Abstract
Mouse models are essential for dissecting disease mechanisms and defining potential drug targets. There are more than 18,500 mouse strains available for research communities in National Resource Center for Mutant Mice (NRCMM) of China, affiliated with Model Animal Research Center of Nanjing University and Gempharmatech Company. In 2019, Gempharmatech launched the Knockout All Project (KOAP) aiming to generate null mutants and gene floxed strains for all protein-coding genes in mouse genome within 5 years. So far, KOAP has generated 8,004 floxed strains and 9,769 KO (knockout) strains (updated to Oct, 2021). NRCMM also created hundreds of Cre transgenic lines, mutant knock-in models, immuno-deficient models, and humanized mouse models. As a member of the international mouse phenotyping consortium (IMPC), NRCMM provides comprehensive phenotyping services for mouse models. In summary, NRCMM will continue to support biomedical community with new mouse models as well as related services.
Similar content being viewed by others
Establishment of NRCMM
House mouse (Mus musculus) has been widely used as laboratory animal in biomedical research since early 1900. Studies found that 99% protein-coding genes display high homology by comparing the mouse and human genome sequences (Lander et al. 2001; Mouse Genome Sequencing et al. 2002). This provides a genetic explanation for mouse and human sharing many common features in embryonic development process, physiology homeostasis regulation, and disease etiology. Genetically engineering mouse models (GEMMs), therefore, have been a crucial tool to understand the function for human genes, probe the pathogenesis of human diseases and test the efficacy of potential drugs.
To support academic and pharmaceutical communities on the use of valuable mouse resources, NRCMM was established at Nanjing, China in 2001. NRCMM is the national mouse resource center established by Ministry of Science and Technology of China, and now are operated jointly by Model Animal Research Center of Nanjing (MARC) and GemPharmatech (GPT). The mouse strains from both parties are enlisted as the resources of NRCMM (shown in Online Resource 1), and MARC and GPT hold independent legal ownerships for their own mouse strains. NRCMM also accepts donations of mouse strains from different institutes/PIs. All mouse strains enlisted in NRCMM repository are available to both non-profit and for-profit organizations. A generic MTA, attached with modified items from respective owners, is required before shipment of these strains. Headquarters of NRCMM also keeps the frozen sperms for all mouse strains. Information of the NRCMM mice can be found at www.nrcmm.cn (the database is undergoing re-construction at this moment). Because the majority mouse strains belong to GPT, GPT created an independent database for people searching for these GPT mice at www.gempharmatech.com.
In the earlier years of the NRCMM, we created the tamoxifen-induced smooth muscle–specific myosin light chain kinase (MLCK) knockout mice. This model, the first cKO (conditional knockout) mouse developed in China, provided the direct evidence in vivo that MLCK play central role in phasic contractions in smooth muscle (He et al. 2008). In neurological research fields, NRCMM also helped to define the role of FSTL1 expression in dorsal root ganglion for pain sensing and role of FGF13 expression in the neocortex and the hippocampus for learning and memory (Li et al. 2011; Wu et al. 2012). Soon after the discovery of CRISPR/Cas9 gene editing technology, we managed to generate knockout mouse models in China in April 2013 (Shen et al. 2013). NRCMM also helped scientists in China to create the first CRISPR/Cas9 based knockout cynomolgus monkey and dog in the world (Ma et al. 2014; Niu et al. 2014).
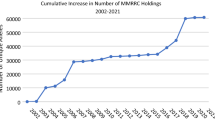
In total, NRCMM, with the effort of all the co-operators, have developed over 18,500 (updated to Oct, 2021) genetically engineered mouse strains, including knockout/conditional knockout mice, Cre transgenic lines, mutant knockin models, immuno-deficient models, and humanized mouse models. Many of these strains may serve as models for metabolic diseases, development defects, cardiovascular diseases and neurodegenerative diseases (Table 1).
NRCMM is the founding member of Asia Mouse Mutagenesis and Resource Association (AMMRA), a pan-Asian organization for promoting and coordinating the development, archiving/distribution, phenotyping, and informatics of mutant mice, and for facilitating access to mice resources in Asia. NRCMM also joined International Mouse Phenotyping Consortium (IMPC) in 2011, which aims to create a comprehensive catalogue of mouse gene function.
CKO/KO mice in Knockout All Project
With the support of sophisticated genetic manipulating technique, interruption or knockout of a specific gene in ES cells and mice have been developed for almost two decades (Goldstein 2001), and knockout mice showed excellent track record as models for elucidation of human gene function and disease mechanism. The systematic targeting of all mouse protein encoding genes would provide a comprehensive and standardized resource, thus benefit the academic and industrial community. Since its complexity and huge expense, such an undertaking usually required the deployment and coordination of resources on a global scale, and three global knockout projects have been launched in the US, Canada and European countries (Austin et al. 2004; Collins et al. 2007). This project is continued by International Mouse Phenotyping consortium (IMPC), an collaborative effort by 21 research institutions to identify the function of every protein-coding gene in the mouse genome. To this end, the IMPC has systematically switched off or ‘knocked out’ 10,125 mouse genes (updated to Oct, 2021), and systematical analysis of these mutant strains are providing transformational insights into the function of these genes (Brown 2021; Cacheiro et al. 2020). The information of IMPC targeted gene can be accessible through GenTar (https://www.gentar.org/tracker/#/).
Although the global knockout mouse projects have achieved great accomplishment, many genes still remain to be knocked out for comprehensively studying gene function (Peterson and Murray 2021). In Gempharmatech, we launched our Knockout All Project (KOAP) in 2018 with ambition to mutagenize all protein-coding gene in mouse genome within 5 years. KOAP creates mouse models via a high throughput CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) based gene editing strategy. Specifically, the CRISPR editing system, including the enzyme and guide RNAs designed to cleave the target genomic loci, and the repair donors carrying loxP sites, were injected into mouse zygotes. Two guide RNAs and two separate donors (each carrying a single loxP site) are used to target each specific gene without incorporating extra cassettes such as selection markers. Typically, generation of both the KO allele (fragment deletion product) and the cKO allele (expected recombination product) could be achieved in one single targeting event.
The schematics for flox sites insertion and QC approaches are presented in Online Resource 2. Briefly, exon(s) encoding the critical functional domain(s) are floxed or deleted in the KOAP mouse strains. The inserting sites of the loxP were carefully selected to avoid disruption the normal expression and function of the targeted gene(s) prior to Cre/loxP excision in the floxed strains. To reduce the risk of disruption of gene expression regulation or initiation of the de novo translation start site when the canonical start codon is deleted, the subsequent 5’ end coding exon(s) right after the 1st coding exon harboring ATG is preferably selected to be floxed. If the critical functional domain(s) are difficult to be floxed, we will introduce a frame-shift into the mRNA transcript after Cre mediated excision to ensure the disruption of the target protein. The size of the floxed region is preferably smaller than 5 kb. In most cases, premature termination codon (PTC) will be produced by frame-shift and the resultant transcripts are predicted to be degraded by the Nonsene-Mediated Decay (NMD) pathway. In certain circumstances, if predicted in-frame start codon that would potentially restart the translation or the main functional domain is included in the first exon, a flox region larger than 5 kb is acceptable in our designs.The Statistics of flox interval was plotted in Online resource 3. However, in some cases, there is only one exon, or the functional domain is encoded by the 1st exon, the whole gene will be floxed. Moreover, according to the published cases using flox mouse strains from our repository, Cre/loxP excision of fragments around 30 kb in size is applicable (Wang et al. 2021; Qu et al. 2021). Finally, we prefer introns larger than 600 bp for inserting of loxP. The predicted functional motifs, such as the splicing branch site (YNCTRAY), will remain intact in our designs. For KOAP project, C57BL/6JGpt (C57BL/6J substrain) was selected as the background strain to facilitate future breeding of the cKO models with established Cre lines. The strategy for each KOAP line could be found in our repository’s website www.gempharmatech.com.
