fsoil.2022.838497
Rillig MC, Aguilar-Trigueros CA, Bergmann J, Verbruggen E, Veresoglou SD, Lehmann A (2015) Plant root and mycorrhizal fungal traits for understanding soil aggregation. New Phyt 205:1385–1388. https://doi.org/10.1111/nph.13045
Rincon-Florez VA, Carvalhais LC, Dang YP, Crawford MH, Schenk PM, Dennis PG (2020) Significant effects on soil microbial communities were not detected after strategic tillage following 44 years of conventional or no-tillage management. Pedobiologia 80:150640. https://doi.org/10.1016/j.pedobi.2020.150640
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584. https://doi.org/10.7717/peerj.2584
Rotthauwe JH, Witzel K-P, Liesack W (1997) The ammonia monooxygenase structural gene amoA as a functional marker: Molecular fine-scale analysis of natural ammonia-oxidizing populations. Appl Env Microbiol 63:4704–4712. https://doi.org/10.1128/aem.63.12.4704-4
Roy J, van Duijnen R, Leifheit EF, Mbedi S, Temperton VM, Rillig MC (2021) Legacy effects of pre-crop plant functional group on fungal root symbionts of barley. Ecol Appl 31:e02378. https://doi.org/10.1002/eap.2378
Russel J (2021) Russel88/MicEco: v0.9.15. Zenodo. https://doi.org/10.5281/zenodo.4733747
Schalamuk S, Cabello M (2010) Arbuscular mycorrhizal fungal propagules from tillage and no-tillage systems: possible effects on Glomeromycota diversity. Mycologia 102:261–268. https://doi.org/10.3852/08-118
Sepehri M, Ghaffari MR, Khayam Nekoui M, Sarhadi E, Moghadam A, Khatabi B, Salekdeh GH (2021) Root endophytic fungus Serendipita indica modulates barley leaf blade proteome by increasing the abundance of photosynthetic proteins in response to salinity. J Appl Microbiol 131:1870–1889. https://doi.org/10.1111/jam.15063
Sharma-Poudyal D, Schlatter D, Yin C, Hulbert S, Paulitz T (2017) Long-term no-till: A major driver of fungal communities in dryland wheat cropping systems. PLoS ONE 12:e0184611. https://doi.org/10.1371/journal.pone.0184611
Smith DP, Peay KG (2014) Sequence depth, not PCR replication, improves ecological inference from next generation DNA sequencing. PLoS ONE 9:e90234. https://doi.org/10.1371/journal.pone.0090234
Smith CR, Blair PL, Boyd C, Cody B, Hazel A, Hedrick A, Kathuria H, Khurana P, Kramer B, Muterspaw K, Peck C, Sells E, Skinner J, Tegeler C, Wolfe Z (2016) Microbial community responses to soil tillage and crop rotation in a corn/soybean agroecosystem. Ecol Evol 6:8075–8084. https://doi.org/10.1002/ece3.2553
Sommermann L, Geistlinger J, Wibberg D, Deubel A, Zwanzig J, Babin D, Schlüter A, Schellenberg I (2018) Fungal community profiles in agricultural soils of a long-term field trial under different tillage, fertilization and crop rotation conditions analyzed by high-throughput ITS-amplicon sequencing. PLoS ONE 13:e0195345. https://doi.org/10.1371/journal.pone.0195345
Stefan L, Hartmann M, Engbersen N, Six J, Schöb C (2021) Positive effects of crop diversity on productivity driven by changes in soil microbial composition. Front Microbio 12:660749. https://doi.org/10.3389/fmicb.2021.660749
Sui P, Li R, Zheng H, Wang H, Luo YY, Y, Zheng J, Liu W, (2022) Long-term conservation tillage practices directly and indirectly affect soil micro-food web in a Chinese Mollisol. Agronomy 12:2356. https://doi.org/10.3390/agronomy12102356
Tosi M, Drummelsmith J, Obregón D, Chahal I, Van Eerd LL, Dunfield KE (2022) Cover crop-driven shifts in soil microbial communities could modulate early tomato biomass via plant-soil feedbacks. Sci Rep 12:9140. https://doi.org/10.1038/s41598-022-11845-x
