Abstract
Purpose
Clear cell, papillary cell, and chromophobe renal cell carcinomas (RCCs) have now been well characterised thanks to large collaborative projects such as The Cancer Genome Atlas (TCGA). Not only has knowledge of the genomic landscape helped inform the development of new drugs, it also promises to fine tune prognostication.
Methods
A literature review was performed summarising the current knowledge on the genetic basis of RCC.
Results
The Von Hippel–Lindau (VHL) tumour suppressor gene undergoes bi-allelic knockout in the vast majority of clear cell RCCs. The next most prevalent aberrations include a cohort of chromatin-modifying genes with diverse roles including PBRM1, SETD2, BAP1, and KMD5C. The most common non-clear cell renal cancers have also undergone genomic profiling and are characterised by distinct genomic landscapes. Many recurrent mutations have prognostic value and show promise in aiding decisions regarding treatment stratification. Intra-tumour heterogeneity appears to hamper the clinical applicability of sparsely sampled tumours. Ways to abrogate heterogeneity will be required to optimise the genomic classification of tumours.
Conclusion
The somatic mutational landscape of the more common renal cancers is well known. Correlation with outcome needs to be more comprehensively furnished, particularly for small renal masses, rarer non-clear cell renal cancers, and for all tumours undergoing targeted therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In this review, we consider what is currently known of the genetic landscape of the commonest subtypes of renal cell cancer (RCC). A glossary has been provided to aid the understanding of specialist terminology (Table 1). Clear cell, papillary, and chromophobe cancers have now been well characterised thanks to the development of sequencing technologies (Table 2) and large collaborative projects such as The Cancer Genome Atlas (TCGA). Not only has knowledge of the genomic landscape helped inform the development of new drugs, this understanding also promises to improve risk stratification of tumours and to determine their sensitivity to systemic therapies. We shall consider each subtype in turn describing genes and pathways of oncogenesis and how these relate to prognosis and treatment response. We finish by discussing limitations of these metrics before widespread clinical applicability may be adopted.
Methods
A non-systematic literature search was conducted using Medline, updated to May 2018. The reference lists of selected manuscripts were checked manually for eligible articles. The most relevant articles summarising existing knowledge on RCC genomics, including tumour cell evolution and progression, were selected for this review. Recurrent aberrations have been defined as those with false discovery rates of < 0.1 and reported in multiple studies in the literature. For prognostic markers, those events that are significant after multiple hypothesis testing have been included (adjusted p value < 0.1). The commonly accepted significance threshold (p < 5 × 10−8) has been used for genome-wide association studies (GWAS).
Results
Clear cell renal cell carcinoma
Epidemiology and genetics
The mutational landscape of clear cell RCC (ccRCC) has been defined most recently through several large-scale whole genome-sequencing studies [1,2,3,4]. These studies reveal that recurrent somatic mutations occur in only a handful of genes, with an overall mutational burden of roughly 1–2 per Mb. In addition, there are only a small number of recurrent copy number aberrations and rare gene fusions. Some insights into clinical risk factors and their genomic correlates have been made. These include patient age, with mutational burden correlating strongly with age via the predominance of the clock-like mutational signature in these genomes [5]. The higher incidence of ccRCC in male patients may partially be accounted by mono-allelic inactivation of the chromatin remodelling gene, KDM5C on the X chromosome [6]. No tobacco-specific mutational process has been detected despite the strong clinical risk factor [5]. The high frequency of A:T-to-T:A transversions consistent with mutational damage as a result of aristolochic acid exposure was detected via sequencing of patients from Eastern Europe [1] and has directly influenced primary prevention strategies.
Genetic risk factors are known to play a role in sporadic RCC development [7, 8]. Patients who have at least one first degree relative with RCC are at an increased risk of developing the disease (OR 1.4, 95% CI 0.71–2.76) [9]. The first renal large-scale GWAS in Europe revealed susceptibility loci at 2p21 and 11q13.3 [10]. The two correlated variants on 2p21 map to EPAS1, a transcription factor previously implicated in RCC, whereas the variant on 11q13.3 contains no characterised genes. An additional susceptibility locus on 12p11.23 was later discovered containing two variants in the ITPR2 gene, though direct functional evidence between ITPR2 and oncogenesis is lacking [11]. Subsequently, a locus on intron 2 of the ZEB2 gene was discovered which may play a role in decreasing regulation of epithelial to mesenchymal transition [12].
Most recently, a variant on 8q24.21 was discovered via interrogation of GWAS from an Icelandic population [13], prior to discovery of an additional seven new loci in the largest such GWAS study to date [14]. More comprehensive details of these loci are shown in Table 3.
Somatic mutations
VHL
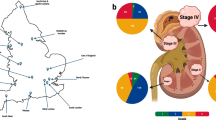
The Von Hippel–Lindau (VHL) tumour suppressor gene, located at 3p25, undergoes bi-allelic knockout in the majority of ccRCCs [15]. Haploinsufficiency of VHL occurs via arm-level loss of chromosome 3p in over 90% of tumours. Astonishingly, this event appears to occur in a handful of cells in childhood or late adolescence, often many decades prior to diagnosis [16] (Fig. 1). The second copy of VHL is lost, usually much later in life, by either non-synonymous mutation or epigenetic down-regulation [1,2,3,4, 16]. VHL inactivation prevents the ubiquitination of hypoxia-inducible factor (HIF) for degradation. Upregulation of HIF is advantageous to tumour cell survival due to increased expression levels of angiogenic factors, lower rates of apoptosis, and higher rates of cellular proliferation (Fig. 2). The ubiquitous nature of upregulated HIF pathways and, therefore, neo-angiogenesis has provided the rationale for treatment with vascular endothelial growth factor (VEGF) inhibitors. Perhaps, unsurprisingly, given its ubiquitous role, no consistent relationship between VHL status and clinical outcome has been found [17, 18].
Mutation frequencies of the most commonly mutated genes in ccRCC and their effect as described by the Hallmarks of Cancer [64]
Chromatin-modifying genes
A cohort of chromatin-modifying genes with diverse roles including PBRM1, SETD2, BAP1, and KMD5C constitutes the next most prevalent somatic mutations. The first three of these genes are also co-located with VHL on chromosome 3p, meaning that after 3p loss, any further non-synonymous mutation will result in complete inactivation of these haploinsufficient genes. PBRM1, a methyltransferase is the second most commonly mutated gene in ccRCC, found in 30–50% of tumours [1, 2]. PBRM1’s inactivation could lead to loss of DNA methylation via reduction of H3K36me3 [2]. SETD2 is mutated in 10–30% of ccRCCs [1, 19,20,21]. SETD2’s intracellular roles are numerous, including the regulation of transcription elongation, RNA processing, and double-stranded DNA break repair [22] that may then activate the p53-mediated checkpoint in the absence of specific p53 mutations [23]. BAP1, a histone deubiquitinase, is mutated in up to 5–15% of ccRCCs [1,2,3, 24, 25]. BAP1’s other roles include control of cellular proliferation and regulation of DNA damage repair [24].
Due to their prevalence, PBRM1, SETD2, and BAP1 have all been investigated as prognostic markers and for possible treatment stratification. A retrospective, validated analysis found that tumours with BAP1 mutations conferred a worse prognosis, higher grade, and worse overall survival when compared to those with PBRM1 mutations or when compared to those without BAP1 mutations [2, 24,25,26]. The presence of BAP1 and PBRM1 mutants appeared anti-correlated, though when co-existing, their presence conferred the worst overall survival. Similarly, the presence of SETD2 confers worse overall survival by a hazard ratio of 1.7 [26].
Genomic profiling of tumours from patients with ccRCC is beginning to illustrate how the presence of mutations in chromatin-modifying genes may aid systemic treatment stratification. For instance, in the RECORD-3 trial [27], different sequences of everolimus (an mTOR inhibitor) and sunitinib (a VEGF inhibitor) appeared to affect progression free survival in metastatic patients according to PBRM1 and BAP1 status. Immuno-oncological agents are now showing increasing promise in metastatic ccRCC settings, where PBRM1 mutations appear to confer clinical benefit after treatment with these agents [28].