The whole project is estimated to be accomplished in 2023. With the highly efficient gene editing technique and straightforward targeting strategies, we have established the high throughput gene targeting pipeline with capacity of creating around 5000 cKO strains each year. Till now, there were in total 9769 KO strains and 8004 cKO strains completed by KOAP (updated to Oct, 2021). Production timeline of KOAP is presented in Fig. 1A
Overview of KOAP (updated to Oct, 2021). A Production timeline of KOAP. B The comparison of alleles between IMPC and KOAP. C The statistics of Cre lines. 165 of 199 Cre strains have been recorded in MGI and 34 strains are novel. CreER versus constitutive, 52 versus 147; H11/R26 KI versus 3' endogenous KI alleles, 49 versus 123
.
We further compared the overlap alleles between IMPC and KOAP null mutant mice. For the IMPC project, there are currently 10,125 genes from either ES cell or CRISPR projects in different states of 'Attempt in Progress', 'Chimeras/Founder Obtained' (if ES cells) or 'Embryo Obtained' (if CRISPR), and 'Genotype Confirmed'. For our KOAP project, there are 11,063 targeted genes for either KO, cKO strains or both (KOAP strains are all in F0 or F1 stage). Therefore, total 15,050 protein-coding genes have been targeted by these two projects. IMPC and KOAP have targeted only 6,138 overlapping genes, indicating both repositories are useful to support the biomedical research community (Fig. 1B).
Transgenic mice that express recombinase in a specific manner are necessary for generation of conditional knockout mouse. Thus, we generated 199 mouse strains expressing Cre recombinase in specific tissues, such as immune system, nervous system, bone, muscle and adipocytes, gastrointestinal tract, cardiovascular system, reproductive system and so on. In order to ensure the appropriate expression of the Cre, we developed comprehensive strategies to generate the Cre lines. When the promoter of the driver gene is well defined, the defined promoter will be selected to direct the specific expression of Cre. Specifically, to avoid affecting other genes, the transgenic element harboring the driver promoter, the Cre coding sequence and a polyA will be introduced into safe harbor sites such as Gt(ROSA)26 (Friedrich and Soriano 1991) or Hipp11 (Hippenmeyer et al. 2010) locus to establish the Cre expressing strains. However, when the promoter and regulatory elements of the driver gene are unknown, we tend to introduce the Cre coding sequence into the endogenous gene to achieve the Cre expressing under the control of inherent regulators, i.e., inserting the Cre coding sequence after the ATG code if the driver gene works well in a heterozygote. Moreover, putting the Cre coding before the stop condon to avoid inactivating the driver gene. As presented in Fig. 1C, 123 strains are created using the in situ insertion strategies; 49 Cre strains are generated via recombinase cassette insertion into ROSA26 or H11 loci, and 27 Cre lines are transgenic strains with random integration of recombinase cassette into mouse genome. Although some of Cre driven genes are not first reported by our work, the strategies we used here (ROSA26/H11 and in situ insertion) to reproduce the Cre lines could provide the information of recombinase location, which will facilitate to make the breeding strategies to generate cKO models. Moreover, all the Cre lines are created in C57BL/6 strains, making them feasible to breed with flox strains created in KOAP to produce cKO mice. Codon-Improved Cre Recombinase (iCre), demonstrating to express at a higher level in mouse, was employed in most GPT Cre lines to allow high efficacy excision (Shimshek et al. 2002). The Cre mouse strains with the driving promoters and the mutation type have been listed in the Online Resource 4. So far, 39 of 199 Cre lines have been bred with flox strains to generate cKO mice or validated with Cre reporters as indicated in Online Resource 4 (updated to Oct, 2021).
Scientists could access the interested mouse strains through our repository’s website www.gempharmatech.com. To better illustrate the interaction network of multiple genes and facilitate scientists to choose the connected genes, we developed the pathway searching option. Pathway searching is a graphic online tool that incorporated most of our KOAP strains into several classic signaling pathway networks. People interested to a specific pathway related genes can easily find the available CKO/KO mouse strains linking with target genes. As exemplified by Akt signaling pathway schematic in Fig. 2, the genes labeled by orange circles are directly linked to the available target strains in our repository. We highly recommend scientists to use the pathway searching to find all the connected genes.
Example of signaling pathway schematic for cKO/KO strains searching. Target gene could be searched through signaling pathways in our website www.gempharmatech.com. The genes highlighted by orange circles link to the available cKO/KO strains
Humanized mouse strains
Tremendous progress was made in understanding human diseases such as cancer, autoimmune disease, and infectious diseases in the last decades. Consequently, great success have been achieved in drug development. However, the species barrier between mouse and human severely limited accurate prediction of drug efficacy and safety study in mouse models. Humanized mice, the chimeric mice with human cells, tissues, organs, or carrying human genes, are pivotal models to fill the translational gap in the preclinical studies (Aartsma-Rus and van Putten 2019; De La Rochere et al. 2018).
Human immune system (HIS) models
Because functional immune system would reject xenograft in mice, various immune-compromised strains were established to ensure the engraftment of human cells, tissues and organs. IL2Rγ is a shared subunit of several interleukin receptors that transduce signals required for differentiation and function of hematopoietic cells. Inactivation of Il2rg gene blocks NK cell development and eventually leads to NK cell defects (Cao et al. 1995). Prkdc is required for the V(D)J recombination process during the B cell and T cell development, and loss-of-function mutation of Prkdc leads to T and B lymphocytes defect (Blunt et al. 1995). The scid mice carrying Prkdc mutation, which led to elimination of mature T and B cells, were broadly used to engraft human tumors and facilitated the drug studies over 50 years (Bosma and Carroll 1991; Pelleitier and Montplaisir 1975). Additionally, NOD strain exhibited reduction of innate immunity, with defective macrophage activity, reduced dendritic cell function, and lack of the hemolytic complement system (Shultz et al. 1995). Thus, we disrupted the Prdkc and Il2rg genes directly in NOD mice and developed the NCG model with enhanced engraftment of human immune cells and tissues. The immune system of NCG was severely impaired due to the defects of functional T, B, NK cells, the complement system, the dendritic cells and macrophages (data not shown), showing quite similar features to that of the previously developed NSG (Ishikawa et al. 2005; Shultz et al. 2005) and NOG (Ito et al. 2002; Yahata et al. 2003) strains. Besides broadly used to constitute cell-derived xenograft (CDX)and patient-derived xenograft (PDX)models (Yang et al. 2019; Zheng et al. 2020), NCG is also an ideal model for generation of human immune system (HIS), as sustainable long-term hematopoiesis of the engrafted human hematopoiesis stem cell (HSC) could be maintained in NCG. This model has been widely distributed over the world and applied in cancer research, regeneration medicine and infectious disease research (Cao et al. 2017; He et al. 2019; Lee et al. 2017; Ludwik et al. 2018; Poggio et al. 2019; Wen et al. 2018; Xia et al. 2020), as well as drug efficacy evaluation in preclinical studies. Although the long-term engraftment of human immune cells, mainly the T cells, was achieved in NCG, other populations of immune cells were challenging to be reconstituted, largely due to the lack of essential cytokines or environment for proliferation, differentiation, and maturation of the human cells. Previous works have reported that expression of human cytokines in immune-compromised mice could improve engraftment of human immune cells. For example, knock-in of human IL2 or IL15 gene in severely immune deficient mice implanted with human HSC cells could enhance human NK cells differentiation as reported (Herndler-Brandstetter et al. 2017; Huntington et al. 2009; Katano et al. 2015; Matsuda et al. 2019); knockout of mouse Flt3 could improve human DC and ILC cells development (Douam et al. 2018; Li et al. 2016; Lopez-Lastra et al. 2017); overexpression of mouse tslp could recover murine lymphatic tissues in BRSG, thus benefit the maturation of reconstituted human T/B cells (Li et al. 2018). Thus, we generate the next generation of NCG, with certain intensive traits such as human cytokines and human HLA expression along with enhanced human cells engraftment, to create more suitable models for immune cells differentiation which are accessible for academic community (Table 2). The enhanced engraftment of human immune cells has been validated in our lab and data could be download in our website when search interested strains.
