Abstract
PIWI-interacting RNAs (piRNAs), small noncoding RNAs 24–35 nucleotides long, are essential for animal fertility. They play critical roles in a range of functions, including transposable element suppression, gene expression regulation, imprinting, and viral defense. In mammals, piRNAs are the most abundant small RNAs in adult testes and the only small RNAs that direct epigenetic modification of chromatin in the nucleus. The production of piRNAs is a complex process from transcription to post-transcription, requiring unique machinery often distinct from the biogenesis of other RNAs. In mice, piRNA biogenesis occurs in specialized subcellular locations, involves dynamic developmental regulation, and displays sexual dimorphism. Furthermore, the genomic loci and sequences of piRNAs evolve much more rapidly than most of the genomic regions. Understanding piRNA biogenesis should reveal novel RNA regulations recognizing and processing piRNA precursors and the forces driving the gain and loss of piRNAs during animal evolution. Such findings may provide the basis for the development of engineered piRNAs capable of modulating epigenetic regulation, thereby offering possible single-dose RNA therapy without changing the genomic DNA. In this review, we focus on the biogenesis of piRNAs in mammalian adult testes that are derived from long non-coding RNAs. Although piRNA biogenesis is believed to be evolutionarily conserved from fruit flies to humans, recent studies argue for the existence of diverse, mammalian-specific RNA-processing pathways that convert precursor RNAs into piRNAs, perhaps associated with the unique features of mammalian piRNAs or germ cell development. We end with the discussion of major questions in the field, including substrate recognition and the birth of new piRNAs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
PIWI-interacting RNAs (piRNAs) are small non-coding RNAs that form 1:1 RNA–protein complexes with PIWI (P-element-induced wimpy testis) proteins. The PIWI gene was first identified in 1997, the disruption of which leads to defects in germ stem cell maintenance in Drosophila (Lin and Spradling 1997). Further studies revealed that a conserved family of PIWI genes with an essential function in germ cells is widely distributed in both vertebrates and invertebrates (Chirn et al. 2015; Murchison et al. 2008; Wynant et al. 2017). Three PIWI paralogs are found in mice, MIWI, MILI, and MIWI2, the deletion of any one of which leads to male infertility (Aravin et al. 2007b; Deng and Lin 2002; Carmell et al. 2007) (Fig. 1). piRNAs were first reported in 2001 in Drosophila as small silencing RNAs with distinct features from known microRNAs (miRNAs) or small interference RNAs (siRNAs) (Aravin et al. 2001). Although research in Tetrahymena in 2002 indicated that PIWI proteins bind small RNAs and the RNA–protein complex functions together in genome rearrangement (Box 1), the name “piRNA” was not coined until a group of studies published in 2006 revealed that a set of germ cell-specific small RNAs associate with PIWI proteins in both Drosophila and mammals(Vagin et al. 2006; Girard et al. 2006; Aravin et al. 2006; Grivna et al. 2006; Lau et al. 2006). These studies uncovered the features that distinguish piRNAs from other small RNAs: (1) piRNAs have a characteristic length of 24–35 nucleotides (nt), which is longer than both miRNAs and endogenous siRNAs; (2) 2′-O-methyl modifications occur at the 3′-end of piRNAs; and (3) piRNAs associate with PIWI proteins, a subclade of Argonaute proteins. While later studies have confirmed the distribution of piRNAs across bilateral animals (Grimson et al. 2008; Lewis et al. 2018), and piRNA biogenesis is believed to be evolutionarily conserved from fruit flies to humans (Czech and Hannon 2016a; Gainetdinov et al. 2018), recent studies argue for the existence of diverse, mammalian-specific RNA-processing pathways that convert precursor RNAs into piRNAs, probably associated with the unique features of mammalian piRNAs or germ cell development. Therefore, this review focuses primarily on mouse piRNAs, although piRNAs from other organisms will be discussed when relevant.
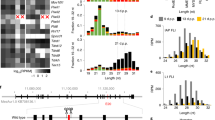
Key events in mouse spermatogenesis and their associated piRNA and PIWI gene expression. The top two panels show germ cell development stages and corresponding key events. The third panel from the top shows piRNA expression levels. piRNA abundance is measured by small RNA sequencing based on (Li et al. 2013) and unpublished data. The bottom panel shows the expression profiles of three PIWI genes in mice: Miwi Mili and Miwi2, DSB Double-strand break, TE transposable element. Dash line represents putative data, IMC intermitochondrial cement, CB chromatoid body
Multi-faceted piRNAs—classifying piRNAs
In mice, piRNAs are divided into three major classes based on their origins (Table 1): transposable element (TE) piRNAs from DNA transposons, endogenous retroviruses, long interspersed nuclear elements (LINEs), or short interspersed nuclear elements (SINEs); intergenic piRNAs from non-coding regions of the genome; and 3′untranslated region (3′UTR) piRNAs that map to the 3´UTRs of protein-coding regions in the sense orientation. TE piRNAs protect the germline genome from TE activation, a function that is evolutionarily conserved in bilateral animals (Kumar and Carmichael 1998; Aravin and Hannon 2008; Farazi et al. 2008; Thomson and Lin 2009; Grimson et al. 2008). TE piRNAs are the dominant piRNA class in Drosophila and zebrafish, whereas intergenic piRNAs are the dominant piRNA class in adult mammalian testes. Both 3′UTR piRNAs and intergenic piRNAs are mostly uniquely mapped to the genome and lack well-identified targets. A small fraction of intergenic piRNAs have been shown to base-pair with and trigger the decay of mRNAs required for sperm maturation (Goh et al. 2015; Gou et al. 2014; Zhang et al. 2015; Wu et al. 2020). 3′UTR piRNAs have been detected in Drosophila, frogs, and diverse mammalian species (Robine et al. 2009; Chirn et al. 2015). These 3′UTR piRNAs are derived from full-length mRNA precursors rather than cryptic transcripts corresponding exclusively to 3′UTRs, and their biogenesis is coupled with translation thus fine-tuning protein synthesis from their mRNA precursors (Sun et al. 2021).
piRNAs can be categorized into primary piRNAs and secondary piRNAs based on their biogenesis pathways, which are principally distinguished by their mechanisms of 5′end formation. Primary piRNAs are produced from long single-stranded RNA precursors that are synthesized from piRNA loci, and their 5′ends are produced during the fragmentation process, likely involving the MitoPLD endonuclease (or PLD6, which has a homolog “Zucchini” in Drosophila) located on mitochondrial outer membranes. Secondary piRNAs are produced from piRNA-targeted transcripts, and their 5′ends are generated by the endonucleolytic activity of PIWI proteins, which cleaves between positions 10 and 11 of the base-pair complementary RNA target relative to the piRNA 5′end. These secondary piRNAs can target the primary piRNA precursor transcripts to generate more secondary piRNAs, resulting in a piRNA-specific “Ping-Pong” loop. This loop is believed to represent an adaptive immune response that enables piRNAs to silence TE transcripts post-transcriptionally, as the loop continues to produce TE piRNAs until the TE transcripts are diminished (Gunawardane et al. 2007; Brennecke et al. 2007). In Drosophila, secondary piRNAs can also trigger a Zucchini-dependent “phased” piRNA biogenic mechanism that resembles primary piRNA biogenesis but only occurs downstream of the initial PIWI-catalyzed cleavage event in a 5′-to-3′ stepwise manner, generating non-overlapping fragments known as pre-piRNAs. After loading onto PIWI proteins, the pre-piRNAs (23–42 nt) are further trimmed and methylated to become mature piRNAs (Mohn et al. 2015; Han et al. 2015; Homolka et al. 2015; Ding et al. 2017; Gainetdinov et al. 2018; Darricarrère et al. 2013; Ishizu et al. 2015).
Based on the dynamics of their expression, mouse piRNAs can be further classified into prenatal piRNAs, pre-pachytene piRNAs, pachytene piRNAs, and hybrid piRNAs. Prenatal piRNAs are TE-rich and associated with MILI and MIWI2 proteins. MILI-bound prenatal piRNAs silence piRNA target transcripts in the cytosol. MIWI2-bound prenatal piRNAs are shuttled to the nucleus to recruit epigenetic machinery to direct the DNA methylation (Schöpp et al. 2020) of TEs around 13.5–15.5 days post coitum (dpc) (Aravin et al. 2008; Kuramochi-Miyagawa et al. 2008). Knockout of MILI or MIWI2 leads to TE desilencing; however, the catalytic activity of MIWI2 is not required(De Fazio et al. 2011), indicating that MILI-bound piRNAs are sufficient to trigger a robust cytosolic Ping-Pong reaction and the formation of MIWI2-bound secondary piRNAs. Thus, the current data suggest that a subset of MILI cleavage products is loaded to MILI staying in the cytosol to trigger a Ping-Pong loop and another subset of cleavage products is loaded to MIWI2, and the MILI cleavage also triggers downstream phased piRNA production loading to MIWI2 (Yang et al. 2016). Pre-pachytene piRNAs, expressed after birth, have the lowest abundance among the four groups and are composed of TE piRNAs and 3′UTR piRNAs. They are associated with MILI and are essential for silencing TEs during the mitotic and meiotic stages of adult spermatogenesis (Di Giacomo et al. 2013). Pachytene piRNAs, along with MIWI proteins, are produced during the pachytene stage of meiosis. They, associated with both MILI and MIWI, are generally poor in TE complementary sequences and are mostly derived from long non-coding RNA (lncRNA) precursors synthesized in intergenic regions. Pachytene piRNAs represent the most abundant class of small RNAs in the adult testis, around 5.7 to 7.2 μM per cell (Gainetdinov et al. 2018). Pachytene piRNAs are produced via a Ping-Pong independent mechanism (Beyret et al. 2012), and, indeed, MIWI endonucleolytic cleavage activity is not required for pachytene piRNA production. Hybrid piRNAs are a class of piRNAs with features of both pachytene piRNAs (being present at increased levels during the pachytene stage and derived from lncRNAs) and pre-pachytene piRNAs (which map to mRNA 3′UTRs) (Li et al. 2013). Hybrid piRNA activation during pachytene stage is driven by the same transcription factor A-MYB as pachytene piRNAs, and their 3′UTRs are embedded with TE sequences, thus producing TE piRNAs that trigger Ping-Pong loops silencing TEs (Sun et al. 2021).
Transcription of piRNA loci
Despite piRNAs being the most heterogenous small non-coding RNA in animals with > 1 million unique sequences detected in individual germlines from most animal species that have been studied (Vagin et al. 2006; Aravin et al. 2006; Lewis et al. 2018), they are produced from discrete genomic loci, historically defined using computational methods and named piRNA clusters (Brennecke et al. 2007; Gainetdinov et al. 2018). Later work has defined the transcriptional start sites, polyA cleavage sites, promoters, and splice sites of the transcriptional units in the piRNA loci in mice (Li et al. 2013). Among all classes of murine piRNAs, we understand the transcriptional regulation of pachytene piRNAs in the most detail. Unlike convergent transcribed dual-stranded piRNA loci in Drosophila and chickens, piRNA loci in mammals are unidirectionally or bidirectionally transcribed similar to the transcription of protein-coding genes (Gould et al. 2012; Sun et al. 2017; Li et al. 2013; Yu et al. 2021). The single-strand pachytene piRNA precursors, ranging from 500 to 80,000 nt (Betel et al. 2007; Li et al. 2013), are generated with the activation of the transcription factor A-MYB (Li et al. 2013) (Fig. 2). A-MYB-mediated transcriptional regulation of pachytene piRNAs is conserved in amniotes (Li et al. 2013), and A-MYB also regulates the mRNAs coding for piRNA biogenic proteins, forming a feedforward loop to ensure the robust activation of piRNA production during pachytene stage. Unlike promoters from protein-coding genes, which often have a high CpG content, the promoters of pachytene piRNA loci have a low CpG content and are heavily methylated (Yu et al. 2021). RNA polymerase II transcribes these pachytene piRNA genes, and the RNAs undergo conventional mRNA processing, including 5′-capping and polyA tailing. Pachytene piRNA precursors often contain introns that are removed by splicing, indicating that splicing occurs during piRNA precursor synthesis. Those precursors that harbor a long first exon require an additional biogenic factor, BTBD18 (BTB Domain Containing 18), to facilitate their transcriptional elongation (Zhou et al. 2017; Yu et al. 2021). Pachytene piRNA precursors also bind THOC1 and THOC2, THO complex subunits, for their nuclear export (Yu et al. 2021) and are eventually localized to the surface of mitochondria for primary piRNA biogenesis (Li et al. 2013; Murano et al. 2019; Fabry et al. 2019) (Fig. 2).
