Abstract
Design and decision-making for marine protected areas (MPAs) on coral reefs require prediction of MPA effects with population models. Modeling of MPAs has shown how the persistence of metapopulations in systems of MPAs depends on the size and spacing of MPAs, and levels of fishing outside the MPAs. However, the pattern of demographic connectivity produced by larval dispersal is a key uncertainty in those modeling studies. The information required to assess population persistence is a dispersal matrix containing the fraction of larvae traveling to each location from each location, not just the current number of larvae exchanged among locations. Recent metapopulation modeling research with hypothetical dispersal matrices has shown how the spatial scale of dispersal, degree of advection versus diffusion, total larval output, and temporal and spatial variability in dispersal influence population persistence. Recent empirical studies using population genetics, parentage analysis, and geochemical and artificial marks in calcified structures have improved the understanding of dispersal. However, many such studies report current self-recruitment (locally produced settlement/settlement from elsewhere), which is not as directly useful as local retention (locally produced settlement/total locally released), which is a component of the dispersal matrix. Modeling of biophysical circulation with larval particle tracking can provide the required elements of dispersal matrices and assess their sensitivity to flows and larval behavior, but it requires more assumptions than direct empirical methods. To make rapid progress in understanding the scales and patterns of connectivity, greater communication between empiricists and population modelers will be needed. Empiricists need to focus more on identifying the characteristics of the dispersal matrix, while population modelers need to track and assimilate evolving empirical results.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In response to the threats to coral reefs (Jackson et al. 2001; Pandolfi et al. 2003; Hughes et al. 2003), marine protected areas (MPAs) have been proposed as a means of strengthening their resilience (e.g., Lubchenco et al. 2003; Hughes et al. 2006; Mumby et al. 2006). MPA design often centers on simply protecting a target fraction of key habitats (reviewed by Sale et al. 2005), rather than the effects of MPAs on metapopulation dynamics (Kritzer and Sale 2006). Management agencies commonly assume the communities naturally occurring in the protected habitats will persist in them, and contribute to unprotected habitats outside MPAs. However, a recent meta-analysis has shown that increases in population density inside MPAs depend on MPA size (Claudet et al. 2008). Furthermore, modeling studies have indicated how larval dispersal and the spatial configurations of MPAs interact to promote population persistence (Crowder et al. 2000; Botsford et al. 2001; Kaplan et al. 2006). However, while these interactions are clear in modeling results, efforts to apply conclusions from these models to assess and design effective MPAs are hindered by uncertainty about larval dispersal (Stockhausen et al. 2000; Botsford et al. 2001).
Fortunately, the nature of demographic connectivity among reef populations is beginning to be described in studies involving genetics, artificial and natural geochemical marks, and biophysical modeling. Those studies are changing the view of the nature of connectivity, notably by increasing the appreciation of the importance of short distance dispersal (Swearer et al. 2002; Taylor and Hellberg 2003; Jones et al. 2005; Cowen et al. 2006; Almany et al. 2007; Jones et al. 2009). However, the information gained through those studies has not been integrated and related to the features of connectivity identified by models as essential to resilience.
Population persistence: replacement over space
The current understanding of the relationship between larval connectivity and the persistence of marine metapopulations has a somewhat technical mathematical basis (Hastings and Botsford 2006), but the intuitive interpretation is merely an extension of the familiar concept of replacement. For single, non-spatial populations, a population will grow if each individual reproduces enough to replace itself in the next generation. This is a familiar characteristic of linear age-structured models (Caswell 2001), but it also applies to age-structured models with density-dependent recruitment (Sissenwine and Shepherd 1987). For human populations, if each couple has an average of slightly more than two children during their lifetime (i.e., lifetime reproduction per individual is just >1.0), the population will stay roughly constant. For marine populations, however, the number of eggs or larvae required to produce one reproductive offspring that survives the larval and early juvenile stage is poorly known. For populations with density-dependent recruitment, the required lower limit on egg production depends on the slope of the egg–recruit relationship at the origin, a poorly known quantity. Because of these difficulties in establishing the actual minimum threshold value of lifetime egg production (LEP) for each species, marine ecologists concerned with fisheries have expressed LEP as a fraction of the unfished, pristine value (fraction of lifetime egg production, or FLEP), and examined empirical information to determine a general safe value of that parameter (e.g., Mace and Sissenwine 1993). Initial meta-analyses suggest that keeping FLEP above 35% ensures adequate replacement over a range of species (Botsford et al. 2008).
While the relationship between FLEP and single population persistence is well known and commonly applied in fishery management, it does not describe the persistence of metapopulations. For metapopulations, the condition for persistence is that the total amount of replacement, through all possible paths be greater than a certain threshold, often assumed to be 35% of the unfished maximum (Hastings and Botsford 2006). A replacement path is the gain (or loss) in replacement at a subpopulation that occurs through the exchange of larvae with other subpopulations over multiple generations. The simplest kind of replacement path consists of larvae returning to their natal subpopulation, similar to the single population description above. The calculated replacement would be the local LEP times the fraction of larvae returning, which is referred to as “local retention” (Paris and Cowen 2004). The next more complex replacement path involves two subpopulations at locations A and B. It would be the LEP at location A times the fraction of larvae leaving that location that reach location B, times the LEP at location B, times the fraction of larvae from B that return to location A. In this path, replacement occurs over multiple generations: larvae spawned at A settle at B, then spawn larvae that return to A and contribute to replacement there. Other, more complex types of replacement paths would involve multiple patches in a similar manner (Hastings and Botsford 2006). A convenient simplification that follows from this description is that calculating the equilibrium population distribution over space requires only a spatial distribution of FLEP and knowledge of the dispersal pattern (e.g., Kaplan et al. 2006, 2008). The critical intuition is that understanding connectivity involves measuring the strength of replacement paths, which are closed loops, not just the amount of recruitment reaching a population or whether a population is a “source” or a “sink.”