Immune checkpoint humanized mice
The promising therapeutic effects of immune checkpoint antibodies such as CTLA4 antibody Ipilimumab (Leach et al. 1996; Weber et al. 2009) and PD1 antibody Pembrolizumab (Syn et al. 2017) have extensively promoted the investment and development of macromolecular drugs in recent years. However, species barriers made the mouse not a suitable model in the preclinical evaluation of antibodies that recognize unique human targets. The human immune system (HIS) models engrafted with human tumors were applicable for the preclinical study of the checkpoint inhibitors (Wang et al. 2018). However, failure of reconstitution of different immune cell populations made such HIS models lack a fully functional immune system. Immune checkpoint gene humanized mouse strains with the competent immune system were developed to address the species obstacles between mouse and human. Over 60 immune checkpoint humanized models, e.g., PD1, CTLA4, SIRPα, TIGIT, and LAG-3, have been established at NRCMM.
Mono- and Combo- therapies against immune checkpoints are under active development in recent years. A clinical study announced in the 2020 ASCO meeting showed meaningful improvement in overall response rates (ORR) and progression-free survival (PFS) in NSCLC with combo-treatment by Tiragolumab (TIGIT antibody) and Atezolizumab (PDL1 antibody) (Rodriguez-Abreu et al. 2020). The outstanding clinical performance of the TIGIT antibody, which is a novel inhibitory immune checkpoint mainly expressed on activated T cells and NK cells in the tumor microenvironment, attracted a great deal of attention (Chauvin and Zarour 2020; Dietze et al. 2013; Solomon and Garrido-Laguna 2018). Treg cells with high TIGIT expression were found to be immune-suppressive in the tumor environment (Joller et al. 2014; Kurtulus et al. 2015). As presented in the 2018 AACR meeting, the anti-TIGIT antibody EOS884448 was proven to kill Tregs in vitro when carrying hIgG1, but not other isotypes. Similarly, the surrogate antibody of EOS884448 with the mIgG2a isotype (the homolog of human IgG1), rather than mIgG1 (the homolog of human IgG4), inhibited the CT26 tumor growth in vivo significantly (Dekkers et al. 2017; LEROY et al. 2018). These results indicated that Fc-mediated Tregs depletion might be a critical mechanism of action (MOA) for therapeutic anti-TIGIT antibodies. To enable the direct in vivo evaluation of human TIGIT antibodies in preclinical animal models, we generated TIGIT mono-humanized and TIGIT/PD1 double-humanized mice in both C57BL/6 and BALB/c background which are prone to Th1 and Th2 predominant immune response respectively (Sellers et al. 2012). Interestingly, the antitumor effect of human TIGIT antibody (Tiragolumab analog, hIgG1 isotype) was recapitulated in humanized mice in BALB/c engrafted with CT26 tumor but not in humanized C57BL/6 mice with MC38 tumor. Furthermore, noticeable Tregs depletion and robust NK cells increase were observed in tumor-infiltrating lymphocytes (TILs) analysis in the BALB/c background models, while no significant change was identified in C57BL/6 background models. Additionally, the potent antitumor effect of Tiragolumab analog was eliminated while using the DANA variant (FcγR-mediated ADCC blockage) to treat tumor bearing mice (data not shown). Together, these results indicated that the antitumor activity of TIGIT antibody was attributed to the Fc-mediated Tregs depletion through NK cells, which was consistent with previous reports (LEROY et al. 2018; Yang et al. 2020). Thus, TIGIT humanized mouse model in BALB/c background not only recapitulated the antitumor efficacy, but also the mechanism of action (MOA), of the therapeutic TIGIT antibodies. Also, we found obviously different responsiveness of PD1 antibodies when tested in PD1 humanized models in BALB/c and C57BL/6 background (data not shown), which might cause by the distinct immune profile, such as proportion of Treg (Chen et al. 2005) and M2 macrophages (Martinez 2011; Sellers et al. 2012), and as well as Th2 cytokines production (Trunova et al. 2011), between BALB/c and C57BL/6 background.
Given the importance of the genetic background of mouse models on the immune responsiveness to cancer treatment, we developed immune checkpoint humanized mouse models on C57BL/6 and BALB/c, respectively. These models, especially humanized models in BALB/c background, would provide tools for better understanding of disease and investigation of therapies for the academic and industrial communities.
ACE2-humanized mouse models
The COVID-19 pandemic outbreak at the end of 2019 is caused by severe acute respiratory syndrome coronavirus 2(SARS-CoV-2) (Ou et al. 2020). SARS-CoV-2 enter host cells via the angiotensin-converting enzyme 2 (ACE2) receptor, which has been identified as common receptor for various viruses, such as SARS-CoV (Kuhn et al. 2004). Since SARS-CoV or SARS-CoV-2 does not bind efficiently to the mouse ACE2 (Qiu et al. 2020), ACE2-humanized mice are indispensable animal models for studying the mechanism of ACE2 related to the COVID-19 outbreak and developing specific drugs, antibodies, and vaccines. We have generated several hACE2 strains based on different strategies and mouse genetic background as listed in Table 3 since COVID-19 outbreak. The strain HuACE2-FL(T037659), with full length human ACE2 CDS in situ replacement of mouse Ace2, showed moderate alveolar septal thickening and inflammatory cell infiltration following SARS-CoV-2 challenge in the work presented by Qiao et al. (2021). In this study, they designed 32 protease inhibitors targeting the main protease (Mpro) of SARS-CoV-2, the one plays a central role in viral replication, and two of these showed strong antiviral activity with significantly reduced lung viral loads and lung lesions in HuACE2-FL(T037659) mice (Qiao et al. 2021). The strain K18-HuACE2 (T037657) was developed by inserting hACE2 CDS driven by the human cytokeratin 18 (K18) promoter targeting epithelial cells. Human ACE2 mRNA was detected with the highest expression level in lung and also in other tissues, including the heart, kidney and intestine of the transgenic mice (data not shown). More severe lung pathologic changes with massive infiltrating cells and obvious alveolar necrosis following influenza A virus (IAV) and SARS-CoV-2 co-infection were presented in this mouse model (Bai et al. 2021). Another strain HuACE2-Chimera (T037630) was generated by in situ replacement of mouse Ace2 with a chimeric ACE2 containing a murine signal peptide, a human extracellular domain, a murine transmembrane domain and intracellular domain. Ma et al. evaluated nanoparticle vaccines efficacy in this transgenic mouse, and reported that high titer of Receptor Binding Domain (RBD) specific IgG after RBD and RBD-HR vaccines immunized were produced, followed by the challenge of authentic SARS-CoV-2 (Ma et al. 2020). Severe inflammation and swelling of the whole lung after infected with SARS-CoV-2S-RBD were also displayed in this strain (Alitongbieke et al. 2020). The information including model strategies could be found in our website www.gempharmatech.com. The users can also access to the ACE2 information as well as the strategies in MGI-Find Mice (IMSR, repository-GPT).
Healthy and genetic quality control
Healthy monitoring is important for providing high quality mouse models. We preformed the strict SPF monitoring program including necropsy, histopathological analysis and serologic, bacteriologic and parasitic tests. Sentinel mice exposed to dirty bedding in each rack are used for periodic pathogens monitoring, and pathogens need to be tested and excluded are listed in Table 4.
Genetic monitoring is another important quality control to confirm strain identity and generate congenic lines. We use single nucleotide polymorphisms (SNPs), the most abundant class of markers in genome, to distinguish genetically close strains (Mekada et al. 2009). Genome-wide SNP scanning panels were developed for 13 inbred strains including a total of 794 SNP loci, and performed every year for genetic monitoring of inbred and genetically engineering strains (Table 4).
Genetic modifications of these mouse strains were confirmed by PCR-based genotyping or southern blot. More validation studies, such as tissue specific expression/knockout analysis and pathological assessment were also performed based on the model types and their application. All validation data, model design strategies and genotyping protocol could be downloaded from website for individual model.