Current model of pachytene piRNA biogenesis in mouse testis. Facilitated by A-MYB and BTBD18, pachytene piRNA precursor transcripts are synthesized by RNA polymerase II, containing a 5′-cap and a poly(A) tail. Introns of these precursors are spliced out, and then these precursors are transported from the nucleus to the cytoplasm and further located to the IMC. Presumably, endonuclease PLD6 on the outer membrane of mitochondria cleaves the piRNA precursors and generates the 5′ ends of future piRNAs. In the first phase of piRNA biogenesis, ribosomes translate the uORF region and piRNAs are produced in a TDRD5-independent manner. In the second phase, ribosomes translocate to the UDR region, facilitated by MOV10L1, and guide piRNA production in a TDRD5-dependent manner. Finally, these cleaved products will be loaded onto MILI or MIWI protein for further 3′ end maturations that PNLDC1 trims and HEN1 adds the 2′-O-methyl group to the end of piRNAs. Moreover, MOV10L1 and TDRD5 bind directly to pachytene piRNA precursors. Figure created with BioRender.com
Post-transcriptional processing of piRNA precursors
The post-transcriptional processing of all primary piRNA precursors can be simplified to three steps: 5′end formation, PIWI loading, and 3′end formation (Fig. 2). The 5′ends of primary piRNAs are formed when piRNA precursors are fragmented, likely by the MitoPLD endonuclease in mice based on structural and biochemical studies on its homologs in Drosophila (Gao and Frohman 2012; Kabayama et al. 2017) and silkworm (Izumi et al. 2020). In mice, MitoPLD, previously shown to be a phospholipid-hydrolyzing enzyme, contains an N-terminal mitochondrial targeting signal and is located on the outer membrane of mitochondria (Gao and Frohman 2012; Kabayama et al. 2017). However, structural studies of Drosophila ZUCCHINI suggests that its active site resembles the bacterial endonuclease Nuc, and Zucchini displays endonuclease activities in vivo (Voigt et al. 2012). During Drosophila phased piRNA biogenesis, ZUCCHINI continuously cleaves the single-stranded RNAs and simultaneously generates the 3′end of the pre-piRNA (before 3′-trimming occurs) and the 5′-end of the next, immediately adjacent pre-pre-piRNA. The cleaved piRNA intermediates, pre-pre-piRNAs, are loaded onto PIWI proteins to trigger the next ZUCCHINI-dependent cleavages between the 3′end of the pre-piRNAs and 5′end of the pre-pre-piRNAs. Recently, the cleavage of single-stranded RNAs by silkworm ZUCCHINI has been recapitulated in vivo (Izumi et al. 2020). MOV10L1 is an RNA helicase required for piRNA biogenesis. It interacts with PIWI proteins stably, while its interaction with pachytene piRNA precursors is transient, requiring crosslink to be detected (Zheng et al. 2010; Vourekas et al. 2015). MOV10L1 is thought to load PIWI proteins onto the pre-pre-piRNAs. The 3′ends of pre-piRNAs undergo trimming by an exonuclease (Trimmer in Drosophila, PNLDC1 in mice) and 2′-O-methylation by HENMT1 (Hayashi et al. 2016; Kirino and Mourelatos 2007; Ohara et al. 2007).
Substrate recognition of piRNA precursors
The mechanisms that identify an RNA for piRNA processing in mice are currently unknown. As 5′end formation is upstream of PIWI loading and 3′end formation, the main question is what specifies the initial cleavages on the piRNA precursors. Based on the current model from Drosophila, the initiation of ZUCCHINI-dependent phased piRNA processing requires an initiator piRNA that has base-pair complementarity to the piRNA precursors. In Drosophila, the initiator piRNAs are provided maternally to the eggs in the germplasm and the cells with germplasm differentiate into germ cells. The maternally deposited initiator RNAs are the secondary piRNAs of the previous (F0) generation, which target the primary piRNA precursors in the germ cells of F1 generation and initiate primary piRNA production in a phased manner. The secondary piRNAs generated from the primary piRNAs will be deposited for primary piRNA production in the following generation (F2) and so on. However, unlike Drosophila germ cells, whose fates are pre-determined by the presence of germplasm where the piRNAs are deposited (Santos and Lehmann 2004), mammalian germ cells are induced de novo from somatic cells during embryogenesis (Nicholls et al. 2019). Thus, the biogenesis of pre-natal piRNAs in mice must have been initiated de novo. Similarly, de novo phased piRNA biogenesis through unknown mechanisms occurs in somatic cells of Drosophila, where piRNAs do not display Ping-Pong signatures (Homolka et al. 2015). Although it has been proposed that pre-pachytene piRNAs could serve as initiator piRNAs for pachytene piRNA biogenesis (Gainetdinov et al. 2018; Czech and Hannon 2016a), PIWI protein is below the detection limit in spermatocytes before pachytene piRNA production commences (Di Giacomo et al. 2013), arguing against the existence of abundant initiator piRNAs for pachytene piRNA biogenesis being provided from the pre-pachytene stage. Thus, it remains unclear whether the phased biogenesis of pachytene piRNAs is initiated by initiator piRNAs or by a de novo mechanism.
Other than the presence or source of initiator piRNAs, two major biogenic differences between mouse pachytene piRNAs and Drosophila germline piRNAs lie with their precursors. First, mouse pachytene piRNA precursors are long and continuous transcripts. Although both types of precursors are single-stranded RNAs, Drosophila germline piRNA precursors are cryptic transcripts that are synthesized in a heterochromatin-dependent and promoter-independent mechanism (Andersen et al. 2017), whereas pachytene piRNA precursors have defined transcriptional initiation sites and polyA cleavage sites and are long continuous RNAs up to 80,000 nt in length. Drosophila germline piRNA precursors are derived from highly repetitive regions, whereas mouse pachytene piRNA precursors are depleted of TE sequences in comparison to the rest of the genome. Given that downstream phased piRNA production declines with distance (Mohn et al. 2015; Han et al. 2015), which may be due to the instability of cleavage products prior to PIWI loading, more frequent cleavage promoted by initiator piRNAs would be required to produce more phased piRNAs in Drosophila (Mohn et al. 2015). However, the lack of repetitive elements and low abundance of piRNA prior to the pachytene stage argue against frequent initiator piRNA-mediated cleavage on pachytene piRNA precursors. Because of the low number of cleavages, if any, promoted by initiator piRNAs, phased pachytene piRNA biogenesis must be highly processive, as single-stranded RNAs are known to form secondary structures and bind to RNA-binding proteins. This requirement for high processivity of phased production may explain why ribosomes are involved in piRNA biogenesis in vertebrates, as elongating ribosomes are strong helicases (Takyar et al. 2005; Qu et al. 2011), unwinding secondary structures or stripping RNA-binding proteins off the precursors.
Second, mouse pachytene piRNA precursors are translated. In Drosophila, the RNA granules where piRNA precursors are processed are in close proximity to the nucleus, allowing direct channeling of the precursors. However, such attachment between the nucleus and RNA granules is not observed in mice. This may cause increased dwell time for piRNA precursors in the cytosol and can promote ribosome translation of the upstream open reading frame (uORF) of piRNA precursors, which are nonetheless annotated as lncRNAs (without long or conserved ORFs) (Sun et al. 2020). After translation of the uORFs, MOV10L1 facilitates the translocation of 80S ribosomes into the uORF downstream regions that produce the majority of piRNAs (Yabuta et al. 2011; Zheng and Wang 2012; Sun et al. 2018; Ding et al. 2018; Guan et al. 2021). Endonucleolytic cleavage occurs on ribosome-bound piRNA precursors near the ribosome E site, generating the pre-pre-piRNAs with ribosomes bound at their 5′-extremities(Sun et al. 2020) (Fig. 2). Given that ribosomes are actively moving along the UDR, based on runoff assays after translation inhibition, the current model proposes that once the ribosomes translocate downstream, PIWI proteins are loaded onto the 5′P end of pre-pre-piRNAs and trigger MitoPLD-dependent cleavage between the ribosomes and PIWI proteins. Although this ribosome-guided piRNA biogenesis mechanism is detected in chickens and lizards, it does not appear to exist in invertebrates (Izumi et al. 2020), nor does it function for regions of uORFs and 5′UTRs of pachytene piRNA precursors in mice. The phenotype of Tdrd5 mutant mice demonstrates the distinct biogenesis pathways that operate 5′ (upstream) and 3′ (downstream) of the stop codon of uORFs, as only the production of piRNAs from uORF downstream regions requires TDRD5, a Tudor domain protein that binds to pachytene piRNA precursors (Ding et al. 2018) (Fig. 2). Thus, it is possible that the ribosome-guided, TDRD5-dependent mechanism has specifically evolved for long continuous piRNA precursors. The differences between mice and Drosophila piRNA precursors suggest that more mammalian-specific biogenic factors involved in substrate recognition and coordination of substrate processing with ribosome translocation have yet to be discovered.
Summary of piRNA biogenic factors
Mouse piRNA biogenic factors, along with their homologs in Drosophila, include transcription factors (A-MYB, BTBD18), PIWI proteins (MIWI2, MILI, MIWI), multiple Tudor domain-containing proteins, ribosomes, endonucleases (MitoPLD), 3′end maturation enzymes (PNLDC1, HENMT1), and other co-factors (MOV10L1, DDX4, GTSF1, MAEL, etc.) (Table 2). Some factors perform functions that are highly conserved between Drosophila to mammalians, such as MitoPLD (PLD6, or Zucchini in Drosophila) and S-adenosylmethionine (SAM)-dependent methyltransferase (Hen1 in Drosophila; HENMT1 in mice) (Table 2). On the other hand, some factors, representing homologous proteins between Drosophila and mice, have gained novel functions through gene duplication, such as MOV10 and MOV10L1, both of which are mouse homologs of Drosophila Armitage. A-MYB, BTBD18, and ribosomes represent factors specific to amniote (including mammals and birds) piRNA biogenesis (Li et al. 2013; Zhou et al. 2017; Sun et al. 2020). Tudor domain-containing proteins (Gan et al. 2019) are essential for piRNA biogenesis in both Drosophila and mice, including TDRD1, TDRD2 (TDRKH), TDRD4 (RNF17), TDRD5, TDRD6, TDRD7, TDRD9, and TDRD12 (ECAT8) (Table 2). The 3′end trimming protein in mice is a poly(A)-specific ribonuclease-like domain-containing 1 (PNLDC1), an ortholog protein missing in Drosophila but present as PARN-1 in C. elegans and silkworm. Drosophila instead uses the miRNA-trimming enzyme Nibbler to shorten piRNAs (Liu et al. 2011; Han et al. 2011; Feltzin et al. 2015), indicating the existence of diverse mechanisms for piRNA 3′end formation (Czech and Hannon 2016b).
