Abstract
Shifts in the microbiome of the intertidal coral Coelastrea aspera (formally known as Goniastrea aspera) from Phuket, Thailand, were noted over the course of a 4-d period of spring tides. During this time, corals were naturally exposed to high temperatures, intense solar radiation, sub-aerial exposure and tidally induced water fluxes. Analysis of the 16S microbiome highlighted that the corals harbored both ‘core or stable’ communities and those which appeared to be more ‘transient or sporadic.’ Only relatively few microbial associates were classified as core microbes; the majority were transient or sporadic. Such transient associates were likely to have been governed by tidally induced variations in mucus thickness and water fluxes. Here we report strong shifts in the bacterial community of C. aspera over a short temporal scale. However, we also show significant differences in the timing of shifts between the two age groups of corals studied. More rapid changes (within 2 d of sub-aerial exposure) occurred within the 4-yr-old colonies, but a slightly delayed response was observed in the 10-yr-old colonies, whereby the microbial associates only changed after 4 d. We hypothesize that these shifts are age related and could be influenced by the observed baseline differences in the microbiome of the 4- and 10-yr-old corals, bacteria–bacteria interactions, and/or host energetics.
Similar content being viewed by others
Explore related subjects
Find the latest articles, discoveries, and news in related topics.Avoid common mistakes on your manuscript.
Introduction
An emerging paradigm of evolutionary theory is that animals and plants should not be considered autonomous entities but rather as biomolecular networks comprising the host together with their associated microbiota (McFall-Ngai et al. 2013). Reef corals are a good example of such a network since they harbor an extensive microbial community. In assessing the limits of the ability of corals to adapt to changing environmental conditions, it is important to understand the mechanisms of co-existence and co-evolution of such communities, often collectively referred to as the ‘holobiont.’ In the past, reef corals have been viewed primarily as a symbiosis between the host animal and the symbiotic unicellular algae (Symbiodinium spp.) (Smith and Douglas 1987), and many studies have focused on the ‘shuffling’ and ‘switching’ of algal types in response to changing thermal environments (van Oppen et al. 2009). However, recently it has been recognized that, in addition to the algal symbionts, corals also host a highly diverse and specific microbiome which includes bacteria, archaea, fungi, protists and viruses (Rohwer et al. 2002). Much less is known about this coral microbiome, particularly with respect to the dynamics of microbial partners through coral development (Thompson et al. 2014), aging (Williams et al. 2015) and their responses to changing environments (Morrow et al. 2012; Hester et al. 2015; Glasl et al. 2016; Röthig et al. 2016).
Much of the current literature examining the microbiome under changing environmental conditions is dominated by studies that have described effects of elevated sea surface temperature (SST) (Ritchie 2006; Rosenberg et al. 2007). In these studies, higher SST led to a reduction in the proportion of the microbial community that produced antimicrobial compounds, that is the microbes that inhibit the growth of potential pathogens. The surface microbial community (SMC) has also been shown to rapidly switch from a healthy to a diseased state under elevated SST, with particular rapidly growing pathogenic bacteria favored (Bourne et al. 2009; Ritchie et al. 2010). In another example, Reshef et al. (2006), using the Vibrio shiloi/Oculina patagonica system, showed that some coral species can adapt rapidly to changing thermal conditions by altering their bacterial community, with symbiotic ‘naturally associated’ bacteria acting to inhibit the growth of other bacterial strains that may be detrimental to coral health.
Few studies have assessed the dynamic changes in coral microbial communities of healthy corals; Koren and Rosenberg (2006) showed that the bacterial community of O. patagonica appeared to vary seasonally. In other studies, coral bacterial associates (Ceh et al. 2012) and bacterio-plankton communities near corals in Kaneohe Bay, Hawaii (Apprill and Rappé 2011), appeared to remain relatively stable before and after coral spawning. Interestingly, in the latter work, slight changes in bacterial load and microbial community composition in the water column were only observed during low tide conditions, leading the authors to conclude that tidal flow and turbulence were likely to be important factors controlling microbial dynamics on coral reefs. A more recent study by Glasl et al. (2016) has also shown that shifts in the composition of the SMC can occur over a period of days, as the coral mucus aged with the prevalence of opportunistic and potentially pathogenic bacteria increasing in abundance in the aging mucus. However, after the release of the aged mucus the microbial community rapidly reverted to its original state. Thus, coral microbial communities are dynamic in both space and time and processes such as cyclic mucus shedding and the tide are important.
In this study, we investigated changes in the microbiome of the intertidal coral Coelastrea aspera over the course of a tidal cycle at a site in Thailand where the tidal range is large (~3 m at springs) and where early afternoon aerial exposure can lead to major solar stresses on the western sides of colonies that face toward the sun (Brown et al. 1994). Such exposure causes severe oxidative damage within coral tissues during extreme low tides (Brown et al. 2002). In recent work, we have also highlighted age-related changes in the microbiome of C. aspera, with the highest microbial diversity in 4- to 12-yr-old corals (Williams et al. 2015). Here we describe the community responses of the coral microbiome in two age classes (4-yr-old and 10-yr-old colonies) over a 4-d spring-tide sequence.
Methods
Field site and sampling regime
The study site is in the Andaman Sea on the west coast of Thailand on the southeast tip of the island of Phuket (7°50′N, 98°25.5′E). This location and its fringing intertidal reefs have been described in detail (Scoffin et al. 1992; Brown et al. 2011). At this site, all colonies of the merulinid coral C. aspera are subaerially exposed to air at low water on the inner reef flat for 2–3 h, and none are found in tidal pools or fully submerged. The large tidal range (>3 m) means that during spring tides, waters leave the reef flat very rapidly (within 20 min), and during the dry season salinity effects induced by the tidal regime are therefore negligible, with the salinity of surface waters varying between 31 and 34‰ (Charuchinda and Hylleberg 1984). Two age classes of C. aspera (as outlined in Williams et al. 2015) were identified on the reef flat to compare microbial communities throughout a tidal cycle in March 2014. Colony ages were determined through alizarin staining, skeleton chronology and by following the demography of this coral population from their initial settlement in the early 1990s (Brown et al. 2014; Williams et al. 2015). The age classes selected were ~4-yr-old and ~10-yr-old corals (Williams et al. 2015). The 4-yr-old corals were hemispherical colonies showing no partial mortality, while the 10-yr-old corals were micro-atolls with ~30–40% mortality. A 10-yr-old colony with approximately 30% partial mortality has about five times the living surface area of a 4-yr-old coral (Babcock 1991).