TERT
Somatic mutations have been detected within the core promoter [27, 29, 30] and 5’UTR [16] of telomerase reverse transcriptase (TERT) in 6–14% of ccRCCs. Their functional corollary appears to include the lengthening of telomeres [16]. Furthermore, the presence of TERT promoter mutations has been shown to decrease cancer-specific survival [29] and increased disease stage [30].
PTEN- and mTOR-signalling pathway
The PTEN gene undergoes both recurrent point mutations (2–12% of samples) and focal deletions (approximately 7% of samples) in ccRCC [2, 3, 27]. The gene encodes a protein and phospholipid phosphatase that controls the balance between cell proliferation, apoptosis, and migration via the PI(3)K/AKT/MTOR pathway. Specific mutations may act in a dominant negative manner, implying that bi-allelic knockout is not always required to suppress function [31]. An interrogation of the TCGA data set revealed that bi-allelic loss of PTEN was uncommon but conferred worse overall survival. Mono-allelic loss was also associated with higher stage and histological grade [32]. Tumours with mutant PTEN status showed a non-significant increase in rates of progression when compared to non-PTEN mutant tumours after either VEGFR or MTOR inhibition in metastatic patients in the RECORD-3 trial [27].
The PI3K-AKT-mTOR signalling axis is directly augmented via MTOR mutations, observed in 4–9% of ccRCC neoplasms [1, 3]. Numerous other genes are involved in the mTOR pathway including PTEN, whose importance is discussed above. Other members of the pathway, such as TSC1/TSC2/PIK3CA, are infrequently mutated. Although FGFR4 undergoes copy gain as part of an arm-level gain of the long-arm of chromosome 5 in approximately 50% of ccRCC, a direct causal link between this event and PI3K-AKT-mTOR has not yet been conclusively found. The PI3K–mTOR pathway is an important growth factor-signalling cascade that alters cellular metabolism. It is, therefore, an attractive target for systemic therapies via compounds collectively named rapalogs that bind to FKBP12 to inhibit the PI3K–AKT–mTOR pathway via abrogated mTORC1 kinase activity [33]. There is some evidence that tumours with mutations in MTOR/TSC1/2 have a better response to rapalogs, although statistical significance has not been reached [27, 34, 35].
TP53
TP53 appears relatively infrequently mutated in ccRCC (2–9% of tumours) [2, 3]. However, aberrations in genes involved in the P53 pathway are relatively common implying that the p53 pathway and cell-cycle checkpoint inhibition play significant roles in ccRCC [3, 23]. TP53 appears to be more frequently mutated in metastases [36] and on survival analysis, confers a worse cancer-specific survival than any other single mutation [37].
Structural variants
Structural variants promote oncogenesis by altering the number of copies of genes in the genome or re-arranging the order of the genome such that either new genes are formed or the expression of a gene is altered. Aside from translocation renal cell carcinoma, gene fusions are uncommon in renal cell cancer. Large-scale copy number variations are common, however, including the almost ubiquitous heterozygous loss of the short arm of chromosome 3. The next most common aberrations are: gain of chromosome 5 (~ 60% of samples) and loss of chromosome 14 (~ 40% of samples), loss of chromosomes 6q, 8p, and 9p, and gain of chromosome 7 (~ 30% of samples) [1,2,3,4, 16, 37, 38]. Multiple authors have investigated copy number aberrations as potential biomarkers, mainly using array comparative genomic hybridisation and cytogenetic studies. Although many of these aberrations have been shown to predict prognosis [38,39,40,41,42,43], only few were repeatedly validated on multivariable analyses. Due to most copy number alterations covering large segments of the genome, it is difficult to uncover the mechanism by which the change confers oncogenic advantage. Some interesting and notable genes within these regions include CDKN2A on chromosome 9p, which has been shown to modulate VEGF expression via its interaction with HIF-1alpha, encoded by HIF1A on 14q [44].
Immunotherapy and mutational burden
Targeted immunotherapy, for instance, in the form of programmed death 1 (PD-1) checkpoint inhibition or cytotoxic T-lymphocyte associated antigen 4 (CTLA-4) inhibition is being increasingly used in both first- and second-line therapies [45]. Prediction of favourable response has not been correlated with PD-1 ligand biomarker expression [46]. In bladder cancer amongst others [47], mutational burden as a surrogate for neoantigen levels has been associated with enhanced response to targeted immunotherapy. Although the total mutational burden is relatively low in renal cancer, it may be a useful adjunct for response prediction to novel immunotherapeutic agents. Recently, a small study correlated total mutational burden with response to targeted immunotherapy in RCC [48]. In this study, estimated tumour mutational burden was similar in those with progressive disease and clinical benefit (10 vs. 11, p = 0.8), as was the duration of therapy in patients with high- and low-tumour mutational burden (71 vs. 70 days, p = 0.39). However, this study was fairly small (n = 31) and included both patients with ccRCC and non-clear cell RCC who received several different targeted immunotherapies.
Heterogeneity
Intratumoural heterogeneity in ccRCC is well understood through bulk tissue DNA sequencing with branching evolution occurring more commonly and earlier than other cancer types [4, 49]. It dominates the evolutionary landscape at the genomic, transcriptomic, and proteomic levels [50]. Phylogenetic analyses led to the trunk-branch model of RCC development (Fig. 1). Somatic mutations that are found in all sampled tumour regions present in the trunk of the phylogenetic tree, including VHL mutations and 3p loss [51, 52]. This finding supports the Knudson two-hit hypothesis [53], where two ‘hits’ (i.e., bi-allelic inactivation of VHL) are required for clonal expansion to yield a clone large enough to stochastically acquire independent branches. Less prevalent mutations appear more commonly on branches with the same gene sometimes mutated on different branches in a fashion consistent with convergent evolution [4, 52]. One direct corollary of these findings is that the estimation of driver gene prevalence based on single regional sequencing significantly under-estimates the true tumour-based estimation. Increased driver prevalence was seen, particularly in PBRM1, BAP1, TP53, PTEN, PIK3CA, and TSC2. Incomplete molecular profiling from single biopsies may hinder accurate prognosis and response to therapy. Extensive multi-regional sequencing appears to demarcate tumour behaviour according to evolutionary subtypes [4].
Clearly, the development of methods to infer tumour behaviour without resorting to exhaustive spatial tissue sampling is vital. Current estimates of the sampling density required to adequately represent tissue biology lie between 3 and 8 [4, 51], making these methods impractical in clinical practice. Alternatively, tumour behaviour and the Darwinian phylogeny may be predicted from other methods of molecular profiling such as transcriptomic analysis or functional imaging.
Papillary renal cell carcinoma
Papillary RCC (pRCC) represents approximately 20% of all kidney cancers and accumulates mutations at a similar rate to ccRCC [54], again with a predominance of a clock-like process. Signature 2, associated with APOBEC family of cytidine deaminases is the next most common signature. Classified as either type I or type II in roughly equal proportions, pRCC occurs either sporadically or as an inherited form [7]. In general, type I cancers are often multifocal and confer a better prognosis than the more aggressive and typically unifocal type II cancers [55]. Although classified as separate entities, types I and II pRCCs share many molecular features, including chromatin modifications seen in ccRCC [56]. Some of the shared genomic features, such as gene fusions involving TFE3 or TFEB, are present in approximately 10% of samples and show no particular disposition to type I or type II cancers [56]. The functional implication of these events remains unclear [57]. Due to the molecular overlap between both types and the fact that prognostic significance of types I vs. II was not confirmed on multivariable analyses, the clinical utility of papillary type has been questioned [55].