Toth EM, Keki Z, Homonnay ZG, Borsodi AK, Marialigeti K, Schumann P (2008) Nocardioides daphniae sp. nov., isolated from Daphnia cucullata (Crustacea: Cladocera). Int JSyst Evol Microbiol 58:78–83. https://doi.org/10.1099/ijs.0.65305-0
Wang LN, Li JL, Zhang GY (2016) Nocardioides rotundus sp. nov., isolated from deep seawater. Int JSyst Evol Microbiol 66:1932–1936. https://doi.org/10.1099/ijsem.0.000966
Wang GZ, Bei SK, Li JP, Bao XG, Zhang JD, Schultz PA, Li HG, Li L, Zhang FS, Bever JD, Zhang JL (2021) Soil microbial legacy drives crop diversity advantage: linking ecological plant-soil feedback with agricultural intercropping. J Appl Ecol 58:496–506. https://doi.org/10.1111/1365-2664.13802
Wang G, Li X, Xi X, Cong W-F (2022) Crop diversification reinforces soil microbiome functions and soil health. Plant Soil 476:375–383. https://doi.org/10.1007/s11104-022-05436-y
Ward NL, Challacombe JF, Janssen PH et al (2009) Three genomes from the phylum Acidobacteria provide insight into the lifestyles of these microorganisms in soils. Appl Environ Microbiol 75:2046–2056. https://doi.org/10.1128/AEM.02294-08
Westfall PH (1997) Multiple testing of general contrasts using logical constraints and correlations. J Am Stat Assoc 92:299–306. https://doi.org/10.1080/01621459.1997.10473627
Wilkinson L (2012) Exact and approximate area-proportional circular Venn and Euler Diagrams. IEEE Trans vis Comput Graph 18:321–331. https://doi.org/10.1109/TVCG.2011.56
Wipf HM-L, Xu L, Gao C, Spinner HB, Taylor J, Lemaux P, Mitchell J, Coleman-Derr D (2021) Agricultural soil management practices differentially shape the bacterial and fungal microbiomes of Sorghum bicolor. Appl Environ Microbiol 87:e02345-20. https://doi.org/10.1128/2FAEM.02345-20
Wooliver R, Kivlin SN, Jagadamma S (2022) Links among crop diversification, microbial diversity, and soil organic carbon: Mini review and case Studies. Front Microbiol 13:854247. https://doi.org/10.3389/fmicb.2022.854247
Wortmann CS, Quincke JA, Drijber JA, Mamo M, Franti T (2008) Soil microbial community change and recovery after one-time tillage of continuous no-till. Agron J 100:1681–1686. https://doi.org/10.2134/agronj2007.0317
Yang Y, Xie H, Mao Z, Bao X, He H, Zhang X, Liang C (2022) Fungi determine increased soil organic carbon more than bacteria through their necromass inputs in conservation tillage croplands. Soil Biol Biochem 167:108587. https://doi.org/10.1016/j.soilbio.2022.108587
Zhang J, Kobert K, Flouri T, Stamatakis A (2014) PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30:614–620. https://doi.org/10.1093/bioinformatics/btt593
Acknowledgements
The work was funded within the project “Crop diversification and low-input farming across Europe: from practitioners’ engagement and ecosystems services to increased revenues and value chain organization” (Diverfarming) under the European Union’s Horizon 2020 Programme for Research & Innovation, grant agreement no. 728003. We thank Eerika Suni, Sirpa Tiikkainen, Anneli Supponen, Katri Leino and other staff members of Natural Resource Institute Finland who provided excellent assistance with laboratory work.
Funding
Open access funding provided by Natural Resources Institute Finland.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fritze, H., Tuomivirta, T., Orrù, L. et al. Effect of no-till followed by crop diversification on the soil microbiome in a boreal short cereal rotation. Biol Fertil Soils 60, 357–374 (2024). https://doi.org/10.1007/s00374-024-01797-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00374-024-01797-x