The age classes were chosen because of their proximity on the intertidal reef flat. All collections were made in an area ~20 m × 20 m at a similar tidal height. Interactions with other organisms and sediment have been shown to cause the onset of coral disease (Nugues et al. 2004; Carlos et al. 2013; Sweet et al. 2013), and sediments have even been proposed as a potential ‘seed bank’ for certain microbial associates of corals (Carlos et al. 2013). With this in mind, we carefully selected colonies to avoid potential contamination from these sources. Each sampled colony was isolated from other colonies by dead reef substrate, and there was no macroalgae adjacent to or close to the individual colonies sampled.
Samples for microbial analysis were taken from the western side of colonies in each of the two age classes using 1.5-cm-diameter hole punches that were carefully cleaned and sterilized between samples. The cores were taken from the middle of the colony, surrounded by living tissue that had no direct contact with the surrounding sediment. Great care was also taken to ensure that the plugs were extracted at similar heights above the reef substrate. A single plug sample was extracted from 48 different colonies at three separate times (Day 0, Day 2 and Day 4) over a 4-d tidal sequence between 28 March and 1 April 2014, resulting in eight replicates per colony for each age group each day. Different colonies were sampled to avoid potential confounding effects due to the effect of sampling itself.
Day 0 sampling was carried out at low tide on the afternoon of 28 March when corals were not aerially exposed. Day 2 sampling was carried out on 30 March, when corals had been aerially exposed in the afternoon for two consecutive days. Day 4 sampling was carried out on 1 April when corals had been aerially exposed in the afternoon for four consecutive days. After collection, all the coral cores were immediately stored in 100% molecular grade ethanol in separate sterile 50-mL Falcon tubes for transport back to the laboratory where they were extracted using Qiagen DNeasy Blood and Tissue Kit following the manufacturers guidelines (Williams et al. 2015).
Physical measurements throughout the tidal cycle
The temperature at the surface of the coral, bulk seawater temperature, solar radiation and tidal height were measured during the sampling period to quantify the major stresses associated with the ebb and flow of the tide.
Coral temperature
The temperature at the surface of a hemispherical colony of C. aspera (approx. 200 mm diameter) was monitored from 16 March to 1 April 2014, encompassing both the sampling period and a spring and neap tide prior to sampling. A 22-mm-diameter × 130-mm-length core angled at 45° from the horizontal was removed from the west-facing surface of a colony using a core drill. A probe from a Seamon mini temperature recorder (factory and laboratory calibrated to ±0.05 °C) was inserted into the coral surface and the core hole finished with silicone sealant. The temperature was logged every 5 min.
Sea water temperature
Bulk sea water temperature was monitored every 30 min using two type U Betatherm 2 kΩ thermistors (absolute accuracy ±0.2 °C) from a long-term station at 2 m below chart datum on a jetty approximately 2 km from the study site. Temperature data were collected using a factory-calibrated Grant Squirrel 2020 logger (±0.05% readings, +0.025% range) and cross-calibrated with the Seamon mini used for the coral temperature measurements.
Solar radiation
Photosynthetically active radiation (PAR, 400–700 nm, μmol m−2 s−1) was recorded on a horizontal plane every minute and logged at 30-min intervals using a 2π cosine-corrected quantum sensor (Macam Photometrics) approximately 1.5 km from the study site. This in situ quantum sensor was cross-calibrated with factory-calibrated quantum sensors prior to the experiment.
Tidal height
Tidal height was obtained from a Royal Thai Navy tide station (Ko Taphao Noi, 07°49.9′N, 98°25.5′E) 1.5 km from the study site. Since 2005, the primary source of tide records has been from a radar gauge recording at 3-min intervals. Hourly data (fast delivery) were used and downloaded from the University of Hawaii Sea Level Center (http://uhslc.soest.hawaii.edu/) for the period prior to and during the experiment.
The tides at Phuket are semi-diurnal with a range of 0.6 m (neap tides) to 3.1 m (spring tides) with relatively little variation in range between successive high and low waters (Ditlev 1978). Because of this pattern, the typical 14-d cyclical increase and decrease in range, coupled with the tidal period, results in low water of the spring tides consistently falling either in the early morning or in the afternoon. It is only on low spring tides that the intertidal coral reefs at Phuket are exposed to air. There is also a seasonal component to sub-aerial exposure with total exposure time greatest on afternoon tides in February–April and on early morning tides in January–February. This seasonal component is in part the result of the timing of the tide and in part due to the marked seasonal depression by up to 20 cm of the mean sea level at this time of year (see Brown et al. 1994).
Bacterial community analysis of coral samples
High-throughput 454 pyrosequencing was conducted on all samples. Bacterial 16S rRNA gene diversity was amplified using standard prokaryotic primers 27F (5′-AGAGTTTGATCMTGGCTCAG-3′) and 518R (5′-ATTACCGCGGCTGCTGG-3′). PCR protocol, product cleaning, quantification and pooling were the same as Williams et al. (2015). Sequences were run on a 454 FLX Titanium pico-titer plate at Newgene in the Centre for Life, Newcastle, UK. Technical control samples were processed by transferring the ethanol used for preservation of the coral samples into Falcon tubes then extracting and sequencing in the same way as the samples. No DNA was detectable either after PCR or from the downstream processing, and so we concluded that all sequences retrieved were true representatives of the coral microbiome (Williams et al. 2015).