Descriptions of dispersal: kernels and matrices
Larval dispersal is commonly described in terms of a dispersal kernel, the probability that a larva released from a particular location will disperse to and successfully settle at other specific locations, if suitable habitat is available (Largier 2003). Note that because larvae may die in transit, the integral of the kernel over space may be <1.0. Larvae that have reached a specific point in space and are competent to settle will settle if there is available habitat at that location. Thus, dispersal kernels are typically continuous functions, representing the two-dimensional spatial distribution of dispersed larvae, but not the presence of available habitat. If the benthic habitat happens to be more or less one-dimensional (such as along a straight coastline or a linear chain of reefs; Fig. 1a), the kernel can also be represented as a one-dimensional function of space (Fig. 1b).
A hypothetical one-dimensional example of a coral reef configuration that demonstrates the elements of the dispersal matrix. a The geographical configuration, b the dispersal kernels for each reef, with varying shape, diffusion, and advection, c a discrete-space version of the dispersal kernels, with each reef being a spatial unit, assuming constant larval survivorship of 0.01, d the corresponding dispersal matrix
Dispersal kernels can vary in a number of fundamental ways (Fig. 2), so the mathematical modeling of marine metapopulations has focused on the effects of this variability on population resilience. They can vary in magnitude (Fig. 2a) reflecting, for example, differences in survival through the larval stage. They can vary in width (Fig. 2b) and displacement from their origin (Fig. 2c). The mean displacement away from the point of release reflects mean dispersal resulting from the interaction of larvae with alongshore advective flow, and the width of the kernel represents stochastic variation about that mean dispersal. This variation may include random diffusion (a function of turbulence and small-scale coherent motions, represented by an eddy diffusivity parameter) (Okubo and Levin 2002), as well as deterministic variation such as current reversals occurring over the course of the larval stage (Largier 2003). Displacement and diffusion are generally expected to be greater for species with longer pelagic larval durations (Largier 2003; Siegel et al. 2003). Dispersal kernels need not be symmetrical about their origin (Fig. 2d), and they may vary spatially and temporally (Fig. 2e, f), possibly differing among spawning events, seasons or years (Fig. 1f) (e.g., Siegel et al. 2008). Particle-tracking simulations incorporating heterogeneous oceanographic data have produced both Gaussian (Siegel et al. 2003) and non-Gaussian, leptokurtic (Aiken et al. 2007) kernels. The shape could also be multimodal (Kaplan and Largier 2006; Fig. 2 g). Finally, while we have represented dispersal kernels as being one-dimensional, they can also be formulated in two dimensions if the habitat pattern is not approximately linear (Cowen et al. 2007; Snäll et al. 2007).
A complete metapopulation model requires a description of the dispersal kernel for all possible locations. To provide that description, the habitat is divided into n discrete habitat patches and dispersal is described by an n × n dispersal matrix, D, which contains the probabilities of larvae dispersing to and from each of the patches (e.g., Cowen et al. 2006). As indicated in Fig. 1d, each matrix entry D ij is the probability of dispersal from patches i to j, i.e., the dispersal kernel for patch i integrated over the area represented by patch j (Fig. 1c). The dispersal matrix D is related to the dispersal kernel in that each row of the matrix is a discrete version of the dispersal kernel for each patch.
Effects of kernel features on metapopulation persistence and MPAs
The characteristic of dispersal receiving the most attention in population modeling is the “dispersal distance,” which can refer to either the width or the advection (displacement) of a dispersal kernel. Assuming the simple scenario of zero advection, no movement of adult organisms, and fairly high fishing rates that do not change with MPA implementation, a species will persist in an MPA so long as the spatial dimensions of the MPA are roughly greater than the width of the dispersal kernel (Botsford et al. 2001). If this criterion is not met (i.e., small MPAs or wider dispersal kernels), the species can still persist, but only if a certain fraction of the coastline is covered with MPAs. That critical fraction is the same as the fraction of natural, unfished lifetime egg production (FLEP) required for a single, non-spatial population of that species to persist (~35%), as described above (Botsford et al. 2001, 2008). This dependence reflects a crucial link between spatial marine resource management with MPAs and conventional fisheries management: they both depend on the highly uncertain critical replacement threshold, the minimum tolerable FLEP for persistence.
Relaxing some of the simplifying assumptions made thus far reveals the effect of other dispersal characteristics, and other factors on population persistence. If fishing is less intense, the fraction of coastline required in MPAs for persistence declines (Botsford et al. 2001; Kaplan et al. 2008). Also, addition of advection to the purely diffusive dispersal kernel (Fig. 2c) reduces self-retention and thus increases the minimum MPA width required for persistence (Botsford et al. 2001; Gaylord and Gaines 2000; Gaines et al. 2003) unless MPA spacing precisely matches the advection of the dispersal kernel (Kaplan 2006). If adult organisms are not sedentary and can move across MPA boundaries, overall MPA effectiveness declines and the persistence is more difficult to achieve.
These general findings are robust to deviations from some common modeling assumptions. Most models use either a Gaussian or Laplacian (two-sided negative exponential) dispersal kernel, but for symmetrical, non-advective kernels, kernel shape (Fig. 2d) does not affect population persistence (Lockwood et al. 2002). Additionally, most models consider coastlines with evenly spaced MPAs, but having a variety of spacings among MPAs does not affect population persistence (Kaplan and Botsford 2005).