Location for piRNA biogenesis and function: RNA granules
Key piRNA biogenic factors, such as PNLDC1, TDRKH, MitoPLD, GASZ, and GPAT2, are found on the outer membrane of mitochondria (Honda et al. 2013; Czech et al. 2013; Ma et al. 2009; Shiromoto et al. 2013; Vagin et al. 2013; Nishimasu et al. 2012; Ipsaro et al. 2012; Haase et al. 2010) where specific RNA granules are located between adjacent mitochondria, called intermitochondrial cement (IMC). A more general term for these RNA- and protein-rich structures in germ cells are germinal granules (or germline granules, or germ granules) (Meikar et al. 2011). These germinal granules are also referred to as a “nuage” (French for “cloud”) due to their amorphous shape, the absence of surrounding membranes, and the abundance of RNA and proteins (Nishida et al. 2007; Seto et al. 2007; Harris and Macdonald 2001; Gunawardane et al. 2007; Brennecke et al. 2007; Meikar et al. 2011). So far, nuages have been found in the germ cells of both vertebrates and invertebrates (Eddy 1975). Nuage morphology, localization, and/or biochemical properties change as germ cells develop (Eddy 1974, 1975; Aravin et al. 2009; Chuma et al. 2009). In mammals, before birth, two types of germinal granules exist: IMC and piP-bodies (Aravin et al. 2009). piP-bodies harbor MIWI2, TDRD9, and MAEL, which are required for secondary piRNAs that will shuttle from the cytosol to nucleus for de novo DNA methylation. piP-bodies are lost after birth, but IMC structures remain until the pachytene stage. Multiple proteins are associated with IMC structures, including MILI, MIWI, TDRD1, TDRD6, TDRD7, TDRD9, MVH (DDX4), and MAEL (Meikar et al. 2011; Castaneda et al. 2014; Soper et al. 2008; Sienski et al. 2012; Aravin et al. 2009; Findley et al. 2003; Matsumoto et al. 2015; Kuramochi-Miyagawa et al. 2010; Xiol et al. 2014; Wenda et al. 2017; Shoji et al. 2009; Tanaka et al. 2011; Patil et al. 2014; Hosokawa et al. 2007; Vasileva et al. 2009; Nishida et al. 2009; Zamparini et al. 2011; Kuramochi-Miyagawa et al. 2004; Deng and Lin 2002). During early pachytene stage, pachytene piRNA precursors are transported to the IMC for piRNA processing. In late pachytene stage, the IMC disappears, and its components diffuse throughout the cytosol (Onohara and Yokota 2012). However, soon after meiosis the components from previous IMC structures aggregate into a single large (∼1 μm) perinuclear granule called a chromatoid body (CB) in haploid round spermatids, which can be clearly observed under the microscope (Meikar et al. 2011).
The CB is recognized by immunofluorescence staining using MVH (DDX4), MILI, and MIWI (Kotaja and Sassone-Corsi 2007; Wang et al. 2009; Meikar et al. 2014, 2011). During spermiogenesis, the CB is initially located near the acrosome and is closely associated with the nuclear envelope (Fawcett et al. 1970). The CB then migrates from the acrosomal region to the caudal pole on the other side of the cell, where it dissociates from the nuclear envelope and remains near the newly grown flagellum (Fawcett et al. 1970). As the sperm flagellum grows, the CB forms a ring structure close to the annulus of flagellum and moves with the annulus, encircling the flagellum (Parvinen 2005; Fawcett et al. 1970). During this process, the CB gradually decreases in size and undergoes progressive disaggregation (Fawcett et al. 1970). In addition to piRNA pathway proteins, the CB also harbors machinery for the miRNA pathway and non-sense mediated mRNA-decay (NMD) pathway (Kotaja and Sassone-Corsi 2007). In sum, for the three types of germinal granules, we propose that IMC is specific for piRNA biogenesis, piP-bodies generate MIWI2-piRNAs in preparation for their nuclear function, and the CB is specific for post-meiosis piRNA function. The functional relevance of processing piRNAs near mitochondria remains unclear, as well as the mechanisms that localize the piRNA precursors to IMC and the significance of CB formation for piRNA function.
Developmental timing for piRNA production
Over 90% of the piRNAs in the adult testis are expressed during the pachytene stage of meiosis. Since these pachytene piRNAs are not required for the completion of meiosis, production at the pachytene stage, rather than a functional necessity, is more likely to be a biogenesis requirement, with the pachytene stage likely providing optimal conditions for the massive production of piRNAs. Meiosis prophase I can be divided into five phases based on chromosome behavior: leptotene, zygotene, pachytene, diplotene, and diakinesis. The pachytene stage, when paternal and maternal chromosomes undergo synapsis and pair with each other, takes the longest time, lasting over a week in mice. The formation of the synaptonemal complex, a ladder-like series of parallel threads that form between homologous chromosomes, is a hallmark of the pachytene stage(Li and Schimenti 2007; Reynolds et al. 2007). Synapsis in mice is coupled with double-strand break repair, resulting in crossing-over. Because male mammals are the heterogametic sex and thus have sex chromosomes of different sizes, the sex chromosomes in male spermatocytes cannot be completely synapsed. A process known as meiotic silencing of unsynapsed chromatin (MSUC) (Turner et al. 2006; Khalil et al. 2004) results in entire sex chromosomes in male spermatocytes being transcriptionally inactive during the pachytene stage. MSUC has two consequences for piRNA biogenesis. First, none of the pachytene piRNA loci reside on sex chromosomes (Li et al. 2013). Second, transcriptional inactivation during synapsis leads to a decoupling of translation and transcription. For example, phosphoglycerate kinase 2 (PGK-2) is transcribed at pachytene stage and is temporally translationally suppressed until the round spermatid stage (Geisinger et al. 2021; Fine et al. 2019; Jamsai et al. 2015; Gold et al. 1983). Thus, piRNA biogenesis factor mRNAs that are translated at the pachytene stage need to be sorted separately from mRNAs experiencing translation repression. While most meiosis research focuses on chromosomal behavior during meiosis prophase, little is known about how transcription, splicing, polyadenylation, RNA exportation, and translation of mRNAs are orchestrated to initiate, promote, and exit meiosis. Our recent studies indicates that ribosome recycling factors as well as NMD pathways are specifically inhibited at pachytene stage. Otherwise, these pathways would compete the substrates with the ribosome-guided piRNA biogenesis (Sun et al. 2021; Shum et al. 2016). It is thus possible the sophisticated temporospatial mRNA regulation that enables pachytene progression also facilitates pachytene piRNA production.
Sexual dimorphism of piRNA pathways
In contrast to flies and zebrafish, where defective piRNA pathways results in sterility in both sexes (Ketting 2011), piRNA pathway disruption in mice only leads to sterile males, whereas females remain fertile (Carmell et al. 2007; Kuramochi-Miyagawa et al. 2004). This sexual dimorphism could be due to oocytes having a silencing mechanism mediated by endogenous siRNAs, which prevents TE activation (Tam et al. 2008; Watanabe et al. 2008). However, the activation of siRNAs in oocytes is rodent specific and not found in bovines nor humans, suggesting that this piRNA-independent defense mechanism is not present in oocytes of other mammalian species (Flemr et al. 2013; Rosenkranz et al. 2015). Moreover, compared to most other mammals with 4 PIWI genes, rodents lack Piwil3, which has been shown to be specifically expressed in oocytes in hamsters, bovines, and humans (Yang et al. 2019; Tan et al. 2020; Ishino et al. 2021), suggesting that the Piwil3-piRNA pathway is specifically missing in the rodent lineage. To test whether the sexual dimorphic requirement for a piRNA pathway is specific for the rodent lineage, three independent groups have disrupted the piRNA pathways in hamsters and revealed that piRNA pathways are indeed required for maintaining germline genome integrity in both sexes (Zhang et al. 2021; Hasuwa et al. 2021; Loubalova et al. 2021). Thus, rodents are likely to be an outlier with the siRNAs replacing the Piwil3-piRNAs in oocytes, and piRNA pathways are generally required for germ cells of both sexes in animals.
Nonetheless, piRNA pathways in the ovaries of humans, bovines, hamster, and macaques do show distinct differences from those in testes with regards to piRNA abundance, piRNA size, modification, piRNA-associated PIWI proteins, and the genomic origins of piRNA species (Rosenkranz et al. 2015; Ishino et al. 2021; Hasuwa et al. 2021; Zhang et al. 2021; Loubalova et al. 2021; Yang et al. 2019; Tan et al. 2020), arguing that piRNA pathways are influenced by the sex of the cell lineage. For instance, in hamsters, compared to testis piRNAs, oocyte piRNAs come from a distinct set of intergenic genomic loci whose transcription are not driven by A-MYB. Piwil3 in hamster oocytes bind to ~ 19 nt piRNAs without 2′-O-methyl modification, and the binding depends on the phosphorylation of Piwil3. In hamsters, while Piwil1 binds to ~ 29 nt piRNAs in testes, they bind to ~ 23 nt and ~ 29 nt piRNA in oocytes, and they only bind to ~ 23 nt piRNAs in 2-cell embryos. The developmentally dependent regulation of piRNA size contradicts the notion that the size of piRNAs is determined by the footprints of the PIWI protein. Therefore, further understanding the route of sexual dimorphism of piRNA pathways, either due to sex chromosome or sex hormone differences, will help to understand sex-dependent biogenic regulation and its impact on piRNA functions.
The evolution of piRNA pathways
Pervasive adaptive evolution among piRNA pathway proteins has been reported in both insect and vertebrate lineages (Yi et al. 2014; Levine et al. 2016, 2012; Simkin et al. 2013; Palmer and Whybrow 2007; Obbard et al. 2009; Kolaczkowski et al. 2011). Two hypotheses have been proposed to explain the force driving this positive selection. The first hypothesis, an arms race with TEs, has been proposed since the discovery of piRNAs (Malone and Hanno 2009; Aravin et al. 2007a). As a prime embodiment of the Red Queen’s race, which originated from studies of host virus battles (Daugherty and Malik 2012), this hypothesis successfully explains why piRNA sequences need to rapidly adapt to keep up with ever-changing TE sequences but fails to provide a satisfactory explanation for the necessity of continuously changing the piRNA pathway proteins. While it is possible that piRNA biogenesis proteins have to constantly adapt to the life history of each recently invaded TE, such as driving the transcription of new piRNA precursors embedded with TEs or targeting new TE transcripts in a different subcellular location, as unlikely viruses, the TEs cannot “fight back” to repress piRNA machinery (Blumenstiel et al. 2016). The second hypothesis is adaptive evolution driven by the ongoing tension between TE silencing and off-target effects (known as genomic autoimmunity) (Blumenstiel et al. 2016). Similar tension has been observed between phage and CRISPR systems (Koonin and Yutin 2020). As the piRNAs have been shown to target mRNAs, off-targeting is the cost for robust TE suppression. The selection for specificity and sensitivity of TE suppression varies at different stages of TE invasion, with high TE activation favoring high specificity, while low TE expression favors sensitivity. Although most studies on the evolution of piRNA pathway proteins were performed on Drosophila, the same principles to suppress TEs and avoid autoimmunity would hold true for mammalians, as TE piRNAs share biogenic protein factors with pachytene piRNAs.
Pachytene piRNA sequences show poor conservation across species, and only 29 of 89 human piRNA loci share synteny conservation (the flanking genes surrounding piRNA clusters are conserved) outside of primates (Özata et al. 2020; Chirn et al. 2015). Despite being essential for male fertility, the function of pachytene piRNAs remains largely unknown. Although some pachytene piRNAs have been shown to trigger the decay of mRNAs required for meiosis or sperm maturation (Wu et al. 2020; Zhang et al. 2015; Gou et al. 2014; Goh et al. 2015), most pachytene piRNAs lack obvious complementary targets. It has also been proposed that PIWI proteins function without piRNAs (Vourekas et al. 2012), and piRNAs may exist to stabilize the PIWI protein without providing any specificity. Several models have been proposed to explain the rapid divergence of pachytene piRNA sequences. One hypothesis proposes that pachytene piRNA evolution drives reproductive isolation (Özata et al. 2020). Another suggests that existing piRNA clusters (Kawaoka et al. 2012) serve as landing pads for TE transposition ‘trapping’ new TE sequences in the cluster (Yamanaka et al. 2014; Zhang et al. 2020) (Fig. 3, middle), a mechanism reminiscent of the acquisition of new sequences in CRISPR loci. While it is known that piRNA loci can autonomously process transcripts harboring insertion sequences into new piRNAs (Muerdter et al. 2012), there is only limited evidence that TE transposition, or other insertion mechanisms, display a preference for targeting piRNA loci. Thus, the adaptive force underlying the rapid divergence of pachytene piRNA sequences currently remains unclear.
