Abstract
Objectives
To use convolutional neural network for fully automated segmentation and radiomics features extraction of hypopharyngeal cancer (HPC) tumor in MRI.
Methods
MR images were collected from 222 HPC patients, among them 178 patients were used for training, and another 44 patients were recruited for testing. U-Net and DeepLab V3 + architectures were used for training the models. The model performance was evaluated using the dice similarity coefficient (DSC), Jaccard index, and average surface distance. The reliability of radiomics parameters of the tumor extracted by the models was assessed using intraclass correlation coefficient (ICC).
Results
The predicted tumor volumes by DeepLab V3 + model and U-Net model were highly correlated with those delineated manually (p < 0.001). The DSC of DeepLab V3 + model was significantly higher than that of U-Net model (0.77 vs 0.75, p < 0.05), particularly in those small tumor volumes of < 10 cm3 (0.74 vs 0.70, p < 0.001). For radiomics extraction of the first-order features, both models exhibited high agreement (ICC: 0.71–0.91) with manual delineation. The radiomics extracted by DeepLab V3 + model had significantly higher ICCs than those extracted by U-Net model for 7 of 19 first-order features and for 8 of 17 shape-based features (p < 0.05).
Conclusion
Both DeepLab V3 + and U-Net models produced reasonable results in automated segmentation and radiomic features extraction of HPC on MR images, whereas DeepLab V3 + had a better performance than U-Net.
Clinical relevance statement
The deep learning model, DeepLab V3 + , exhibited promising performance in automated tumor segmentation and radiomics extraction for hypopharyngeal cancer on MRI. This approach holds great potential for enhancing the radiotherapy workflow and facilitating prediction of treatment outcomes.
Key Points
• DeepLab V3 + and U-Net models produced reasonable results in automated segmentation and radiomic features extraction of HPC on MR images.
• DeepLab V3 + model was more accurate than U-Net in automated segmentation, especially on small tumors.
• DeepLab V3 + exhibited higher agreement for about half of the first-order and shape-based radiomics features than U-Net.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hypopharyngeal cancer (HPC) has the worst prognosis among all head and neck squamous cell cancers (HNSCC). The incidence and mortality of HPC continues to increase over the past decades [1]. Most patients with HPC have advanced cancer staging at presentation, and chemoradiation is currently considered as the mainstay of organ-preservation therapy. Previous studies have shown that magnetic resonance imaging (MRI) provides anatomic, qualitative, and quantitative functional information in clinical practice for staging, treatment planning, and response assessment of HNSCC [2,3,4,5].
Determining tumor contour is essential for radiation therapy planning, and MRI is widely applied for pretreatment delineation of tumor extension [6]. In addition, tumor contouring plays an essential role for radiomics analysis. Magnetic resonance (MR) radiomics can serve as a biomarker in the prediction of treatment outcomes [7], and survival in HNSCC [8]. However, manual contouring of tumors in a slice-by-slice fashion is labor intensive and prone to interobserver variations [9]. An effective tumor auto-segmentation tool is expected to improve the efficiency of the radiotherapy workflow.
In recent years, convolutional neural networks (CNNs) have been extensively used for semantic segmentation on medical image [10, 11]; however, its applications on HNSCC have been studied on computed tomography (CT) or PET/CT [12, 13], with the dice similarity coefficient (DSC) being 0.66 on CT and 0.74 on PET/CT, respectively. A few studies have focused on MRI; however, the accuracies of segmentation were rather low with the DSC ranging between 0.49 and 0.65 [14, 15].
U-Net, a CNN-based method for biomedical image segmentation, has been widely used recently [16,17,18]. However, one of the challenges of U-Net is the excessive downsizing by consecutive pooling layers, which makes it sensitive to the size of mask for training. To this end, DeepLab V3 + [19] architecture was introduced which utilized the atrous spatial pyramidal pooling (ASPP) to obtain multi-scale context information to improve the limit of consecutive pooling layers. DeepLab V3 + has been applied in many segmentation tasks, where the performances were reported to be superior to many state-of-the-art networks [20, 21].
The aim of this study was to evaluate the performance of DeepLab V3 + for automated segmentation and radiomics extraction of HPC primary tumors in MRI and compare with that of U-Net.