While the presence of both spatial (Fig. 2e) and temporal (Fig. 2f) variability in dispersal is readily acknowledged, neither has been explored in modeling to a sufficient degree to achieve a good understanding of their effects. It is typically assumed that dispersal kernels integrating spawning over the spawning season of a given species are temporally constant. There are a few examples of studies that accounted for temporal variation in dispersal patterns, including rare disturbance events such as cyclones (James et al. 2002; Wolanski et al. 2004; Bode et al. 2006). However, none of these have addressed population persistence with MPAs directly.
Finally, the magnitude of dispersal has been difficult to explore systematically because of the lack of empirical information (Fig. 2a). Most modeling studies that explore the effects of dispersal width maintain the total larval production per egg produced as a constant, leaving out any differences in reproduction with larval dispersal distance (e.g., James et al. 2002; Aiken et al. 2007). For example, in the studies above comparing the effects of larval dispersal distance on persistence, the total number of successful larvae produced did not vary with larval dispersal distance, rather it was held constant. While this assumption was parsimonious, it has no empirical basis. There is mounting evidence, however, that even within species, not all larvae are equal (e.g., Berkeley et al. 2004; Hamilton et al. 2008).
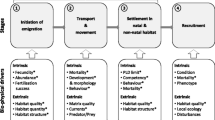
Empirical approaches to measuring dispersal
Information quantifying larval dispersal is accumulating from studies involving population genetics, biophysical circulation models, paternity analysis, tagging/natural markers, and larval/settlement observations. These methods differ in their empirical bases, and in the characteristics of the dispersal matrix that they can quantify. The differences in empirical information can be best understood in terms of steps in connectivity, as illustrated by the general form of a metapopulation model (Fig. 3). Population connectivity begins with the spatial distribution of egg or larval production, which depends on adult age structure and fecundity (Fig. 3a). Larval production is multiplied (matrix multiplication) by the dispersal matrix (Fig. 1d) to obtain the distribution of potential settlers (Fig. 3b). Actual settlement and recruitment then depends on the presence of suitable habitat and can depend on the density of potential settlers and adults (Fig. 3c, d).
An example of one step in the connectivity process for the metapopulation occupying the reefs depicted in Fig. 1. a The total egg production on each reef. Reef B has lower output due to low quality habitat. Reefs D and E have lower output due to low population density. b Egg production multiplied by the dispersal kernel gives the spatial distribution of potential settlers (assuming homogenous larval mortality). c Integrating the settler distribution over the area on each reef gives the total settlers in each location. d Settler densities are reduced due to habitat- and density-dependent mortality. Settlers at each reef experience density-dependent Beverton–Holt mortality with density-independent survivorship of 0.8 and an asymptotic maximum recruit density of 1 recruit m−2. On reef B, poor habitat causes per-capita fecundity and density-independent survivorship to be 50% of that on the other reefs
The definition of the dispersal kernel suggests an idealized approach to empirically estimating the values of each component of the dispersal matrix, i.e., the values of all the D ij ’s. Since each row of that matrix represents a discrete version of the dispersal kernel for that source location (Fig. 1c, d), a simple frequentist interpretation of probability would lead one to estimate each D ij by releasing a known number, say N i , of larvae from each location, recording the number of those that settled at each location, n ij , then computing D ij = n ij /N i . This simple idealized sampling scheme, which may be difficult to accomplish in practice, proves useful below in the interpretation of existing empirical methods.
Note that local retention as defined above is distinct from the quantity reported as “self-recruitment” in most empirical studies. Self-recruitment is typically calculated as the proportion of settlers at a location that were spawned locally (e.g., Jones et al. 1999; Swearer et al. 1999; Almany et al. 2007). This quantity is a measure of how isolated a focal population is (see also Jones et al. 2009). The high levels of percent self-recruitment reported recently have drawn attention to how small the scale of dispersal can be (Jones et al. 1999, 2005; Swearer et al. 1999). However, percent self-recruitment cannot be used directly to estimate population persistence. For example, compare reefs C and E in Fig. 3. Reef C has <50% self-recruitment, because it receives many larvae from reef D, yet reef C still has high enough local retention to be self-persistent. Reef E, however, has 100% self-recruitment (it receives no larvae from elsewhere) but also has very low local retention and is not self-persistent in this scenario. Replacement depends on the fraction of locally produced larvae that return home (local retention), not the fraction of settling larvae that were spawned locally (self-recruitment). While local retention is clearly an important parameter, in practice it is difficult to measure, as the reproductive output and the fate of all juveniles sourced from a particular sub-population must be known. If almost all sub-populations are sampled and all exhibit high self-recruitment estimates, then it is clear that they will also be characterized by high local retention and narrow dispersal kernels.
Population genetics
Population genetics describes connectivity by comparing allele frequencies among spatially discrete subpopulations. High levels of genetic similarity between populations suggest gene flow over time, usually through larval dispersal; whereas significant differentiation between populations indicates significant and persistent barriers to larval exchange. Approaches determining population genetic structure within reef metapopulations are valuable in assessing patterns and degrees of connectivity when methods to directly track larvae are not possible.