Current models of new piRNA acquisition. (1) Duplication of piRNA loci. piRNA clusters are duplicated or deleted in the genome and generate more piRNAs. (2) Insertion into pre-existing piRNA loci. New piRNAs can be generated by inserting sequences into pre-existing piRNA clusters. (3) Activation of a provirus for piRNA production. A provirus was first activated for piRNA production with sense orientation, and then the transcription template of the piRNA locus was switched. The direction of piRNA cluster transcription may change, and piRNAs can be generated from antisense piRNA loci. Figure created with BioRender.com
Two mutational modes are known to give rise to the birth of new piRNA loci. The first mode is copy number duplication or deletion of existing piRNA loci (Assis and Kondrashov 2009; Chirn et al. 2015) (Fig. 3, left). For example, mammalian piRNA loci have been reported to display elevated rates of copy number variation (Gould et al. 2012). However, it remains unclear what is the mutational mechanism leading to the copy number variations of piRNA loci. The second mode is activation of novel TE insertions into piRNA loci (Sun et al. 2017; Yu et al. 2019). For example, in koalas and chickens, proviruses that recently invaded the germline gained the ability to produce piRNAs. In koalas, piRNAs that target the KoRV-A retrovirus were originally produced from the sense strand of KoRV-A but then, through an unknown mechanism, shifted toward the production of piRNAs antisense to the retrovirus (Yu et al. 2019) (Fig. 3, right). Domesticated chickens were found to activate piRNA production from a truncated proviral locus against avian leukosis viruses, whereas undomesticated chickens did not produce piRNAs from the same locus (Sun et al. 2017). According to the “out of testis” hypothesis (Kaessmann 2010), the permissive chromatin state of spermatocytes and spermatids makes testes an ideal tissue for the emergence of new genes. Thus, it is not surprising that proviral loci are transcribed in testes, but it remains unclear what genomic and/or epigenetic mechanisms led to their ability to produce piRNAs.
Perspective
piRNAs constitute a unique and rapidly evolving class of small non-coding RNAs with a wide diversity of functions and various biogenesis pathways. Developments in genetics and sequencing techniques have advanced piRNA research through the identification of multiple key players during the various phases of piRNA processing; however, the absence of cell culture systems, or in vitro systems, that can recapitulate mammalian piRNA production has hindered the mechanistic studies of piRNA biogenic pathways. Furthermore, studying mammalian piRNA function, especially pachytene piRNAs, using conventional genetics remains a challenge. As pachytene piRNAs share key piRNA biogenic factors with pre-pachytene piRNAs, animals with defective piRNA biogenesis exhibit de-silenced TEs that trigger arrest during early spermatogenesis, which consequently masks their function during spermiogenesis. Furthermore, A-MYB, the key pachytene piRNA transcription factor, also regulates the expression of (protein-coding) mRNAs such that A-Myb mutants also display early meiotic arrest (Li et al. 2013). Pachytene piRNAs are primarily derived from 100 intergenic genomic loci (Li et al. 2013) that are likely functionally redundant. As a result, deleting a single piRNA-producing locus often has no obvious phenotype (Wu et al. 2020; Homolka et al. 2015). Recently, sperm maturation defects were observed upon knockout of the promoter of a pair of piRNA clusters with only subtle changes of the entire transcriptome (Choi et al. 2021; Wu et al. 2020). In conclusion, significant advances have been made in our understanding of piRNA biology over the past several years; however, a number of key questions surrounding piRNA biogenesis remain:
-
(1)
Why are some transcripts processed to piRNAs while others are not? piRNA precursors could be either marked epigenetically during transcription or recognized post-transcriptionally in the cytosol. Unlike Drosophila germline piRNA loci, which are marked with the chromatin-bound protein Rhino (Klattenhoff et al. 2009), currently no epigenetic factors have been identified that specially bind piRNA loci in mammals. It is also unlikely that there would be unique splicing structures in mammals, as has been proposed for Drosophila piRNA biogenesis, because mammalian piRNA precursors are canonically spliced (Li et al. 2013; Sun et al. 2021). Whether pachytene piRNA precursors have unique secondary or tertiary structures, RNA modifications, or translation intermediates remains to be determined.
-
(2)
How are piRNA precursors recruited to mitochondria? What is the significance of the connection between piRNA biogenesis and mitochondrial biology? Either the piRNA precursors are channeled directly to the IMC from the nucleus without interacting with ribosomes prior to IMC localization or they are translated first, and the precursor translation intermediates are then transported to mitochondria. Given that the IMC is not close to the nucleus, if the former is the case, a special mechanism, such as packing piRNA precursors using RNA-binding proteins, would be required to prevent ribosome access prior to IMC localization. Alternatively, specific precursor translation intermediates could be recognized by the piRNA-processing machinery, or the translation products could be recognized by mitochondrion-localizing chaperones.
-
(3)
Is phased piRNA biogenesis initiated de novo or by initiator piRNAs? If phased biogenesis is triggered by base-pair complementary initiator piRNAs, the pairing rules between piRNAs and targets are likely to tolerant mismatches to allow sufficient cleavage events on the long single-stranded piRNA precursors. Alternatively, piRNA precursors may recruit an endonuclease or the conventional RNA decay machinery for de novo piRNA processing. One possibility is that piRNA biogenesis may be initiated by an endonuclease that specifically targets ribosomes outside of ORFs, as translation-mediated RNA decay is common, involving mechanisms such as NMD, no-go decay, no-stop decay, or co-translational Exo1-mediated decay (Shoemaker and Green 2012; Graille and Séraphin 2012; Kervestin and Jacobson 2012; Schoenberg and Maquat 2012).
-
(4)
How does the first nucleotide uridine (1U) bias arise? 1U bias is the most noticeable sequence signature on primary piRNAs. Crystal structures of the silkworm PIWI protein Siwi(Matsumoto et al. 2016) and Drosophila Piwi (Yamaguchi et al. 2020), as well as in vitro biochemical assays from silkworm(Matsumoto et al. 2016), revealed that the 5′-uridine fits well into the PIWI/Siwi MID domain. However, in Pnldc1 mutant mice, the un-trimmed piRNAs form a head-to-tail pattern with no gaps between them, indicating that the fragmentation process must generate the 1U bias prior to PIWI loading. Since MitoPLD/ZUCCHINI does not show any preference for cleaving before uridines in vitro, nor do ribosomes display any binding bias, either MitoPLD is not the main endonuclease fragmenting the pachytene piRNA precursors (that show a 1U bias) or another co-factor exists that works coordinately with MitoPLD and ribosomes to generate the 1U bias.
-
(5)
Why are large quantities of piRNAs produced at the pachytene stage? We are yet to understand the biological significance of the burst of piRNA production at the pachytene stage. This production is due either to a functional or biogenic requirement. As discussed previously, while a biogenic requirement seems more likely, the disruption of pachytene piRNA biogenesis in hamsters leads to pachytene arrest (Hasuwa et al. 2021; Loubalova et al. 2021; Zhang et al. 2021), distinct from round spermatid arrest in mice, suggesting that pachytene piRNAs may be generally required for meiosis progression with rodent as an outlier. As most studies on meiosis have focused on chromosome behavior or epigenetic regulation, we know little about how RNA metabolism, such as translation and RNA granule movement, coordinates with the complex choreography occurring in nuclei. Such studies may shed light on the biogenic requirements that facilitate the coordinate production of 3.8–8.4 million piRNA molecules in each spermatocyte during the pachytene stage (Gainetdinov et al. 2018).
-
(6)
How is a new piRNA locus born and how does it die during evolution? Pachytene piRNA loci can rise and disappear rapidly over short evolutionary timescales (80 million years between mice and humans) in mammals. This rapid divergence could be due to elevated mutation rates and/or adaptive selection. The current notion that pachytene piRNAs function to target mRNAs required for sperm maturation does not seem sufficient to provide such a selective force. The idea that pachytene piRNAs and their mRNA targets behave like a toxicant and anti-toxicant is an attractive model that needs further investigation (Aravin 2020). The possibilities that piRNA loci exhibit elevated mutation rates or that piRNA loci represent preferential TE landing sites have also not been fully explored.
-
(7)
Are there somatic piRNAs in vertebrates? The notion is that somatic piRNA pathways widely exist in invertebrates, but was lost in vertebrates (Lewis et al. 2018). However, there are a number of reports in vertebrates regarding the existence of piRNA-like molecules or the expression of PIWI proteins (Galton et al. 2021; Mai et al. 2020; Moyano and Stefani 2015; Keam et al. 2014; Yan et al. 2011; Mei et al. 2015; Zhao et al. 2015; Nandi et al. 2016; Perera et al. 2019; Sharma et al. 2001; Freedman et al. 2016; Martinez et al. 2015; Lee et al. 2011; Cheng et al. 2014). These reports failed to provide a satisfactory answer as to why there is no phenotype beyond reproduction when disrupting piRNA pathways in mice. It is an interesting area for further investigation: either somatic piRNAs exist in low abundance with under-explored function, or mice are not a good model to study somatic piRNAs.
Here, we have provided an overview of piRNA biogenesis in mice and have referenced a range of other organisms where relevant. We discuss the current progress in our understanding of piRNA conservation and evolution. Finally, we outline several key questions in the field regarding piRNA biogenesis. With the rapid advances in sequencing technologies and development of new techniques and model organisms for multi-omics and comparative studies, we envision that significant strides will be made in the next few years. A complete understanding of piRNA biogenesis, evolution, and function may facilitate the development and application of artificial piRNAs as tools for epigenetic regulation and a wide variety of other possible uses.
