Abstract
Objectives
The analysis of myocardial deformation using feature tracking in cardiovascular MR allows for the assessment of global and segmental strain values. The aim of this study was to compare strain values derived from artificial intelligence (AI)–based contours with manually derived strain values in healthy volunteers and patients with cardiac pathologies.
Materials and methods
A cohort of 136 subjects (60 healthy volunteers and 76 patients; of those including 46 cases with left ventricular hypertrophy (LVH) of varying etiology and 30 cases with chronic myocardial infarction) was analyzed. Comparisons were based on quantitative strain analysis and on a geometric level by the Dice similarity coefficient (DSC) of the segmentations. Strain quantification was performed in 3 long-axis slices and short-axis (SAX) stack with epi- and endocardial contours in end-diastole. AI contours were checked for plausibility and potential errors in the tracking algorithm.
Results
AI-derived strain values overestimated radial strain (+ 1.8 ± 1.7% (mean difference ± standard deviation); p = 0.03) and underestimated circumferential (− 0.8 ± 0.8%; p = 0.02) and longitudinal strain (− 0.1 ± 0.8%; p = 0.54). Pairwise group comparisons revealed no significant differences for global strain. The DSC showed good agreement for healthy volunteers (85.3 ± 10.3% for SAX) and patients (80.8 ± 9.6% for SAX). In 27 cases (27/76; 35.5%), a tracking error was found, predominantly (24/27; 88.9%) in the LVH group and 22 of those (22/27; 81.5%) at the insertion of the papillary muscle in lateral segments.
Conclusions
Strain analysis based on AI-segmented images shows good results in healthy volunteers and in most of the patient groups. Hypertrophied ventricles remain a challenge for contouring and feature tracking.
Clinical relevance statement
AI-based segmentations can help to streamline and standardize strain analysis by feature tracking.
Key Points
• Assessment of strain in cardiovascular magnetic resonance by feature tracking can generate global and segmental strain values.
• Commercially available artificial intelligence algorithms provide segmentation for strain analysis comparable to manual segmentation.
• Hypertrophied ventricles are challenging in regards of strain analysis by feature tracking.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Myocardial strain allows for a quantitative measurement of myocardial deformation. Analysis of strain using cardiovascular magnetic resonance (CMR) can be obtained either by tissue tagging or by direct feature tracking (FT) on standard cine images. While CMR tagging has been validated and has further advanced into various different techniques (e.g., (fast) Strain Encoding Magnetic Resonance imaging (SENC) or displacement encoding with stimulated echoes (DENSE)) [1,2,3,4], it still has the drawback of requiring special sequences and scan time. In contrast to this, FT is a promising tool as it allows for the assessment of segmental and global strains in longitudinal, circumferential, and radial directions (LS, CS, RS, GLS, GCS, and GRS, respectively) from standard cine images which are usually acquired during clinical routine [5]. Left ventricular (LV) strain analysis applying CMR has been implied in a wide array of clinical diseases ranging from chemotherapy-induced cardiotoxicity [6, 7] to ischemic heart disease [8,9,10] and even non-ischemic heart diseases like hypertrophic cardiomyopathies (HCMs) [11, 12] or cases of acute myocarditis [13, 14]. Despite its wide utility and power to detect myocardial changes even in states with preserved function, FT still lacks standardization and consensus about the methodological process. In a previous study, different factors, like post-processing software used, slice selection, and 2D or 3D analysis, which all have the potential to influence strain values, were analyzed [15]. Additionally, one must take the time-consuming manual contouring process as well as the reader’s level of expertise and training into consideration, which also impact strain evaluation [16]. One potential approach to reduce influence of manually derived contours is the use of artificial intelligence (AI)–derived contours. AI-based segmentation and strain evaluation have been previously applied and validated in a large cohort with commercially available software [17] as well as commercially unavailable software [18]. These advances may further streamline strain assessment and help to reach a consensus about a standardized approach to feature tracking in clinical routine and “big data” studies.
Our study aimed at evaluating and comparing manual and AI-based approaches regarding quantitative strain metrics used in clinical routine as well as on a contour level for strain assessment by FT in healthy volunteers and patients with different cardiac diseases in order to identify strengths and weaknesses of these methods.
Materials and methods
The ethics review board approved all studies and all participants gave written informed consent.
Study population
In the healthy volunteers cohort, 67 subjects, retrospectively recruited in a previous study [15], were included. For the final analysis, 7 volunteers had to be excluded due to lack of a short-axis (SAX) stack covering the entire ventricle and one due to significant respiratory artifacts, which ultimately resulted to a healthy cohort consisting of 60 subjects. For the clinical validation, a cohort of 76 patients, chosen from previous studies [19,20,21], with cases including left ventricular hypertrophy (LVH) (n = 46 consisting of 8 patients with arterial hypertension (AHT)), 24 with aortic stenosis (AS), 14 with HCM, and chronic myocardial infarction (CMI) (N = 30 consisting of 10 patients with preserved left ventricular ejection fraction (LVEF), no wall motion abnormalities (WMA) and focal fibrosis (CMI-F), 10 with reduced LVEF, regional WMA, focal fibrosis (CMI-WF), and 10 with reduced LVEF with dilated LVs, global WMA, and focal fibrosis (CMI-EWF)), was constructed.
Imaging protocol
CMR was performed either at a 1.5-T scanner (MAGNETOM Avanto−FIT, Siemens Healthineers) or a 3-T scanner (MAGNETOM Verio, Siemens Healthineers). Steady-state free precession-based cine images were acquired for 3 long-axis (LAX) views including a 2 chamber view (cv), a 3 cv, and a 4 cv as well as one SAX stack covering the entire left ventricle. Sequence parameters for the SAX stack at the 1.5-T scanner were as follows: time of repetition 2.8–3.31 ms, slice thickness 7 mm with no gap, flip angle 80°, echo time 1.2–1.44 ms, field of view 340–380 × 276– 308, 75 mm2, matrix 192 × 156, voxel size 1.4–2.0 × 1.4–2.0, 30 cardiac phases; and for the 3-T scanner: time of repetition 3.1 ms, slice thickness 6 mm with no gap, flip angle 45°, echo time 1.3 ms, field of view 340 × 276 mm2, matrix 192 × 156, voxel size 1.4 × 1.4, 30 cardiac phases.