Type I
Hereditary papillary renal cancer (HPRC) predisposes to type I pRCC via autosomal dominant inheritance of a mutation in the MET proto-oncogene [7]. Increased MET mRNA expression is commonly observed in the sporadic form of the disease [56]. This increased expression is potentially directly driven by whole chromosomal copy number aberrations, in particular trisomy 7 which is present in the majority of type I pRCC tumours. In addition, approximately fifteen per cent of sporadic cases harbour activating mutations in the tyrosine kinase domain or contain splice-site variants [56, 58]. The MET pathway interacts with other key oncological pathways such as RAS and PIK3 causes increased angiogenesis and increased cell dissociation and is, therefore, the subject of interest for targeted inhibition. A multicentre phase II study investigated foretinib, an inhibitor of MET, VEGFR2, RON, and AXL tyrosine kinase in sporadic and HPRC-associated papillary RCC [59]. Although overall response rates were moderate, half of patients with germline MET mutations had a partial response. Unfortunately, no other pathological or molecular review was undertaken, but improved treatment stratification by type I or MET mutational status shows promise.
Additional genes recurrently mutated in type I pRCC include KDM6A, SMARCB1 and NFE2L2 [56]. Despite widening the net to discover other candidate driver mutations through known-cancer associated genes, one-third of tumours had no clearly discernible driver, other than trisomy of broad copy number alterations, most commonly chromosome 7.
Type II
The inherited form of the more aggressive, type II pRCC tumours is caused by germline mutation of the gene encoding fumarate hydratase (FH) [7]. Sporadic FH mutations are rarely found; however, mutations of genes in the downstream NRF2–antioxidant response element (ARE) pathway such as NFE2L2 are recurrently detected.
CDKN2A alterations are present in 25% of type II pRCC tumours when loss of heterogeneity, promoter hypermethylation, and somatic mutations are considered together [56]. Increased expression of cell-cycle related genes was seen, most likely via retinoblastoma protein. The presence of CDKN2A alterations was also adversely associated survival in univariate analysis of the whole cohort and when limited to the more aggressive type II phenotype [56].
A small subset of type II pRCC; CpG island methylator phenotype (CIMP) had universal hypermethylation of the CDKN2A promoter and also a high prevalence of germline or somatic mutations in FH [56]. These tumours expressed increased levels of hypoxia-related genes and evidence of a Warburg-like metabolic shift. These effects underpin the rationale to trial agents such as bevacizumab (VEGF inhibition) and erlotinib (TKI), in the phase II setting (NCT01130519).
The chromatin-modifying genes SETD2, BAP1, and PBRM1 are recurrently mutated in the absence of consistent loss of heterozygosity or promoter hypermethylation [56].
Chromophobe renal cell carcinoma
Chromophobe RCC (chRCC) accounts for 5% of renal carcinomas, but this figure is higher amongst young women. It is relatively indolent, although sarcomatoid differentiation renders it highly aggressive [55]. These tumours derived from the distal nephron accumulate mutations at a low rate (~ 0.4 per Mb) [54]. The most characteristic feature is extensive whole chromosomal loss of heterozygosity involving chromosomes 1, 2, 6, 10, 13, 17, and 21 [60]. The TP53 and PTEN genes were recurrently mutated almost exclusively in classic (i.e., non-eosinophilic) variants [60, 61]. Recurrent aberrations in the TERT gene were also detected with some harbouring the canonical 228T mutations, but mainly via structural variants that correlated strongly with increased TERT expression [60]. The eosinophilic subtype, describing an eosinophilic cytoplasm with densely packed mitochondria, harboured cases that were devoid of copy number aberrations and some that were enriched for the mitochondrial MT-ND5 gene mutations [60, 61]. The causal mechanism between this mutation and the histopathological phenotype has not yet been ascertained.
There are little data on genomic correlates with patient outcomes. Sun et al. analysed 66 patients from the TCGA database. TP53 mutations were found in 33% of tumours, while loss of HNF1B was seen in 88%. Prevalence of both TP53 mutations and loss of HNF1b increased with tumour stage and were linked with poor survival [62]. Casuscelli et al. [63] studied genomic outcome correlates of 79 chRCCs of all stages. TP53 mutations (58%), PTEN mutations (24%), and imbalanced chromosome duplication (duplication of ≥ 3 chromosomes, 25%) were enriched in patients with metastatic disease. While each feature was associated with inferior survival, the combination of all three changes yielded the worst prognosis.
Oncocytoma
Oncocytomas are benign tumours that share many features with eosinophilic chRCCs, including derivation from the distal tubule and recurrent mutations in mitochondria-encoded proteins [61]. Type 2 oncocytomas also contains recurrent whole chromosome losses that resemble those seen in eosinophilic chRCC [61]. The absence of PT53 mutations and activation of the p53 pathway in oncocytomas highlights p53 as a barrier to oncocytoma progression.
Conclusions
Scientific literature provides a detailed view of the genomic landscapes for each of the more common renal cancers. The genomic archaeology of clear cell tumours is particularly well characterised through exhaustive multi-regional sequencing. Through this knowledge, there is the potential to better stratify the risk of progression and survival for kidney cancer. Emerging evidence is showing that the presence or the absence of certain mutations may relate to therapeutic sensitivity or resistance. There are a number of gaps in our knowledge; these particularly relate to the behaviour of small renal masses, rarer subtypes of renal cell cancer, and the response of tumours to newer targeted agents.
References