Typically, genetic structure of a metapopulation is characterized by sampling adults from various subpopulations. These individuals represent the distribution of successful recruits within the metapopulation model (Fig. 3d). Adult populations represent an accumulation of genetic signals from larval sources over time, with influence of ecological (e.g., selection) and evolutionary (e.g., mutation and drift) forces acting on individuals comprising the population. For reef organisms with relatively short lifespans (<multiple decades) and that reproduce exclusively through sexual means (e.g., most reef fish) estimates of gene flow between populations are an adequate approximation of contemporary levels of genetic connectivity. On the other hand, levels of connectivity among organisms with considerable overlap across generations, such as corals and sponges, may not be appropriate indicators of present-day connectivity patterns. The age of sampled individuals can be decades to centuries and propagation through asexual means can produce virtually immortal genotypes, thus adult population structure of these organisms represents connectivity processes that occurred decades to centuries ago. Significant demographic changes in populations over time, for example, population declines in coral reef ecosystems (Hughes et al. 2003; Pandolfi et al. 2003), may result in significant differences in larval production, influencing the magnitude of the dispersal kernel and overall connectivity levels over time. Traditional population genetic statistics (F-statistics) have been used to estimate gene flow between populations and viewed as a proxy for dispersal over evolutionary time scales, but are not sensitive to recent changes in gene flow and genetic structure of these long-lived organisms may retain the signature of past events rather than reveal present connectivity patterns (Bossart and Pashley Prowell 1998; Benzie 1999). Alternative population genetics statistics involve assignment of individuals to putative natal populations based on the frequencies of multilocus genotypes in these populations (e.g., Paetkau et al. 1995; Waser and Strobeck 1998; Pritchard et al. 2000, Wilson and Rannala 2003) using frequency probabilities, likelihood methods or Bayesian analyses (reviewed in Manel et al. 2005). A primary advantage of assignment methods is the ability to evaluate contemporary connectivity rates without the unrealistic assumptions required by traditional methods.
If settlers and natal sources are sampled over a broad enough spatial scale, it should be possible to estimate the width, advection, and shape of the kernel using assignment techniques. If larval production data for each source were also available, kernel magnitude and the dispersal matrix could also be estimated. However, some caution is required when applying genetic methods. A single migrating larva per generation can genetically homogenize populations (Spieth 1974), so genetically similar populations may have demographically negligible larval exchange. Conversely, population genetics will be inadequate to resolve connectivity between populations that exchange many larvae each generation or to estimate local retention.
Parentage analysis
While genetic assignment techniques can link a settler to its natal reef, parentage analysis can identify its actual parent. Paternity or parentage assignation can be achieved by any type of genetic marker, provided it is sufficiently polymorphic. Microsatellites have been used for paternity analysis in terrestrial animal and plant species, but this approach is rarely used to estimate connectivity, especially in marine ecosystems (Jones et al. 2005 provide a promising first example).
Because parentage analysis relates offspring to parents directly, it yields information on the critical features of dispersal. Parentage analysis can identify kernel width, advection, shape, and temporal variation, so long as settlers from all possible destinations are sampled. It is also possible to determine the amount of successful larval output (if actual abundance of settled genotypes is estimated), and the magnitude of the kernel (if the larval output from each parent can be estimated).
There are limitations to this approach. Parentage analysis requires a high number of variable genetic markers (Jones and Ardren 2003), but genetic screening of a few thousand individuals for multiple markers, such as microsatellites, is no longer a technological restriction. Instead, the main limitation is performing adequate sampling. It is necessary to sample a significant proportion of the “source” adult population, and then sample new recruits over the entire potential range of dispersal. Sampling recruits is the less critical aspect of this approach, as it is reasonable to sample some proportion of recruits in several locations throughout the range of potential dispersal. It is more critical to collect a significant proportion of the adult source population, because a failure to collect most parents will lead to underestimation of the representation of the parental genotypes in recruit samples. This requirement will certainly limit the locations and taxa that can be investigated.
Natural and artificial tags in calcified structures
Geochemical tags in the calcified structures of marine organisms may provide information on larval dispersal that is difficult to obtain using conventional marking approaches. This technique relies on variability in physico-chemical properties of ambient environments to generate geochemical signatures that are recorded in calcified structures such as otoliths, statoliths, and shells. Many of these structures also contain a detailed chronological record of location in the form of daily or annual increments. Reconstruction of environmental conditions can be achieved by measuring the elemental or isotopic composition of these structures. This technique can be particularly powerful because every individual from a population with a unique geochemical signature is indelibly tagged, and therefore recapturing marked individuals is relatively easy.
Geochemical signatures are typically used to determine the origins of individuals settling into juvenile habitats (Becker et al. 2007), or to examine the natal homing of spawning adults that migrate significant distances after settlement (e.g., Thorrold et al. 2001). The spatial precision of the technique is likely to be relatively coarse because the scale of variability of temperature and water chemistry in coral reef environments is likely greater than the scale of spatial subdivision in metapopulations (e.g., Ruttenberg and Warner 2006). As such, natural geochemical signatures are probably not going to be particularly useful for estimating dispersal kernels. The approach may, however, provide sufficient information to estimate the larval dispersal matrix for cases in which adult population abundances are known (or can be assumed) for each of the subpopulations and the subpopulations have well-defined spatial boundaries and geochemical signatures (e.g., whole estuaries, Thorrold et al. 1998, 2001).
Natural tags have also been used in a binary sense to determine whether individuals are retained in a particular location with a well-defined signature or have come from elsewhere (Swearer et al. 1999; Standish et al. 2008). Unfortunately, this type of information is of limited use in characterizing dispersal matrices, because it provides an estimate of self-recruitment, not local retention (see Fig. 2d).
Transgenerational isotope labeling
Calcified structures have also been used as a place to locate artificial chemical tags. The approach relies on the observation that, unlike most tissues, otoliths are metabolically inert and therefore a chemical tag will remain in the structure throughout the lifetime of an individual. While artificial chemical tags have been used in hatchery and field situations for many years (Levin 1990; Jones et al. 1999), the potential power of the approach has increased significantly with the discovery of a method for transgenerational tagging of larvae using enriched stable isotopes (Thorrold et al. 2006). The transgenerational isotope labeling (TRAIL) approach is based on the maternal transmission of stable isotopes from spawning females to the otoliths of embryos produced by an individual after exposure to the isotope. The unique isotope signature permanently encodes an indelible tag in the otoliths of offspring than can be detected using laser ablation mass spectrometry.