Manual segmentations
Manual segmentation was performed with dedicated software (circle CVI42 version 5.14.7, Circle Cardiovascular Imaging Inc.). Manual endo- and epicardial contours were drawn in end-diastole (ED), determined by the phase with the largest LV volume in SAX as well as in the 2 cv, 3 cv, and 4 cv. We were particularly attentive during segmentation to avoid contouring phases with the left ventricular outflow tract (LVOT) still visible in diastole and/or systole. Papillary muscles were not separately contoured as recently published [15]. Reference points for the delineation of segments were manually placed at the subepicardial border at the anterior intersection of the left and right ventricle.
AI-generated segmentations
Similarly to the manual strain assessment, AI contours were derived in ED. The AI segmented slices with visible LVOT using open LV endo- and epicardial contours, which were disregarded for strain analysis. Reference points were automatically set by the AI; however, each point was manually validated to obtain comparable segmental values. The AI segmentation algorithms employed in the Circle CVI42 software are comprised of different deep convolutional neural network models trained to perform SAX and LAX CMR image segmentation. A similar model architecture as that of the standard U-Net is adopted for this purpose, along with various data augmentation techniques to enhance the generalizability of the trained model. The model was trained on the UK Biobank data as well as datasets that include patient data with pathological conditions including tetralogy, cardiomyopathy, and hypertension [22]. These models operate solely on image pixel data and image header information such as image dimensions and pixel spacing.
Strain assessment
After segmentation, a FT algorithm provided strain values. The algorithm uses myocardial points and tracks them along the cardiac cycle [23, 24]. On a quantitative level, the manual and the AI approach were compared for strain assessment in CS and RS retrieved from SAX and LS retrieved from LAX views. All strain values were derived for global as well as segmental values according to the 17-segment model of the American heart association (AHA) for CMR without the apical segment [25]. Correct FT was assessed by either mesh analysis or by tracking the myocardial points through the phases. Improper tracking was defined as mesh overlay or myocardial points not following the extent of the contours [15, 26]. To enable comparability between the segmentations regarding the strain analysis, we verified that the AI algorithm chose the proper ED phase.
Statistical analysis
All continuous variables are presented by mean and standard deviation (SD). Normal distribution was visualized by QQ plots. A mixed model was used to assess measurement differences segmentally and globally between the modalities for healthy volunteers and patients combined. In the mixed model, a global test was applied to test for any differences. In the case of a significant global test, pairwise comparisons were performed. Additionally, we tested whether a difference found between the AI and manual segmentations was homogenous over all groups or whether a certain group showed major deviations.
Additionally, both segmentation approaches were compared using the “Lazy Luna” (LL) tool which allows to assess the similarity those of an experienced reader and an AI on the contour level, via reproducibility validation metrics [27]. We chose the Dice similarity coefficient (DSC) and the Hausdorff distance (HD) to compare the consensus of the manual contours and the AI approach. DSC scores were calculated based on myocardial class, which was derived from the intersection of the endo- and epicardial contours placed manually or by the AI. High DSC numbers signifying a substantial overlap of the segmented areas and low numbers indicate incongruences. Vice versa holds true for the HD metric. In order to compare the proper placement of the insertion point, the LL tool additionally compared the manual- and AI-placed insertion point based on an angular difference to the left ventricular centroid. As some SAX acquisitions were acquired after contrast media application (post-CM), GRS and GCS as well as DSC and HD metrics were compared with acquisitions pre-contrast media application (pre-CM). Statistical analysis was performed using dedicated software (SPSS version 26, International Business Machines and SAS version 9.4, SAS Institute Inc.). The segmentation comparison tool “Lazy Luna” and the bulls-eye plots were created in Python (Version3.8, Python Software Foundation) [27].
Results
Study population
In the healthy cohort, 67 subjects were re-analyzed and in the clinical cohort 76 patients. Details concerning the healthy volunteer cohort and the patients are given in Table 1. Overall, AI-derived strain values showed a trend towards overestimation of RS values ((mean difference % (± SD)) + 1.8 (± 1.7)) and an underestimation of CS (− 0.8 (± 0.8)) and LS values (− 0.1 (± 0.8)) (Fig. 1). Regarding the entire studied cohort, including the healthy probands and the ones with pathologies, global testing revealed significant differences for GCS (p = 0.03), GRS (p = 0.03), and RS AHA segments 5 (p = 0.045), 10 (p = 0.04), 11 (p = 0.002), and 12 (p = 0.03). Pairwise testing revealed no significant differences if manual and AI approaches were compared for the specific cohort for GCS, GRS, and RS AHA segment 12. Statistically significant results were found between AI and manual strain values for the subgroups AS and HCM for RS AHA segment 5 (p = 0.01 and p = 0.03, respectively), 10 (p = 0.01 and p = 0.02, respectively), and 11 (p = 0.02 and p = 0.01, respectively) (Supplementary material 1). Additionally, we found no significant interaction between the methods and the examined subgroups except for AHA segment 10 for CS (p = 0.02) as well as RS (p = 0.02) (Supplementary Material 2). Overall, in 83/136 cases (61%), SAX images were acquired post-CM. For the subgroups, the following percentages of SAX were acquired after post-CM: healthy cohort 37/60 (61.7%), CMI-F 10/10 (100%), CMI-WF 10/10 (100%), CMI-EWF 10/10 (100%), AHT 7/8 (87.5%), AS 8/24 (33.3%), HCM 1/12 (8.3%).