Methods and materials
Patients
This study retrospectively analyzed the dataset of patients with HPC stage T3 and T4 during 2006 to 2017. The Institutional Review Board approved the study, and informed consent was waved. The inclusion criteria were (a) newly biopsy-proven stage T3 and T4 (American Joint Committee on Cancer, AJCC) hypopharyngeal squamous cell carcinoma and (b) having complete our uniform chemoradiation scheme. The exclusion criteria were inaccessibility for follow-up, contraindications to MRI, history of previous head or neck cancers, second malignancies, renal failure, and mental health–related inability to cooperate.
We included 242 consecutive patients, of whom 12 were excluded because they had indistinct margins and 8 were excluded because their images had considerable artifacts. Accordingly, 222 patients were included in the final study sample, and their MR images were used as the dataset for analysis. We separated the dataset obtained from our 178 patients (80.2%) as a training dataset and those from the remaining 44 patients (19.8%) as a testing dataset. The training dataset was further divided into training and validation subsets by using fivefold cross-validation (executed using 35 or 36 images per fold). This step was implemented to ensure that the models’ performance would not be affected by data splitting, thus preventing overfitting.
Treatment
All patients received intensity-modulated radiotherapy using 6-MV photon beams. The initial prophylactic field included gross tumor with at least 1-cm margins and neck lymphatics at risk for 46–56 Gy, then cone-down boost to the gross tumor area up to 72 Gy. Concurrent chemotherapy consisted of intravenous cisplatin, oral tegafur plus oral leucovorin.
MRI data and image annotation
MRI data were acquired using three MRI scanners (Siemens, GE, and Philips). All patients underwent the standard MRI protocol in our institute. T1-weighted turbo spin echo (TSE) images (TR/TE: 400–760/11–16 ms) and T2-weighted fat-saturated (T2w-fs) TSE images (TR/TE: 4100–5300/80–92 ms) were acquired in axial and coronary planes. After an intravenous administration of gadolinium at a dose of 0.1 mmol/kg, a fat-saturated contrast-enhanced T1-weighted (cT1w-fs) TSE sequence (TR/TE: 400–680 ms) was obtained in the axial, sagittal, and coronal planes. The cT1w-fs images obtained in the axial plane were used for model training and testing, and the parameters were outlined as follows: field of view, 220 mm; slice thickness, 4 mm; acquisition matrix, 256 × 256 or 384 × 320; reconstruction matrix, 512 × 512; number of slices, 30–36; and scan time, 200–280 s.
Regions of interest (ROIs) capturing tumor contours throughout the primary tumor volume were segmented slice by slide on cT1w-fs MR images by two neuroradiologists (S.H.N., with 27 years of experience in head and neck imaging; C.H.Y., with 12 years of experience in head and neck imaging) via consensus, with the aid of T2-weighted MR images, using an in-house interface developed through MATLAB (Mathworks). Both neuroradiologists were blinded to the clinical outcomes. The labeled ROIs were used as the ground truth for the model training.
Image preprocessing
Data augmentation was performed on each training image set such that the training model can be robust against the degree of enlargement, rotation, and parallel shift. Each image was randomly rotated within the range −10 to 10°. The range of the random shift was 0.9–1.1 of the image length and width. The image was zoomed by the range of 0.9–1.1.
Network and training
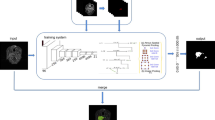
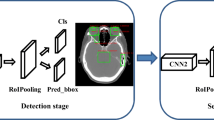
Two types of network architectures were used for training and testing the performances of tumor segmentation: the U-Net as previous studies [18, 22], and DeepLab V3 + (Fig. 1), a decoder module with a depth separable convolution for ASPP layers to improve the object boundary detection [23]. The networks were trained with weight randomization and stochastic gradient descent Adam optimizer method [24]. The signal intensities of all images were normalized to a mean = 0 and standard deviation = 1 [25]. The learning rate was set to 0.001, and the number of epochs until convergence was 100 with batch sizes of 2. The network was trained using Keras 2.1.4 written in Python 3.5.4 and TensorFlow 1.5.0. The code for the DeepLab V3 + model is available at https://github.com/bonlime/ keras-deeplab-v3-plus.
Performance evaluation
Each model’s segmentation performance was assessed using the following metrics [26]: (1) the DSC, a measure of spatial overlap calculated using the formula 2TP/(FP + 2TP + FN), where TP, FP, and FN represent true-positive, false-positive, and false-negative detections, respectively; (2) Jaccard index, calculated using the formula TP/(TP + FP + FN); and (3) average surface distance (ASD), calculated as the mean Euclidean distance between surface voxels in the segmented object and ground truth.