Pyrosequences were processed using the QIIME pipeline (version 1.5.0) in the same way as Williams et al. (2015). From the 48 individual samples, a total of 789,643 raw nucleotide reads were produced with an average length of 56 bp, corresponding to 179 Mb. After filtering, a total of 456,322 quality reads were acquired. The length of the remaining sequences varied from 151 to 491 bp, with an average length of 400 bp. All metadata have been submitted to GenBank under the SRA submission accession number SRP100822.
In QIIME, each of the samples was rarefied to the sample that exhibited the lowest number of reads (597 sequences) for both taxonomy and diversity analyses. Rarefaction curves were generated for Chao1, phylogenetic diversity and observed species. Alpha rarefaction analyses were performed by computing the average richness metric value from five subplots for each of the samples (Turlapati et al. 2013). Analysis showed that the distributions of the diversity indices were substantially skewed and variances were heterogeneous among groupings (ages and days).
An analysis of similarity (ANOSIM) was used to test differences in 16S rRNA gene bacterial assemblage using PRIMER 6.0 (Clarke and Gorley 2006). Patterns of the 16S rRNA gene bacterial assemblages were represented on a non-metric multidimensional scaling (nMDS) plot. All remaining analyses were conducted in the statistical programming language R (R Core Team 2016). Abundance data were square root transformed and normalized to relative abundances prior to multivariate analysis to satisfy assumptions of homogeneity of variance. Bray–Curtis dissimilarity matrices were used to generate an nMDS plot (using the isoMDS function in the ‘vegan’ library; Oksanen et al. 2016). nMDS plots were also developed for data collected on each day of sampling to illustrate the relative changes in community patterns with time. Differences in microbial community variability for coral age–day combinations were assessed as described by Hester et al. (2015). The variation within each microbial community was estimated by embedding the dissimilarity measures (Bray–Curtis distance matrix) within Euclidean space, resulting in the two-dimensional distances between individual samples approximating compositional dissimilarity. This also enabled visualization of host microbial community variability and grouping within principal coordinate space. Group centroids were then calculated for each coral age–day combination (i.e., across replicates), and each replicate’s distance to its respective group centroid was calculated. The sample-centroid distances were used to test for differences in variability among groups (coral age–day combinations) with permutation tests using the ‘betadisper’ function in the ‘vegan’ library (Oksanen et al. 2016). Permutational MANOVA (PERMANOVA; Anderson and Walsh 2013) was used for nonparametric assessment of multivariate differences in microbial community composition among coral age–day combinations based on relative abundances of the microbial communities. Permutation tests of differences in community variability demonstrated that the assumption of homogeneity of dispersions in PERMANOVA was met. Pairwise comparison tests between coral age–day combinations were conducted, with Bonferroni corrections applied to compensate for the increased risk of type I errors.
We also adopted a Monte Carlo approach for assessing differences in diversity indices among groups, following Roff (2006). Differences in diversity between ~4- and ~10-yr-old corals (ignoring potential effects of day) were assessed by randomly reassigning the diversity indices to each age group and calculating the difference between the resulting mean diversities. This was repeated 10,000 times to generate a distribution based on the null hypothesis of no difference in diversity between the groups. The actual difference in mean diversity was then calculated and compared with this distribution to estimate the associated p value. This was done for each diversity index. The process was then repeated to compare diversities between days (pairwise) within coral age groups.
Results
Physical measurements
Bulk sea water temperature, coral temperature and solar irradiance
Bulk sea water temperature from 16 March to 1 April warmed by ~1.2 °C. Superimposed on this, there was a cyclical diurnal warming in the afternoon and cooling at night (range 2 °C) under the influence of solar radiation (Fig. 1a).
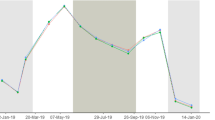
Physical parameters recorded on the southeast tip of Phuket, Thailand (7°50′N, 98°25.5′E), between a 16 March to 1 April 2014, and b the coral sampling period 27 March to 1 April 2014 with coral temperature record (black line), bulk sea water temperature (blue line) and solar irradiance (magenta line). Green dots represent the times when the water level fell to expose the reef flat and temperature recorder and when they were re-submerged. Red dots represent sampling times
The coral temperature record displayed a similar overall warming of ~1.7 °C throughout the period. The dynamic range of ~15 °C (24.33–39.48 °C) between the highest and lowest temperature experienced by the coral was likely driven by (1) the influence of warming due to solar heating of the sea water on the shallow reef flat as the tide fell during the middle of the day, and the direct heating effect of solar irradiance on the western side of the coral during sub-aerial exposure, and (2) cooling during periods of sub-aerial exposure at night. The largest daily range (25.73–39.48 °C = 13.75 °C) was recorded on 16 March 2014, during the first spring-tide period, while the smallest range (27.8–30.93 °C = 3.13 °C) was during the neap tide on 23 March 2014 (Fig. 1a).
The period from 16 March to 1 April was one of generally clear sky days with solar irradiance peaking at solar noon (Fig. 1a). Clouds reduced irradiance around midday on 31 March and throughout most of 1 April (Fig. 1b).
Coral temperature during tidal exposure
A pattern of coral temperature variation can be seen in the more detailed temperature record between 27 March and 1 April (Fig. 1). This pattern involved an overnight cooling of coral temperature and bulk sea temperature of ~0.5 °C. During low spring tides when the coral was subaerially exposed in the morning, the coral temperature cooled rapidly by up to 4 °C following exposure, possibly because air temperature was lower than the sea temperature and/or due to evaporative cooling. Once re-submerged, the coral temperature quickly returned to that of the seawater over the reef flat which was warmer than the bulk sea temperature by ~1 °C.