Global strain values for the healthy and the disease cohorts for circumferential, radial, and longitudinal strains in %. Overall, AI-derived strain values showed a trend towards overestimation of RS values and underestimation of CS and LS values. Global significant differences were found for circumferential and radial global strain values, with pairwise comparison revealing no significant differences. LVH, left ventricular hypertrophy; AHT, arterial hypertension; AS, aortic stenosis; HCM, hypertrophic cardiomyopathy; CMI, chronic myocardial infarction; F, cases with focal fibrosis, no wall motion abnormalities, and preserved left ventricular function; WF, cases with focal fibrosis, wall motion abnormalities, and reduced left ventricular function; EWF, cases with fibrosis, global wall motion abnormalities, and reduced left ventricular function and dilated left ventricles
Strain analysis—healthy cohort
Strain analysis was feasible in all 60 cases for CS, RS, and LS. Global values were as follows: manual (mean % (± SD)) − 16.2 (± 2.2) GCS; 25.5 (± 4.9) GRS; − 17.5 (± 1.8) GLS; and for the AI approach: − 16.7 (± 2.2) GCS; 26.6 (± 5.0) GRS; − 17.3 (± 1.7) GLS. Segmental strain values with standard deviations are presented in Fig. 2. The DSC (in %) showed good agreement between the manual and AI-derived contours with 85.3 ± 10.3 for SAX contours, 85.8 ± 2.9 for 2cv contours, 83.1 ± 5.1 for 3cv contours, and 84.1 ± 4.1 for 4cv contours (Fig. 3). Comparison of the insertion points revealed a mean angle difference of 5.1 ± 10.9° (in relative values 1.4 ± 3.0% difference). Regarding both approaches, the highest CS and LS values as well as the lowest RS values were confined to segments 11 or 12 (insertion point of the papillary muscle) of the AHA model.
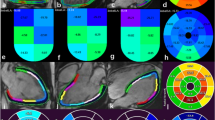
Segmental strain values for the healthy cohort for circumferential, radial and longitudinal strains in %. Segmental strain values, according to the American heart association model, are presented from left to right for circumferential, radial, and longitudinal strains in %. Pairwise comparisons showed no statistically significant differences between the manual (top) and the AI-based (bottom) segmentations
Dice metric comparison between manual and AI segmentations. Comparison of manual (second column from the left) and AI-based segmentations (third column from the left). Third column from the right depicts myocardial class annotations for manual segmentations (red) and the rightest column for AI segmentations (blue). The second column from the right demonstrates the spatial overlap between the contours (green area). Discrepancies are in the corresponding colors of manual (red) or AI contours (blue). DICE values ranged from 83% in the 3-chamber view to 94% in the short axis for the exemplary case
Strain analysis—patients with various cardiac diseases
AI-derived contour generation and strain analysis was possible in all clinical cases. Global and segmental values are presented in Supplementary Material 1. The Dice metric values showed good agreement between the manual and AI-derived contours with 80.8 ± 9.6 for SAX contours, 85.1 ± 4.6 for 2cv contours, 85.9 ± 6.7 for 3cv contours, and 85.5 ± 4.4 for 4cv contours on average (Table 2). Angular differences for the insertion points were on average 4.4 ± 9.4 degrees (1.2 ± 2.6%) for the cohort with cardiac disease. For the individual pathologies, the results were the following: AHT 3.3 ± 6.4° (0.9 ± 1.8%), AS 2.0 ± 6.5° (0.7 ± 1.8%), HCM 0.3 ± 12.7° (0.01 ± 3.5%), CMI-F 8.3 ± 7.7° (2.3 ± 2.1%), CMI-WF 5.9 ± 8.6° (1.6 ± 2.4%), and CMI-EWF 9.6 ± 9.2° (2.7 ± 2.6%). Figure 4 presents examples with proper tracking and corresponding pathologic features. Tracking errors were found in 27 cases (35.5%; 27/76) of which 3 were from CMI-WF and 24 from the LVH group with 4 in the AHT group (16.7%; 4/24), 16 in the AS group 66.7%, 16/24) and 4 in the HCM group (16.7%; 4/24). Further analysis revealed that tracking errors in the LVH group were mostly confined to the basal, midventricular anterolateral, and inferolateral segments at the insertion point of the papillary muscle (81.5%; 22/24), leading to higher LS strain values in these segments. The inferior midventricular segment was the third most affected region (8.3%; 2/24). In one case from the CMI group, despite proper epicardial contours, tracking features were identified within the right ventricle (3.7%; 1/76). Figure 5 shows examples of tracking errors encountered in the clinical cohort.
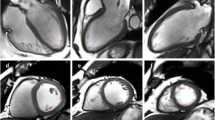
Examples of proper AI contours with strain values and underlying pathologies. Top row represents a patient with a chronic myocardial infarction and focal subendocardial scar at the inferolateral wall (late gadolinium enhancement (LGE)) images in a 2 chamber views (top row, left image) and short axis (top row, second from the left) with subtle wall motion abnormalities. Global longitudinal strain was reduced (− 15.9%). Middle row represents a patient with chronic myocardial infarction and microvascular obstruction on LGE imaging (middle row left and second from the left). Left ventricular function and global longitudinal strains were severely impaired. Bottom row represents a patient with left ventricular hypertrophy due to long-standing arterial hypertension. On LGE imaging a diffuse fibrotic process is visible (bottom row left and second from the left). Global longitudinal strain is mildly reduced with no focal accentuation. The second and third columns from the left represent manual and AI segmentations of two chamber views, respectively
Examples of tracking errors. The left and the middle column show tracking errors in the 3-chamber view at the insertion point of the papillary muscle in the anterolateral segment. Middle and bottom row represent manual and AI segmentations respectively. The right column shows a tracking error in the 2-chamber view in the inferior midventricular segment. Middle and bottom row represent manual and AI segmentations, respectively
Analysis of images pre- and post-contrast media application
GRS and GCS were lower post-CM for manual segmentations (GRS pre-CM 25.7 ± 6.9% vs. 20.1 ± 7.8 post-CM application; p < 0.001; GCS pre-CM − 15.0 ± 3.0 vs. − 13.3 ± 4.2 post-CM; p < 0.001) and AI segmentations (GRS pre-CM 28.2 ± 6.8% vs. 21.4 ± 8.1 post-CM; p < 0.001; GCS pre-CM − 17.1 ± 2.8 vs. − 13.0 ± 4.3 post-CM; p < 0.001).