Scelo G, Riazalhosseini Y, Greger L, Letourneau L, Gonzalez-Porta M, Wozniak MB, Bourgey M, Harnden P, Egevad L, Jackson SM, Karimzadeh M, Arseneault M, Lepage P, How-Kit A, Daunay A, Renault V, Blanche H, Tubacher E, Sehmoun J, Viksna J, Celms E, Opmanis M, Zarins A, Vasudev NS, Seywright M, Abedi-Ardekani B, Carreira C, Selby PJ, Cartledge JJ, Byrnes G, Zavadil J, Su J, Holcatova I, Brisuda A, Zaridze D, Moukeria A, Foretova L, Navratilova M, Mates D, Jinga V, Artemov A, Nedoluzhko A, Mazur A, Rastorguev S, Boulygina E, Heath S, Gut M, Bihoreau MT, Lechner D, Foglio M, Gut IG, Skryabin K, Prokhortchouk E, Cambon-Thomsen A, Rung J, Bourque G, Brennan P, Tost J, Banks RE, Brazma A, Lathrop GM (2014) Variation in genomic landscape of clear cell renal cell carcinoma across Europe. Nat Commun 5:5135. https://doi.org/10.1038/ncomms6135
Cancer Genome Atlas Research N (2013) Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 499(7456):43–49. https://doi.org/10.1038/nature12222
Sato Y, Yoshizato T, Shiraishi Y, Maekawa S, Okuno Y, Kamura T, Shimamura T, Sato-Otsubo A, Nagae G, Suzuki H, Nagata Y, Yoshida K, Kon A, Suzuki Y, Chiba K, Tanaka H, Niida A, Fujimoto A, Tsunoda T, Morikawa T, Maeda D, Kume H, Sugano S, Fukayama M, Aburatani H, Sanada M, Miyano S, Homma Y, Ogawa S (2013) Integrated molecular analysis of clear-cell renal cell carcinoma. Nat Genet 45(8):860–867. https://doi.org/10.1038/ng.2699
Turajlic S, Xu H, Litchfield K, Rowan A, Horswell S, Chambers T, O’Brien T, Lopez JI, Watkins TBK, Nicol D, Stares M, Challacombe B, Hazell S, Chandra A, Mitchell TJ, Au L, Eichler-Jonsson C, Jabbar F, Soultati A, Chowdhury S, Rudman S, Lynch J, Fernando A, Stamp G, Nye E, Stewart A, Xing W, Smith JC, Escudero M, Huffman A, Matthews N, Elgar G, Phillimore B, Costa M, Begum S, Ward S, Salm M, Boeing S, Fisher R, Spain L, Navas C, Gronroos E, Hobor S, Sharma S, Aurangzeb I, Lall S, Polson A, Varia M, Horsfield C, Fotiadis N, Pickering L, Schwarz RF, Silva B, Herrero J, Luscombe NM, Jamal-Hanjani M, Rosenthal R, Birkbak NJ, Wilson GA, Pipek O, Ribli D, Krzystanek M, Csabai I, Szallasi Z, Gore M, McGranahan N, Van Loo P, Campbell P, Larkin J, Swanton C, Consortium TRR (2018) Deterministic evolutionary trajectories influence primary tumor growth: TRACERx renal. Cell 173(3):595–610 e511. https://doi.org/10.1016/j.cell.2018.03.043
Alexandrov LB, Jones PH, Wedge DC, Sale JE, Campbell PJ, Nik-Zainal S, Stratton MR (2015) Clock-like mutational processes in human somatic cells. Nat Genet 47(12):1402–1407. https://doi.org/10.1038/ng.3441
Ricketts CJ, Linehan WM (2015) Gender specific mutation incidence and survival associations in clear cell renal cell carcinoma (CCRCC). PLoS One 10(10):e0140257. https://doi.org/10.1371/journal.pone.0140257
Maher ER (2018) Hereditary renal cell carcinoma syndromes: diagnosis, surveillance and management. World J Urol. https://doi.org/10.1007/s00345-018-2288-5 (in press)
Rossi SH, Klatte T, Usher-Smith J, Stewart GD (2018) Epidemiology and screening for renal cancer. World J Urol. https://doi.org/10.1007/s00345-018-2286-7 (in press)
Hung RJ, Moore L, Boffetta P, Feng BJ, Toro JR, Rothman N, Zaridze D, Navratilova M, Bencko V, Janout V, Kollarova H, Szeszenia-Dabrowska N, Mates D, Chow WH, Brennan P (2007) Family history and the risk of kidney cancer: a multicenter case-control study in Central Europe. Cancer Epidemiol Biomark Prev 16(6):1287–1290. https://doi.org/10.1158/1055-9965.EPI-06-0963
Purdue MP, Johansson M, Zelenika D, Toro JR, Scelo G, Moore LE, Prokhortchouk E, Wu X, Kiemeney LA, Gaborieau V, Jacobs KB, Chow WH, Zaridze D, Matveev V, Lubinski J, Trubicka J, Szeszenia-Dabrowska N, Lissowska J, Rudnai P, Fabianova E, Bucur A, Bencko V, Foretova L, Janout V, Boffetta P, Colt JS, Davis FG, Schwartz KL, Banks RE, Selby PJ, Harnden P, Berg CD, Hsing AW, Grubb RL 3rd, Boeing H, Vineis P, Clavel-Chapelon F, Palli D, Tumino R, Krogh V, Panico S, Duell EJ, Quiros JR, Sanchez MJ, Navarro C, Ardanaz E, Dorronsoro M, Khaw KT, Allen NE, Bueno-de-Mesquita HB, Peeters PH, Trichopoulos D, Linseisen J, Ljungberg B, Overvad K, Tjonneland A, Romieu I, Riboli E, Mukeria A, Shangina O, Stevens VL, Thun MJ, Diver WR, Gapstur SM, Pharoah PD, Easton DF, Albanes D, Weinstein SJ, Virtamo J, Vatten L, Hveem K, Njolstad I, Tell GS, Stoltenberg C, Kumar R, Koppova K, Cussenot O, Benhamou S, Oosterwijk E, Vermeulen SH, Aben KK, van der Marel SL, Ye Y, Wood CG, Pu X, Mazur AM, Boulygina ES, Chekanov NN, Foglio M, Lechner D, Gut I, Heath S, Blanche H, Hutchinson A, Thomas G, Wang Z, Yeager M, Fraumeni JF Jr, Skryabin KG, McKay JD, Rothman N, Chanock SJ, Lathrop M, Brennan P (2011) Genome-wide association study of renal cell carcinoma identifies two susceptibility loci on 2p21 and 11q13.3. Nat Genet 43(1):60–65. https://doi.org/10.1038/ng.723
Wu X, Scelo G, Purdue MP, Rothman N, Johansson M, Ye Y, Wang Z, Zelenika D, Moore LE, Wood CG, Prokhortchouk E, Gaborieau V, Jacobs KB, Chow WH, Toro JR, Zaridze D, Lin J, Lubinski J, Trubicka J, Szeszenia-Dabrowska N, Lissowska J, Rudnai P, Fabianova E, Mates D, Jinga V, Bencko V, Slamova A, Holcatova I, Navratilova M, Janout V, Boffetta P, Colt JS, Davis FG, Schwartz KL, Banks RE, Selby PJ, Harnden P, Berg CD, Hsing AW, Grubb RL 3rd, Boeing H, Vineis P, Clavel-Chapelon F, Palli D, Tumino R, Krogh V, Panico S, Duell EJ, Quiros JR, Sanchez MJ, Navarro C, Ardanaz E, Dorronsoro M, Khaw KT, Allen NE, Bueno-de-Mesquita HB, Peeters PH, Trichopoulos D, Linseisen J, Ljungberg B, Overvad K, Tjonneland A, Romieu I, Riboli E, Stevens VL, Thun MJ, Diver WR, Gapstur SM, Pharoah PD, Easton DF, Albanes D, Virtamo J, Vatten L, Hveem K, Fletcher T, Koppova K, Cussenot O, Cancel-Tassin G, Benhamou S, Hildebrandt MA, Pu X, Foglio M, Lechner D, Hutchinson A, Yeager M, Fraumeni JF Jr, Lathrop M, Skryabin KG, McKay JD, Gu J, Brennan P, Chanock SJ (2012) A genome-wide association study identifies a novel susceptibility locus for renal cell carcinoma on 12p11.23. Hum Mol Genet 21(2):456–462. https://doi.org/10.1093/hmg/ddr479
Henrion M, Frampton M, Scelo G, Purdue M, Ye Y, Broderick P, Ritchie A, Kaplan R, Meade A, McKay J, Johansson M, Lathrop M, Larkin J, Rothman N, Wang Z, Chow WH, Stevens VL, Ryan Diver W, Gapstur SM, Albanes D, Virtamo J, Wu X, Brennan P, Chanock S, Eisen T, Houlston RS (2013) Common variation at 2q22.3 (ZEB2) influences the risk of renal cancer. Hum Mol Genet 22(4):825–831. https://doi.org/10.1093/hmg/dds489