Future perspective
The GEMMs for rare disease research initiative
Rare diseases (RDs), the one do not yet have universal definition, refer to any disease with low prevalence, which comprise more than 7000 different conditions (Montserrat Moliner and Waligora 2013). They are usually life-threatening, chronic and clinically complex, affecting 3.5–5.9% of the world’s population (Murillo-Cuesta et al. 2020). With the lacking of larger enough market and economic gains, they are also known as orphan diseases, meaning difficulty to gain support and resources for discovering treatment. Although greater attention being paid to RDs worldwide, leading to increasing number of studies of RDs and new drug development, more than 95% of the RDs currently have no effective treatment (Kaufmann et al. 2018). Accordingly, RDs are considered a public health priority, and specific research programs like the International Rare Diseases Research Consortium (IRDiRC) have been established, with the goal of accelerating diagnosis and approving 1000 new therapies for RDs by 2027 (Gahl et al. 2016; Murillo-Cuesta et al. 2020). NRCMM joined China Alliance for Rare Disease in 2020. The China Alliance for Rare Disease (CHARD, https://www.chard.org.cn/) is a national, non-profit, cooperative exchange platform for rare diseases, approved by the national health authority, and supported by Peking Union Medical College Hospital, PhiRDA, the Chinese Medical Association, the Chinese Hospital Association and the Chinese Research Hospital Association, as well as multiple universities, scientific research institutes and enterprises. Alliance aims to improve standardized rare disease diagnosis, treatment and education for patients and the medical community in China.
RDs are excellent candidates for generation of GEMMs, as most RDs involve mutations in a single gene. GEMMs are indispensable to identify the genetic bases and molecular mechanisms of RDs, as well as to understand their physiopathology, clinical heterogeneity and genotype–phenotype correlations, and ultimately, to develop intervention or treatment of RDs (Boat and Field 2011). Usually, medical translation moves from bench to clinic, but in the study of RDs, we need change direction to work from clinic to bench. To this end, NRCMM launched another large scale mission, “The GEMMs for Rare Disease Research Initiative” in 2020. The initiative aims to provide high quality and cost effective genetically engineered mouse models (GEMMs) for previously underfunded biomedical research areas. With high throughput pipeline successfully practicing in KOAP and abundant of clinical RDs findings, the initiative is with ambitious to cover all human rare diseases models caused by point mutation, which will serve to illustrate the phenotypic variability of RDs to fill the knowledge gaps in this area. We hope to provide a set of models for clinicians and scientists to create an optimal pathway for translating clinic to bench and then back to clinic, and contribute to drug discovery and development process. We are enthusiastic to collaborate with clinicians and scientists to develop the point mutation models. So far, “The GEMMs for Rare Disease Research Initiative” has generated 87 GEMMs, which are of great value in supporting biomedical research and novel drug research and development in this area. In 2021, we continue to encourage applications for “The GEMMs for Rare Disease Research Initiative” to promote the research and novel therapeutic development for the treatment of rare diseases. The introduction of “The GEMMs for Rare Disease Research Initiative” is linked below
https://www.gempharmatech.us/pages/gem_funding_opportunities.php
We are dedicated to keep expanding our repository by developing various new GEMMs models, e.g. knockout/conditional knockout mouse models, drug target gene humanized models, human disease mouse models. With the high throughput pipelines for mouse phenotyping, we contributed to the phenotyping project of knockout mice initiated by IMPC followed standardized phenotyping protocols from international mouse phenotyping resource of standardized screens (IMPRess). The integration of massive GEMMs resource with standardized phenotyping platform in our center will help academic and industrial community to harnessing the power of the genome to drive biomedical discovery. We are willing to keep contributing to the basic research and drug preclinical studies. By creating a publicly available resource of GEMMs and phenotyping platform, NRCMM will continue to make effort to knock down barriers for biologists to use mouse models in their research.
Data availability
Addition datasets used or analyzed for the current study are available from the corresponding authors on reasonable request.
Code availability
Not applicable.
References
Aartsma-Rus A, van Putten M (2019) The use of genetically humanized animal models for personalized medicine approaches. Dis Model Mech 13:041673
Alitongbieke G, Li X-M, Wu Q-C, Lin Z-C, Huang J-F, Xue Y, Liu J-N, Lin J-M, Pan T, Chen Y-X (2020) Study on β-Chitosan against the binding of SARS-CoV-2S-RBD/ACE2. bioRxiv
Ashizawa T, Iizuka A, Nonomura C, Kondou R, Maeda C, Miyata H, Sugino T, Mitsuya K, Hayashi N, Nakasu Y, Maruyama K, Yamaguchi K, Katano I, Ito M, Akiyama Y (2017) Antitumor effect of programmed death-1 (PD-1) blockade in humanized the NOG-MHC double knockout mouse. Clin Cancer Res 23:149–158
Austin CP, Battey JF, Bradley A, Bucan M, Capecchi M, Collins FS, Dove WF, Duyk G, Dymecki S, Eppig JT (2004) The knockout mouse project. Nat Genet 36:921
Bai L, Zhao Y, Dong J, Liang S, Guo M, Liu X, Wang X, Huang Z, Sun X, Zhang Z, Dong L, Liu Q, Zheng Y, Niu D, Xiang M, Song K, Ye J, Zheng W, Tang Z, Tang M, Zhou Y, Shen C, Dai M, Zhou L, Chen Y, Yan H, Lan K, Xu K (2021) Coinfection with influenza A virus enhances SARS-CoV-2 infectivity. Cell Res 31:395–403
Bao X, Zhang X, Wang L, Wang Z, Huang J, Zhang Q, Ye Y, Liu Y, Chen D, Zuo Y, Liu Q, Xu P, Huang B, Fang J, Lao J, Feng X, Li Y, Kurita R, Nakamura Y, Yu W, Ju C, Huang C, Mohandas N, Li D, Zhao C, Xu X (2021) Epigenetic inactivation of ERF reactivates gamma-globin expression in beta-thalassemia. Am J Hum Genet 108:709–721
Billerbeck E, Barry WT, Mu K, Dorner M, Rice CM, Ploss A (2011) Development of human CD4+FoxP3+ regulatory T cells in human stem cell factor-, granulocyte-macrophage colony-stimulating factor-, and interleukin-3-expressing NOD-SCID IL2Rgamma(null) humanized mice. Blood 117:3076–3086
Blunt T, Finnie NJ, Taccioli GE, Smith GC, Demengeot J, Gottlieb TM, Mizuta R, Varghese AJ, Alt FW, Jeggo PA, Jackson SP (1995) Defective DNA-dependent protein kinase activity is linked to V(D)J recombination and DNA repair defects associated with the murine scid mutation. Cell 80:813–823
Boat TF, Field MJ (2011) Rare diseases and orphan products: Accelerating research and development.