The DSC and HD metrics were in the same range for pre-CM and post-CM SAX segmentations; however, a lower score post-CM were noted (DSC: 83.6 ± 9.3% for pre-CM vs. 80.5 ± 14.3% post-CM; HD: 3.3 ± 1.4 mm pre-CM vs. 3.6 ± 1.7 mm post-CM).
Discussion
The main results of our studies are as follows: strain analysis by FT on cine images based on AI-derived contours is feasible and results in equivalent global and segmental strain values with the exception of lateral segments in hypertrophied ventricles. The difference however is attributable to tracking errors as the spatial overlap metric shows good agreement of the methods.
CMR has become the gold standard for LV and right ventricular volume and mass quantification [28] with a standardized approach for analysis and post-processing of images [29]. Yet, there are no consensus recommendations on how to quantify LV myocardial tissue dynamics and deformation by applying CMR. Therefore, we explored how strain assessment by FT can potentially become more standardized involving AI-powered approaches. In our study, we could demonstrate that AI-generated contours for strain assessment by FT are reliable and result in equivalent global and segmental values. In a previous study by Ruijsink et al, DSC between manual and AI-based segmentation was 93% for the endocardial segmentation and 84% for the epicardial segmentation [18]. We found a similar DSC for the myocardial class in our study. Other segmentation algorithms which were tested on the “Automatic Cardiac Diagnosis Challenge” (ACDC) dataset, achieved DSC scores up to 96% for the LV [30]. These scores are higher than the one presented here; however, the underlying dataset in the challenge plays an important factor. The ACDC dataset included similar pathologies as in this study, such as HCM, CMI, and dilated cardiomyopathies; however, whether scans were carried out after contrast administration was not clearly depicted. Images obtained after contrast media application pose an additional challenge as myocardial fibrosis might be mistaken for the blood pool by AI algorithms as well as human readers. In this study, we found the overall lowest segmentation overlap, indicated by DSC and HD metrics, in the CMI group. As all scans in the CMI group were carried out after contrast administration and each case included at least one focal fibrosis; these scans posed a challenge for the algorithm. This was also evidenced by the lower DSC and HD in the comparison of pre-CM and post-CM images. In the subset of LVH, we found higher DSC scores in comparison to the healthy and CMI cohorts. We believe that this paradoxon can be explained by the larger LVM in the LVH cohort as the overall differences in segmentations are divided by a larger area. When considering the HD metric, the healthy cohort showed the lowest value regarding the SAX segmentations.
When segmental values, which are defined according to the AHA model, are compared, the insertion point has to be taken into consideration. In order to verify the proper insertion, visual analysis can be carried out; however, for large data, this is tedious. We propose therefore a comparison based on angular differences as outlined in the methods. The highest angular differences (8.7°) were seen in the CMI-F cohort. Comparing this to the general division of the AHA segments (60°), we feel that these are neglectable; however, the relevance of angular differences and their impact on AHA segments ought to be investigated further.
Interestingly, the study by Ruijsink et al used 3 SAX slices as well as 2cv and 4cv for strain assessment with diastolic contours. This might potentially impact the strain analysis of CS and RS values [15]. Previous studies reported normal values for FT on the full SAX coverage [5, 31]. To additionally achieve a more streamlined post-processing of CMR images, AI algorithms, with the placement of contours in ED and ES for functional assessment, can “recycle” these contours for strain assessment by FT. Further studies employing this approach, potentially in clinical routine scenarios, are needed to verify whether this approach is feasible in a real-world setting. We analyzed not only a healthy population but also one with different clinical entities, which were chosen as either to be a challenge regarding the segmentation (post-contrast cines with fibrosis) or for the FT algorithm (wall motion abnormalities) or both (LVH subgroup, CMI-EWF). In addition, we compared all cases 1:1 and not only selected cases, which minimizes the possibility of errors. The LVH group, especially AS and HCM cases, was burdensome for the AI on a contour and tracking level. Studies about AS and CMR FT have been previously published but none covered AI-derived contours or analysis of tracking errors [32,33,34,35,36,37]. As the majority of tracking issues in the LVH group was related to the anterolateral and inferolateral segments, papillary muscle hypertrophy could have played a role [38, 39]. As the FT algorithm relies on recognition of voxel-based features, which are in general sparse or even absent in the normal myocardium [40], hypertrophy of the myocardial tissue can further deteriorate tracking. We found the highest CS and LS as well as the lowest RS value in the abovementioned segments in the healthy cohort. Andre et al presented similar results, with the highest variance in these segments. Potentially, the movement of the papillary muscle can have an impact on the values in these segments even in non-pathologic states, which would be further exacerbated with hypertrophy of the myocardium and its appendages. A previous study found statistically significant differences between healthy male and female volunteers in AHA segment 5 of LS analysis [15]. One other study reported a significantly larger papillary muscle mass in males compared to females [41], which might possibly explain the previous findings and the ones presented here. In concordance with the identified segments that pose a challenge for the FT algorithm, the AI-derived strain values show the only significant differences compared to manual contours in these segments. Interestingly, the differences between the methods were due to strain values for AS and HCM cases (Supplementary Material 2). These differences however are not based on a contour level as evidenced by the DSC. Potential influencing factors might relate to the LAX extent or the acquisition itself. In comparison to the other pathologies, we found the 3cv slice location in this group frequently at a narrower angle. This finding might be related to a prominent and dilated ascending aortic root in AS patients impairing proper 3cv acquisitions [42, 43].