Gudmundsson J, Sulem P, Gudbjartsson DF, Masson G, Petursdottir V, Hardarson S, Gudjonsson SA, Johannsdottir H, Helgadottir HT, Stacey SN, Magnusson OT, Helgason H, Panadero A, van der Zanden LF, Aben KK, Vermeulen SH, Oosterwijk E, Kong A, Mayordomo JI, Sverrisdottir A, Jonsson E, Gudbjartsson T, Einarsson GV, Kiemeney LA, Thorsteinsdottir U, Rafnar T, Stefansson K (2013) A common variant at 8q24.21 is associated with renal cell cancer. Nat Commun 4:2776. https://doi.org/10.1038/ncomms3776
Scelo G, Purdue MP, Brown KM, Johansson M, Wang Z, Eckel-Passow JE, Ye Y, Hofmann JN, Choi J, Foll M, Gaborieau V, Machiela MJ, Colli LM, Li P, Sampson JN, Abedi-Ardekani B, Besse C, Blanche H, Boland A, Burdette L, Chabrier A, Durand G, Le Calvez-Kelm F, Prokhortchouk E, Robinot N, Skryabin KG, Wozniak MB, Yeager M, Basta-Jovanovic G, Dzamic Z, Foretova L, Holcatova I, Janout V, Mates D, Mukeriya A, Rascu S, Zaridze D, Bencko V, Cybulski C, Fabianova E, Jinga V, Lissowska J, Lubinski J, Navratilova M, Rudnai P, Szeszenia-Dabrowska N, Benhamou S, Cancel-Tassin G, Cussenot O, Baglietto L, Boeing H, Khaw KT, Weiderpass E, Ljungberg B, Sitaram RT, Bruinsma F, Jordan SJ, Severi G, Winship I, Hveem K, Vatten LJ, Fletcher T, Koppova K, Larsson SC, Wolk A, Banks RE, Selby PJ, Easton DF, Pharoah P, Andreotti G, Freeman LEB, Koutros S, Albanes D, Mannisto S, Weinstein S, Clark PE, Edwards TL, Lipworth L, Gapstur SM, Stevens VL, Carol H, Freedman ML, Pomerantz MM, Cho E, Kraft P, Preston MA, Wilson KM, Michael Gaziano J, Sesso HD, Black A, Freedman ND, Huang WY, Anema JG, Kahnoski RJ, Lane BR, Noyes SL, Petillo D, Teh BT, Peters U, White E, Anderson GL, Johnson L, Luo J, Buring J, Lee IM, Chow WH, Moore LE, Wood C, Eisen T, Henrion M, Larkin J, Barman P, Leibovich BC, Choueiri TK, Mark Lathrop G, Rothman N, Deleuze JF, McKay JD, Parker AS, Wu X, Houlston RS, Brennan P, Chanock SJ (2017) Genome-wide association study identifies multiple risk loci for renal cell carcinoma. Nat Commun 8:15724. https://doi.org/10.1038/ncomms15724
Gnarra JR, Tory K, Weng Y, Schmidt L, Wei MH, Li H, Latif F, Liu S, Chen F, Duh FM et al (1994) Mutations of the VHL tumour suppressor gene in renal carcinoma. Nat Genet 7(1):85–90. https://doi.org/10.1038/ng0594-85
Mitchell TJ, Turajlic S, Rowan A, Nicol D, Farmery JHR, O’Brien T, Martincorena I, Tarpey P, Angelopoulos N, Yates LR, Butler AP, Raine K, Stewart GD, Challacombe B, Fernando A, Lopez JI, Hazell S, Chandra A, Chowdhury S, Rudman S, Soultati A, Stamp G, Fotiadis N, Pickering L, Au L, Spain L, Lynch J, Stares M, Teague J, Maura F, Wedge DC, Horswell S, Chambers T, Litchfield K, Xu H, Stewart A, Elaidi R, Oudard S, McGranahan N, Csabai I, Gore M, Futreal PA, Larkin J, Lynch AG, Szallasi Z, Swanton C, Campbell PJ, Consortium TRR (2018) Timing the landmark events in the evolution of clear cell renal cell cancer: TRACERx renal. Cell 173 (3):611–623 e617. https://doi.org/10.1016/j.cell.2018.02.020
Smits KM, Schouten LJ, van Dijk BA, Hulsbergen-van de Kaa CA, Wouters KA, Oosterwijk E, van Engeland M, van den Brandt PA (2008) Genetic and epigenetic alterations in the von hippel-lindau gene: the influence on renal cancer prognosis. Clin Cancer Res 14(3):782–787. https://doi.org/10.1158/1078-0432.ccr-07-1753
Patard JJ, Fergelot P, Karakiewicz PI, Klatte T, Trinh QD, Rioux-Leclercq N, Said JW, Belldegrun AS, Pantuck AJ (2008) Low CAIX expression and absence of VHL gene mutation are associated with tumor aggressiveness and poor survival of clear cell renal cell carcinoma. Int J Cancer 123(2):395–400. https://doi.org/10.1002/ijc.23496
Dalgliesh GL, Furge K, Greenman C, Chen L, Bignell G, Butler A, Davies H, Edkins S, Hardy C, Latimer C, Teague J, Andrews J, Barthorpe S, Beare D, Buck G, Campbell PJ, Forbes S, Jia M, Jones D, Knott H, Kok CY, Lau KW, Leroy C, Lin ML, McBride DJ, Maddison M, Maguire S, McLay K, Menzies A, Mironenko T, Mulderrig L, Mudie L, O’Meara S, Pleasance E, Rajasingham A, Shepherd R, Smith R, Stebbings L, Stephens P, Tang G, Tarpey PS, Turrell K, Dykema KJ, Khoo SK, Petillo D, Wondergem B, Anema J, Kahnoski RJ, Teh BT, Stratton MR, Futreal PA (2010) Systematic sequencing of renal carcinoma reveals inactivation of histone modifying genes. Nature 463(7279):360–363. https://doi.org/10.1038/nature08672
Varela I, Tarpey P, Raine K, Huang D, Ong CK, Stephens P, Davies H, Jones D, Lin ML, Teague J, Bignell G, Butler A, Cho J, Dalgliesh GL, Galappaththige D, Greenman C, Hardy C, Jia M, Latimer C, Lau KW, Marshall J, McLaren S, Menzies A, Mudie L, Stebbings L, Largaespada DA, Wessels LF, Richard S, Kahnoski RJ, Anema J, Tuveson DA, Perez-Mancera PA, Mustonen V, Fischer A, Adams DJ, Rust A, Chan-on W, Subimerb C, Dykema K, Furge K, Campbell PJ, Teh BT, Stratton MR, Futreal PA (2011) Exome sequencing identifies frequent mutation of the SWI/SNF complex gene PBRM1 in renal carcinoma. Nature 469(7331):539–542. https://doi.org/10.1038/nature09639
Guo G, Gui Y, Gao S, Tang A, Hu X, Huang Y, Jia W, Li Z, He M, Sun L, Song P, Sun X, Zhao X, Yang S, Liang C, Wan S, Zhou F, Chen C, Zhu J, Li X, Jian M, Zhou L, Ye R, Huang P, Chen J, Jiang T, Liu X, Wang Y, Zou J, Jiang Z, Wu R, Wu S, Fan F, Zhang Z, Liu L, Yang R, Liu X, Wu H, Yin W, Zhao X, Liu Y, Peng H, Jiang B, Feng Q, Li C, Xie J, Lu J, Kristiansen K, Li Y, Zhang X, Li S, Wang J, Yang H, Cai Z, Wang J (2012) Frequent mutations of genes encoding ubiquitin-mediated proteolysis pathway components in clear cell renal cell carcinoma. Nat Genet 44(1):17–19. https://doi.org/10.1038/ng.1014
Li J, Duns G, Westers H, Sijmons R, van den Berg A, Kok K (2016) SETD2: an epigenetic modifier with tumor suppressor functionality. Oncotarget. https://doi.org/10.18632/oncotarget.9368
Carvalho S, Vitor AC, Sridhara SC, Martins FB, Raposo AC, Desterro JM, Ferreira J, de Almeida SF (2014) SETD2 is required for DNA double-strand break repair and activation of the p53-mediated checkpoint. eLife 3:e02482. https://doi.org/10.7554/elife.02482