References
Andersen PR, Tirian L, Vunjak M, Brennecke J (2017) A heterochromatin-dependent transcription machinery drives piRNA expression. Nature 549:54–59
Aravin A, Gaidatzis D, Pfeffer S, Lagos-Quintana M, Landgraf P, Iovino N, Morris P, Brownstein MJ, Kuramochi-Miyagawa S, Nakano T, Chien M, Russo JJ, Ju J, Sheridan R, Sander C, Zavolan M, Tuschl T (2006) A novel class of small RNAs bind to MILI protein in mouse testes. Nature 442:203–207
Aravin AA (2020) Pachytene piRNAs as beneficial regulators or a defense system gone rogue. Nat Genet 52:644–645
Aravin AA, Hannon GJ (2008) Small RNA silencing pathways in germ and stem cells. Cold Spring Harb Symp Quant Biol 73:283–290
Aravin AA, Hannon GJ, Brennecke J (2007a) The Piwi-piRNA pathway provides an adaptive defense in the transposon arms race. Science 318:761–764
Aravin AA, Sachidanandam R, Bourchis D, Schaefer C, Pezic D, Toth KF, Bestor T, Hannon GJ (2008) A piRNA pathway primed by individual transposons is linked to de novo DNA methylation in mice. Mol Cell 31:785–799
Aravin AA, Sachidanandam R, Girard A, Fejes-Toth K, Hannon GJ (2007b) Developmentally regulated piRNA clusters implicate MILI in transposon control. Science 316:744–747
Aravin AA, van Derheijden GW, Castañeda J, Vagin VV, Hannon GJ, Bortvin A (2009) Cytoplasmic compartmentalization of the fetal piRNA pathway in mice. PLoS Genet 5:1000764
Aravin AA, Naumova NM, Tulin AV, Vagin VV, Rozovsky YM, Gvozdev VA (2001) Double-stranded RNA-mediated silencing of genomic tandem repeats and transposable elements in the D. melanogaster germline. Curr Biol 11:1017–1027
Assis R, Kondrashov AS (2009) Rapid repetitive element-mediated expansion of piRNA clusters in mammalian evolution. Proc Natl Acad Sci USA 106:7079–7082
Betel D, Sheridan R, Marks DS, Sander C (2007) Computational analysis of mouse piRNA sequence and biogenesis. PLoS Comput Biol 3:222
Beyret E, Liu N, Lin H (2012) piRNA biogenesis during adult spermatogenesis in mice is independent of the ping-pong mechanism. Cell Res 22:1429–1439
Blumenstiel JP, Erwin AA, Hemmer LW (2016) What drives positive selection in the drosophila piRNA machinery? The genomic autoimmunity hypothesis. Yale J Biol Med 89:499–512
Brennecke J, Aravin AA, Stark A, Dus M, Kellis M, Sachidanandam R, Hannon GJ (2007) Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128:1089–1103
Carmell MA, Girard A, van de Kant HJ, Bourchis D, Bestor TH, de Rooij DG, Hannon GJ (2007) MIWI2 is essential for spermatogenesis and repression of transposons in the mouse male germline. Dev Cell 12:503–514
Carmo-Fonseca M, Tollervey D, Pepperkok R, Barabino SM, Merdes A, Brunner C, Zamore PD, Green MR, Hurt E, Lamond AI (1991) Mammalian nuclei contain foci which are highly enriched in components of the pre-mRNA splicing machinery. EMBO J 10:195–206
Castaneda J, Genzor P, van der Heijden GW, Sarkeshik A, Yates JR, Ingolia NT, Bortvin A (2014) Reduced pachytene piRNAs and translation underlie spermiogenic arrest in Maelstrom mutant mice. EMBO J 33:1999–2019
Chen C, Jin J, James DA, Adams-Cioaba MA, Park JG, Guo Y, Tenaglia E, Xu C, Gish G, Min J, Pawson T (2009) Mouse Piwi interactome identifies binding mechanism of Tdrkh Tudor domain to arginine methylated Miwi. Proc Natl Acad Sci USA 106:20336–20341
Cheng EC, Kang D, Wang Z, Lin H (2014) PIWI proteins are dispensable for mouse somatic development and reprogramming of fibroblasts into pluripotent stem cells. PLoS ONE 9:97821
Chirn GW, Rahman R, Sytnikova YA, Matts JA, Zeng M, Gerlach D, Yu M, Berger B, Naramura M, Kile BT, Lau NC (2015) Conserved piRNA Expression from a Distinct Set of piRNA Cluster Loci in Eutherian Mammals. PLoS Genet 11:e1005652
Choi H, Wang Z, Dean J (2021) Sperm acrosome overgrowth and infertility in mice lacking chromosome 18 pachytene piRNA. PLoS Genet 17:e1009485
Chuma S, Hosokawa M, Kitamura K, Kasai S, Fujioka M, Hiyoshi M, Takamune K, Noce T, Nakatsuji N (2006) Tdrd1/Mtr-1, a tudor-related gene, is essential for male germ-cell differentiation and nuage/germinal granule formation in mice. Proc Natl Acad Sci USA 103:15894–15899
Chuma S, Hosokawa M, Tanaka T, Nakatsuji N (2009) Ultrastructural characterization of spermatogenesis and its evolutionary conservation in the germline: germinal granules in mammals. Mol Cell Endocrinol 306:17–23
Czech B, Hannon GJ (2016a) One loop to rule them all: The ping-pong cycle and piRNA-guided silencing. Trends Biochem Sci 41:324–337
Czech B, Hannon GJ (2016b) A Happy 3’ Ending to the piRNA Maturation Story. Cell 164:838–840
Czech B, Preall JB, McGinn J, Hannon GJ (2013) A transcriptome-wide RNAi screen in the Drosophila ovary reveals factors of the germline piRNA pathway. Mol Cell 50:749–761
Darricarrère N, Liu N, Watanabe T, Lin H (2013) Function of Piwi, a nuclear Piwi/Argonaute protein, is independent of its slicer activity. Proc Natl Acad Sci U S A 110:1297–1302
Daugherty MD, Malik HS (2012) Rules of engagement: molecular insights from host-virus arms races. Annu Rev Genet 46:677–700
De Fazio S, Bartonicek N, Di Giacomo M, Abreu-Goodger C, Sankar A, Funaya C, Antony C, Moreira PN, Enright AJ, O’Carroll D (2011) The endonuclease activity of Mili fuels piRNA amplification that silences LINE1 elements. Nature 480:259–263
Deng W, Lin H (2002) miwi, a murine homolog of piwi, encodes a cytoplasmic protein essential for spermatogenesis. Dev Cell 2:819–830
Di Giacomo M, Comazzetto S, Saini H, De Fazio S, Carrieri C, Morgan M, Vasiliauskaite L, Benes V, Enright AJ, O’Carroll D (2013) Multiple epigenetic mechanisms and the piRNA pathway enforce LINE1 silencing during adult spermatogenesis. Mol Cell 50:601–608
Ding D, Liu J, Dong K, Melnick AF, Latham KE, Chen C (2019) Mitochondrial membrane-based initial separation of MIWI and MILI functions during pachytene piRNA biogenesis. Nucleic Acids Res 47:2594–2608
Ding D, Liu J, Dong K, Midic U, Hess RA, Xie H, Demireva EY, Chen C (2017) PNLDC1 is essential for piRNA 3’ end trimming and transposon silencing during spermatogenesis in mice. Nat Commun 8:819
Ding D, Liu J, Midic U, Wu Y, Dong K, Melnick A, Latham KE, Chen C (2018) TDRD5 binds piRNA precursors and selectively enhances pachytene piRNA processing in mice. Nat Commun 9:127
Eddy EM (1974) Fine structural observations on the form and distribution of nuage in germ cells of the rat. Anat Rec 178:731–757
Eddy EM (1975) Germ plasm and the differentiation of the germ cell line. Int Rev Cytol 43:229–280
Fabry MH, Ciabrelli F, Munafò M, Eastwood EL, Kneuss E, Falciatori I, Falconio FA, Hannon GJ, Czech B (2019) piRNA-guided co-transcriptional silencing coopts nuclear export factors. Elife 8:e47999
Farazi, T. A., Juranek, S. A., and Tuschl, T. (2008). The growing catalog of small RNAs and their association with distinct Argonaute/Piwi family members. Development
Fawcett DW, Eddy EM, Phillips DM (1970) Observations on the fine structure and relationships of the chromatoid body in mammalian spermatogenesis. Biol Reprod 2:129–153
Feltzin VL, Khaladkar M, Abe M, Parisi M, Hendriks GJ, Kim J, Bonini NM (2015) The exonuclease Nibbler regulates age-associated traits and modulates piRNA length in Drosophila. Aging Cell 14:443–452
Findley SD, Tamanaha M, Clegg NJ, Ruohola-Baker H (2003) Maelstrom, a Drosophila spindle-class gene, encodes a protein that colocalizes with Vasa and RDE1/AGO1 homolog, Aubergine, in nuage. Development 130:859–871
Fine AD, Ball RL, Fujiwara Y, Handel MA, Carter GW (2019) Uncoupling of transcriptomic and cytological differentiation in mouse spermatocytes with impaired meiosis. Mol Biol Cell 30:717–728
Flemr M, Malik R, Franke V, Nejepinska J, Sedlacek R, Vlahovicek K, Svoboda P (2013) A retrotransposon-driven dicer isoform directs endogenous small interfering RNA production in mouse oocytes. Cell 155:807–816
Freedman JE, Gerstein M, Mick E, Rozowsky J, Levy D, Kitchen R, Das S, Shah R, Danielson K, Beaulieu L, Navarro FC, Wang Y, Galeev TR, Holman A, Kwong RY, Murthy V, Tanriverdi SE, Koupenova-Zamor M, Mikhalev E, Tanriverdi K (2016) Diverse human extracellular RNAs are widely detected in human plasma. Nat Commun 7:11106
Frost RJ, Hamra FK, Richardson JA, Qi X, Bassel-Duby R, Olson EN (2010) MOV10L1 is necessary for protection of spermatocytes against retrotransposons by Piwi-interacting RNAs. Proc Natl Acad Sci USA 107:11847–11852
Gainetdinov I, Colpan C, Arif A, Cecchini K, Zamore PD (2018) A Single Mechanism of Biogenesis, Initiated and Directed by PIWI Proteins, Explains piRNA Production in Most Animals. Mol Cell 71:775-790.e5
Galton, R., Fejes-Toth, K., and Bronner, M. E. (2021). A somatic piRNA pathway regulates epithelial-to-mesenchymal transition of chick neural crest cells.
Gan B, Chen S, Liu H, Min J, Liu K (2019) Structure and function of eTudor domain containing TDRD proteins. Crit Rev Biochem Mol Biol 54:119–132
Gao Q, Frohman MA (2012) Roles for the lipid-signaling enzyme MitoPLD in mitochondrial dynamics, piRNA biogenesis, and spermatogenesis. BMB Rep 45:7–13
Geisinger A, Rodríguez-Casuriaga R, Benavente R (2021) Transcriptomics of Meiosis in the Male Mouse. Front Cell Dev Biol 9:626020
Girard A, Sachidanandam R, Hannon GJ, Carmell MA (2006) A germline-specific class of small RNAs binds mammalian Piwi proteins. Nature 442:199–202
Goh WS, Falciatori I, Tam OH, Burgess R, Meikar O, Kotaja N, Hammell M, Hannon GJ (2015) piRNA-directed cleavage of meiotic transcripts regulates spermatogenesis. Genes Dev 29:1032–1044
Gold B, Fujimoto H, Kramer JM, Erickson RP, Hecht NB (1983) Haploid accumulation and translational control of phosphoglycerate kinase-2 messenger RNA during mouse spermatogenesis. Dev Biol 98:392–399
Gou LT, Dai P, Yang JH, Xue Y, Hu YP, Zhou Y, Kang JY, Wang X, Li H, Hua MM, Zhao S, Hu SD, Wu LG, Shi HJ, Li Y, Fu XD, Qu LH, Wang ED, Liu MF (2014) Pachytene piRNAs instruct massive mRNA elimination during late spermiogenesis. Cell Res 24:680–700
Gould DW, Lukic S, Chen KC (2012) Selective constraint on copy number variation in human piwi-interacting RNA Loci. PLoS ONE 7:e46611
Graille M, Séraphin B (2012) Surveillance pathways rescuing eukaryotic ribosomes lost in translation. Nat Rev Mol Cell Biol 13:727–735
Grimson A, Srivastava M, Fahey B, Woodcroft BJ, Chiang HR, King N, Degnan BM, Rokhsar DS, Bartel DP (2008) Early origins and evolution of microRNAs and Piwi-interacting RNAs in animals. Nature 455:1193–1197