During the daytime, the temperature of submerged coral rose steadily until early afternoon, due to solar radiation heating the increasingly shallow water covering the reef flat. This pattern was invariant between neap and spring tides, although the magnitude of the warming was greater as the water depth over the reef flat reduced during spring tides. During the sampling period, the coral experienced high prolonged temperatures on the afternoon spring tide of 27 March, between 1410 and 1700 hrs when, although the reef flat was not exposed, the reef flat seawater temperature was above 34 °C for a 3-h period. On 28 March, the reef flat was not exposed but temperatures stayed above 34 °C from 1410 to 1635 hrs, peaking at 36.88 °C. On the following day when corals were aerially exposed between 1510 and 1645 hrs, temperatures were again above 34 °C with a maximum of 35.13 °C. Physical conditions were extreme on the afternoon of the first day of aerial exposure with the west sides of corals exposed to high air temperatures, high solar radiation and desiccation stresses. Similar conditions were encountered on the second day of afternoon aerial exposure (Day 2). However, on days following the spring-tide sequence, physical stresses in the afternoon decreased with the later timing of low tide and reduced solar radiation (Fig. 1b).
Bacterial diversity in coral samples
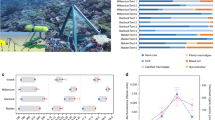
In total, 3183 OTUs were detected across the 48 samples at a sequence depth of 597. These consisted of 173 unique phylotypes (Table 1; Electronic supplementary material, ESM Table S1). Of these, 30 phylotypes were found in ≥30% of all the samples regardless of age (Fig. 2). Splitting the corals by age meant that the number of phylotypes found across all samples of the same group increased to 32 for ~4-yr-old colonies and 35 for ~10-yr-old colonies (Fig. 2). Using a more stringent cutoff of ≥70% (i.e., bacteria present in 70% or more of the samples assessed), the number of phylotypes dropped to 16, found across all samples regardless of age. This increased to 19 in ~4-yr-old colonies, but dropped to 13 in ~10-yr-old colonies (Fig. 2). Using a cutoff of ≥90%, the core microbes were further refined. At this level, only eight phylotypes were consistently found regardless of age (Fig. 2). These included phylotypes identified as Rothia (B46; ESM Table S1); Propionibacterium (B54); Streptococcus (B163); Neisseria (B265); Methylobacterium (B213); Pseudomonas (B303) and two from the Phylum Cyanobacterium (B125, B138) (Table 2).
From these phylotypes, only five were found in all samples regardless of age or time period sampled (i.e., found in 100% of the samples assessed; Fig. 2). These included Pseudomonas (B303), two cyanobacteria (B125, B138), Propionibacterium (B54) and Rothia (B46). Again, this number was higher when we assessed only ~4-yr-old coral colonies (i.e., 8 phylotypes), which included the addition of Streptococcus (B163), Methylobacterium (B213) and Kiloniellales (B202). However, only five members were present in 10-yr-old colonies. Interestingly, there were no specific or obvious patterns in relative abundances of these core bacteria across the different age classes or days sampled (Table 3; ESM Table S1).
There were no significant differences between ~4- and ~10-yr-old corals in any of the diversity indices for total bacterial diversity (ignoring potential effects of day) (Table 4; ESM Fig. S1), although note that all p values were marginal. There was also no variation in the overall community associated with particular coral age–day combinations (Fig. 3; ESM Fig. S1a). Making allowance for the differentiation of data points according to day and age, there were significant differences in the microbial community composition between both day (pseudo-F = 3.72, p = 0.014) and age (pseudo-F = 1.95, p < 0.001) (ESM Fig. S1b, c, d). Pairwise analyses, corrected for potential increase in type I errors, showed that this significance held true for all comparisons (Day 0 vs Day 2: pseudo-F = 3.34, p = 0.003; Day 0 vs Day 4: pseudo-F = 4.28, p = 0.003; Day 2 vs Day 4: pseudo-F = 3.68, p = 0.003).
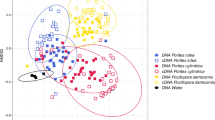
a Variability in bacterial communities associated with particular coral age–day combinations of Coelastrea aspera, based on Bray–Curtis dissimilarity distances embedded in Euclidean space to estimate compositional dissimilarity visualized in principal coordinate space. b Boxplot of the Euclidean distances of samples from their respective centroid based on all samples of coral age–day combinations. Black line is the median, box boundaries represent upper and lower inter-quartile ranges, whiskers represent either the most extreme points or one and half times the inter-quartile range, dots are points beyond the one and half inter-quartile range c Principal coordinates analysis (PCoA) for ~4-yr-old colonies only and d PCoA for ~10-yr-old colonies. Samples were taken from two ages of corals (~4-yr-old or ~10-yr-old colonies), at three different times (Day 0, Day 2 and Day 4)
ANOSIM analysis corroborated these differences for ~4-yr-old colonies (Figs. 3c, 4a). However, for ~10-yr-old colonies there were significant differences only for Day 0 versus Day 4 and Day 2 versus Day 4 (p < 0.003), and no significance was found between Day 0 and Day 2 (p = 0.119; Figs. 3d, 4b).
Interestingly, comparisons of the diversity between days within ages indicated that more significant differences occurred in the ~10-yr-old corals than the ~4-yr-old corals (Table 4). Only the comparison of the alpha diversity between Day 0 (lower) and Day 2 (higher) was significant at the 0.05 level within the ~4-yr-old corals (Table 4). This was also true for the ~10-yr-old corals and was supported by a corresponding significant difference in the Chao index (Table 4).
Due to the complex patterns identified, and the indication that age is important when assessing the microbiome of corals, at least for C. aspera, the rest of the results are reported by age class.
Four-year-old corals
The microbial communities associated with the corals separated into clear groups by day (Fig. 4a). Evenness was highest at Day 0 and decreased throughout the tidal period (Day 2 to Day 4) (Table 1). In contrast, alpha diversity and species richness (Chao1) were lowest on Day 0, peaked during Day 2 and reduced again on Day 4 (Table 1). The full list of bacterial communities detected and their relative abundances are shown in ESM Table S1. Here we outline the specific phylotypes which appear to be responsible for the shifts in profile between time periods.