Radiomics
To assess the reliability of predicted ROIs by the established models, we examined the radiomics features of tumor ROIs extracted by manual and automatic segmentation models. The radiomics features of tumors were calculated using Pyradiomics software [27] based on the 3D volumes of ROIs on the images (https://pyradiomics.readthedocs.io/en/latest/features.html). A total of 105 radiomics were extracted, which were divided into the following classes: first-order statistics (19 features), shape-based (17 features), gray-level co-occurrence matrix (GLCM, 24 features), gray-level run length matrix (GLRLM, 14 features), gray-level size zone matrix (GLSZM, 13 features), neighboring gray-tone difference matrix (NGTDM, 4 features), and gray-level dependence matrix (GLDM, 14 features).
Statistics
Statistical analysis was performed using GraphPad Prism software version 8.0 for Mac. The two models’ performance metrics were compared using the Wilcoxon rank test. The differences in DSCs among the MRI scanners were assessed using the Kruskal–Wallis test. Bland–Altman plots were used to assess the agreement between the tumor volumes obtained from the automated models and those obtained by manual segmentation. Pearson’s correlation analysis was used to determine the correlations between the tumor volumes obtained from the automated models and those obtained by manual segmentation. The reliability of radiomics features of tumor ROIs was evaluated using intraclass correlation coefficient (ICC) obtained by manual and automatic segmentation models. The differences in DSCs with regard to tumor size and tumor histology were assessed using the Mann–Whitney U test.
Results
Patient characteristics
Table 1 presents the clinical and demographic features of the training (n = 178) and testing datasets (n = 44). The median patient age was 53 years (range: 33–83 years). No statistically significant differences were found regarding the clinical or demographic characteristics between the training and testing datasets.
Model performance
Table 2 presents a summary of the DeepLab V3 + and U-Net models’ performance. DeepLab V3 + significantly outperformed U-Net in all metrics. DeepLab V3 + had a higher DSC (p < 0.05) and Jaccard index (p < 0.001) but a lower ASD (p < 0.001) than did U-Net. The models’ performance levels were similar across the images obtained from the 3 different MRI scanners employed in this study (p = 0.78 and 0.83 for U-Net and DeepLab V3 + models, respectively) (Supplementary Table 1). Representative figures (Fig. 2) show delineation of primary tumors in a large HPC and a small HPC using U-Net and DeepLab V3 + models respectively. After analyzing the 421 image slices in the testing dataset, we found that U-Net yielded 41 (9.7%) FPs, which occurred most frequently in the adjacent lymph nodes (n = 30), followed by the submandibular gland (n = 14) and the tongue base (n = 1). DeepLab V3 + yielded only 2 (0.5%) FPs in the submandibular gland.
Examples of tumor segmentation in two patients with a large hypopharyngeal tumor (upper row) and a small hypopharyngeal tumor (lower row) on cT1w-fs images. The tumors are delineated in red for manual segmentation, in blue by U-Net model, and in yellow by DeepLab V3 + . The numbers indicate the DSC scores. The white arrows indicate the false positives in the predicted segmentation. The large white arrow points to a large metastatic lymph node, while the small white arrow points to left submandibular gland
Impact of tumor volume to prediction accuracy
Figure 3 shows that the predicted tumor volumes by U-Net and DeepLab V3 + models were highly correlated with those delineated manually (r = 0.93 and 0.96 for U-Net and DeepLab V3 + models, respectively, p < 0.001 for both). The Bland–Altman plots demonstrated that the U-Net model had a higher mean difference bias compared to the DeepLab V3 + model (bias:16.6% vs 7.69%; 95% limit of agreement: −30 to 72.6% vs −40.4 to 50.7% for U-Net and DeepLab V3 + models, respectively).