Artificial chemical tags have some significant advantages over natural geochemical signatures. First, the spatial scale over which unique tags can be identified is determined by the researcher rather than the ambient environment. Therefore, the technique may be able to generate a dispersal kernel from a single point source, assuming most of the settlement sites are sampled. Females continue to produce tagged larvae for several months after initial exposure to the enriched isotope (Thorrold et al. 2006), thereby facilitating examination of variability in the kernel over scales of several months. As with any mark-recapture study, TRAIL requires an estimate of the percentage of total larval production tagged. In some situations, it is possible to assume that 100% of larvae released from a location will be tagged (Almany et al. 2007). In other situations, it will be necessary to assume that females producing tagged larvae are a random sample of all females present at a single location. To obtain the magnitude of the kernel, not just the shape, width, and advection, one must also estimate the total number of tagged larvae spawned, not just the percentage. While it is possible to tag a number of different locations with a unique-enriched isotope, in reality the approach is likely too labor-intensive to be feasible for parameterizing a full dispersal matrix.
Modeling approaches to measuring dispersal
In the past decade, substantial progress has been made in the use of coupled biophysical modeling to understand the degree of connectivity between populations (reviewed by Werner et al. 2007). Biophysical modeling studies differ from the above empirical methods in that their results can include any of the characteristics of larval dispersal depicted in Fig. 2. By seeding the model with a large number of particles (“larvae”), it is possible to assemble dispersal kernels and matrices from the start (spawning) and the end point (settlement) of individual particle trajectories. Just as in the empirical methods that determine larval origins as discussed above, any one run is a stochastic realization of a probabilistic process. Therefore, estimating the full kernel requires averaging over many dispersal events within the spectrum of model behavior (Cowen et al. 2006).
Numerical models can be used to evaluate the temporal variability of larval dispersal (Fig. 2f) and settlement at different time scales so long as the time-scale relevant to the target organism is resolved by the forcing of the ocean circulation model (Paris et al. 2002). At present, spatially explicit models forced by realistic currents coupled with demographic parameters produce dispersal kernels for a range of spatial scales over which dispersal (and perhaps also survival, see Cowen et al. 2000) is practically unquantifiable by current empirical methods (James et al. 2002; Cowen et al. 2006).
The rich detail afforded by biophysical models is balanced by a less direct empirical connection to the real world, relative to the other methods considered in this paper. The empirical basis for the circulation model typically involves boundary conditions and environmental forcing from an atmospheric model. Circulation models can also be validated by, or assimilate data from, a variety of physical observations such as hydrographic data and drifter observations. The biological part of these biophysical models may be based on the laboratory or field observations of larval traits to ensure fidelity between actual larval dispersal and model predictions (Werner et al. 2007; Gallego et al. 2007).
Numerical models can explicitly include species-specific ontogenetic behavior to better understand the characteristics of larval dispersal illustrated in Fig. 2. For example, when larval behavior is included, advection (Fig. 2c) decreases significantly while diffusion (Fig. 2b) typically remains similar (Paris et al. 2007). Diffusion may increase with random (i.e., non-oriented) larval swimming, with larvae possibly reaching more habitat patches (Armsworth and Roughgarden 2005). However, increased diffusion dilutes the number of settling larvae at any given location while directed movement enhances spatial partitioning.
Modeling has shown that vertical larval movement influences their dispersal direction and may represent a survival mechanism by which larvae balance the risks of encountering predators and starving (Armsworth et al. 2001; Vikebø et al. 2007). This links overall larval success (Fig. 2a) with advection (Fig. 2c) and diffusion (Fig. 2b). To date, only a laboratory study has determined survivorship as a function of ontogeny for coral reef larvae (Graham et al. 2008).
Biophysical models have also been used to study the interaction between dispersing larvae and habitat, which establishes the spawning location and production (i.e., initial conditions) as well as a halo of cues that attract competent larvae. Thus, models of larval exchange that integrate habitat along individual trajectories can improve estimates of dispersal kernels (Paris et al. 2007).
Discussion
At a time when recommendations for and implementation of MPAs are increasing, there remains some mismatch between information needed to predict population persistence, and the empirical efforts to gather that information. An understanding of current levels of connectivity (i.e., the total number of larvae currently exchanged among reefs) is not sufficient for projecting future behavior after the spatial distribution of larval production or habitat is changed (by fishing or bleaching, respectively, for example). Rather, projection of future responses to disturbances will require knowing connectivity per larva released, i.e., the dispersal matrix. This problem in the science of MPAs is a subset of the more general problems addressed in recent attention to “movement ecology” (Nathan et al. 2008), which is aimed at a better understanding of dispersal of individuals, but not necessarily their population consequences.
The empirical approaches differ in the assumptions underpinning their estimates of the dispersal kernel. Traditional genetic approaches that gauge dispersal from the dependence of genetic difference on physical distance require a number of assumptions about the nature of genetic differentiation, and will be estimates of both current and past connectivity. Approaches that actually identify natal origins such as genetic assignment methods, parentage analysis, and natural tagging are more precise in identifying dispersal patterns, but a full estimation of the kernel still depends on the spatial distribution of larval production (i.e., adult abundance). Methods such as parentage analysis and TRAIL that estimate numbers of each type of larvae released and numbers available for settlement at each location make an additional step toward estimating the dispersal matrix. All of these methods use observations made after settlement, so as estimates of dispersal during the larval phase they are confounded to varying degrees by density-dependence in recruitment and the spatial pattern of habitat.