In general, specific phylotypes were present during some periods and absent or rare in others. Some bacteria in ~4-yr-old coral colonies (Corynebacterium sp. and a member of the Family Oxalobacteraceae) showed a marked decline between Day 0 and Day 2, whereas the remainder showed an increased frequency between Day 0 and Day 2 (Table 5). By Day 4, three bacteria retained a high frequency of occurrence in coral colonies (Fulvivirga sp., a member of the Phylum Cyanobacterium and one from the Family Amoebophilaceae), while the remainder either reverted to a low incidence or to the frequency observed on Day 0.
Ten-year-old corals
In contrast to younger corals, the bacterial communities associated with ~10-yr-old colonies clustered into only two clear groups. Those from Day 0 and Day 2 showed no clear separation but were separate from Day 4 (Figs. 3d, 4b). Evenness was highest on Day 4 and lowest on Day 2 (Table 1). Alpha diversity increased throughout the sampling days (Table 1) while species richness (Chao1) showed a similar pattern to that of ~4-yr-old colonies, peaking on Day 2 (Table 1). In the ~10-yr-old corals, diversity was also higher on Day 4 than on Day 2 (log H′) and greater on Day 4 than on Day 0 (Alpha and log H′) (Table 1).
Five of the 11 phylotypes highlighted in Table 6 increased in frequency in ~10-yr-old corals, five decreased in frequency, and one remained stable between Day 2 and Day 4. Those that increased in frequency included one member of each of the genera Veillonella and Fulvivirga, one member of each of the Families Spirochaetaceae and Piscirickettsiaceae, and an unidentified member of the class Alphaprotebacterium. Those decreasing in frequency included one member each from the genera Kocuria and Granulicatella, one each from the Families Amoebophilaceae, Flavobacteriaceae and Phyllobacteriaceae, and an unidentified member from the Phylum Cyanobacterium. All phylotypes showed a similar frequency of occurrence in the individual colonies throughout the sampling period.
Discussion
This study shows that the microbiota of corals can be dynamic over short and long temporal scales, with major changes in the microbiome of C. aspera over a 4-d spring-tide period. The physical conditions experienced by corals at the study site during low spring tides included elevated temperature, intense solar radiation, sub-aerial exposure and extreme tidal water movements. Temperature and solar radiation effects were maximal during the first two days of sub-aerial exposure, and their negative effects on the physiology of the western sides of colonies have been previously documented in C. aspera at this site (Brown and Dunne 2008; Brown et al. 2002). The effects of sub-aerial exposure on increased mucus production have also been highlighted (Jatkar et al. 2010). A temporal study of waterborne bacteria over the reef flat at Heron Island on the GBR revealed no tidal influence (Sweet et al. 2010). However, the reef setting in the present study is in marked contrast to the clear waters of Heron Island. The study site is subject to very turbid waters and strong sediment fluxes on large spring tides (Scoffin et al. 1997) similar to other sedimentary environments (de Jonge and van Beusekom 1995). Moreover, C. aspera experiences intimate contact with sediments, particularly during large tidal water movements on and off the reef flat (Scoffin et al. 1997).
All of the above physical factors have the potential to affect the composition of coral-associated bacteria. High temperatures and solar radiation are known to induce oxidative stress in C. aspera with high levels of oxidative damage and increased antioxidant enzyme levels (CuZnSOD) being produced under such conditions (Brown et al. 2002). It has been suggested that high reactive oxygen concentrations are a barrier to microbial colonization of intracellular sites and that they may also play a role in colonization of micro-organisms at other locations in the coral where active oxygen is produced (Thompson et al. 2014). The production of mucus in response to sub-aerial exposure is another important factor affecting colonization by bacterial communities. Mucus thickness in C. aspera colonies increased by almost 50% during the course of sub-aerial exposure during a series of low spring tides at the site in an earlier study (Jatkar et al. 2010), an effect which could potentially influence the accessibility and dynamics of more transient members of the coral’s microbial assemblages. In addition, extensive mucus flocs are evident on the flooding tide (BE Brown pers. obs.) which could also affect the coral microbiome. In the present study, the frequency of mucus-associated bacteria such as Granulicatella and Veillonella (Pride et al. 2012) increased in ~10-yr-old corals on Days 2 and 4 during the spring-tide period. Fluctuations in other bacteria commonly associated with the water column, sediment or fish and shellfish farms, suggest that water fluxes during this period may also have played an important role in modulating the coral microbial community. The nearby coastline supports both oyster and fish farms, and two of the bacteria detected with a high frequency on Day 2 in ~4-yr-old corals have been formerly associated with shellfish farms (Fulvivirga) and fish diseases (Piscirickettsiaceae) (Nedashkovskaya et al. 2007; Rozas and Enríquez 2014).