Pena-Llopis S, Vega-Rubin-de-Celis S, Liao A, Leng N, Pavia-Jimenez A, Wang S, Yamasaki T, Zhrebker L, Sivanand S, Spence P, Kinch L, Hambuch T, Jain S, Lotan Y, Margulis V, Sagalowsky AI, Summerour PB, Kabbani W, Wong SW, Grishin N, Laurent M, Xie XJ, Haudenschild CD, Ross MT, Bentley DR, Kapur P, Brugarolas J (2012) BAP1 loss defines a new class of renal cell carcinoma. Nat Genet 44(7):751–759. https://doi.org/10.1038/ng.2323
Kapur P, Pena-Llopis S, Christie A, Zhrebker L, Pavia-Jimenez A, Rathmell WK, Xie XJ, Brugarolas J (2013) Effects on survival of BAP1 and PBRM1 mutations in sporadic clear-cell renal-cell carcinoma: a retrospective analysis with independent validation. Lancet Oncol 14(2):159–167. https://doi.org/10.1016/S1470-2045(12)70584-3
Hakimi AA, Ostrovnaya I, Reva B, Schultz N, Chen YB, Gonen M, Liu H, Takeda S, Voss MH, Tickoo SK, Reuter VE, Russo P, Cheng EH, Sander C, Motzer RJ, Hsieh JJ, cc RCCCGARNi (2013) Adverse outcomes in clear cell renal cell carcinoma with mutations of 3p21 epigenetic regulators BAP1 and SETD2: a report by MSKCC and the KIRC TCGA research network. Clin Cancer Res 19(12):3259–3267. https://doi.org/10.1158/1078-0432.CCR-12-3886
Hsieh JJ, Chen D, Wang PI, Marker M, Redzematovic A, Chen YB, Selcuklu SD, Weinhold N, Bouvier N, Huberman KH, Bhanot U, Chevinsky MS, Patel P, Pinciroli P, Won HH, You D, Viale A, Lee W, Hakimi AA, Berger MF, Socci ND, Cheng EH, Knox J, Voss MH, Voi M, Motzer RJ (2016) Genomic biomarkers of a randomized trial comparing first-line everolimus and sunitinib in patients with metastatic renal cell carcinoma. Eur Urol. https://doi.org/10.1016/j.eururo.2016.10.007
Miao D, Margolis CA, Gao W, Voss MH, Li W, Martini DJ, Norton C, Bosse D, Wankowicz SM, Cullen D, Horak C, Wind-Rotolo M, Tracy A, Giannakis M, Hodi FS, Drake CG, Ball MW, Allaf ME, Snyder A, Hellmann MD, Ho T, Motzer RJ, Signoretti S, Kaelin WG Jr, Choueiri TK, Van Allen EM (2018) Genomic correlates of response to immune checkpoint therapies in clear cell renal cell carcinoma. Science 359(6377):801–806. https://doi.org/10.1126/science.aan5951
Hosen I, Rachakonda PS, Heidenreich B, Sitaram RT, Ljungberg B, Roos G, Hemminki K, Kumar R (2015) TERT promoter mutations in clear cell renal cell carcinoma. Int J Cancer 136(10):2448–2452. https://doi.org/10.1002/ijc.29279
Wang K, Liu T, Liu L, Liu J, Liu C, Wang C, Ge N, Ren H, Yan K, Hu S, Bjorkholm M, Fan Y, Xu D (2014) TERT promoter mutations in renal cell carcinomas and upper tract urothelial carcinomas. Oncotarget 5(7):1829–1836. https://doi.org/10.18632/oncotarget.1829
Papa A, Wan L, Bonora M, Salmena L, Song MS, Hobbs RM, Lunardi A, Webster K, Ng C, Newton RH, Knoblauch N, Guarnerio J, Ito K, Turka LA, Beck AH, Pinton P, Bronson RT, Wei W, Pandolfi PP (2014) Cancer-associated PTEN mutants act in a dominant-negative manner to suppress PTEN protein function. Cell 157(3):595–610. https://doi.org/10.1016/j.cell.2014.03.027
Lee HJ, Lee HY, Lee JH, Lee H, Kang G, Song JS, Kang J, Kim JH (2014) Prognostic significance of biallelic loss of PTEN in clear cell renal cell carcinoma. J Urol 192(3):940–946. https://doi.org/10.1016/j.juro.2014.03.097
Kang SA, Pacold ME, Cervantes CL, Lim D, Lou HJ, Ottina K, Gray NS, Turk BE, Yaffe MB, Sabatini DM (2013) mTORC1 phosphorylation sites encode their sensitivity to starvation and rapamycin. Science 341(6144):1236566. https://doi.org/10.1126/science.1236566
Kwiatkowski DJ, Choueiri TK, Fay AP, Rini BI, Thorner AR, de Velasco G, Tyburczy ME, Hamieh L, Albiges L, Agarwal N, Ho TH, Song J, Pignon JC, Barrios PM, Michaelson MD, Van Allen EM, Krajewski KM, Porta C, Pal SK, Bellmunt J, McDermott DF, Heng DY, Gray KP, Signoretti S (2016) Mutations in TSC1, TSC2, and MTOR are associated with response to rapalogs in patients with metastatic renal cell carcinoma. Clin Cancer Res 22(10):2445–2452. https://doi.org/10.1158/1078-0432.CCR-15-2631
Voss MH, Hakimi AA, Pham CG, Brannon AR, Chen YB, Cunha LF, Akin O, Liu H, Takeda S, Scott SN, Socci ND, Viale A, Schultz N, Sander C, Reuter VE, Russo P, Cheng EH, Motzer RJ, Berger MF, Hsieh JJ (2014) Tumor genetic analyses of patients with metastatic renal cell carcinoma and extended benefit from mTOR inhibitor therapy. Clin Cancer Res 20(7):1955–1964. https://doi.org/10.1158/1078-0432.CCR-13-2345
de Velasco G, Wankowicz SA, Madison R, Ali SM, Norton C, Duquette A, Ross JS, Bosse D, Lalani AA, Miller VA, Stephens PJ, Young L, Hakimi AA, Signoretti S, Pal SK, Choueiri TK (2018) Targeted genomic landscape of metastases compared to primary tumours in clear cell metastatic renal cell carcinoma. Br J Cancer 118(9):1238–1242. https://doi.org/10.1038/s41416-018-0064-3
Gulati S, Martinez P, Joshi T, Birkbak NJ, Santos CR, Rowan AJ, Pickering L, Gore M, Larkin J, Szallasi Z, Bates PA, Swanton C, Gerlinger M (2014) Systematic evaluation of the prognostic impact and intratumour heterogeneity of clear cell renal cell carcinoma biomarkers. Eur Urol 66(5):936–948. https://doi.org/10.1016/j.eururo.2014.06.053
Turajlic S, Xu H, Litchfield K, Rowan A, Chambers T, Lopez JI, Nicol D, O’Brien T, Larkin J, Horswell S, Stares M, Au L, Jamal-Hanjani M, Challacombe B, Chandra A, Hazell S, Eichler-Jonsson C, Soultati A, Chowdhury S, Rudman S, Lynch J, Fernando A, Stamp G, Nye E, Jabbar F, Spain L, Lall S, Guarch R, Falzon M, Proctor I, Pickering L, Gore M, Watkins TBK, Ward S, Stewart A, DiNatale R, Becerra MF, Reznik E, Hsieh JJ, Richmond TA, Mayhew GF, Hill SM, McNally CD, Jones C, Rosenbaum H, Stanislaw S, Burgess DL, Alexander NR, Swanton C, Peace, Consortium TRR (2018) Tracking cancer evolution reveals constrained routes to metastases: TRACERx renal. Cell 173(3):581–594 e512. https://doi.org/10.1016/j.cell.2018.03.057
El-Mokadem I, Fitzpatrick J, Bondad J, Rauchhaus P, Cunningham J, Pratt N, Fleming S, Nabi G (2014) Chromosome 9p deletion in clear cell renal cell carcinoma predicts recurrence and survival following surgery. Br J Cancer 111(7):1381–1390. https://doi.org/10.1038/bjc.2014.420
Shen C, Beroukhim R, Schumacher SE, Zhou J, Chang M, Signoretti S, Kaelin WG Jr (2011) Genetic and functional studies implicate HIF1alpha as a 14q kidney cancer suppressor gene. Cancer Discov 1(3):222–235. https://doi.org/10.1158/2159-8290.CD-11-0098