Biophysical models can directly calculate dispersal matrices and explore their dependence on circulation and larval behavior. However, they differ from the empirical approaches in their underlying assumptions, depending not on the accuracy of direct observations of abundance during the connectivity process, but rather on how well they represent the relevant physical processes, the boundary conditions of the physical model, the atmospheric forcing, and the ability of the biological component to represent larval buoyancy and behavior realistically.
While empirical information has not yet provided all of the elements of a dispersal matrix that is a high goal and may be difficult to achieve. To put this in perspective, one must consider that the question of adequate replacement is still highly uncertain even for single non-spatial populations, as evidenced in contemporary fishery management. In particular, estimation of the critical replacement level required for persistence requires observations of total egg production over enough years, and at low enough levels, to estimate the slope of the egg–recruit relationship at the origin. For most marine fisheries, the latter is not known.
Population modeling has indicated how important the width of dispersal kernels is to population dynamics, since kernel width (or, in a discrete framework, the probability of local retention) essentially determines whether persistence will depend on self-replacement or a network effect involving replacement over a number of generations. This is one area in which empirical methods are just beginning to measure the relevant parameter. To date, because sampling is usually focused on a single population, estimates have been limited to self-recruitment, the proportion of settlers exhibiting a natal signature (Jones et al. 1999, 2005; Swearer et al. 1999; Almany et al. 2007; Carreras-Carbonell et al. 2007), rather than the more informative local retention (=number returning home/total number released, e.g., Paris and Cowen 2004). Monitoring larval production and estimating local retention is a direction ripe for empirical exploration. Additionally, fully parameterizing the dispersal kernel requires sampling settlers from multiple potential destinations and examining their natal signatures. This effort will be necessary to characterize larval exchange among subpopulations and understand the patterns of persistence due to network effects.
The overview presented here reveals a surprisingly low level of communication and integration between: (1) modeling efforts to determine the population consequences of larval dispersal on coral reefs and (2) empirical efforts to estimate the characteristics of larval dispersal. There is an obvious need for the ability to translate empirical findings into their consequences for spatial management of coral reef resources. This will require greater communication between population modelers and those generating empirical estimates of dispersal, especially in light of the rapid propagation of MPAs. On the empirical side, greater effort to quantify dispersal from multiple origins to multiple destinations is needed. The unparalleled potential of the biophysical approach could benefit by more studies in which there are empirical confirmation of model characteristics of simulated larvae through some of the more direct methods. Conversely, the multiple random outcomes generated by biophysical circulation models provide a basis for needed exploration of the response of metapopulation models to interannual temporal and spatial stochasticity in the context of MPAs. There is also a need for a general understanding by decision makers of the results from these fields. There are still too many under the false impression that setting aside an MPA will recreate or preserve a pristine ecosystem, no matter how small the MPA is relative to the spatial scale of larval dispersal distances of a species.
References
Aiken CM, Navarrete SA, Castillo MI, Castilla JC (2007) Along-shore larval dispersal kernels in a numerical ocean model of the central Chilean coast. Mar Ecol Prog Ser 339:13–24
Almany GR, Berumen ML, Thorrold SR, Planes S, Jones GP (2007) Local replenishment of coral reef fish populations in a marine reserve. Science 316:742–744
Armsworth PR, Roughgarden JE (2005) The impact of directed versus random movement on population dynamics and biodiversity patterns. Am Nat 165:449–465
Armsworth PR, James MK, Bode L (2001) When to press on or turn back: dispersal strategies for reef fish larvae. Am Nat 157:434–450
Becker BJ, Levin LA, Fodrie FJ, McMillan PA (2007) Complex larval connectivity patterns among marine invertebrate populations. Proc Natl Acad Sci 104:3267–3272
Benzie JAH (1999) Genetic structure of coral reef organisms: ghosts of dispersal past. Am Zool 39:131–145
Berkeley SA, Chapman C, Sogard SM (2004) Maternal age as a determinant of larval growth and survival in a marine fish, Sebastes melanops. Ecology 85:1258–1264
Bode M, Bode L, Armsworth PR (2006) Larval dispersal reveals regional sources and sinks in the Great Barrier Reef. Mar Ecol Prog Ser 308:17–25
Bossart JL, Pashley Prowell D (1998) Genetic estimates of population structure and gene flow: limitations, lessons, and new directions. Trends Ecol Evol 13:202–206
Botsford LW, Hastings A, Gaines SG (2001) Dependence of sustainability on the configuration of marine reserves and larval dispersal distance. Ecol Lett 4:144–150