It has been proposed that the microbiome can be separated into microbes that can be classed as ‘transient’ and more sporadic versus those that are more stable or sometimes referred to as ‘core’ members (Ainsworth et al. 2015; Hester et al. 2015; Sweet and Bulling 2017). A large proportion of the bacterial phylotypes found in this study fall into the category of ‘transient’ members (i.e., only present in relatively few samples with little to no consistency among replicate colonies). Interestingly, there were marked differences in the timing of changes in ‘transient’ bacteria between ~4- and ~10-yr-old corals during the spring-tide period, despite the fact that both age classes were living adjacent to each other and samples were extracted from colonies at the same height above the substrate. Microbial associates of the younger corals changed rapidly by Day 2 of the tidal sequence in response to altered environmental conditions, but those of older corals remained relatively stable until Day 4, changing only when challenging physical conditions had started to ameliorate. Explanations for such observations may involve variations in the initial composition of the more stable or ‘core’ microbiome of the different age classes (Williams et al. 2015), bacteria–bacteria interactions (Long and Azam 2001; Reshef et al. 2006; Ritchie 2006), and host energetics (Anthony et al. 2002). Coelastrea aspera reaches maximum reproductive output at 4–5 yr, and this continues without decline for older colonies (ESM Fig. S2). Older corals (>5 yr) are therefore likely to have higher reproductive costs than colonies aged 4 yr or younger, i.e., the younger colonies would be less fecund (ESM Fig. S2). Additional energetic costs in older corals are incurred from sediment shedding (Junjie et al. 2014) and lesion healing as colonies grow and adopt a micro-atoll growth form (Scoffin et al. 1997). The mucus layer is a critical, dynamic habitat for surface-colonizing bacteria (Ritchie 2006; Nakajima et al. 2009), and such factors may have a profound effect on the energy available for mucus production. Although these explanations are plausible and consistent with current evidence, they remain largely speculative, as the microbiome of the water column adjacent to the corals was not sampled in this study. This was because the original purpose of the study was to determine whether there were rapid changes in the coral microbiome during large tidal water exchanges but not to identify the origins of transient members of the microbiome. However, having established that such rapid changes in microbial community structure occur, it will be important for future studies to map the flow of microbes between corals and the local environment to understand drivers of microbial community dynamics.
The dominant, consistent and stable microbial associates of C. aspera (i.e., ‘core’ members of the microbiome) contained significantly fewer microbes than those classed as transients. However, the exact definition of a ‘core microbiome’ is currently relatively arbitrary and lacks consistency among studies (reviewed in Sweet and Bulling 2017). Yet the definition can have important consequences when comparing studies. For example, the ‘core’ microbiome of the human gut ranges from 57 to 75 shared microbial species depending on the cutoff used (90 and 50%, respectively; Turnbaugh and Gordon 2009). However, in the same study there was no single abundant bacterial species (defined as >0.5% of the community) shared by all 154 individuals studied. The core microbiome of sponges has also recently been characterized and, again, core members depended on the cutoff used (Schmitt et al. 2012). At 97% cutoff only three core phylotypes were identified, increasing to eight when a 95% cutoff was used, and 18 at 90% (Schmitt et al. 2012). In this study, only five phylotypes were identified as core using the 90% cutoff (Pseudomonas, two cyanobacteria, Propionibacterium and Rothia). However, this number increased to eight phylotypes in the ~4-yr-old coral colonies.
Few studies have attempted to split the microbial community of corals into transient and core members. However, Ainsworth et al. (2015), for example, showed that the core microbiome of Acropora granulosa consisted of 159 phylotypes from a total of 1508 detected (10.5%), while the Leptoseris spp. microbiome was dominated by 204 phylotypes from 1424 (14.3%) and Montipora capitata had a core microbiome consisting of 350 phylotypes out of 1433 (24.4%). These values for ‘core’ members are considerably higher than the examples of other organisms given above. However, Ainsworth et al. (2015) used a 30% cutoff for describing the core coral microbiome (i.e., for a bacterial phylotype to be described as a core member it had to be in at least 30% of all samples). If we use the same cutoff, the percentage of core members in the community (with relation to the total number of phylotypes sequenced) is similar, i.e., there is an increase to 32 out of 162 (19.8%) and 34 out of 175 (19.4%) in ~4- and ~10-yr-old colonies, respectively. This example illustrates the issues associated with the core microbiome concept. Other studies have argued that defining microbiomes as either ‘stable’ and/or ‘sporadic’ symbiotic communities would be more beneficial (Hester et al. 2015; Glasl et al. 2016). Regardless of the exact definition, with an increase in studies reporting changes in microbial communities, a meaningful consensus on definitions may emerge if consistencies can be found between systems. Interestingly, we identified two phylotypes (Ralstonia and Pseudomonas) that warrant specific attention. Ralstonia has been highlighted as a ‘core’ member in many other coral species and has been shown to be associated with coral host cells containing dinoflagellates (Ainsworth et al. 2015). Although this phylotype was dominant and stable in the majority of corals sampled in this study, it would have not fitted into the classification of ‘core’ based on the 90% cutoff and so may have been missed if we were solely reliant on the importance of core associates. In contrast, Pseudomonas was highlighted as a core member across both age groups regardless of day sampled. Interestingly, members of this genus have also been described as core microbes associated with other organisms such as the nematode Caenorhabditis elegans (Dirksen et al. 2016). In nematodes, the bacterium has been linked to host health, acting as an effective barrier against fungal pathogens (Dirksen et al. 2016) and this bacterial genus may play a similar role in corals.
Although the significance of the ‘core’ microbiome of corals and its relationship with more ‘transient’ members of the holobiont remains to be fully understood, there are striking parallels between the results of this study and those found in other systems (Shade and Handelsman 2012; Li et al. 2013; Dirksen et al. 2016). These consistencies may have particular relevance to the link between the stability of the microbiome and its role in overall health status. If the ‘core’ biome is disrupted, for example during periods of stress, ‘transient’ microbiota may then be able to establish and increase in abundance, albeit temporarily. Such transitions may mark a key moment when normally transient potentially pathogenic organisms become established and diseases begin to manifest themselves (Cho and Blaser 2012; Krediet et al. 2013). Indeed, such shifts are routinely noted in the majority of studies assessing the microbiota of both healthy and diseased coral tissues at any given time (reviewed in Bourne et al. 2009).
While it is now recognized that coral-associated microbiota are likely to be linked to the capacity of corals to adapt to changing environmental conditions (Thompson et al. 2014), we have little understanding of either the timescales or the processes involved. The current study and related work (Thompson et al. 2014; Williams et al. 2015) show that coral microbial diversity can shift on temporal scales ranging from days (within a tidal cycle) to years (when comparing different age classes of the same coral species). While at this stage we may only speculate on the specific drivers involved and the underlying mechanisms responsible for such changes, this study highlights the highly dynamic relationships between the coral microbiome and its surrounding environment.