The intersegmental differences are a drawback for FT-derived strain values. In a head-to-head comparison between fast-SENC, tagging, and FT, all techniques had good reproducibility; however, in a segmental inter-study comparison, FT showed the lowest agreement [3]. This was confirmed by other studies reporting a rather large variation across segments rendering comparisons rather unfeasible and additionally demonstrating that segmental analysis with FT is complex and clinical implications uncertain [31, 44]. A potential solution might be the use of regional instead of segmental values [45]. In contrast to FT, other techniques such as DENSE have a higher reproducibility of segmental strain values [46, 47]. In addition, segmental strain values provided by DENSE have been shown to carry a prognostic implication in patients after an acute myocardial infarction [48]. Regarding the segmental approach, SENC-derived strain values similarly show a better intersegmental agreement [3, 49]. The fast SENC technique seems highly reproducible, even across different sites [50, 51]. Clinical application of this technique has shown clinical merit; however, more research is needed regarding segmental values [52]. In general, all strain values derived from commercially available software are potentially limited in their comparability as new versions provide new values; hence, providing the software version applied is of great importance.
Lastly, we want to comment on the effect of contrast media application on FT-derived strain values. On the one hand, we noticed that the segmentation becomes more challenging for the AI algorithm; on the other hand, post-CM strain values are lower. This is in line with previous literature [53]. As SAX acquisition are now most of the time acquired after contrast media application, challenges and AI segmentation networks have to take this into consideration.
Limitations
This is a single-center study with a limited number of cases but reflects different disease entities. This limits potential statistical power in the detection of significant differences in the pairwise comparisons. Another limitation is that we did not compare cardiac contours along the cardiac cycle in order to depict the proper tracking of the FT algorithm. Furthermore, we want to point out that we did not use another vendor, which reduces generalizability.
Conclusions
Our study shows that application of AI-derived contours for feature tracking and strain analysis in CMR yields results comparable to manual segmentations. Attention should be taken while evaluating left ventricle hypertrophy cases especially in patients with aortic stenosis independent of the use of manual or AI-derived contours.
Abbreviations
- ACDC:
-
Automatic Cardiac Diagnosis Challenge
- AHA:
-
American Heart Association
- AHT:
-
Arterial hypertension
- AI:
-
Artificial intelligence
- AS:
-
Aortic stenosis
- CMI:
-
Chronic myocardial infarction
- CMI-EWF:
-
CMI with reduced LVEF, dilated LV, and global WMA
- CMI-F:
-
CMI with focal fibrosis
- CMI-WF:
-
CMI with focal fibrosis, reduced LVEF, and regional WMA
- CMR:
-
Cardiovascular magnetic resonance
- CNN:
-
Convolutional neural network
- CS:
-
Circumferential strain
- cv:
-
Chamber view
- DENSE:
-
Displacement encoding with stimulated echoes
- DSC:
-
Dice similarity coefficient
- ED:
-
End-diastole
- FT:
-
Feature tracking
- GCS:
-
Global circumferential strain
- GLS:
-
Global longitudinal strain
- GRS:
-
Global radial strain
- HCM:
-
Hypertrophic cardiomyopathy
- HD:
-
Hausdorff distance
- LAX:
-
Long axis
- LL:
-
“Lazy Luna” tool
- LS:
-
Longitudinal strain
- LV:
-
Left ventricle
- LVEF:
-
Left ventricular ejection fraction
- LVH:
-
Left ventricular hypertrophy
- LVOT:
-
Left ventricular outflow tract
- post-CM:
-
Post-contrast media application
- pre-CM:
-
Pre-contrast media application
- RS:
-
Radial strain
- SAX:
-
Short axis
- SD:
-
Standard deviation
- SENC:
-
Strain Encoding Magnetic Resonance imaging
- WMA:
-
Wall motion abnormalities
References
Korosoglou G, Giusca S, Hofmann NP et al (2019) Strain-encoded magnetic resonance: a method for the assessment of myocardial deformation. ESC Heart Fail 6:584–602. https://doi.org/10.1002/ehf2.12442
Kim D, Gilson WD, Kramer CM, Epstein FH (2004) Myocardial tissue tracking with two-dimensional cine displacement-encoded MR imaging: development and initial evaluation. Radiology 230:862–871. https://doi.org/10.1148/radiol.2303021213
Bucius P, Erley J, Tanacli R et al (2020) Comparison of feature tracking, fast-SENC, and myocardial tagging for global and segmental left ventricular strain. ESC Heart Fail 7:523–532. https://doi.org/10.1002/ehf2.12576
Lapinskas T, Zieschang V, Erley J et al (2019) Strain-encoded cardiac magnetic resonance imaging: a new approach for fast estimation of left ventricular function. BMC Cardiovasc Disord 19:52. https://doi.org/10.1186/s12872-019-1031-5
Liu B, Dardeer AM, Moody WE et al (2018) Reference ranges for three-dimensional feature tracking cardiac magnetic resonance: comparison with two-dimensional methodology and relevance of age and gender. Int J Cardiovasc Imaging 34:761–775. https://doi.org/10.1007/s10554-017-1277-x
Jolly M-P, Jordan JH, Meléndez GC et al (2017) Automated assessments of circumferential strain from cine CMR correlate with LVEF declines in cancer patients early after receipt of cardio-toxic chemotherapy. J Cardiovasc Magn Reson 19:59. https://doi.org/10.1186/s12968-017-0373-3