A, B Correlations of tumor volumes between the manual and predicted tumor volumes using DeepLab and U-Net models. C, D Blade-Altman plots of the tumor volumes between the manual and the model-derived volumes. The percentage difference was calculated by the difference of model-derived volume minus the mean of the two measurements. The blue dot line indicated the bias of difference. The gray shade area indicated the 95% limits of agreement
Figure 4 plots the relations of DSC of U-Net and DeepLab V3 + models to the tumor volume of ground truth. The U-Net model showed lower DSC than DeepLab V3 + in small tumors. Through a stepwise approach, we subdivided our dataset into subgroups by using various thresholds and then compared the performance of these thresholds in producing differences in DSCs between tumors. We observed that a threshold of 10 cm3 produced the most pronounced differences in DSCs between large and small tumors. We conducted a subgroup analysis by dividing the test dataset into two subgroups according to tumor volume: small (volume < 10 cm3; n = 21) and large (volume > 10 cm3; n = 23) groups. In the small group, DeepLab V3 + had significantly a higher DSC (0.74, 95% CI: 0.72–0.76) than did U-Net (0.70, 95% CI: 0.68–0.72; p < 0.001); however, in the large group, the models did not differ significantly in DSCs (DeepLab V3 + : 0.8, 95% CI: 0.78–0.82; U-Net: 0.79, 95% CI: 0.76–0.82; p = 0.72; Fig. 4C).
Reproducibility of radiomics features
Figure 5 shows the ICC values of radiomics features extracted by manual and predicted ROIs using U-Net and DeepLab V3 + models. For the first-order features, both models exhibited high agreement (ICC range: 0.71–0.89 for U-Net model and 0.71–0.91 for DeepLab V3 + model). The DeepLab V3 + model had significantly higher ICCs for the following 7 of the 19 features: mean, median, 10 percentile, 90 percentile, skewness, total energy, uniformity (p < 0.05). For the shape-based features, both models exhibited moderate to good agreement (ICC range: 0.63–0.81 for U-Net model and 0.63–0.85 for DeepLab V3 + model). The DeepLab V3 + model had significantly higher ICCs for the following 8 of the 17 features: elongation, major axis length, mesh volume, spherical disproportion, sphericity, surface area, surface volume ratio, voxel volume (p < 0.05). The ICCs of the other radiomics features were moderate or low for both models (ICC range, 0.41 to 0.71 vs 0.43 to 0.73 for U-Net vs DeepLab V3 + model, respectively.)
Discussion
The use of deep learning for assessing head and neck cancer is an evolving field with considerable scientific and clinical potential. However, only a few studies have investigated the use of deep learning models for specific types of head and neck cancer on MR images. Our study used CNN-based models, namely DeepLab V3 + and U-Net, for automatic segmentation and radiomic features extraction of HPC on MR images. DeepLab V3 + exhibited superior segmentation performance to U-Net and other CNN models [14, 15], providing a promising model for improving the accuracy of head and neck cancer auto-segmentation using AI networks. We also demonstrated the generalizability of radiomic features derived by the models, which can benefit future research. We recommend the use of modern MRI scanners in facilities that handle high volumes of hypopharyngeal cancer patients. These scanners offer superior soft tissue contrast, as well as strong performance potential to support DL models in automated tumor segmentation and radiomics feature extraction.
We used a cT1w-fs sequence for model training because it clearly depicts tumor boundaries and is preferred for tumor extent identification in clinical practice. A T2w-fs sequence is also valuable for lesion extent evaluation; hence, we draw ROI on cT1w-fs images with the aid of T2-fs images in this study. Wong et al reported that U-Net exhibited similar performance levels in segmenting nasopharyngeal carcinoma tumors on T2w-fs and cT1-fs images; accordingly, T2w-fs images may be suitable for automatic tumor segmentation in clinical scenarios that require avoiding contrast agents [28].
Manually tumor delineation in radiotherapy planning is time-consuming and variable based on the level of expertise. While auto-segmentation algorithms may potentially save effort and time of a radiotherapist, the challenges in accurate tumor definition remain. Bielak et al [14] investigated the HNSCCs using a 3D DeepMedic CNN architecture with 7 MRI channel input and scored the average DSC of 0.65. Schouten et al [15] performed multi-view CNN for various HNSCCs, and presented an average DSC of 0.49 for all cases and 0.57 for HPC. In the current study, we specifically trained on HPC patients with U-Net and DeepLab V3 + models, and achieved DSCs of 0.72 and 0.77, respectively. The results suggest that training a dedicated CNN model with U-Net or DeepLab V3 + for HPC would attain better prediction of the target tumor contour, and may offer the potential improvement of artificial intelligence (AI)–based auto-segmentation.