Grivna ST, Beyret E, Wang Z, Lin H (2006) A novel class of small RNAs in mouse spermatogenic cells. Genes Dev 20:1709–1714
Guan Y, Keeney S, Jain D, Wang PJ (2021) yama, a mutant allele of Mov10l1, disrupts retrotransposon silencing and piRNA biogenesis. PLoS Genet 17:e1009265
Gunawardane LS, Saito K, Nishida KM, Miyoshi K, Kawamura Y, Nagami T, Siomi H, Siomi MC (2007) A slicer-mediated mechanism for repeat-associated siRNA 5′ end formation in Drosophila. Science 315:1587–1590
Haase AD (2016) A Small RNA-Based Immune System Defends Germ Cells against Mobile Genetic Elements. Stem Cells Int 2016:7595791
Haase AD, Fenoglio S, Muerdter F, Guzzardo PM, Czech B, Pappin DJ, Chen C, Gordon A, Hannon GJ (2010) Probing the initiation and effector phases of the somatic piRNA pathway in Drosophila. Genes Dev 24(22):2499–2504
Han BW, Hung JH, Weng Z, Zamore PD, Ameres SL (2011) The 3’-to-5’ exoribonuclease Nibbler shapes the 3’ ends of microRNAs bound to Drosophila Argonaute1. Curr Biol 21:1878–1887
Han BW, Wang W, Li C, Weng Z, Zamore PD (2015) Noncoding RNA. piRNA-guided transposon cleavage initiates Zucchini-dependent, phased piRNA production. Science 348:817–821
Handler D, Olivieri D, Novatchkova M, Gruber FS, Meixner K, Mechtler K, Stark A, Sachidanandam R, Brennecke J (2011) A systematic analysis of Drosophila TUDOR domain-containing proteins identifies Vreteno and the Tdrd12 family as essential primary piRNA pathway factors. EMBO J 30:3977–3993
Harris AN, Macdonald PM (2001) aubergine encodes a Drosophila polar granule component required for pole cell formation and related to eIF2C. Development 128:2823–2832
Hasuwa, H., Iwasaki, Y. W., Au Yeung, W. K., Ishino, K., Masuda, H., Sasaki, H., and Siomi, H. (2021). Production of functional oocytes requires maternally expressed PIWI genes and piRNAs in golden hamsters. Nat Cell Biol
Hayashi R, Schnabl J, Handler D, Mohn F, Ameres SL, Brennecke J (2016) Genetic and mechanistic diversity of piRNA 3’-end formation. Nature 539:588–592
Homolka D, Pandey RR, Goriaux C, Brasset E, Vaury C, Sachidanandam R, Fauvarque MO, Pillai RS (2015) PIWI slicing and RNA elements in precursors instruct directional primary piRNA biogenesis. Cell Rep 12:418–428
Honda S, Kirino Y, Maragkakis M, Alexiou P, Ohtaki A, Murali R, Mourelatos Z, Kirino Y (2013) Mitochondrial protein BmPAPI modulates the length of mature piRNAs. RNA 19:1405–1418
Horwich MD, Li C, Matranga C, Vagin V, Farley G, Wang P, Zamore PD (2007) The Drosophila RNA methyltransferase, DmHen1, modifies germline piRNAs and single-stranded siRNAs in RISC. Curr Biol 17:1265–1272
Hosokawa M, Shoji M, Kitamura K, Tanaka T, Noce T, Chuma S, Nakatsuji N (2007) Tudor-related proteins TDRD1/MTR-1, TDRD6 and TDRD7/TRAP: domain composition, intracellular localization, and function in male germ cells in mice. Dev Biol 301:38–52
Ipsaro JJ, Haase AD, Knott SR, Joshua-Tor L, Hannon GJ (2012) The structural biochemistry of Zucchini implicates it as a nuclease in piRNA biogenesis. Nature 491:279–283
Ishino K, Hasuwa H, Yoshimura J, Iwasaki YW, Nishihara H, Seki NM, Hirano T, Tsuchiya M, Ishizaki H, Masuda H, Kuramoto T, Saito K, Sakakibara Y, Toyoda A, Itoh T, Siomi MC, Morishita S, Siomi H (2021) Hamster PIWI proteins bind to piRNAs with stage-specific size variations during oocyte maturation. Nucleic Acids Res 49:2700–2720
Ishizu H, Iwasaki YW, Hirakata S, Ozaki H, Iwasaki W, Siomi H, Siomi MC (2015) Somatic primary piRNA biogenesis driven by cis-acting RNA elements and trans-acting Yb. Cell Rep 12:429–440
Izumi N, Shoji K, Suzuki Y, Katsuma S, Tomari Y (2020) Zucchini consensus motifs determine the mechanism of pre-piRNA production. Nature 578:311–316
Jahn CL, Klobutcher LA (2002) Genome remodeling in ciliated protozoa. Annu Rev Microbiol 56:489–520
Jamsai D, O’Connor AE, Odonnell L, Lo JC, O’Bryan MK (2015) Uncoupling of transcription and translation of Fanconi anemia (FANC) complex proteins during spermatogenesis. Spermatogenesis 5:979061
Kabayama Y, Toh H, Katanaya A, Sakurai T, Chuma S, Kuramochi-Miyagawa S, Saga Y, Nakano T, Sasaki H (2017) Roles of MIWI, MILI and PLD6 in small RNA regulation in mouse growing oocytes. Nucleic Acids Res 45:5387–5398
Kaessmann H (2010) Origins, evolution, and phenotypic impact of new genes. Genome Res 20:1313–1326
Kawaoka S, Mitsutake H, Kiuchi T, Kobayashi M, Yoshikawa M, Suzuki Y, Sugano S, Shimada T, Kobayashi J, Tomari Y, Katsuma S (2012) A role for transcription from a piRNA cluster in de novo piRNA production. RNA 18:265–273
Keam SP, Young PE, McCorkindale AL, Dang TH, Clancy JL, Humphreys DT, Preiss T, Hutvagner G, Martin DI, Cropley JE, Suter CM (2014) The human Piwi protein Hiwi2 associates with tRNA-derived piRNAs in somatic cells. Nucleic Acids Res 42:8984–8995
Kervestin S, Jacobson A (2012) NMD: a multifaceted response to premature translational termination. Nat Rev Mol Cell Biol 13:700–712
Ketting RF (2011) The many faces of RNAi. Dev Cell 20:148–161
Khalil AM, Boyar FZ, Driscoll DJ (2004) Dynamic histone modifications mark sex chromosome inactivation and reactivation during mammalian spermatogenesis. Proc Natl Acad Sci USA 101:16583–16587
Kim M, Ki BS, Hong K, Park SP, Ko JJ, Choi Y (2016) Tudor Domain Containing Protein TDRD12 Expresses at the Acrosome of Spermatids in Mouse Testis. Asian-Australas J Anim Sci 29:944–951
Kirino Y, Mourelatos Z (2007) Mouse Piwi-interacting RNAs are 2´-O-methylated at their 3′ termini. Nat Struct Mol Biol 14:347–348
Klattenhoff C, Xi H, Li C, Lee S, Xu J, Khurana JS, Zhang F, Schultz N, Koppetsch BS, Nowosielska A, Seitz H, Zamore PD, Weng Z, Theurkauf WE (2009) The Drosophila HP1 homolog Rhino is required for transposon silencing and piRNA production by dual-strand clusters. Cell 138:1137–1149
Kolaczkowski B, Hupalo DN, Kern AD (2011) Recurrent adaptation in RNA interference genes across the Drosophila phylogeny. Mol Biol Evol 28:1033–1042
Koonin EV, Yutin N (2020) The crAss-like Phage group: how metagenomics reshaped the human virome. Trends Microbiol 28:349–359
Kotaja N, Sassone-Corsi P (2007) The chromatoid body: a germ-cell-specific RNA-processing centre. Nat Rev Mol Cell Biol 8:85–90
Kumar M, Carmichael GG (1998) Antisense RNA: function and fate of duplex RNA in cells of higher eukaryotes. Microbiol Mol Biol Rev 62:1415–1434
Kuramochi-Miyagawa S, Kimura T, Ijiri TW, Isobe T, Asada N, Fujita Y, Ikawa M, Iwai N, Okabe M, Deng W, Lin H, Matsuda Y, Nakano T (2004) Mili, a mammalian member of piwi family gene, is essential for spermatogenesis. Development 131:839–849
Kuramochi-Miyagawa S, Watanabe T, Gotoh K, Takamatsu K, Chuma S, Kojima-Kita K, Shiromoto Y, Asada N, Toyoda A, Fujiyama A, Totoki Y, Shibata T, Kimura T, Nakatsuji N, Noce T, Sasaki H, Nakano T (2010) MVH in piRNA processing and gene silencing of retrotransposons. Genes Dev 24:887–892
Kuramochi-Miyagawa S, Watanabe T, Gotoh K, Totoki Y, Toyoda A, Ikawa M, Asada N, Kojima K, Yamaguchi Y, Ijiri TW, Hata K, Li E, Matsuda Y, Kimura T, Okabe M, Sakaki Y, Sasaki H, Nakano T (2008) DNA methylation of retrotransposon genes is regulated by Piwi family members MILI and MIWI2 in murine fetal testes. Genes Dev 22:908–917
Lau NC, Seto AG, Kim J, Kuramochi-Miyagawa S, Nakano T, Bartel DP, Kingston RE (2006) Characterization of the piRNA complex from rat testes. Science 313:363–367
Lee EJ, Banerjee S, Zhou H, Jammalamadaka A, Arcila M, Manjunath BS, Kosik KS (2011) Identification of piRNAs in the central nervous system. RNA 17:1090–1099
Levine MT, McCoy C, Vermaak D, Lee YC, Hiatt MA, Matsen FA, Malik HS (2012) Phylogenomic analysis reveals dynamic evolutionary history of the Drosophila heterochromatin protein 1 (HP1) gene family. PLoS Genet 8:e1002729
Levine MT, Vander Wende HM, Hsieh E, Baker EP, Malik HS (2016) Recurrent gene duplication diversifies genome defense repertoire in drosophila. Mol Biol Evol 33:1641–1653
Lewis SH, Quarles KA, Yang Y, Tanguy M, Frézal L, Smith SA, Sharma PP, Cordaux R, Gilbert C, Giraud I, Collins DH, Zamore PD, Miska EA, Sarkies P, Jiggins FM (2018) Pan-arthropod analysis reveals somatic piRNAs as an ancestral defence against transposable elements. Nat Ecol Evol 2:174–181
Li XC, Schimenti JC (2007) Mouse pachytene checkpoint 2 (trip13) is required for completing meiotic recombination but not synapsis. PLoS Genet 3:e130
Li XZ, Roy CK, Dong X, Bolcun-Filas E, Wang J, Han BW, Xu J, Moore MJ, Schimenti JC, Weng Z, Zamore PD (2013) An ancient transcription factor initiates the burst of piRNA production during early meiosis in mouse testes. Mol Cell 50:67–81
Lim SL, Qu ZP, Kortschak RD, Lawrence DM, Geoghegan J, Hempfling AL, Bergmann M, Goodnow CC, Ormandy CJ, Wong L, Mann J, Scott HS, Jamsai D, Adelson DL, Obryan MK (2015) HENMT1 and piRNA stability are required for adult male germ cell transposon repression and to define the spermatogenic program in the mouse. PLoS Genet 11:e1005620
Lin H, Spradling AC (1997) A novel group of pumilio mutations affects the asymmetric division of germline stem cells in the Drosophila ovary. Development 124:2463–2476
Liu L, Qi H, Wang J, Lin H (2011) PAPI, a novel TUDOR-domain protein, complexes with AGO3, ME31B and TRAL in the nuage to silence transposition. Development 138:1863–1873
Loubalova Z, Fulka H, Horvat F, Pasulka J, Malik R, Hirose M, Ogura A, Svoboda P (2021) Formation of spermatogonia and fertile oocytes in golden hamsters requires piRNAs. Nat Cell Biol
Ma L, Buchold GM, Greenbaum MP, Roy A, Burns KH, Zhu H, Han DY, Harris RA, Coarfa C, Gunaratne PH, Yan W, Matzuk MM (2009) GASZ is essential for male meiosis and suppression of retrotransposon expression in the male germline. PLoS Genet 5:e1000635
Mai D, Zheng Y, Guo H, Ding P, Bai R, Li M, Ye Y, Zhang J, Huang X, Liu D, Sui Q, Pan L, Su J, Deng J, Wu G, Li R, Deng S, Bai Y, Ligu Y, Tan W, Wu C, Wu T, Zheng J, Lin D (2020) Serum piRNA-54265 is a new biomarker for early detection and clinical surveillance of Human Colorectal Cancer. Theranostics 10:8468–8478
Malone CD, Hannon GJ (2009) Small RNAs as guardians of the genome. Cell 136:656–668
Mao Y, Qian SB (2020) Ribosome-guided piRNA production. Nat Cell Biol 22:141–142