Botsford LW, Brumbaugh DR, Grimes C, Kellner JB, Largier J, O’Farrell MR, Ralston S, Soulanille E, Wepestad V (2008) Connectivity, sustainability and yield: bridging the gap between conventional fishery management and marine protected areas. Rev Fish Biol Fish. doi:10.1007/s11160-008-9092-z
Carreras-Carbonell J, Macpherson E, Pascual M (2007) High self-recruitment levels in a Mediterranean littoral fish population revealed by microsatellite markers. Mar Biol 151:719–727
Caswell H (2001) Matrix population models. Sinauer, Sunderland, MA
Claudet J, Osenberg CW, Benedetti-Cecchi L, Domenici P, Garciá-Charton JA, Pérez-Ruzafa A, Badalamenti F, Bayle-Sempere J, Brito A, Bulleri F, Culioli JM, Dimech M, Falcón JM, Guala I, Milazzo M, Sánchez-Meca, Somerfield PJ, Stobart B, Vandeperre F, Valle C, Planes S (2008) Marine reserves: size and age do matter. Ecol Lett 11:481–489
Cowen RK, Lwiza KMM, Sponaugle S, Paris CB, Olson DB (2000) Connectivity of marine populations: open or closed? Science 287:857–859
Cowen RK, Paris CB, Srinivasan A (2006) Scaling of connectivity in marine populations. Science 311:522–527
Cowen RK, Gawarkievicz PJ, Thorrold SR, Werner FE (2007) Population connectivity in marine systems: an overview. Oceanography 20:14–21
Crowder LB, Lyman SJ, Figueira WF, Priddy J (2000) Source-sink population dynamics and the problem of siting marine reserves. Bull Mar Sci 66:799–820
Gaines SD, Gaylord B, Largier JL (2003) Avoiding current oversights in marine reserve design. Ecol Appl 13:S32–S46
Gallego A, North EW, Petitgas P (2007) Introduction: status and future of modelling physical–biological interactions during the early life of fishes. Mar Ecol Prog Ser 347:121–126
Gaylord B, Gaines SD (2000) Temperature or transport? Range limits in marine species mediated solely by flow. Am Nat 155:769–789
Graham EM, Baird AH, Connolly SR (2008) Survival dynamics of scleractinian coral larvae and implication for dispersal. Coral Reefs. doi 10.2007/s00338-008-0361-z
Hamilton SL, Regetz J, Warner RR (2008) Postsettlement survival linked to larval life in a marine fish. Proc Natl Acad Sci USA 105:1561–1566
Hastings A, Botsford LW (2006) Persistence of spatial populations depends on returning home. Proc Natl Acad Sci USA 103:6067–6072
Hughes TP, Baird AH, Bellwood DR, Card M, Connolly SR, Folke C, Grosberg R, Hoegh-Guldberg O, Jackson JBC, Kleypas J, Lough JM, Marshall P, Nyström M, Palumbi SR, Pandolfi JM, Rosen B, Roughgarden J (2003) Climate change, human impacts, and the resilience of coral reefs. Science 301:929–933
Hughes TP, Bellwood DR, Folke CS, McCook LJ, Pandolfi JM (2006) No-take areas, herbivory and coral reef resilience. Trends Ecol Evol 22:1–3
Jackson JBC, Kirby MX, Berger WH, Bjorndal KA, Botsford LW, Bourque BJ, Bradbury RH, Cooke R, Erlandson J, Estes JA, Hughes TP, Kidwell S, Lange CB, Lenihan HS, Pandolfi JM, Peterson CH, Steneck RS, Tegner MJ, Warner RR (2001) Historical overfishing and the recent collapse of coastal ecosystems. Science 293:629–638
James MK, Armsworth PR, Mason LB, Bode L (2002) The structure of reef fish metapopulations: modelling larval dispersal and retention patterns. Proc R Soc Lond B 269:2079–2086
Jones AG, Ardren WR (2003) Methods of parentage analysis in natural populations. Mol Ecol 12(10):2511–2523
Jones GP, Milicich MJ, Emslie MJ, Lunow C (1999) Self-recruitment in a coral reef fish population. Nature 402:802–804
Jones GP, Planes S, Thorrold SR (2005) Coral reef fish larvae settle close to home. Curr Biol 15:1314–1318
Jones GP, Almany G, Russ G, Sale P, Steneck R, van Oppen M, Willis B (2009) Larval retention and connectivity among populations of corals and reef fishes: history, advances and challenges (this issue). doi:10.1007/s00338-009-0469-9
Kaplan DM (2006) Alongshore advection and marine reserves: consequences for modeling and management. Mar Ecol Prog Ser 309:11–24
Kaplan DM, Botsford LW (2005) Effects of variability in spacing of marine reserve on fisheries yield and sustainability. Can J Fish Aquat Sci 62:905–912
Kaplan DM, Largier J (2006) HF radar-derived origin and destination of surface waters off Bodega Bay, California. Deep-Sea Res Pt II 53:2906–2930
Kaplan DM, Botsford LW, Jorgensen S (2006) Dispersal-per-recruit: an efficient method for assessing sustainability in networks of marine reserves. Ecol Appl 16:2248–2263
Kaplan DM, LW Botsford, S D Gaines and SJ Jorgensen (2008) Model-based assessment of persistence in proposed marine protected area designs for the central California coast. Ecol Appl (in press)
Kritzer JP, Sale PF (2006) The metapopulation ecology of coral reef fishes. In: Kritzer JP, Sale PF (eds) Marine metapopulations. Elsevier Academic Press, Burlington, MA, pp 31–67
Largier JL (2003) Considerations in estimating larval dispersal distances from oceanographic data. Ecol Appl 13:S71–S89
Levin LA (1990) A review of methods for labeling and tracking marine invertebrate larvae. Ophelia 32:115–144
Lockwood DR, Hastings A, Botsford LW (2002) The effects of dispersal patterns on marine reserves: does the tail wag the dog? Theor Pop Biol 61:297–309
Lubchenco J, Palumbi SR, Gaines SD, Andelman S (2003) Plugging a hole in the ocean: the emerging science of marine reserves. Ecol Appl 13:S3–S7
Mace PM, Sissenwine MP (1993) How much spawning per recruit is enough? Can Spec Publ Fish Aquat Sci 120:101–118
Manel S, Gaggiotti OE, Waples RS (2005) Assignment methods: matching biological questions with appropriate techniques. Trends Ecol Evol 20:136–142
Mumby PJ, Dahlgren CP, Harbone AR, Kappel CV, Micheli F, Brumbaugh DR, Holmes KE, Mendes JM, Broad K, Sanchirico JN, Buch K, Box S, Stoffle RW, Gill AB (2006) Fishing, trophic cascades, and the process of grazing on coral reefs. Science 311:98–101