Barbosa MF, Fusco DR, Gaiolla RD et al (2021) Characterization of subclinical diastolic dysfunction by cardiac magnetic resonance feature-tracking in adult survivors of non-Hodgkin lymphoma treated with anthracyclines. BMC Cardiovasc Disord 21:170. https://doi.org/10.1186/s12872-021-01996-6
Eitel I, Stiermaier T, Lange T et al (2018) Cardiac magnetic resonance myocardial feature tracking for optimized prediction of cardiovascular events following myocardial infarction. JACC Cardiovasc Imaging 11:1433–1444. https://doi.org/10.1016/j.jcmg.2017.11.034
Lange T, Stiermaier T, Backhaus SJ et al (2021) Functional and prognostic implications of cardiac magnetic resonance feature tracking-derived remote myocardial strain analyses in patients following acute myocardial infarction. Clin Res Cardiol 110:270–280. https://doi.org/10.1007/s00392-020-01747-1
Wamil M, Borlotti A, Liu D et al (2019) Combined T1-mapping and tissue tracking analysis predicts severity of ischemic injury following acute STEMI—an Oxford Acute Myocardial Infarction (OxAMI) study. Int J Cardiovasc Imaging 35:1297–1308. https://doi.org/10.1007/s10554-019-01542-8
Nucifora G, Muser D, Gianfagna P et al (2015) Systolic and diastolic myocardial mechanics in hypertrophic cardiomyopathy and their link to the extent of hypertrophy, replacement fibrosis and interstitial fibrosis. Int J Cardiovasc Imaging 31:1603–1610. https://doi.org/10.1007/s10554-015-0720-0
Bogarapu S, Puchalski MD, Everitt MD et al (2016) Novel cardiac magnetic resonance feature tracking (CMR-FT) analysis for detection of myocardial fibrosis in pediatric hypertrophic cardiomyopathy. Pediatr Cardiol 37:663–673. https://doi.org/10.1007/s00246-015-1329-8
Baeßler B, Schaarschmidt F, Dick A et al (2016) Diagnostic implications of magnetic resonance feature tracking derived myocardial strain parameters in acute myocarditis. Eur J Radiol 85:218–227. https://doi.org/10.1016/j.ejrad.2015.11.023
Weigand J, Nielsen JC, Sengupta PP et al (2016) Feature tracking-derived peak systolic strain compared to late gadolinium enhancement in troponin-positive myocarditis: a case-control study. Pediatr Cardiol 37:696–703. https://doi.org/10.1007/s00246-015-1333-z
Lim C, Blaszczyk E, Riazy L et al (2020) Quantification of myocardial strain assessed by cardiovascular magnetic resonance feature tracking in healthy subjects-influence of segmentation and analysis software. Eur Radiol. https://doi.org/10.1007/s00330-020-07539-5
Backhaus SJ, Metschies G, Billing M et al (2019) Cardiovascular magnetic resonance imaging feature tracking: impact of training on observer performance and reproducibility. PLoS One 14:e0210127. https://doi.org/10.1371/journal.pone.0210127
Backhaus SJ, Aldehayat H, Kowallick JT et al (2022) Artificial intelligence fully automated myocardial strain quantification for risk stratification following acute myocardial infarction. Sci Rep 12:12220. https://doi.org/10.1038/s41598-022-16228-w
Ruijsink B, Puyol-Antón E, Oksuz I et al (2020) Fully automated, quality-controlled cardiac analysis from CMR: validation and large-scale application to characterize cardiac function. JACC Cardiovasc Imaging 13:684–695. https://doi.org/10.1016/j.jcmg.2019.05.030
von Knobelsdorff-Brenkenhoff F, Hennig P, Menza M et al (2016) Myocardial dysfunction in patients with aortic stenosis and hypertensive heart disease assessed by MR tissue phase mapping. J Magn Reson Imaging 44:168–177. https://doi.org/10.1002/jmri.25125
Prothmann M, von Knobelsdorff-Brenkenhoff F, Töpper A et al (2016) High spatial resolution cardiovascular magnetic resonance at 7.0 Tesla in patients with hypertrophic cardiomyopathy - first experiences: lesson learned from 7.0 Tesla. PLoS One 11:e0148066. https://doi.org/10.1371/journal.pone.0148066
Gröschel J, Ammann C, Zange L et al (2022) Fast acquisition of left and right ventricular function parameters applying cardiovascular magnetic resonance in clinical routine - validation of a 2-shot compressed sensing cine sequence. Scand Cardiovasc J 56:266–275. https://doi.org/10.1080/14017431.2022.2099010
Petersen SE, Matthews PM, Francis JM et al (2016) UK Biobank’s cardiovascular magnetic resonance protocol. J Cardiovasc Magn Reson 18:8. https://doi.org/10.1186/s12968-016-0227-4
Bistoquet A, Oshinski J, Skrinjar O (2007) Left ventricular deformation recovery from cine MRI using an incompressible model. IEEE Trans Med Imaging 26:1136–1153. https://doi.org/10.1109/TMI.2007.903693
Bistoquet A, Oshinski J, Skrinjar O (2008) Myocardial deformation recovery from cine MRI using a nearly incompressible biventricular model. Med Image Anal 12:69–85. https://doi.org/10.1016/j.media.2007.10.009
Selvadurai BSN, Puntmann VO, Bluemke DA et al (2018) Definition of left ventricular segments for cardiac magnetic resonance imaging. JACC Cardiovasc Imaging 11:926–928. https://doi.org/10.1016/j.jcmg.2017.09.010
Heinke R, Pathan F, Le M et al (2019) Towards standardized postprocessing of global longitudinal strain by feature tracking - OptiStrain CMR-FT study. BMC Cardiovasc Disord 19:267. https://doi.org/10.1186/s12872-019-1255-4
Hadler T, Wetzl J, Lange S et al (2022) Introduction of Lazy Luna an automatic software-driven multilevel comparison of ventricular function quantification in cardiovascular magnetic resonance imaging. Sci Rep 12:6629. https://doi.org/10.1038/s41598-022-10464-w