The accuracies of segmentation of CNN have been reported to be positively related with tumor size [14, 15, 18]. Schouten et al [15] reported that multi-view CNN produced improved segmentation results with larger tumor volumes, and tended to overestimate the tumor volume with FPs around the primary tumor or in adjacent enlarged lymph nodes. In our study, U-Net had significantly lower DSCs than DeepLab V3 + , particularly for small-size tumors. A reason could be attributed that U-Net utilized the general fully convolutional network, which is sensitive to the mask size of input data due to excessive downsizing by consecutive pooling layers [29], while DeepLab network with ASPP can capture context information at multiple spatial scales and produce accurate segmentation on input images with different size. Our Bland–Altman plots revealed that U-Net had a higher mean difference bias than did DeepLab V3 + (16.6% vs 7.69%), probably because U-Net had more FPs in the adjacent lymph nodes and submandibular gland. In radiotherapy, FP findings can lead to erroneous treatment plans with undesirable toxicity. The highest tolerable error for tumor segmentation in radiotherapy may vary depending on the patient’s medical status, tumor type, location, and prescribed radiation dose, and should be kept as low as possible. Although U-net has been showing potential in segmenting medial images, radiotherapy physicians should recognize that it may yield TPs near the primary lesions, as shown in our cases and as reported by studies applying it to segmentation of the knee meniscus [30] and lung nodules [31].
Our results demonstrated that the radiomics parameters of HPC extracted by both U-Net and DeepLab V3 + models were robust for the first-order and the shape-based features. This finding suggests that such CNN-based automated segmentation approach could improve the high-throughput extraction of radiomics features in a timely manner. Xue et al [32] showed that the first-order radiomics features had significantly higher ICCs than the other texture features. Our study proved that the first-order features remained reliable using the CNN-based approach with U-Net or DeepLab V3 + model, in line with high reproducibility of first-order radiomics features of diffusion-weighted imaging for cervical cancer using U-Net [18]. In addition, our results also demonstrated that DeepLab V3 + exhibited shape-based features with higher reproducibility than U-Net. Since the shape features depend solely on the segmentation masks related to the accuracy of tumor contours, this finding suggests that DeepLab V3 + would be a feasible extraction method for MRI radiomics features that would potentially help reliable outcome prediction of HPC specifically.
Despite the abovementioned strengths, our study had some limitations. First, this study demonstrated that DeepLab V3 + can accurately segment HPC on MR images. However, further research is necessary to define the clinical acceptability of the model and estimate how much time it could save in the radiotherapy workflow. Second, our study showed that DeepLab V3 + extracted MRI radiomics features of HPC with good reproducibility, but to what extent such extracted feature aid reliable outcome prediction would be the subject of further investigation. Third, the findings of a single-center study may not be generalizable to other hospitals. Although our sample size of 220 patients was relatively small, a total of 2085 tumor slices were used. Nonetheless, larger multicenter studies are warranted for external validation and prospective testing. Our results suggest that the trained models exhibited similar performance levels when tested on images from three different MRI scanners. Therefore, the models may be applicable to images from MRI scanners with similar imaging parameters in other hospitals.
In conclusion, our study revealed that both DeepLab V3 + and U-Net models produced reasonable results in automated segmentation and radiomic features extraction of HPC on MR images, whereas DeepLab V3 + had a better performance than U-Net.