Kroeger N, Klatte T, Chamie K, Rao PN, Birkhauser FD, Sonn GA, Riss J, Kabbinavar FF, Belldegrun AS, Pantuck AJ (2013) Deletions of chromosomes 3p and 14q molecularly subclassify clear cell renal cell carcinoma. Cancer 119(8):1547–1554. https://doi.org/10.1002/cncr.27947
Klatte T, Kroeger N, Rampersaud EN, Birkhauser FD, Logan JE, Sonn G, Riss J, Rao PN, Kabbinavar FF, Belldegrun AS, Pantuck AJ (2012) Gain of chromosome 8q is associated with metastases and poor survival of patients with clear cell renal cell carcinoma. Cancer 118(23):5777–5782. https://doi.org/10.1002/cncr.27607
Klatte T, Rao PN, de Martino M, LaRochelle J, Shuch B, Zomorodian N, Said J, Kabbinavar FF, Belldegrun AS, Pantuck AJ (2009) Cytogenetic profile predicts prognosis of patients with clear cell renal cell carcinoma. J Clin Oncol 27(5):746–753. https://doi.org/10.1200/JCO.2007.15.8345
Zhang J, Lu A, Li L, Yue J, Lu Y (2010) p16 modulates VEGF expression via its interaction with HIF-1alpha in breast cancer cells. Cancer Invest 28(6):588–597. https://doi.org/10.3109/07357900903286941
Bedke J, Gauler T, Grünwald V, Hegele A, Herrmann E, Hinz S, Janssen J, Schmitz S, Schostak M, Tesch H, Zastrow S, Miller K (2017) Systemic therapy in metastatic renal cell carcinoma. World J Urol 35(2):179–188. https://doi.org/10.1007/s00345-016-1868-5
Motzer RJ, Escudier B, McDermott DF, George S, Hammers HJ, Srinivas S, Tykodi SS, Sosman JA, Procopio G, Plimack ER, Castellano D, Choueiri TK, Gurney H, Donskov F, Bono P, Wagstaff J, Gauler TC, Ueda T, Tomita Y, Schutz FA, Kollmannsberger C, Larkin J, Ravaud A, Simon JS, Xu LA, Waxman IM, Sharma P, CheckMate I (2015) Nivolumab versus everolimus in advanced renal-cell carcinoma. N Engl J Med 373(19):1803–1813. https://doi.org/10.1056/NEJMoa1510665
Rosenberg JE, Hoffman-Censits J, Powles T, van der Heijden MS, Balar AV, Necchi A, Dawson N, O’Donnell PH, Balmanoukian A, Loriot Y, Srinivas S, Retz MM, Grivas P, Joseph RW, Galsky MD, Fleming MT, Petrylak DP, Perez-Gracia JL, Burris HA, Castellano D, Canil C, Bellmunt J, Bajorin D, Nickles D, Bourgon R, Frampton GM, Cui N, Mariathasan S, Abidoye O, Fine GD, Dreicer R (2016) Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet 387(10031):1909–1920. https://doi.org/10.1016/S0140-6736(16)00561-4
Maia MC, Almeida L, Bergerot PG, Dizman N, Pal SK (2018) Relationship of tumor mutational burden (TMB) to immunotherapy response in metastatic renal cell carcinoma (mRCC). J Clin Oncol 36(6_suppl):662. https://doi.org/10.1200/jco.2018.36.6_suppl.662
Gerlinger M, Rowan AJ, Horswell S, Larkin J, Endesfelder D, Gronroos E, Martinez P, Matthews N, Stewart A, Tarpey P, Varela I, Phillimore B, Begum S, McDonald NQ, Butler A, Jones D, Raine K, Latimer C, Santos CR, Nohadani M, Eklund AC, Spencer-Dene B, Clark G, Pickering L, Stamp G, Gore M, Szallasi Z, Downward J, Futreal PA, Swanton C (2012) Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med 366(10):883–892. https://doi.org/10.1056/NEJMoa1113205
Soultati A, Stares M, Swanton C, Larkin J, Turajlic S (2015) How should clinicians address intratumour heterogeneity in clear cell renal cell carcinoma? Curr Opin Urol 25(5):358–366. https://doi.org/10.1097/mou.0000000000000204
Sankin A, Hakimi AA, Mikkilineni N, Ostrovnaya I, Silk MT, Liang Y, Mano R, Chevinsky M, Motzer RJ, Solomon SB, Cheng EH, Durack JC, Coleman JA, Russo P, Hsieh JJ (2014) The impact of genetic heterogeneity on biomarker development in kidney cancer assessed by multiregional sampling. Cancer Med 3(6):1485–1492. https://doi.org/10.1002/cam4.293
Gerlinger M, Horswell S, Larkin J, Rowan AJ, Salm MP, Varela I, Fisher R, McGranahan N, Matthews N, Santos CR, Martinez P, Phillimore B, Begum S, Rabinowitz A, Spencer-Dene B, Gulati S, Bates PA, Stamp G, Pickering L, Gore M, Nicol DL, Hazell S, Futreal PA, Stewart A, Swanton C (2014) Genomic architecture and evolution of clear cell renal cell carcinomas defined by multiregion sequencing. Nat Genet 46(3):225–233. https://doi.org/10.1038/ng.2891
Knudson AG Jr (1971) Mutation and cancer: statistical study of retinoblastoma. Proc Natl Acad Sci USA 68(4):820–823
Alexandrov LB, Nik-Zainal S, Wedge DC, Aparicio SA, Behjati S, Biankin AV, Bignell GR, Bolli N, Borg A, Borresen-Dale AL, Boyault S, Burkhardt B, Butler AP, Caldas C, Davies HR, Desmedt C, Eils R, Eyfjord JE, Foekens JA, Greaves M, Hosoda F, Hutter B, Ilicic T, Imbeaud S, Imielinski M, Jager N, Jones DT, Jones D, Knappskog S, Kool M, Lakhani SR, Lopez-Otin C, Martin S, Munshi NC, Nakamura H, Northcott PA, Pajic M, Papaemmanuil E, Paradiso A, Pearson JV, Puente XS, Raine K, Ramakrishna M, Richardson AL, Richter J, Rosenstiel P, Schlesner M, Schumacher TN, Span PN, Teague JW, Totoki Y, Tutt AN, Valdes-Mas R, van Buuren MM, van ‘t Veer L, Vincent-Salomon A, Waddell N, Yates LR, Australian Pancreatic Cancer Genome I, Consortium IBC, Consortium IM-S, PedBrain I, Zucman-Rossi J, Futreal PA, McDermott U, Lichter P, Meyerson M, Grimmond SM, Siebert R, Campo E, Shibata T, Pfister SM, Campbell PJ, Stratton MR (2013) Signatures of mutational processes in human cancer. Nature 500(7463):415–421. https://doi.org/10.1038/nature12477
Klatte T, Rossi SH, Stewart GD (2018) Prognostic factors and prognostic models for renal cell carcinoma: a literature review. World J Urol. https://doi.org/10.1007/s00345-018-2309-4 (in press)