Bosma MJ, Carroll AM (1991) The SCID mouse mutant: definition, characterization, and potential uses. Annu Rev Immunol 9:323–350
Brehm MA, Cuthbert A, Yang C, Miller DM, DiIorio P, Laning J, Burzenski L, Gott B, Foreman O, Kavirayani A, Herlihy M, Rossini AA, Shultz LD, Greiner DL (2010) Parameters for establishing humanized mouse models to study human immunity: analysis of human hematopoietic stem cell engraftment in three immunodeficient strains of mice bearing the IL2rgamma(null) mutation. Clin Immunol 135:84–98
Brehm MA, Kenney LL, Wiles MV, Low BE, Tisch RM, Burzenski L, Mueller C, Greiner DL, Shultz LD (2019) Lack of acute xenogeneic graft- versus-host disease, but retention of T-cell function following engraftment of human peripheral blood mononuclear cells in NSG mice deficient in MHC class I and II expression. FASEB J 33:3137–3151
Brown SDM (2021) Advances in mouse genetics for the study of human disease. Hum Mol Genet 30:R274–R284
Cacheiro P, Munoz-Fuentes V, Murray SA, Dickinson ME, Bucan M, Nutter LMJ, Peterson KA, Haselimashhadi H, Flenniken AM, Morgan H, Westerberg H, Konopka T, Hsu CW, Christiansen A, Lanza DG, Beaudet AL, Heaney JD, Fuchs H, Gailus-Durner V, Sorg T, Prochazka J, Novosadova V, Lelliott CJ, Wardle-Jones H, Wells S, Teboul L, Cater H, Stewart M, Hough T, Wurst W, Sedlacek R, Adams DJ, Seavitt JR, Tocchini-Valentini G, Mammano F, Braun RE, McKerlie C, Herault Y, de Angelis MH, Mallon AM, Lloyd KCK, Brown SDM, Parkinson H, Meehan TF, Smedley D, Genomics England Research C, International Mouse Phenotyping C (2020) Human and mouse essentiality screens as a resource for disease gene discovery. Nat Commun 11:655
Cao X, Shores EW, Hu-Li J, Anver MR, Kelsall BL, Russell SM, Drago J, Noguchi M, Grinberg A, Bloom ET et al (1995) Defective lymphoid development in mice lacking expression of the common cytokine receptor gamma chain. Immunity 2:223–238
Cao Z, Ye T, Sun Y, Ji G, Shido K, Chen Y, Luo L, Na F, Li X, Huang Z, Ko JL, Mittal V, Qiao L, Chen C, Martinez FJ, Rafii S, Ding BS (2017) Targeting the vascular and perivascular niches as a regenerative therapy for lung and liver fibrosis. Sci Transl Med 9
Chauvin JM, Zarour HM (2020) TIGIT in cancer immunotherapy. J Immunother Cancer 8:e000957
Chen X, Oppenheim JJ, Howard OM (2005) BALB/c mice have more CD4+CD25+ T regulatory cells and show greater susceptibility to suppression of their CD4+CD25- responder T cells than C57BL/6 mice. J Leukoc Biol 78(1):114–121. https://doi.org/10.1189/jlb.0604341
Collins FS, Rossant J, Wurst W (2007) A mouse for all reasons. Cell 128:9–13
De La Rochere P, Guil-Luna S, Decaudin D, Azar G, Sidhu SS, Piaggio E (2018) Humanized mice for the study of immuno-oncology. Trends Immunol 39:748–763
Dekkers G, Bentlage AEH, Stegmann TC, Howie HL, Lissenberg-Thunnissen S, Zimring J, Rispens T, Vidarsson G (2017) Affinity of human IgG subclasses to mouse Fc gamma receptors. Mabs 9:767–773
Dietze KK, Zelinskyy G, Liu J, Kretzmer F, Schimmer S, Dittmer U (2013) Combining regulatory T cell depletion and inhibitory receptor blockade improves reactivation of exhausted virus-specific CD8+ T cells and efficiently reduces chronic retroviral loads. PLoS Pathog 9:e1003798
Douam F, Ziegler CGK, Hrebikova G, Fant B, Leach R, Parsons L, Wang W, Gaska JM, Winer BY, Heller B, Shalek AK, Ploss A (2018) Selective expansion of myeloid and NK cells in humanized mice yields human-like vaccine responses. Nat Commun 9:5031
Friedrich G, Soriano P (1991) Promoter traps in embryonic stem cells: a genetic screen to identify and mutate developmental genes in mice. Genes Dev 5:1513–1523
Gahl WA, Mulvihill JJ, Toro C, Markello TC, Wise AL, Ramoni RB, Adams DR, Tifft CJ (2016) The NIH Undiagnosed Diseases Program and Network: Applications to modern medicine. Mol Genet Metab 117:393–400
Goldstein J (2001) Knockout mice and test-tube babies. Nat Med 7:1079–1080
He WQ, Peng YJ, Zhang WC, Lv N, Tang J, Chen C, Zhang CH, Gao S, Chen HQ, Zhi G, Feil R, Kamm KE, Stull JT, Gao X, Zhu MS (2008) Myosin light chain kinase is central to smooth muscle contraction and required for gastrointestinal motility in mice. Gastroenterology 135:610–620
He J, Yang Q, Xiao Q, Lei A, Li X, Zhou P, Liu T, Zhang L, Shi K, Yang Q, Dong J, Zhou J (2019) IRF-7 is a critical regulator of Type 2 innate lymphoid cells in allergic airway inflammation. Cell Rep 29, 2718–2730 e2716
Herndler-Brandstetter D, Shan L, Yao Y, Stecher C, Plajer V, Lietzenmayer M, Strowig T, de Zoete MR, Palm NW, Chen J, Blish CA, Frleta D, Gurer C, Macdonald LE, Murphy AJ, Yancopoulos GD, Montgomery RR, Flavell RA (2017) Humanized mouse model supports development, function, and tissue residency of human natural killer cells. Proc Natl Acad Sci USA 114:E9626–E9634
Hippenmeyer S, Youn YH, Moon HM, Miyamichi K, Zong H, Wynshaw-Boris A, Luo L (2010) Genetic mosaic dissection of Lis1 and Ndel1 in neuronal migration. Neuron 68:695–709
Huntington ND, Legrand N, Alves NL, Jaron B, Weijer K, Plet A, Corcuff E, Mortier E, Jacques Y, Spits H, Di Santo JP (2009) IL-15 trans-presentation promotes human NK cell development and differentiation in vivo. J Exp Med 206:25–34
Ishikawa F, Yasukawa M, Lyons B, Yoshida S, Miyamoto T, Yoshimoto G, Watanabe T, Akashi K, Shultz LD, Harada M (2005) Development of functional human blood and immune systems in NOD/SCID/IL2 receptor gamma chain(null) mice. Blood 106:1565–1573
Ito M, Hiramatsu H, Kobayashi K, Suzue K, Kawahata M, Hioki K, Ueyama Y, Koyanagi Y, Sugamura K, Tsuji K, Heike T, Nakahata T (2002) NOD/SCID/gamma(c)(null) mouse: an excellent recipient mouse model for engraftment of human cells. Blood 100:3175–3182
Joller N, Lozano E, Burkett PR, Patel B, Xiao S, Zhu C, Xia J, Tan TG, Sefik E, Yajnik V, Sharpe AH, Quintana FJ, Mathis D, Benoist C, Hafler DA, Kuchroo VK (2014) Treg cells expressing the coinhibitory molecule TIGIT selectively inhibit proinflammatory Th1 and Th17 cell responses. Immunity 40:569–581
Katano I, Takahashi T, Ito R, Kamisako T, Mizusawa T, Ka Y, Ogura T, Suemizu H, Kawakami Y, Ito M (2015) Predominant development of mature and functional human NK cells in a novel human IL-2-producing transgenic NOG mouse. J Immunol 194:3513–3525
Kaufmann P, Pariser AR, Austin C (2018) From scientific discovery to treatments for rare diseases—the view from the National Center for Advancing Translational Sciences—Office of Rare Diseases Research. Orphanet J Rare Dis 13:196