References
Ainsworth T, Krause L, Bridge T, Torda G, Raina J-B, Zakrzewski M, Gates RD, Padilla-Gamiño JL, Spalding HL, Smith C, Woolsey ES, Bourne DG, Bongaerts P, Hoegh-Guldberg O, Leggat W (2015) The coral core microbiome identifies rare bacterial taxa as ubiquitous endosymbionts. ISME J 9:2261–2274
Anderson MJ, Walsh DC (2013) PERMANOVA, ANOSIM, and the Mantel test in the face of heterogeneous dispersions: what null hypothesis are you testing? Ecol Monogr 83:557–574
Anthony KRN, Connolly SR, Willis BL (2002) Comparative analysis of energy allocation to tissue and skeletal growth in corals. Limnol Oceanogr 47:1417–1429
Apprill A, Rappé MS (2011) Response of the microbial community to coral spawning in lagoon and reef flat environments of Hawaii, USA. Aquat Microb Ecol 62:251–266
Babcock RC (1991) Comparative demography of three species of scleractinian corals using age- and size-dependent classifications. Ecol Monogr 61:225–244
Bourne DG, Garren M, Work TM, Rosenberg E, Smith GW, Harvell CD (2009) Microbial disease and the coral holobiont. Trends Microbiol 17:554–562
Brown BE, Dunne RP (2008) Solar radiation modulates bleaching and damage protection in a shallow water coral. Mar Ecol Prog Ser 362:99–107
Brown BE, Dunne RP, Scoffin TP, Letissier MDA (1994) Solar damage in intertidal corals. Mar Ecol Prog Ser 105:219–230
Brown BE, Downs CA, Dunne RP, Gibb SW (2002) Exploring the basis of thermotolerance in the reef coral Goniastrea aspera. Mar Ecol Prog Ser 242:119–129
Brown BE, Dunne RP, Phongsuwan N, Somerfield PJ (2011) Increased sea level promotes coral cover on shallow reef flats in the Andaman Sea, eastern Indian Ocean. Coral Reefs 30:867–878
Brown BE, Dunne RP, Phongsuwan N, Patchim L, Hawkridge JM (2014) The reef coral Goniastrea aspera: a “winner” becomes a “loser” during a severe bleaching event in Thailand. Coral Reefs 33:395–401
Carlos C, Torres TT, Ottoboni LMM (2013) Bacterial communities and species-specific associations with the mucus of Brazilian coral species. Sci Rep 3:1–7
Ceh J, Raina JB, Soo RM, van Keulen M, Bourne DG (2012) Coral–bacterial communities before and after a coral mass spawning event on Ningaloo Reef. PLoS One 7:e36920
Charuchinda M, Hylleberg J (1984) Skeletal extension of Acropora formosa at a fringing reef in the Andaman Sea. Coral Reefs 3:215–219
Cho I, Blaser M (2012) The human microbiome: at the interface of health and disease. Nat Rev Genet 13:260–270
Clarke KR, Gorley RN (2006) PRIMER v6: user manual/tutorial. PRIMER-E Ltd, Plymouth, UK
de Jonge VN, van Beusekom JEE (1995) Wind- and tide-induced resuspension of sediment and microphytobenthos from tidal flats in the Ems estuary. Limnol Oceanogr 40:776–778
Dirksen P, Marsh SA, Braker I, Heitland N, Wagner S, Nakad R, Mader S, Petersen C, Kowallik V, Rosenstiel P, Félix M-A, Schulenburg H (2016) The native microbiome of the nematode Caenorhabditis elegans: gateway to a new host–microbiome model. BMC Biol 14:38
Ditlev H (1978) Zonation of corals (Scleractinia: Coelenterata) on intertidal reef flats at Ko Phuket, Eastern Indian Ocean. Mar Biol 47:29–39
Glasl B, Herndl GJ, Frade PR (2016) The microbiome of coral surface mucus has a key role in mediating holobiont health and survival upon disturbance. ISME J 10:2280–2292
Hester ER, Barott KL, Nulton J, Vermeij MJ, Rohwer FL (2015) Stable and sporadic symbiotic communities of coral and algal holobionts. ISME J 10:1157–1169
Jatkar AA, Brown BE, Bythell JC, Guppy R, Morris NJ, Pearson JP (2010) Coral mucus: the properties of its constituent mucins. Biomacromolecules 11:883–888
Junjie RK, Browne NK, Erftemeijer PLA, Todd PA (2014) Impacts of sediments on coral energetics: partitioning the effects of turbidity and settling particles. PLoS One 9:e107195
Koren O, Rosenberg E (2006) Bacteria associated with mucus and tissues of the coral Oculina patagonica in summer and winter. Appl Environ Microbiol 72:5254–5259
Krediet CJ, Ritchie KB, Paul VJ, Teplitski M (2013) Coral-associated micro-organisms and their roles in promoting coral health and thwarting diseases. Proc R Soc Lond B Biol Sci 280:20122328
Li K, Bihan M, Methe BA (2013) Analyses of the stability and core taxonomic memberships of the human microbiome. PLoS One 8:e63139
Long RA, Azam F (2001) Antagonistic interactions among marine bacteria. Appl Environ Microbiol 67:4875–4983
McFall-Ngai M, Hadfield MG, Bosch TCG, Carey HV, Domazet-Lošo T, Douglas AE, Dubilier N, Eberl G, Fukami T, Gilbert SF, Hentschel U, King N, Kjelleberg S, Knoll AH, Kremer N, Mazmanian SK, Metcalf JL, Nealson K, Pierce NE, Rawls JF, Reid A, Ruby EG, Rumpho M, Sanders JG, Tautz D, Wernegreen JJ (2013) Animals in a bacterial world, a new imperative for the life sciences. Proc Natl Acad Sci U S A 110:3229–3236
Morrow KM, Moss AG, Chadwick NE, Liles MR (2012) Bacterial associates of two Caribbean coral species reveal species-specific distribution and geographic variability. Appl Environ Microbiol 78:6438–6449