Martinez VD, Vucic EA, Thu KL, Hubaux R, Enfield KS, Pikor LA, Becker-Santos DD, Brown CJ, Lam S, Lam WL (2015) Unique somatic and malignant expression patterns implicate PIWI-interacting RNAs in cancer-type specific biology. Sci Rep 5:10423
Matsumoto N, Nishimasu H, Sakakibara K, Nishida KM, Hirano T, Ishitani R, Siomi H, Siomi MC, Nureki O (2016) Crystal structure of silkworm PIWI-clade argonaute Siwi bound to piRNA. Cell 167:484-497.e9
Matsumoto N, Sato K, Nishimasu H, Namba Y, Miyakubi K, Dohmae N, Ishitani R, Siomi H, Siomi MC, Nureki O (2015) Crystal structure and activity of the endoribonuclease domain of the piRNA pathway factor maelstrom. Cell Rep 11:366–375
Mei Y, Wang Y, Kumari P, Shetty AC, Clark D, Gable T, MacKerell AD, Ma MZ, Weber DJ, Yang AJ, Edelman MJ, Mao L (2015) A piRNA-like small RNA interacts with and modulates p-ERM proteins in human somatic cells. Nat Commun 6:7316
Meikar O, Da Ros M, Korhonen H, Kotaja N (2011) Chromatoid body and small RNAs in male germ cells. Reproduction 142:195–209
Meikar O, Vagin VV, Chalmel F, Sostar K, Lardenois A, Hammell M, Jin Y, Da Ros M, Wasik KA, Toppari J, Hannon GJ, Kotaja N (2014) An atlas of chromatoid body components. RNA 20:483–495
Mochizuki K, Fine NA, Fujisawa T, Gorovsky MA (2002) Analysis of a piwi-related gene implicates small RNAs in genome rearrangement in tetrahymena. Cell 110:689–699
Mochizuki K, Gorovsky MA (2004) Small RNAs in genome rearrangement in Tetrahymena. Curr Opin Genet Dev 14:181–187
Mohn F, Handler D, Brennecke J (2015) Noncoding RNA. piRNA-guided slicing specifies transcripts for Zucchini-dependent, phased piRNA biogenesis. Science 348:812–817
Moyano M, Stefani G (2015) piRNA involvement in genome stability and human cancer. J Hematol Oncol 8:38
Muerdter F, Olovnikov I, Molaro A, Rozhkov NV, Czech B, Gordon A, Hannon GJ, Aravin AA (2012) Production of artificial piRNAs in flies and mice. RNA 18:42–52
Murano K, Iwasaki YW, Ishizu H, Mashiko A, Shibuya A, Kondo S, Adachi S, Suzuki S, Saito K, Natsume T, Siomi MC, Siomi H (2019) Nuclear RNA export factor variant initiates piRNA-guided co-transcriptional silencing. EMBO J 38:e102870
Murchison EP, Kheradpour P, Sachidanandam R, Smith C, Hodges E, Xuan Z, Kellis M, Grutzner F, Stark A, Hannon GJ (2008) Conservation of small RNA pathways in platypus. Genome Res 18:995–1004
Nandi S, Chandramohan D, Fioriti L, Melnick AM, Hébert JM, Mason CE, Rajasethupathy P, Kandel ER (2016) Roles for small noncoding RNAs in silencing of retrotransposons in the mammalian brain. Proc Natl Acad Sci USA 113:12697–12702
Nicholls PK, Schorle H, Naqvi S, Hu YC, Fan Y, Carmell MA, Dobrinski I, Watson AL, Carlson DF, Fahrenkrug SC, Page DC (2019) Mammalian germ cells are determined after PGC colonization of the nascent gonad. Proc Natl Acad Sci U S A 116:25677–25687
Nishida KM, Okada TN, Kawamura T, Mituyama T, Kawamura Y, Inagaki S, Huang H, Chen D, Kodama T, Siomi H, Siomi MC (2009) Functional involvement of Tudor and dPRMT5 in the piRNA processing pathway in Drosophila germlines. EMBO J 28:3820–3831
Nishida KM, Saito K, Mori T, Kawamura Y, Nagami-Okada T, Inagaki S, Siomi H, Siomi MC (2007) Gene silencing mechanisms mediated by Aubergine piRNA complexes in Drosophila male gonad. RNA 13:1911–1922
Nishimasu H, Ishizu H, Saito K, Fukuhara S, Kamatani MK, Bonnefond L, Matsumoto N, Nishizawa T, Nakanaga K, Aoki J, Ishitani R, Siomi H, Siomi MC, Nureki O (2012) Structure and function of Zucchini endoribonuclease in piRNA biogenesis. Nature 491:284–287
Nishimura T, Nagamori I, Nakatani T, Izumi N, Tomari Y, Kuramochi-Miyagawa S, Nakano T (2018) PNLDC1, mouse pre-piRNA Trimmer, is required for meiotic and post-meiotic male germ cell development. EMBO Rep 19:e44957
Obbard DJ, Welch JJ, Kim KW, Jiggins FM (2009) Quantifying adaptive evolution in the Drosophila immune system. PLoS Genet 5:e1000698
Ohara T, Sakaguchi Y, Suzuki T, Ueda H, Miyauchi K, Suzuki T (2007) The 3′ termini of mouse Piwi-interacting RNAs are 2´-O-methylated. Nat Struct Mol Biol 14:349–350
Olivieri D, Senti KA, Subramanian S, Sachidanandam R, Brennecke J (2012) The cochaperone shutdown defines a group of biogenesis factors essential for all piRNA populations in Drosophila. Mol Cell 47:954–969
Onohara Y, Yokota S (2012) Nuage components and their contents in mammalian spermatogenic cells, as revealed by immunoelectron microscopy. Meiosis 40:217–240
Özata DM, Yu T, Mou H, Gainetdinov I, Colpan C, Cecchini K, Kaymaz Y, Wu PH, Fan K, Kucukural A, Weng Z, Zamore PD (2020) Evolutionarily conserved pachytene piRNA loci are highly divergent among modern humans. Nat Ecol Evol 4:156–168
Palmer, S., and Whybrow, A. (2007). Handbook of coaching psychology
Pan J, Goodheart M, Chuma S, Nakatsuji N, Page DC, Wang PJ (2005) RNF17, a component of the mammalian germ cell nuage, is essential for spermiogenesis. Development 132:4029–4039
Pandey RR, Homolka D, Olotu O, Sachidanandam R, Kotaja N, Pillai RS (2018) Exonuclease domain-containing 1 enhances miwi2 pirna biogenesis via its interaction with TDRD12. Cell Rep 24:3423-3432.e4
Pandey RR, Tokuzawa Y, Yang Z, Hayashi E, Ichisaka T, Kajita S, Asano Y, Kunieda T, Sachidanandam R, Chuma S, Yamanaka S, Pillai RS (2013) Tudor domain containing 12 (TDRD12) is essential for secondary PIWI interacting RNA biogenesis in mice. Proc Natl Acad Sci USA 110:16492–16497
Parvinen M (2005) The chromatoid body in spermatogenesis. Int J Androl 28:189–201
Patil VS, Anand A, Chakrabarti A, Kai T (2014) The Tudor domain protein Tapas, a homolog of the vertebrate Tdrd7, functions in the piRNA pathway to regulate retrotransposons in germline of Drosophila melanogaster. BMC Biol 12:61
Perera BPU, Tsai ZT, Colwell ML, Jones TR, Goodrich JM, Wang K, Sartor MA, Faulk C, Dolinoy DC (2019) Somatic expression of piRNA and associated machinery in the mouse identifies short, tissue-specific piRNA. Epigenetics 14:504–521
Preall JB, Czech B, Guzzardo PM, Muerdter F, Hannon GJ (2012) shutdown is a component of the Drosophila piRNA biogenesis machinery. RNA 18:1446–1457
Qu X, Wen JD, Lancaster L, Noller HF, Bustamante C, Tinoco I (2011) The ribosome uses two active mechanisms to unwind messenger RNA during translation. Nature 475:118–121
Reuter M, Berninger P, Chuma S, Shah H, Hosokawa M, Funaya C, Antony C, Sachidanandam R, Pillai RS (2011) Miwi catalysis is required for piRNA amplification-independent LINE1 transposon silencing. Nature 480:264–267
Reynolds N, Collier B, Bingham V, Gray NK, Cooke HJ (2007) Translation of the synaptonemal complex component Sycp3 is enhanced in vivo by the germ cell specific regulator Dazl. RNA 13:974–981
Robine N, Lau NC, Balla S, Jin Z, Okamura K, Kuramochi-Miyagawa S, Blower MD, Lai EC (2009) A broadly conserved pathway generates 3’UTR-directed primary piRNAs. Curr Biol 19:2066–2076
Rosenkranz D, Han CT, Roovers EF, Zischler H, Ketting RF (2015) Piwi proteins and piRNAs in mammalian oocytes and early embryos: From sample to sequence. Genom Data 5:309–313
Saito K, Sakaguchi Y, Suzuki T, Suzuki T, Siomi H, Siomi MC (2007) Pimet, the Drosophila homolog of HEN1, mediates 2′-O-methylation of Piwi- interacting RNAs at their 3′ ends. Genes Dev 21:1603–1608
Santos AC, Lehmann R (2004) Germ cell specification and migration in Drosophila and beyond. Curr Biol 14:R578–R589
Saxe JP, Chen M, Zhao H, Lin H (2013) Tdrkh is essential for spermatogenesis and participates in primary piRNA biogenesis in the germline. EMBO J 32:1869–1885
Schoenberg DR, Maquat LE (2012) Regulation of cytoplasmic mRNA decay. Nat Rev Genet 13:246–259
Schöpp T, Zoch A, Berrens RV, Auchynnikava T, Kabayama Y, Vasiliauskaitė L, Rappsilber J, Allshire RC, O’Carroll D (2020) TEX15 is an essential executor of MIWI2-directed transposon DNA methylation and silencing. Nat Commun 11:3739
Seto AG, Kingston RE, Lau NC (2007) The coming of age for Piwi proteins. Mol Cell 26:603–609
Sharma AK, Nelson MC, Brandt JE, Wessman M, Mahmud N, Weller KP, Hoffman R (2001) Human CD34(+) stem cells express the hiwi gene, a human homologue of the Drosophila gene piwi. Blood 97:426–434
Shiromoto Y, Kuramochi-Miyagawa S, Daiba A, Chuma S, Katanaya A, Katsumata A, Nishimura K, Ohtaka M, Nakanishi M, Nakamura T, Yoshinaga K, Asada N, Nakamura S, Yasunaga T, Kojima-Kita K, Itou D, Kimura T, Nakano T (2013) GPAT2, a mitochondrial outer membrane protein, in piRNA biogenesis in germline stem cells. RNA 19:803–810
Shoemaker CJ, Green R (2012) Translation drives mRNA quality control. Nat Struct Mol Biol 19:594–601
Shoji M, Tanaka T, Hosokawa M, Reuter M, Stark A, Kato Y, Kondoh G, Okawa K, Chujo T, Suzuki T, Hata K, Martin SL, Noce T, Kuramochi-Miyagawa S, Nakano T, Sasaki H, Pillai RS, Nakatsuji N, Chuma S (2009) The TDRD9-MIWI2 complex is essential for piRNA-mediated retrotransposon silencing in the mouse male germline. Dev Cell 17:775–787
Shum EY, Jones SH, Shao A, Dumdie J, Krause MD, Chan WK, Lou CH, Espinoza JL, Song HW, Phan MH, Ramaiah M, Huang L, McCarrey JR, Peterson KJ, De Rooij DG, Cook-Andersen H, Wilkinson MF (2016) The antagonistic gene paralogs Upf3a and Upf3b govern nonsense-mediated RNA decay. Cell 165:382–395
Sienski G, Dönertas D, Brennecke J (2012) Transcriptional silencing of transposons by Piwi and maelstrom and its impact on chromatin state and gene expression. Cell 151:964–980
Simkin A, Wong A, Poh YP, Theurkauf WE, Jensen JD (2013) Recurrent and recent selective sweeps in the piRNA pathway. Evolution 67:1081–1090
Smith JM, Bowles J, Wilson M, Teasdale RD, Koopman P (2004) Expression of the tudor-related gene Tdrd5 during development of the male germline in mice. Gene Expr Patterns 4:701–705
Soper SF, van der Heijden GW, Hardiman TC, Goodheart M, Martin SL, de Boer P, Bortvin A (2008) Mouse maelstrom, a component of nuage, is essential for spermatogenesis and transposon repression in meiosis. Dev Cell 15:285–297
Specchia V, Piacentini L, Tritto P, Fanti L, D’Alessandro R, Palumbo G, Pimpinelli S, Bozzetti MP (2010) Hsp90 prevents phenotypic variation by suppressing the mutagenic activity of transposons. Nature 463:662–665
Sun YH, Xie LH, Zhuo X, Chen Q, Ghoneim D, Zhang B, Jagne J, Yang C, Li XZ (2017) Domestic chickens activate a piRNA defense against avian leukosis virus. Elife 6:48