Cancer Genome Atlas Research N, Linehan WM, Spellman PT, Ricketts CJ, Creighton CJ, Fei SS, Davis C, Wheeler DA, Murray BA, Schmidt L, Vocke CD, Peto M, Al Mamun AA, Shinbrot E, Sethi A, Brooks S, Rathmell WK, Brooks AN, Hoadley KA, Robertson AG, Brooks D, Bowlby R, Sadeghi S, Shen H, Weisenberger DJ, Bootwalla M, Baylin SB, Laird PW, Cherniack AD, Saksena G, Haake S, Li J, Liang H, Lu Y, Mills GB, Akbani R, Leiserson MD, Raphael BJ, Anur P, Bottaro D, Albiges L, Barnabas N, Choueiri TK, Czerniak B, Godwin AK, Hakimi AA, Ho TH, Hsieh J, Ittmann M, Kim WY, Krishnan B, Merino MJ, Mills Shaw KR, Reuter VE, Reznik E, Shelley CS, Shuch B, Signoretti S, Srinivasan R, Tamboli P, Thomas G, Tickoo S, Burnett K, Crain D, Gardner J, Lau K, Mallery D, Morris S, Paulauskis JD, Penny RJ, Shelton C, Shelton WT, Sherman M, Thompson E, Yena P, Avedon MT, Bowen J, Gastier-Foster JM, Gerken M, Leraas KM, Lichtenberg TM, Ramirez NC, Santos T, Wise L, Zmuda E, Demchok JA, Felau I, Hutter CM, Sheth M, Sofia HJ, Tarnuzzer R, Wang Z, Yang L, Zenklusen JC, Zhang J, Ayala B, Baboud J, Chudamani S, Liu J, Lolla L, Naresh R, Pihl T, Sun Q, Wan Y, Wu Y, Ally A, Balasundaram M, Balu S, Beroukhim R, Bodenheimer T, Buhay C, Butterfield YS, Carlsen R, Carter SL, Chao H, Chuah E, Clarke A, Covington KR, Dahdouli M, Dewal N, Dhalla N, Doddapaneni HV, Drummond JA, Gabriel SB, Gibbs RA, Guin R, Hale W, Hawes A, Hayes DN, Holt RA, Hoyle AP, Jefferys SR, Jones SJ, Jones CD, Kalra D, Kovar C, Lewis L, Li J, Ma Y, Marra MA, Mayo M, Meng S, Meyerson M, Mieczkowski PA, Moore RA, Morton D, Mose LE, Mungall AJ, Muzny D, Parker JS, Perou CM, Roach J, Schein JE, Schumacher SE, Shi Y, Simons JV, Sipahimalani P, Skelly T, Soloway MG, Sougnez C, Tam A, Tan D, Thiessen N, Veluvolu U, Wang M, Wilkerson MD, Wong T, Wu J, Xi L, Zhou J, Bedford J, Chen F, Fu Y, Gerstein M, Haussler D, Kasaian K, Lai P, Ling S, Radenbaugh A, Van Den Berg D, Weinstein JN, Zhu J, Albert M, Alexopoulou I, Andersen JJ, Auman JT, Bartlett J, Bastacky S, Bergsten J, Blute ML, Boice L, Bollag RJ, Boyd J, Castle E, Chen YB, Cheville JC, Curley E, Davies B, DeVolk A, Dhir R, Dike L, Eckman J, Engel J, Harr J, Hrebinko R, Huang M, Huelsenbeck-Dill L, Iacocca M, Jacobs B, Lobis M, Maranchie JK, McMeekin S, Myers J, Nelson J, Parfitt J, Parwani A, Petrelli N, Rabeno B, Roy S, Salner AL, Slaton J, Stanton M, Thompson RH, Thorne L, Tucker K, Weinberger PM, Winemiller C, Zach LA, Zuna R (2016) Comprehensive molecular characterization of papillary renal-cell carcinoma. N Engl J Med 374(2):135–145. https://doi.org/10.1056/nejmoa1505917
Kauffman EC, Ricketts CJ, Rais-Bahrami S, Yang Y, Merino MJ, Bottaro DP, Srinivasan R, Linehan WM (2014) Molecular genetics and cellular features of TFE3 and TFEB fusion kidney cancers. Nat Rev Urol 11(8):465–475. https://doi.org/10.1038/nrurol.2014.162
Kovac M, Navas C, Horswell S, Salm M, Bardella C, Rowan A, Stares M, Castro-Giner F, Fisher R, de Bruin EC, Kovacova M, Gorman M, Makino S, Williams J, Jaeger E, Jones A, Howarth K, Larkin J, Pickering L, Gore M, Nicol DL, Hazell S, Stamp G, O’Brien T, Challacombe B, Matthews N, Phillimore B, Begum S, Rabinowitz A, Varela I, Chandra A, Horsfield C, Polson A, Tran M, Bhatt R, Terracciano L, Eppenberger-Castori S, Protheroe A, Maher E, El Bahrawy M, Fleming S, Ratcliffe P, Heinimann K, Swanton C, Tomlinson I (2015) Recurrent chromosomal gains and heterogeneous driver mutations characterise papillary renal cancer evolution. Nat Commun 6:6336. https://doi.org/10.1038/ncomms7336
Choueiri TK, Vaishampayan U, Rosenberg JE, Logan TF, Harzstark AL, Bukowski RM, Rini BI, Srinivas S, Stein MN, Adams LM, Ottesen LH, Laubscher KH, Sherman L, McDermott DF, Haas NB, Flaherty KT, Ross R, Eisenberg P, Meltzer PS, Merino MJ, Bottaro DP, Linehan WM, Srinivasan R (2013) Phase II and biomarker study of the dual MET/VEGFR2 inhibitor foretinib in patients with papillary renal cell carcinoma. J Clin Oncol 31(2):181–186. https://doi.org/10.1200/JCO.2012.43.3383
Davis CF, Ricketts CJ, Wang M, Yang L, Cherniack AD, Shen H, Buhay C, Kang H, Kim SC, Fahey CC, Hacker KE, Bhanot G, Gordenin DA, Chu A, Gunaratne PH, Biehl M, Seth S, Kaipparettu BA, Bristow CA, Donehower LA, Wallen EM, Smith AB, Tickoo SK, Tamboli P, Reuter V, Schmidt LS, Hsieh JJ, Choueiri TK, Hakimi AA, Cancer Genome Atlas Research N, Chin L, Meyerson M, Kucherlapati R, Park WY, Robertson AG, Laird PW, Henske EP, Kwiatkowski DJ, Park PJ, Morgan M, Shuch B, Muzny D, Wheeler DA, Linehan WM, Gibbs RA, Rathmell WK, Creighton CJ (2014) The somatic genomic landscape of chromophobe renal cell carcinoma. Cancer Cell 26(3):319–330. https://doi.org/10.1016/j.ccr.2014.07.014
Joshi S, Tolkunov D, Aviv H, Hakimi AA, Yao M, Hsieh JJ, Ganesan S, Chan CS, White E (2015) The genomic landscape of renal oncocytoma identifies a metabolic barrier to tumorigenesis. Cell Rep 13(9):1895–1908. https://doi.org/10.1016/j.celrep.2015.10.059
Sun M, Tong P, Kong W, Dong B, Huang Y, Park IY, Zhou L, Liu XD, Ding Z, Zhang X, Bai S, German P, Powell R, Wang Q, Tong X, Tannir NM, Matin SF, Rathmell WK, Fuller GN, McCutcheon IE, Walker CL, Wang J, Jonasch E (2017) HNF1B loss exacerbates the development of chromophobe renal cell carcinomas. Cancer Res 77(19):5313–5326. https://doi.org/10.1158/0008-5472.can-17-0986
Casuscelli J, Weinhold N, Gundem G, Wang L, Zabor EC, Drill E, Wang PI, Nanjangud GJ, Redzematovic A, Nargund AM, Manley BJ, Arcila ME, Donin NM, Cheville JC, Thompson RH, Pantuck AJ, Russo P, Cheng EH, Lee W, Tickoo SK, Ostrovnaya I, Creighton CJ, Papaemmanuil E, Seshan VE, Hakimi AA, Hsieh JJ (2017) Genomic landscape and evolution of metastatic chromophobe renal cell carcinoma. JCI Insight. https://doi.org/10.1172/jci.insight.92688
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144(5):646–674. https://doi.org/10.1016/j.cell.2011.02.013
Acknowledgements
We would like to acknowledge The Urology Foundation, who kindly provided a research grant for SHR.
Author information
Authors and Affiliations
Contributions
TJM project development, data collection, data analysis, and manuscript writing and editing. SHR data analysis and manuscript editing. TK data analysis and manuscript editing. GDS data analysis and manuscript editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no relevant conflict of interest.
Research involving human participants and/or animals
The following manuscript is a review of existing data. Therefore, this article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
For this type of study (review), formal consent is not required.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Mitchell, T.J., Rossi, S.H., Klatte, T. et al. Genomics and clinical correlates of renal cell carcinoma. World J Urol 36, 1899–1911 (2018). https://doi.org/10.1007/s00345-018-2429-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00345-018-2429-x