Nathan R, Getz WM, Revilla E, Holyoak M, Kadmon R, Saltz D, Smouse PE (2008) A movement ecology paradigm for unifying organismal movement research. Proc Natl Acad Sci USA 105:19052–19059
Okubo A, Levin SA (2002) The basics of diffusion. In: Okubo A, Levin SA (eds) Diffusion and ecological problems: modern perspectives, 2nd edn. Springer, New York, pp 10–30
Paetkau D, Calvert W, Stirling I, Strobeck C (1995) Microsatellite analysis of population structure in Canadian polar bears. Mol Ecol 4:347–354
Pandolfi JM, Bradbury RH, Sala E, Hughes TP, Bjorndal KA, Cooke RG, McArdle D, McClenachan L, Newman MJH, Paredes G, Warner RR, Jackson JBC (2003) Global trajectories of the long-term decline of coral reef ecosystems. Science 301:955–958
Paris CB, Cowen RK (2004) Direct evidence of a biophysical retention mechanism for coral reef fish larvae. Limnol Oceanogr 49(6):1964–1979
Paris CB, Cowen RK, Lwiza KMM, Wang D-P, Olson DB (2002) Multivariate objective analysis of the coastal circulation of Barbados, West Indies: implications for larval transport. Deep-Sea Res Pt I 49:1363–1386
Paris CB, Chérubin LM, Cowen RK (2007) Surfing, spinning, or diving from reef to reef: effects on population connectivity. Mar Ecol Prog Ser 347:285–300
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Ruttenberg BI, Warner RR (2006) Variation in the chemical composition of natal otoliths from a reef fish in the Galápagos Islands. Mar Ecol Prog Ser 328:225–236
Sale PF, Cowen RK, Danilowicz BS, Jones GP, Kritzer JP, Lindeman KC, Planes S, Polunin NVC, Russ GR, Sadovy Y, Steneck RS (2005) Critical science gaps impede use of no-take fishery reserves. Trends Ecol Evol 20:74–80
Siegel DA, Kinlan BP, Gaylord B, Gaines SD (2003) Lagrangian descriptions of marine larval dispersion. Mar Ecol Prog Ser 260:83–96
Siegel DA, Mitarai S, Costello CJ, Gaines SD, Kendall BE, Warner RR, Winters KB (2008) The stochastic nature of larval connectivity among nearshore marine populations. Proc Natl Acad Sci USA 105:8974–8979
Sissenwine MP, Shepherd JG (1987) An alternative perspective on recruitment overfishing and biological reference points. Can J Fish Aquat Sci 44:913–918
Snäll T, O’Hara RB, Arjas E (2007) A mathematical and statistical framework for modelling dispersal. Oikos 116:1037–1050
Spieth PT (1974) Gene flow and genetic differentiation. Genetics 78:961–965
Standish JD, Sheehy M, Warner RR (2008) Use of otolith natal elemental signatures as natural tags to evaluate connectivity among open-coast fish populations. Mar Ecol Prog Ser 356:259–268
Stockhausen WT, Lipcius RN, Hickey BM (2000) Joint effects of larval dispersal, population regulation, marine reserve design, and exploitation on production and recruitment in the Caribbean spiny lobster. Bull Mar Sci 66:957–990
Swearer SE, Caselle JE, Lea DW, Warner RR (1999) Larval retention and recruitment in an island population of a coral-reef fish. Nature 402:799–802
Swearer SE, Shima JS, Hellberg ME, Thorrold SR, Jones GP, Robertson DR, Morgan SG, Selkoe KA, Ruiz GM, Warner RR (2002) Evidence of self-recruitment in demersal marine populations. Bull Mar Sci 70:251–271
Taylor MS, Hellberg ME (2003) Genetic evidence for local retention of pelagic larvae in a Caribbean reef fish. Science 299:107–109
Thorrold S, Jones CM, Swart PK, Targett TE (1998) Accurate classification of juvenile weakfish Cynoscion regalis to estuarine nursery areas based on chemical signatures in otoliths. Mar Ecol Prog Ser 173:253–265
Thorrold SR, Latkoczy C, Swart PK, Jones CM (2001) Natal homing in a marine fish metapopulation. Science 291:297–299
Thorrold SR, Jones GP, Planes S, Hare JA (2006) Transgenerational marking of embryonic otoliths in marine fishes using barium stable isotopes. Can J Fish Aquat Sci 63:1193–1197
Vikebø F, Jørgensen C, Kristiansen T, Fiksen Ø (2007) Drift, growth, and survival of larval Northeast Arctic cod with simple rules of behaviour. Mar Ecol Prog Ser 347:207–219
Waser PM, Strobeck C (1998) Genetic signatures of interpopulation dispersal. Trends Ecol Evol 13:43–44
Werner FE, Cowen RK, Paris CB (2007) Coupled biological and physical models: present capabilities and necessary developments for future studies of population connectivity. Oceanography 20:54–69
Wilson GA, Rannala B (2003) Bayesian inference of recent migration rates using multilocus genotypes. Genetics 163:1177–1191
Wolanski E, Richmond RH, McCook L (2004) A model of the effects of land-based, human activities on the health of coral reefs in the Great Barrier Reef and in Fouha Bay, Guam, Micronesia. J Mar Syst 46:133–144
Acknowledgments
Work by CB Paris was supported by the National Science Foundation grant NSF-OCE 0550732. Work by M-A Coffroth and SR Thorrold was supported by the National Science Foundation grant NSF-OCE 0424688. Work by TL Shearer was supported by an International Cooperative Biodiversity Group grant R21 TW006662-01 from the Fogarty International Center at the National Institutes of Health
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Ecology Editor Prof. Peter Mumby
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Botsford, L.W., White, J.W., Coffroth, M.A. et al. Connectivity and resilience of coral reef metapopulations in marine protected areas: matching empirical efforts to predictive needs. Coral Reefs 28, 327–337 (2009). https://doi.org/10.1007/s00338-009-0466-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00338-009-0466-z