Petersen SE, Aung N, Sanghvi MM et al (2017) Reference ranges for cardiac structure and function using cardiovascular magnetic resonance (CMR) in Caucasians from the UK Biobank population cohort. J Cardiovasc Magn Reson 19:18. https://doi.org/10.1186/s12968-017-0327-9
Schulz-Menger J, Bluemke DA, Bremerich J et al (2020) Standardized image interpretation and post-processing in cardiovascular magnetic resonance - 2020 update. J Cardiovasc Magn Reson 22:19. https://doi.org/10.1186/s12968-020-00610-6
Bernard O, Lalande A, Zotti C et al (2018) Deep learning techniques for automatic MRI cardiac multi-structures segmentation and diagnosis: is the problem solved? IEEE Trans Med Imaging 37:2514–2525. https://doi.org/10.1109/TMI.2018.2837502
Andre F, Steen H, Matheis P et al (2015) Age- and gender-related normal left ventricular deformation assessed by cardiovascular magnetic resonance feature tracking. J Cardiovasc Magn Reson 17:25. https://doi.org/10.1186/s12968-015-0123-3
Kim MY, Park EA, Lee W, Lee SP (2020) Cardiac magnetic resonance feature tracking in aortic stenosis: exploration of strain parameters and prognostic value in asymptomatic patients with preserved ejection fraction. Korean J Radiol 21:268–279. https://doi.org/10.3348/kjr.2019.0441
Miskinyte E, Bucius P, Erley J et al (2019) Assessment of global longitudinal and circumferential strain using computed tomography feature tracking: intra-individual comparison with CMR feature tracking and myocardial tagging in patients with severe aortic stenosis. J Clin Med 8:E1423. https://doi.org/10.3390/jcm8091423
Al Musa T, Uddin A, Swoboda PP et al (2017) Myocardial strain and symptom severity in severe aortic stenosis: insights from cardiovascular magnetic resonance. Quant Imaging Med Surg 7:38–47. https://doi.org/10.21037/qims.2017.02.05
Spath NB, Gomez M, Everett RJ et al (2019) Global Longitudinal strain analysis using cardiac MRI in aortic stenosis: comparison with left ventricular remodeling, myocardial fibrosis, and 2-year clinical outcomes. Radiol Cardiothorac Imaging 1:e190027. https://doi.org/10.1148/ryct.2019190027
Neisius U, Myerson L, Fahmy AS et al (2019) Cardiovascular magnetic resonance feature tracking strain analysis for discrimination between hypertensive heart disease and hypertrophic cardiomyopathy. PLoS One 14:e0221061. https://doi.org/10.1371/journal.pone.0221061
Gastl M, Lachmann V, Christidi A et al (2021) Cardiac magnetic resonance T2 mapping and feature tracking in athlete’s heart and HCM. Eur Radiol 31:2768–2777. https://doi.org/10.1007/s00330-020-07289-4
Harrigan CJ, Appelbaum E, Maron BJ et al (2008) Significance of papillary muscle abnormalities identified by cardiovascular magnetic resonance in hypertrophic cardiomyopathy. Am J Cardiol 101:668–673. https://doi.org/10.1016/j.amjcard.2007.10.032
Roberts WC, Cohen LS (1972) Left ventricular papillary muscles. Description of the normal and a survey of conditions causing them to be abnormal. Circulation 46:138–154. https://doi.org/10.1161/01.cir.46.1.138
Pedrizzetti G, Claus P, Kilner PJ, Nagel E (2016) Principles of cardiovascular magnetic resonance feature tracking and echocardiographic speckle tracking for informed clinical use. J Cardiovasc Magn Reson 18:51. https://doi.org/10.1186/s12968-016-0269-7
Vogel-Claussen J, Finn JP, Gomes AS et al (2006) Left ventricular papillary muscle mass: relationship to left ventricular mass and volumes by magnetic resonance imaging. J Comput Assist Tomogr 30:426–432. https://doi.org/10.1097/00004728-200605000-00013
Wassmuth R, von Knobelsdorff-Brenkenhoff F, Gruettner H et al (2014) Cardiac magnetic resonance imaging of congenital bicuspid aortic valves and associated aortic pathologies in adults. Eur Heart J - Cardiovasc Imaging 15:673–679. https://doi.org/10.1093/ehjci/jet275
Rossi A, van der Linde D, Yap SC et al (2013) Ascending aorta dilatation in patients with bicuspid aortic valve stenosis: a prospective CMR study. Eur Radiol 23:642–649. https://doi.org/10.1007/s00330-012-2651-7
Morton G, Schuster A, Jogiya R et al (2012) Inter-study reproducibility of cardiovascular magnetic resonance myocardial feature tracking. J Cardiovasc Magn Reson 14:43. https://doi.org/10.1186/1532-429X-14-43
Schmidt B, Dick A, Treutlein M et al (2017) Intra- and inter-observer reproducibility of global and regional magnetic resonance feature tracking derived strain parameters of the left and right ventricle. Eur J Radiol 89:97–105. https://doi.org/10.1016/j.ejrad.2017.01.025
Auger DA, Ghadimi S, Cai X et al (2022) Reproducibility of global and segmental myocardial strain using cine DENSE at 3 T: a multicenter cardiovascular magnetic resonance study in healthy subjects and patients with heart disease. J Cardiovasc Magn Reson 24:23. https://doi.org/10.1186/s12968-022-00851-7
Lin K, Meng L, Collins JD et al (2017) Reproducibility of cine displacement encoding with stimulated echoes (DENSE) in human subjects. Magn Reson Imaging 35:148–153. https://doi.org/10.1016/j.mri.2016.08.009
Mangion K, Carrick D, Clerfond G et al (2019) Predictors of segmental myocardial functional recovery in patients after an acute ST-Elevation myocardial infarction. Eur J Radiol 112:121–129. https://doi.org/10.1016/j.ejrad.2019.01.010
Neizel M, Lossnitzer D, Korosoglou G et al (2009) Strain-encoded (SENC) magnetic resonance imaging to evaluate regional heterogeneity of myocardial strain in healthy volunteers: comparison with conventional tagging. J Magn Reson Imaging 29:99–105. https://doi.org/10.1002/jmri.21612