Kuhn JH, Li W, Choe H, Farzan M (2004) Angiotensin-converting enzyme 2: a functional receptor for SARS coronavirus. Cell Mol Life Sci 61:2738–2743
Kurtulus S, Sakuishi K, Ngiow SF, Joller N, Tan DJ, Teng MW, Smyth MJ, Kuchroo VK, Anderson AC (2015) TIGIT predominantly regulates the immune response via regulatory T cells. J Clin Invest 125:4053–4062
Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, Devon K, Dewar K, Doyle M, FitzHugh W, Funke R, Gage D, Harris K, Heaford A, Howland J, Kann L, Lehoczky J, LeVine R, McEwan P, McKernan K, Meldrim J, Mesirov JP, Miranda C, Morris W, Naylor J, Raymond C, Rosetti M, Santos R, Sheridan A, Sougnez C, Stange-Thomann Y, Stojanovic N, Subramanian A, Wyman D, Rogers J, Sulston J, Ainscough R, Beck S, Bentley D, Burton J, Clee C, Carter N, Coulson A, Deadman R, Deloukas P, Dunham A, Dunham I, Durbin R, French L, Grafham D, Gregory S, Hubbard T, Humphray S, Hunt A, Jones M, Lloyd C, McMurray A, Matthews L, Mercer S, Milne S, Mullikin JC, Mungall A, Plumb R, Ross M, Shownkeen R, Sims S, Waterston RH, Wilson RK, Hillier LW, McPherson JD, Marra MA, Mardis ER, Fulton LA, Chinwalla AT, Pepin KH, Gish WR, Chissoe SL, Wendl MC, Delehaunty KD, Miner TL, Delehaunty A, Kramer JB, Cook LL, Fulton RS, Johnson DL, Minx PJ, Clifton SW, Hawkins T, Branscomb E, Predki P, Richardson P, Wenning S, Slezak T, Doggett N, Cheng JF, Olsen A, Lucas S, Elkin C, Uberbacher E, Frazier M, Gibbs RA, Muzny DM, Scherer SE, Bouck JB, Sodergren EJ, Worley KC, Rives CM, Gorrell JH, Metzker ML, Naylor SL, Kucherlapati RS, Nelson DL, Weinstock GM, Sakaki Y, Fujiyama A, Hattori M, Yada T, Toyoda A, Itoh T, Kawagoe C, Watanabe H, Totoki Y, Taylor T, Weissenbach J, Heilig R, Saurin W, Artiguenave F, Brottier P, Bruls T, Pelletier E, Robert C, Wincker P, Smith DR, Doucette-Stamm L, Rubenfield M, Weinstock K, Lee HM, Dubois J, Rosenthal A, Platzer M, Nyakatura G, Taudien S, Rump A, Yang H, Yu J, Wang J, Huang G, Gu J, Hood L, Rowen L, Madan A, Qin S, Davis RW, Federspiel NA, Abola AP, Proctor MJ, Myers RM, Schmutz J, Dickson M, Grimwood J, Cox DR, Olson MV, Kaul R, Raymond C, Shimizu N, Kawasaki K, Minoshima S, Evans GA, Athanasiou M, Schultz R, Roe BA, Chen F, Pan H, Ramser J, Lehrach H, Reinhardt R, McCombie WR, de la Bastide M, Dedhia N, Blocker H, Hornischer K, Nordsiek G, Agarwala R, Aravind L, Bailey JA, Bateman A, Batzoglou S, Birney E, Bork P, Brown DG, Burge CB, Cerutti L, Chen HC, Church D, Clamp M, Copley RR, Doerks T, Eddy SR, Eichler EE, Furey TS, Galagan J, Gilbert JG, Harmon C, Hayashizaki Y, Haussler D, Hermjakob H, Hokamp K, Jang W, Johnson LS, Jones TA, Kasif S, Kaspryzk A, Kennedy S, Kent WJ, Kitts P, Koonin EV, Korf I, Kulp D, Lancet D, Lowe TM, McLysaght A, Mikkelsen T, Moran JV, Mulder N, Pollara VJ, Ponting CP, Schuler G, Schultz J, Slater G, Smit AF, Stupka E, Szustakowki J, Thierry-Mieg D, Thierry-Mieg J, Wagner L, Wallis J, Wheeler R, Williams A, Wolf YI, Wolfe KH, Yang SP, Yeh RF, Collins F, Guyer MS, Peterson J, Felsenfeld A, Wetterstrand KA, Patrinos A, Morgan MJ, de Jong P, Catanese JJ, Osoegawa K, Shizuya H, Choi S, Chen YJ, Szustakowki J, International Human Genome Sequencing C (2001) Initial sequencing and analysis of the human genome. Nature 409:860–921
Leach DR, Krummel MF, Allison JP (1996) Enhancement of antitumor immunity by CTLA-4 blockade. Science 271:1734–1736
Lee S, Kivimae S, Szoka FC (2017) Clodronate improves survival of transplanted Hoxb8 myeloid progenitors with constitutively active GMCSFR in immunocompetent mice. Mol Ther Methods Clin Dev 7:60–73
Leroy X, Hoofd C, Cuende J, Denies S, Rabolli V, Preillon J, Lambolez F, Prasad S, Mercier M, Nyawouame F, Bodo V, Wald N, Driessens G, Detheux M (2018) Abstract LB-114: a-TIGIT antagonist antibody EOS884448 shows dual mechanism of action by restoration of T cell effector functions and preferential depletion of Treg. Cancer Res 78:LB114
Li KC, Zhang FX, Li CL, Wang F, Yu MY, Zhong YQ, Zhang KH, Lu YJ, Wang Q, Ma XL, Yao JR, Wang JY, Lin LB, Han M, Zhang YQ, Kuner R, Xiao HS, Bao L, Gao X, Zhang X (2011) Follistatin-like 1 suppresses sensory afferent transmission by activating Na+, K+-ATPase. Neuron 69:974–987
Li Y, Mention JJ, Court N, Masse-Ranson G, Toubert A, Spits H, Legrand N, Corcuff E, Strick-Marchand H, Di Santo JP (2016) A novel Flt3-deficient HIS mouse model with selective enhancement of human DC development. Eur J Immunol 46:1291–1299
Li Y, Masse-Ranson G, Garcia Z, Bruel T, Kok A, Strick-Marchand H, Jouvion G, Serafini N, Lim AI, Dusseaux M, Hieu T, Bourgade F, Toubert A, Finke D, Schwartz O, Bousso P, Mouquet H, Di Santo JP (2018) A human immune system mouse model with robust lymph node development. Nat Methods 15:623–630
Lopez-Lastra S, Masse-Ranson G, Fiquet O, Darche S, Serafini N, Li Y, Dusseaux M, Strick-Marchand H, Di Santo JP (2017) A functional DC cross talk promotes human ILC homeostasis in humanized mice. Blood Adv 1:601–614
Ludwik KA, McDonald OG, Brenin DR, Lannigan DA (2018) ERalpha-mediated nuclear sequestration of RSK2 is required for ER(+) breast cancer tumorigenesis. Cancer Res 78:2014–2025
Ma Y, Zhang X, Shen B, Lu Y, Chen W, Ma J, Bai L, Huang X, Zhang L (2014) Generating rats with conditional alleles using CRISPR/Cas9. Cell Res 24:122–125
Ma X, Zou F, Yu F, Li R, Yuan Y, Zhang Y, Zhang X, Deng J, Chen T, Song Z, Qiao Y, Zhan Y, Liu J, Zhang J, Zhang X, Peng Z, Li Y, Lin Y, Liang L, Wang G, Chen Y, Chen Q, Pan T, He X, Zhang H (2020) Nanoparticle vaccines based on the receptor binding domain (RBD) and heptad repeat (HR) of SARS-CoV-2 elicit robust protective immune responses. Immunity 53(1315–1330):e1319
Madsen L, Labrecque N, Engberg J, Dierich A, Svejgaard A, Benoist C, Mathis D, Fugger L (1999) Mice lacking all conventional MHC class II genes. Proc Natl Acad Sci USA 96:10338–10343
Martinez FO (2011) Regulators of macrophage activation. Eur J Immunol 41(6):1531–1534. https://doi.org/10.1002/eji.201141670
Masse-Ranson G, Dusseaux M, Fiquet O, Darche S, Boussand M, Li Y, Lopez-Lastra S, Legrand N, Corcuff E, Toubert A, Centlivre M, Bruel T, Spits H, Schwartz O, Levy Y, Strick-Marchand H, Di Santo JP (2019) Accelerated thymopoiesis and improved T-cell responses in HLA-A2/-DR2 transgenic BRGS-based human immune system mice. Eur J Immunol 49:954–965