Nakajima R, Yoshida T, Azman BAR, Zaleha K, Othman BHR, Toda T (2009) In situ release of coral mucus by Acropora and its influence on the heterotrophic bacteria. Aquat Ecol 43:815–823
Nedashkovskaya OI, Kim SB, Shin DS, Beleneva IA, Mikhailov VV (2007) Fulvivirga kasyanovii gen. nov., sp. nov., a novel member of the phylum Bacteroidetes isolated from seawater in a mussel farm. Int J Syst Evol Microbiol 57:1046–1049
Nugues MM, Smith GW, van Hooidonk RJ, Seabra MI, Bak RPM (2004) Algal contact as a trigger for coral disease. Ecol Lett 7:919–923
Oksanen JF, Blanchet G, Friendly M, Kindt R, Legendre P, McGlinn D, Minchin PR, O’Hara PR, Simpson GL, Solymos P, Stevens H, Szoecs E, Wagner H (2016) vegan: community ecology package. R package version 2.4-0
Pride DT, Salzman J, Haynes M, Rohwer F, Davis-Long C, White RA, Loomer P, Armitage GC, Relman DA (2012) Evidence of a robust resident bacteriophage population revealed through analysis of the human salivary virome. ISME J 6:915–926
R Core Team (2016) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Reshef L, Koren O, Loya Y, Zilber-Rosenberg I, Rosenberg E (2006) The coral probiotic hypothesis. Environ Microbiol 8:2068–2073
Ritchie KB (2006) Regulation of microbial populations by coral surface mucus and mucus-associated bacteria. Mar Ecol Prog Ser 322:1–14
Ritchie KB, Jones LE, Ellner SP, Ritchie KB, Jones LE, Ellner SP (2010) How microbial community composition regulates coral disease development. Plos Biol 8:e1000345
Roff DA (2006) Introduction to computer-intensive methods of data analysis in biology. Cambridge University Press, Cambridge, UK
Rohwer F, Seguritan V, Azam F, Knowlton N (2002) Diversity and distribution of coral-associated bacteria. Mar Ecol Prog Ser 243:1–10
Rosenberg E, Koren O, Reshef L, Efrony R, Zilber-Rosenberg I (2007) The role of microorganisms in coral health, disease and evolution. Nat Rev Microbiol 5:355–362
Röthig T, Ochsenkühn MA, Roik A, van der Merwe R, Voolstra CR (2016) Long-term salinity tolerance is accompanied by major restructuring of the coral bacterial microbiome. Mol Ecol 25:1308–1323
Rozas M, Enríquez R (2014) Piscirickettsiosis and Piscirickettsia salmonis in fish: a review. J Fish Dis 37:163–188
Schmitt S, Tsai P, Bell J, Fromont J, Ilan M, Lindquist N, Perez T, Rodrigo A, Schupp PJ, Vacelet J, Webster N, Hentschel U, Taylor MW (2012) Assessing the complex sponge microbiota: core, variable and species-specific bacterial communities in marine sponges. ISME J 6:564–576
Scoffin TP, Tudhope AW, Brown BE, Chansang H, Cheeney RF (1992) Patterns and possible environmental controls of skeletogenesis of Porites lutea, South Thailand. Coral Reefs 11:1–11
Scoffin TP, Brown BE, Dunne RP, LeTissier MDA (1997) The controls on growth form of intertidal massive corals, Phuket, south Thailand. Palaios 12:237–248
Shade A, Handelsman J (2012) Beyond the Venn diagram: the hunt for a core microbiome. Environ Microbiol 14:4–12
Smith DC, Douglas AE (1987) The biology of symbiosis. Edward Arnold (Publishers) Ltd
Sweet MJ, Bulling MT (2017) On the importance of the microbiome and pathobiome in coral health and disease. Front Mar Sci 4:9
Sweet MJ, Croquer A, Bythell JC (2010) Temporal and spatial patterns in waterborne bacterial communities of an island reef system. Aquat Microb Ecol 61:1–11
Sweet MJ, Bythell JC, Nugues MM (2013) Algae as reservoirs for coral pathogens. PLoS One 8:e69717
Thompson JR, Rivera HE, Closek CJ, Medina M (2014) Microbes in the coral holobiont: partners through evolution, development, and ecological interactions. Front Cell Infect Microbiol 4:176
Turlapati SA, Minocha R, Bhiravarasa PS, Tisa LS, Thomas WK, Minocha SC (2013) Chronic N-amended soils exhibit an altered bacterial community structure in Harvard Forest, MA, USA. FEMS Microbiol Ecol 83:478–493
Turnbaugh PJ, Gordon JI (2009) The core gut microbiome, energy balance and obesity. J Physiol 587:4153–4158
van Oppen M, Baker A, Coffroth M, Willis B (2009) Bleaching resistance and the role of algal endosymbionts. In: van Oppen M, Lough J (eds) Coral bleaching: patterns, processes, causes and consequences. Springer, Berlin, Heidelberg, pp 83–102
Williams AD, Brown BE, Putchim L, Sweet MJ (2015) Age-relateds shifts in bacterial diversity in a reef coral. PLoS One 10:e0144902
Acknowledgements
We acknowledge the Leverhulme Trust for support of BEB and the Director and staff of the Phuket Marine Biological Center for their continued assistance in our research. Finally, we thank the reviewers and the topic editor for their thoughtful, detailed and constructive feedback which has greatly improved this manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Biology Editor Dr. Line K. Bay
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Sweet, M.J., Brown, B.E., Dunne, R.P. et al. Evidence for rapid, tide-related shifts in the microbiome of the coral Coelastrea aspera . Coral Reefs 36, 815–828 (2017). https://doi.org/10.1007/s00338-017-1572-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00338-017-1572-y