Giusca S, Korosoglou G, Zieschang V et al (2018) Reproducibility study on myocardial strain assessment using fast-SENC cardiac magnetic resonance imaging. Sci Rep 8:14100. https://doi.org/10.1038/s41598-018-32226-3
Erley J, Zieschang V, Lapinskas T et al (2020) A multi-vendor, multi-center study on reproducibility and comparability of fast strain-encoded cardiovascular magnetic resonance imaging. Int J Cardiovasc Imaging 36:899–911. https://doi.org/10.1007/s10554-020-01775-y
Korosoglou G, Giusca S, Montenbruck M et al (2021) Fast strain-encoded cardiac magnetic resonance for diagnostic classification and risk stratification of heart failure patients. JACC Cardiovasc Imaging 14:1177–1188. https://doi.org/10.1016/j.jcmg.2020.10.024
Kuetting DLR, Dabir D, Homsi R et al (2016) The effects of extracellular contrast agent (Gadobutrol) on the precision and reproducibility of cardiovascular magnetic resonance feature tracking. J Cardiovasc Magn Reson 18:30. https://doi.org/10.1186/s12968-016-0249-y
Acknowledgements
We gratefully thank our CMR technicians Kerstin Kretschel, Denise Kleindienst, and Martina Kohla for the technical assistance as well as our study nurses Annette Köhler and Elke Nickel-Szczech. The whole CMR Working Group was involved, and we are thankful for their input on individual levels. We thank circle CVI42 for providing the AI prototype. We also thank Carsten Schwenke for the input regarding the statistical evaluation. We thank Natalie Kaban for proofreading the manuscript.
Funding
Open Access funding enabled and organized by Projekt DEAL. This study has received funding in means of support of the work of Jan Gröschel by a grant of the European Union (18HLT05 QUIERO). Professor Dr. Schulz-Menger holds institutional grants of the Charité Medical University.
Author information
Authors and Affiliations
Contributions
JG participated on the study design, applied for ethic board approval, read the images, performed the statistical analyses, and drafted the manuscript with input from JK, DV, TH, SH, PB, JSM, and EB. JK assisted with the design of the study and statistical analysis and was involved in manuscript writing. DV assisted with the data analysis and statistical analysis and was involved in manuscript writing. TH assisted with developing the comparison tool and graphics and was involved in manuscript writing. SH assisted with the design of the study and statistical analysis and was involved in manuscript writing. PB assisted with developing the analysis software and the AI algorithm and was involved in manuscript writing. JSM conceived and designed the study, supported the statistical analysis, supervised the manuscript writing, and provided continuous guidance throughout the study as the head of the working group. EB conceived and designed the study, read the images, supported the statistical analysis, and supervised the manuscript writing. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Guarantor
The scientific guarantor of this publication is Edyta Blaszczyk.
Conflict of interest
Phillip Barckow is an employee of Circle Cardiovascular Imaging Inc. The authors of this manuscript declare no other relationships with any companies, whose products or services may be related to the subject matter of the article.
Statistics and biometry
Carsten Schwenke kindly provided statistical advice for this manuscript.
Informed consent
Written informed consent was obtained from all subjects (patients) in the previous studies in which the patients were recruited.
Ethical approval
Institutional Review Board approval was obtained.
Study subjects or cohorts overlap
Some study subjects or cohorts have been previously reported in:
• Lim C, Blaszczyk E, Riazy L, et al (2020) Quantification of myocardial strain assessed by cardiovascular magnetic resonance feature tracking in healthy subjects-influence of segmentation and analysis software. Eur Radiol. https://doi.org/10.1007/s00330-020-07539-5
• von Knobelsdorff-Brenkenhoff F, Hennig P, Menza M, et al (2016) Myocardial dysfunction in patients with aortic stenosis and hypertensive heart disease assessed by MR tissue phase mapping. J Magn Reson Imaging 44:168–177. https://doi.org/10.1002/jmri.25125
• Prothmann M, von Knobelsdorff-Brenkenhoff F, Töpper A, et al (2016) High Spatial Resolution Cardiovascular Magnetic Resonance at 7.0 Tesla in Patients with Hypertrophic Cardiomyopathy—First Experiences: Lesson Learned from 7.0 Tesla. PLoS One 11:e0148066. https://doi.org/10.1371/journal.pone.0148066
• Gröschel J, Ammann C, Zange L, et al (2022) Fast acquisition of left and right ventricular function parameters applying cardiovascular magnetic resonance in clinical routine—validation of a 2-shot compressed sensing cine sequence. Scand Cardiovasc J 56:266–275. https://doi.org/10.1080/14017431.2022.2099010
Methodology
• Retrospective.
• cross-sectional study.
• multicenter study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gröschel, J., Kuhnt, J., Viezzer, D. et al. Comparison of manual and artificial intelligence based quantification of myocardial strain by feature tracking—a cardiovascular MR study in health and disease. Eur Radiol 34, 1003–1015 (2024). https://doi.org/10.1007/s00330-023-10127-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-023-10127-y