Matsuda M, Ono R, Iyoda T, Endo T, Iwasaki M, Tomizawa-Murasawa M, Saito Y, Kaneko A, Shimizu K, Yamada D, Ogonuki N, Watanabe T, Nakayama M, Koseki Y, Kezuka-Shiotani F, Hasegawa T, Yabe H, Kato S, Ogura A, Shultz LD, Ohara O, Taniguchi M, Koseki H, Fujii SI, Ishikawa F (2019) Human NK cell development in hIL-7 and hIL-15 knockin NOD/SCID/IL2rgKO mice. Life Sci Alliance 2:e201800195
McIntosh BE, Brown ME, Duffin BM, Maufort JP, Vereide DT, Slukvin II, Thomson JA (2015) Nonirradiated NOD, B6.SCID Il2rgamma-/- Kit(W41/W41) (NBSGW) mice support multilineage engraftment of human hematopoietic cells. Stem Cell Rep 4:171–180
Mekada K, Abe K, Murakami A, Nakamura S, Nakata H, Moriwaki K, Obata Y, Yoshiki A (2009) Genetic differences among C57BL/6 substrains. Exp Anim 58:141–149
Montserrat Moliner A, Waligora J (2013) The European union policy in the field of rare diseases. Public Health Genomics 16:268–277
Mouse Genome Sequencing C, Waterston RH, Lindblad-Toh K, Birney E, Rogers J, Abril JF, Agarwal P, Agarwala R, Ainscough R, Alexandersson M, An P, Antonarakis SE, Attwood J, Baertsch R, Bailey J, Barlow K, Beck S, Berry E, Birren B, Bloom T, Bork P, Botcherby M, Bray N, Brent MR, Brown DG, Brown SD, Bult C, Burton J, Butler J, Campbell RD, Carninci P, Cawley S, Chiaromonte F, Chinwalla AT, Church DM, Clamp M, Clee C, Collins FS, Cook LL, Copley RR, Coulson A, Couronne O, Cuff J, Curwen V, Cutts T, Daly M, David R, Davies J, Delehaunty KD, Deri J, Dermitzakis ET, Dewey C, Dickens NJ, Diekhans M, Dodge S, Dubchak I, Dunn DM, Eddy SR, Elnitski L, Emes RD, Eswara P, Eyras E, Felsenfeld A, Fewell GA, Flicek P, Foley K, Frankel WN, Fulton LA, Fulton RS, Furey TS, Gage D, Gibbs RA, Glusman G, Gnerre S, Goldman N, Goodstadt L, Grafham D, Graves TA, Green ED, Gregory S, Guigo R, Guyer M, Hardison RC, Haussler D, Hayashizaki Y, Hillier LW, Hinrichs A, Hlavina W, Holzer T, Hsu F, Hua A, Hubbard T, Hunt A, Jackson I, Jaffe DB, Johnson LS, Jones M, Jones TA, Joy A, Kamal M, Karlsson EK, Karolchik D, Kasprzyk A, Kawai J, Keibler E, Kells C, Kent WJ, Kirby A, Kolbe DL, Korf I, Kucherlapati RS, Kulbokas EJ, Kulp D, Landers T, Leger JP, Leonard S, Letunic I, Levine R, Li J, Li M, Lloyd C, Lucas S, Ma B, Maglott DR, Mardis ER, Matthews L, Mauceli E, Mayer JH, McCarthy M, McCombie WR, McLaren S, McLay K, McPherson JD, Meldrim J, Meredith B, Mesirov JP, Miller W, Miner TL, Mongin E, Montgomery KT, Morgan M, Mott R, Mullikin JC, Muzny DM, Nash WE, Nelson JO, Nhan MN, Nicol R, Ning Z, Nusbaum C, O’Connor MJ, Okazaki Y, Oliver K, Overton-Larty E, Pachter L, Parra G, Pepin KH, Peterson J, Pevzner P, Plumb R, Pohl CS, Poliakov A, Ponce TC, Ponting CP, Potter S, Quail M, Reymond A, Roe BA, Roskin KM, Rubin EM, Rust AG, Santos R, Sapojnikov V, Schultz B, Schultz J, Schwartz MS, Schwartz S, Scott C, Seaman S, Searle S, Sharpe T, Sheridan A, Shownkeen R, Sims S, Singer JB, Slater G, Smit A, Smith DR, Spencer B, Stabenau A, Stange-Thomann N, Sugnet C, Suyama M, Tesler G, Thompson J, Torrents D, Trevaskis E, Tromp J, Ucla C, Ureta-Vidal A, Vinson JP, Von Niederhausern AC, Wade CM, Wall M, Weber RJ, Weiss RB, Wendl MC, West AP, Wetterstrand K, Wheeler R, Whelan S, Wierzbowski J, Willey D, Williams S, Wilson RK, Winter E, Worley KC, Wyman D, Yang S, Yang SP, Zdobnov EM, Zody MC, Lander ES (2002) Initial sequencing and comparative analysis of the mouse genome. Nature 420:520–562
Murillo-Cuesta S, Artuch R, Asensio F, de la Villa P, Dierssen M, Enriquez JA, Fillat C, Fourcade S, Ibanez B, Montoliu L, Oliver E, Pujol A, Salido E, Vallejo M, Varela-Nieto I (2020) The value of mouse models of rare diseases: a Spanish experience. Front Genet 11:583932
Niu Y, Shen B, Cui Y, Chen Y, Wang J, Wang L, Kang Y, Zhao X, Si W, Li W, Xiang AP, Zhou J, Guo X, Bi Y, Si C, Hu B, Dong G, Wang H, Zhou Z, Li T, Tan T, Pu X, Wang F, Ji S, Zhou Q, Huang X, Ji W, Sha J (2014) Generation of gene-modified cynomolgus monkey via Cas9/RNA-mediated gene targeting in one-cell embryos. Cell 156:836–843
Ou X, Liu Y, Lei X, Li P, Mi D, Ren L, Guo L, Guo R, Chen T, Hu J, Xiang Z, Mu Z, Chen X, Chen J, Hu K, Jin Q, Wang J, Qian Z (2020) Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat Commun 11:1620
Pelleitier M, Montplaisir S (1975) The nude mouse: a model of deficient T-cell function. Methods Achiev Exp Pathol 7:149–166
Peterson KA, Murray SA (2021) Progress towards completing the mutant mouse null resource. Mamm Genome
Poggio M, Hu T, Pai CC, Chu B, Belair CD, Chang A, Montabana E, Lang UE, Fu Q, Fong L, Blelloch R (2019) Suppression of exosomal PD-L1 induces systemic anti-tumor immunity and memory. Cell 177, 414–427 e413
Qiao J, Li YS, Zeng R, Liu FL, Luo RH, Huang C, Wang YF, Zhang J, Quan B, Shen C, Mao X, Liu X, Sun W, Yang W, Ni X, Wang K, Xu L, Duan ZL, Zou QC, Zhang HL, Qu W, Long YH, Li MH, Yang RC, Liu X, You J, Zhou Y, Yao R, Li WP, Liu JM, Chen P, Liu Y, Lin GF, Yang X, Zou J, Li L, Hu Y, Lu GW, Li WM, Wei YQ, Zheng YT, Lei J, Yang S (2021) SARS-CoV-2 M(pro) inhibitors with antiviral activity in a transgenic mouse model. Science 371:1374–1378
Qiu Y, Zhao YB, Wang Q, Li JY, Zhou ZJ, Liao CH, Ge XY (2020) Predicting the angiotensin converting enzyme 2 (ACE2) utilizing capability as the receptor of SARS-CoV-2. Microbes Infect 22:221–225
Qu H, Miao T, Wang Y, Tan L, Huang B, Zhang L, Liu X, Long M, Zhang R, Liao X, Gong X, Wang J, Xiong X, Liu J, Li X, Yu J, Yang G, Zhu Z, Zheng H, Zheng Y (2021) Dedicator of cytokinesis 5 regulates keratinocyte function and promotes diabetic wound healing. Diabetes 70:1170–1184
Rahmig S, Kronstein-Wiedemann R, Fohgrub J, Kronstein N, Nevmerzhitskaya A, Bornhauser M, Gassmann M, Platz A, Ordemann R, Tonn T, Waskow C (2016) Improved human erythropoiesis and platelet formation in humanized NSGW41 mice. Stem Cell Rep 7:591–601
Rodriguez-Abreu D, Johnson ML, Hussein MA, Cobo M, Patel AJ, Secen NM, Lee KH, Massuti B, Hiret S, Yang JC-H (2020) Primary analysis of a randomized, double-blind, phase II study of the anti-TIGIT antibody tiragolumab (tira) plus atezolizumab (atezo) versus placebo plus atezo as first-line (1L) treatment in patients with PD-L1-selected NSCLC (CITYSCAPE). American Society of Clinical Oncology
Schmidt MR, Appel MC, Giassi LJ, Greiner DL, Shultz LD, Woodland RT (2008) Human BLyS facilitates engraftment of human PBL derived B cells in immunodeficient mice. PLoS ONE 3:e3192