Sun YH, Zhu J, Xie LH, Li Z, Meduri R, Zhu X, Song C, Chen C, Ricci EP, Weng Z, Li XZ (2020) Ribosomes guide pachytene piRNA formation on long intergenic piRNA precursors. Nat Cell Biol 22:200–212
Sun YH, Jiang F, Li XZ (2018) Disruption of Tdrd5 decouples the stepwise processing of long precursor transcripts during pachytene PIWI-interacting RNA biogenesis. Biol Reprod 99:684–685
Sun YH, Wang RH, Du K, Zheng J, Xie LH, Pereira AA, Zhang C, Ricci EP, Li XZ (2021) Coupled protein synthesis and ribosome-guided piRNA processing on mRNAs. Nat Commun 2(1):5970
Takyar S, Hickerson RP, Noller HF (2005) mRNA helicase activity of the ribosome. Cell 120:49–58
Tam OH, Aravin AA, Stein P, Girard A, Murchison EP, Cheloufi S, Hodges E, Anger M, Sachidanandam R, Schultz RM, Hannon GJ (2008) Pseudogene-derived small interfering RNAs regulate gene expression in mouse oocytes. Nature 453:534–538
Tan M, Tol HTAV, Rosenkranz D, Roovers EF, Damen MJ, Stout TAE, Wu W, Roelen BAJ (2020) PIWIL3 forms a complex with TDRKH in mammalian oocytes. Cells 9:48
Tanaka T, Hosokawa M, Vagin VV, Reuter M, Hayashi E, Mochizuki AL, Kitamura K, Yamanaka H, Kondoh G, Okawa K, Kuramochi-Miyagawa S, Nakano T, Sachidanandam R, Hannon GJ, Pillai RS, Nakatsuji N, Chuma S (2011) Tudor domain containing 7 (Tdrd7) is essential for dynamic ribonucleoprotein (RNP) remodeling of chromatoid bodies during spermatogenesis. Proc Natl Acad Sci USA 108:10579–10584
Thomson T, Lin H (2009) The biogenesis and function of PIWI proteins and piRNAs: progress and prospect. Annu Rev Cell Dev Biol 25:355–376
Turner JM, Mahadevaiah SK, Ellis PJ, Mitchell MJ, Burgoyne PS (2006) Pachytene asynapsis drives meiotic sex chromosome inactivation and leads to substantial postmeiotic repression in spermatids. Dev Cell 10:521–529
Vagin VV, Sigova A, Li C, Seitz H, Gvozdev V, Zamore PD (2006) A distinct small RNA pathway silences selfish genetic elements in the germline. Science 313:320–324
Vagin VV, Yu Y, Jankowska A, Luo Y, Wasik KA, Malone CD, Harrison E, Rosebrock A, Wakimoto BT, Fagegaltier D, Muerdter F, Hannon GJ (2013) Minotaur is critical for primary piRNA biogenesis. RNA 19:1064–1077
Vasileva A, Tiedau D, Firooznia A, Muller-Reichert T, Jessberger R (2009) Tdrd6 is required for spermiogenesis, chromatoid body architecture, and regulation of miRNA expression. Curr Biol 19:630–639
Voigt F, Reuter M, Kasaruho A, Schulz EC, Pillai RS, Barabas O (2012) Crystal structure of the primary piRNA biogenesis factor Zucchini reveals similarity to the bacterial PLD endonuclease Nuc. RNA 18:2128–2134
Vourekas A, Zheng K, Fu Q, Maragkakis M, Alexiou P, Ma J, Pillai RS, Mourelatos Z, Wang PJ (2015) The RNA helicase MOV10L1 binds piRNA precursors to initiate piRNA processing. Genes Dev 29:617–629
Vourekas A, Zheng Q, Alexiou P, Maragkakis M, Kirino Y, Gregory BD, Mourelatos Z (2012) Mili and Miwi target RNA repertoire reveals piRNA biogenesis and function of Miwi in spermiogenesis. Nat Struct Mol Biol 19:773–781
Wang H, Ma Z, Niu K, Xiao Y, Wu X, Pan C, Zhao Y, Wang K, Zhang Y, Liu N (2016) Antagonistic roles of Nibbler and Hen1 in modulating piRNA 3’ ends in Drosophila. Development 143:530–539
Wang J, Saxe JP, Tanaka T, Chuma S, Lin H (2009) Mili interacts with tudor domain-containing protein 1 in regulating spermatogenesis. Curr Biol 19:640–644
Wasik KA, Tam OH, Knott SR, Falciatori I, Hammell M, Vagin VV, Hannon GJ (2015) RNF17 blocks promiscuous activity of PIWI proteins in mouse testes. Genes Dev 29:1403–1415
Watanabe T, Totoki Y, Toyoda A, Kaneda M, Kuramochi-Miyagawa S, Obata Y, Chiba H, Kohara Y, Kono T, Nakano T, Surani MA, Sakaki Y, Sasaki H (2008) Endogenous siRNAs from naturally formed dsRNAs regulate transcripts in mouse oocytes. Nature 453:539–543
Wenda JM, Homolka D, Yang Z, Spinelli P, Sachidanandam R, Pandey RR, Pillai RS (2017) Distinct roles of RNA helicases MVH and TDRD9 in PIWI slicing-triggered mammalian piRNA biogenesis and function. Dev Cell 41:623-637.e9
Wu PH, Fu Y, Cecchini K, Özata DM, Arif A, Yu T, Colpan C, Gainetdinov I, Weng Z, Zamore PD (2020) The evolutionarily conserved piRNA-producing locus pi6 is required for male mouse fertility. Nat Genet 52:728–739
Wynant N, Santos D, Vanden Broeck J (2017) The evolution of animal Argonautes: evidence for the absence of antiviral AGO Argonautes in vertebrates. Sci Rep 7:9230
Xiol J, Cora E, Koglgruber R, Chuma S, Subramanian S, Hosokawa M, Reuter M, Yang Z, Berninger P, Palencia A, Benes V, Penninger J, Sachidanandam R, Pillai RS (2012) A role for Fkbp6 and the chaperone machinery in piRNA amplification and transposon silencing. Mol Cell 47:970–979
Xiol J, Spinelli P, Laussmann MA, Homolka D, Yang Z, Cora E, Couté Y, Conn S, Kadlec J, Sachidanandam R, Kaksonen M, Cusack S, Ephrussi A, Pillai RS (2014) RNA clamping by Vasa assembles a piRNA amplifier complex on transposon transcripts. Cell 157:1698–1711
Yabuta Y, Ohta H, Abe T, Kurimoto K, Chuma S, Saitou M (2011) TDRD5 is required for retrotransposon silencing, chromatoid body assembly, and spermiogenesis in mice. J Cell Biol 192:781–795
Yamaguchi S, Oe A, Nishida KM, Yamashita K, Kajiya A, Hirano S, Matsumoto N, Dohmae N, Ishitani R, Saito K, Siomi H, Nishimasu H, Siomi MC, Nureki O (2020) Crystal structure of Drosophila Piwi. Nat Commun 11:858
Yamanaka S, Siomi MC, Siomi H (2014) piRNA clusters and open chromatin structure. Mob DNA 5:22
Yan Z, Hu HY, Jiang X, Maierhofer V, Neb E, He L, Hu Y, Hu H, Li N, Chen W, Khaitovich P (2011) Widespread expression of piRNA-like molecules in somatic tissues. Nucleic Acids Res 39:6596–6607
Yang Q, Li R, Lyu Q, Hou L, Liu Z, Sun Q, Liu M, Shi H, Xu B, Yin M, Yan Z, Huang Y, Liu M, Li Y, Wu L (2019) Single-cell CAS-seq reveals a class of short PIWI-interacting RNAs in human oocytes. Nat Commun 10:3389
Yang Z, Chen KM, Pandey RR, Homolka D, Reuter M, Janeiro BK, Sachidanandam R, Fauvarque MO, McCarthy AA, Pillai RS (2016) PIWI Slicing and EXD1 Drive Biogenesis of Nuclear piRNAs from Cytosolic Targets of the Mouse piRNA Pathway. Mol Cell 61:138–152
Yao MC, Fuller P, Xi X (2003) Programmed DNA deletion as an RNA-guided system of genome defense. Science 300:1581–1584
Yi M, Chen F, Luo M, Cheng Y, Zhao H, Cheng H, Zhou R (2014) Rapid evolution of piRNA pathway in the teleost fish: implication for an adaptation to transposon diversity. Genome Biol Evol 6:1393–1407
Yoshimura T, Toyoda S, Kuramochi-Miyagawa S, Miyazaki T, Miyazaki S, Tashiro F, Yamato E, Nakano T, Miyazaki J (2009) Gtsf1/Cue110, a gene encoding a protein with two copies of a CHHC Zn-finger motif, is involved in spermatogenesis and retrotransposon suppression in murine testes. Dev Biol 335:216–227
Yoshimura T, Watanabe T, Kuramochi-Miyagawa S, Takemoto N, Shiromoto Y, Kudo A, Kanai-Azuma M, Tashiro F, Miyazaki S, Katanaya A, Chuma S, Miyazaki JI (2018) Mouse GTSF1 is an essential factor for secondary piRNA biogenesis. EMBO Rep 19:e42054
Yu T, Fan K, Özata DM, Zhang G, Fu Y, Theurkauf WE, Zamore PD, Weng Z (2021) Long first exons and epigenetic marks distinguish conserved pachytene piRNA clusters from other mammalian genes. Nat Commun 12:73
Yu T, Koppetsch BS, Pagliarani S, Johnston S, Silverstein NJ, Luban J, Chappell K, Weng Z, Theurkauf WE (2019) The piRNA Response to Retroviral Invasion of the Koala Genome. Cell 179:632-643.e12
Zamparini AL, Davis MY, Malone CD, Vieira E, Zavadil J, Sachidanandam R, Hannon GJ, Lehmann R (2011) Vreteno, a gonad-specific protein, is essential for germline development and primary piRNA biogenesis in Drosophila. Development 138:4039–4050
Zhang, H., Zhang, F., Chen, Q., Li, M., Lv, X., Xiao, Y., Zhang, Z., Hou, L., Lai, Y., Zhang, Y., Zhang, A., Gao, S., Fu, H., Xiao, W., Zhou, J., Diao, F., Shi, A., Su, Y.-Q., Zeng, W., Wu, L., and Li, J. (2021). The piRNA pathway is essential for generating functional oocytes in golden hamsters. Nat Cell Biol
Zhang P, Kang JY, Gou LT, Wang J, Xue Y, Skogerboe G, Dai P, Huang DW, Chen R, Fu XD, Liu MF, He S (2015) MIWI and piRNA-mediated cleavage of messenger RNAs in mouse testes. Cell Res 25:193–207
Zhang S, Pointer B, Kelleher ES (2020) Rapid evolution of piRNA-mediated silencing of an invading transposable element was driven by abundant de novo mutations. Genome Res 30:566–575
Zhang Y, Guo R, Cui Y, Zhu Z, Zhang Y, Wu H, Zheng B, Yue Q, Bai S, Zeng W, Guo X, Zhou Z, Shen B, Zheng K, Liu M, Ye L, Sha J (2017) An essential role for PNLDC1 in piRNA 3’ end trimming and male fertility in mice. Cell Res 27:1392–1396
Zhang Z, Koppetsch BS, Wang J, Tipping C, Weng Z, Theurkauf WE, Zamore PD (2014) Antisense piRNA amplification, but not piRNA production or nuage assembly, requires the Tudor-domain protein Qin. EMBO J 33:536–539
Zhao PP, Yao MJ, Chang SY, Gou LT, Liu MF, Qiu ZL, Yuan XB (2015) Novel function of PIWIL1 in neuronal polarization and migration via regulation of microtubule-associated proteins. Mol Brain 8:39
Zheng K, Wang PJ (2012) Blockade of pachytene piRNA biogenesis reveals a novel requirement for maintaining post-meiotic germline genome integrity. PLoS Genet 8:e1003038
Zheng K, Xiol J, Reuter M, Eckardt S, Leu NA, McLaughlin KJ, Stark A, Sachidanandam R, Pillai RS, Wang PJ (2010) Mouse MOV10L1 associates with Piwi proteins and is an essential component of the Piwi-interacting RNA (piRNA) pathway. Proc Natl Acad Sci USA 107:11841–11846
Zhou L, Canagarajah B, Zhao Y, Baibakov B, Tokuhiro K, Maric D, Dean J (2017) BTBD18 Regulates a Subset of piRNA-Generating Loci through Transcription Elongation in Mice. Dev Cell 40:453-466.e5
Acknowledgements
We thank Guy Riddihough, John Roy Lozada, and other members in the Li lab for editing and providing constructive suggestions on the manuscript. This work was supported in part by National Institutes of Health Grant R35GM128782 and Agriculture and Food Research Initiative Competitive Grant No. 2018-67015-27615 from the USDA National Institute of Food and Agriculture.
Author information
Authors and Affiliations
Contributions
YHS and XZL wrote the manuscript. BL proofread the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sun, Y.H., Lee, B. & Li, X.Z. The birth of piRNAs: how mammalian piRNAs are produced, originated, and evolved. Mamm Genome 33, 293–311 (2022). https://doi.org/10.1007/s00335-021-09927-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00335-021-09927-8