Sellers RS, Clifford CB, Treuting PM, Brayton C (2012) Immunological variation between inbred laboratory mouse strains: points to consider in phenotyping genetically immunomodified mice. Vet Pathol 49:32–43
Shen B, Zhang J, Wu H, Wang J, Ma K, Li Z, Zhang X, Zhang P, Huang X (2013) Generation of gene-modified mice via Cas9/RNA-mediated gene targeting. Cell Res 23:720–723
Shultz LD, Lyons BL, Burzenski LM, Gott B, Chen X, Chaleff S, Kotb M, Gillies SD, King M, Mangada J, Greiner DL, Handgretinger R (2005) Human lymphoid and myeloid cell development in NOD/LtSz-scid IL2R gamma null mice engrafted with mobilized human hemopoietic stem cells. J Immunol 174:6477–6489
Shimshek DR, Kim J, Hübner MR, Spergel DJ, Buchholz F, Casanova E, Stewart AF, Seeburg PH, Sprengel R (2002) Codon-improved Cre recombinase (iCre) expression in the mouse. Genesis 32(1):19–26
Shultz LD, Schweitzer PA, Christianson SW, Gott B, Schweitzer IB, Tennent B, McKenna S, Mobraaten L, Rajan TV, Greiner DL et al (1995) Multiple defects in innate and adaptive immunologic function in NOD/LtSz-scid mice. J Immunol 154:180–191
Solomon BL, Garrido-Laguna I (2018) TIGIT: a novel immunotherapy target moving from bench to bedside. Cancer Immunol Immunother 67:1659–1667
Syn NL, Teng MWL, Mok TSK, Soo RA (2017) De-novo and acquired resistance to immune checkpoint targeting. Lancet Oncol 18:e731–e741
Trunova GV, Makarova OV, Diatroptov ME, Bogdanova IM, Mikchailova LP, Abdulaeva SO (2011) Morphofunctional characteristic of the immune system in BALB/c and C57Bl/6 mice. Bull Exp Biol Med 151(1):99–102. https://doi.org/10.1007/s10517-011-1268-1
van Lent AU, Dontje W, Nagasawa M, Siamari R, Bakker AQ, Pouw SM, Maijoor KA, Weijer K, Cornelissen JJ, Blom B, Di Santo JP, Spits H, Legrand N (2009) IL-7 enhances thymic human T cell development in “human immune system” Rag2-/-IL-2Rgammac-/- mice without affecting peripheral T cell homeostasis. J Immunol 183:7645–7655
Wang M, Yao LC, Cheng M, Cai D, Martinek J, Pan CX, Shi W, Ma AH, De Vere White RW, Airhart S, Liu ET, Banchereau J, Brehm MA, Greiner DL, Shultz LD, Palucka K, Keck JG (2018) Humanized mice in studying efficacy and mechanisms of PD-1-targeted cancer immunotherapy. FASEB J 32:1537–1549
Wang Y, Su Y, Yu G, Wang X, Chen X, Yu B, Cheng Y, Li R, Saez JC, Yi C, Xiao L, Niu J (2021) Reduced Oligodendrocyte Precursor Cell Impairs Astrocytic Development in Early Life Stress. Adv Sci (weinh) 8:e2101181
Waskow C, Madan V, Bartels S, Costa C, Blasig R, Rodewald HR (2009) Hematopoietic stem cell transplantation without irradiation. Nat Methods 6:267–269
Weber J, Thompson JA, Hamid O, Minor D, Amin A, Ron I, Ridolfi R, Assi H, Maraveyas A, Berman D, Siegel J, O’Day SJ (2009) A randomized, double-blind, placebo-controlled, phase II study comparing the tolerability and efficacy of ipilimumab administered with or without prophylactic budesonide in patients with unresectable stage III or IV melanoma. Clin Cancer Res 15:5591–5598
Wen H, Ma H, Cai Q, Lin S, Lei X, He B, Wu S, Wang Z, Gao Y, Liu W, Liu W, Tao Q, Long Z, Yan M, Li D, Kelley KW, Yang Y, Huang H, Liu Q (2018) Recurrent ECSIT mutation encoding V140A triggers hyperinflammation and promotes hemophagocytic syndrome in extranodal NK/T cell lymphoma. Nat Med 24:154–164
Wu QF, Yang L, Li S, Wang Q, Yuan XB, Gao X, Bao L, Zhang X (2012) Fibroblast growth factor 13 is a microtubule-stabilizing protein regulating neuronal polarization and migration. Cell 149:1549–1564
Xia K, Ma Y, Feng X, Deng R, Ke Q, Xiang AP, Deng C (2020) Endosialin defines human stem Leydig cells with regenerative potential. Hum Reprod 35:2197–2212
Yahata T, Ando K, Sato T, Miyatake H, Nakamura Y, Muguruma Y, Kato S, Hotta T (2003) A highly sensitive strategy for SCID-repopulating cell assay by direct injection of primitive human hematopoietic cells into NOD/SCID mice bone marrow. Blood 101:2905–2913
Yang W, Fan WS, Ye MX, Li Z, Gu CL, Zhu YP, Hao YP, Wang ZQ, Wang L, Meng YG (2019) Establishment of the PDTX model of gynecological tumors. Am J Transl Res 11:3779–3789
Yang F, Zhao L, Wei Z, Yang Y, Liu J, Li Y, Tian X, Liu X, Lu X, Sui J (2020) A cross-species reactive TIGIT-blocking antibody Fc dependently confers potent antitumor effects. J Immunol 205:2156–2168
Yu H, Borsotti C, Schickel JN, Zhu S, Strowig T, Eynon EE, Frleta D, Gurer C, Murphy AJ, Yancopoulos GD, Meffre E, Manz MG, Flavell RA (2017) A novel humanized mouse model with significant improvement of class-switched, antigen-specific antibody production. Blood 129:959–969
Zhang S, Liu Y, Wang X, Yang L, Li H, Wang Y, Liu M, Zhao X, Xie Y, Yang Y, Zhang S, Fan Z, Dong J, Yuan Z, Ding Z, Zhang Y, Hu L (2020) SARS-CoV-2 binds platelet ACE2 to enhance thrombosis in COVID-19. J Hematol Oncol 13:120
Zheng Y, Li C, Xin P, Peng Q, Zhang W, Liu S, Zhu X (2020) Calreticulin increases growth and progression of natural killer/T-cell lymphoma. Aging (albany, NY) 12:23822–23835
Acknowledgements
We appreciate the assistance of Luis Santos and his colleagues in IMPC for providing the null mutant alleles information of IMPC project. This project is supported partially by the grants to National Resource Center of Mutant Mice of China from Ministry of Science and Technology.
Funding
NRCMM is partially supported by Ministry of Science and Technology of China. This work is also funded by Collaborative Innovation of Industry, University and Research Institute Major Program of Guangzhou (201604046020).
Author information
Authors and Affiliations
Contributions
XG, CJ and JL developed the concept of this manuscript. JL, MZ, JZ, LL, SC performed the literature research, data collection and analysis. JL, MZ drafted the manuscript and XG, CJ, JZ and SC critically revised the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures involving animals were approved by Institutional Animal Care and Use Committee (IACUC) of GemPharmatech Co.Ltd.
Consent to participate
Informed consent was obtained from all individual participants included in this work.
Consent for publication
Consent for publication was obtained from all authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ju, C., Liang, J., Zhang, M. et al. The mouse resource at National Resource Center for Mutant Mice. Mamm Genome 33, 143–156 (2022). https://doi.org/10.1007/s00335-021-09940-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00335-021-09940-x