Abbreviations
- ASPP:
-
Atrous spatial pyramid pooling
- CNN:
-
Convolutional neural network
- CT:
-
Computed tomography
- cT1w-fs:
-
Fat-saturated contrast-enhanced T1-weighted
- DSC:
-
Dice similarity coefficient
- HNSCC:
-
Head and neck squamous cell cancers
- HPC:
-
Hypopharyngeal cancer
- ICC:
-
Intraclass correlation coefficients
- MRI:
-
Magnetic resonance imaging
- PET:
-
Positron emission tomography
- ROI:
-
Region of interest
- T2w-fs:
-
Fat-saturated T2 weighted
References
Lin Z, Lin H, Lin C (2020) Dynamic prediction of cancer-specific survival for primary hypopharyngeal squamous cell carcinoma. Int J Clin Oncol 25:1260–1269
Wong CK, Chan SC, Ng SH et al (2019) Textural features on 18F-FDG PET/CT and dynamic contrast-enhanced MR imaging for predicting treatment response and survival of patients with hypopharyngeal carcinoma. Medicine (Baltimore) 98:e16608
Ng SH, Liao CT, Lin CY et al (2016) Dynamic contrast-enhanced MRI, diffusion-weighted MRI and 18F-FDG PET/CT for the prediction of survival in oropharyngeal or hypopharyngeal squamous cell carcinoma treated with chemoradiation. Eur Radiol 26:4162–4172
Alfieri S, Romano R, Bologna M et al (2021) Prognostic role of pre-treatment magnetic resonance imaging (MRI)-based radiomic analysis in effectively cured head and neck squamous cell carcinoma (HNSCC) patients. Acta Oncol 60:1192–1200
Becker M, Monnier Y, de Vito C (2022) MR imaging of laryngeal and hypopharyngeal cancer. Magn Reson Imaging Clin N Am 30:53–72
Ma DJ, Zhu JM, Grigsby PW (2011) Tumor volume discrepancies between FDG-PET and MRI for cervical cancer. Radiother Oncol 98:139–142
Chen J, Lu S, Mao Y et al (2022) An MRI-based radiomics-clinical nomogram for the overall survival prediction in patients with hypopharyngeal squamous cell carcinoma: a multi-cohort study. Eur Radiol 32:1548–1557
Mes SW, van Velden FHP, Peltenburg B et al (2020) Outcome prediction of head and neck squamous cell carcinoma by MRI radiomic signatures. Eur Radiol 30:6311–6321
Dimopoulos JC, De Vos V, Berger D et al (2009) Inter-observer comparison of target delineation for MRI-assisted cervical cancer brachytherapy: application of the GYN GEC-ESTRO recommendations. Radiother Oncol 91:166–172
Perkuhn M, Stavrinou P, Thiele F et al (2018) Clinical evaluation of a multiparametric deep learning model for glioblastoma segmentation using heterogeneous magnetic resonance imaging data from clinical routine. Invest Radiol 53:647–654
Tian Z, Liu L, Zhang Z, Fei B (2018) PSNet: prostate segmentation on MRI based on a convolutional neural network. J Med Imaging (Bellingham) 5:021208
Zhang Z, Zhao T, Gay H, Zhang W, Sun B (2021) Weaving attention U-net: a novel hybrid CNN and attention-based method for organs-at-risk segmentation in head and neck CT images. Med Phys 48:7052–7062
Groendahl AR, Skjei Knudtsen I, Huynh BN et al (2021) A comparison of methods for fully automatic segmentation of tumors and involved nodes in PET/CT of head and neck cancers. Phys Med Biol 66:065012
Bielak L, Wiedenmann N, Berlin A et al (2020) Convolutional neural networks for head and neck tumor segmentation on 7-channel multiparametric MRI: a leave-one-out analysis. Radiat Oncol 15:181
Schouten JPE, Noteboom S, Martens RM et al (2022) Automatic segmentation of head and neck primary tumors on MRI using a multi-view CNN. Cancer Imaging 22:8
Ronneberger O, Fischer P, Brox T (2015) U-Net: convolutional networks for biomedical image segmentation. Springer International Publishing, Cham, pp 234–241
Yin XX, Sun L, Fu Y, Lu R, Zhang Y (2022) U-Net-based medical image segmentation. J Healthc Eng 2022:4189781
Lin YC, Lin CH, Lu HY et al (2020) Deep learning for fully automated tumor segmentation and extraction of magnetic resonance radiomics features in cervical cancer. Eur Radiol 30:1297–1305
Chen L-C, Zhu Y, Papandreou G, Schroff F, Adam H (2018) Encoder-decoder with atrous separable convolution for semantic image segmentation. https://doi.org/10.48550/arXiv.1802.02611
EL-Bana S, Al-Kabbany A, Sharkas M (2020) A two-stage framework for automated malignant pulmonary nodule detection in CT scans. Diagnostics 10:131
Wang J, Liu X (2021) Medical image recognition and segmentation of pathological slices of gastric cancer based on Deeplab v3+ neural network. Comput Methods Programs Biomed 207:106210
Liu X, Han C, Wang H et al (2021) Fully automated pelvic bone segmentation in multiparameteric MRI using a 3D convolutional neural network. Insights Imaging 12:93
Zhang X, Jiang L, Yang D, Yan J, Lu X (2019) Urine sediment recognition method based on multi-view deep residual learning in microscopic image. J Med Syst 43:325
Arcos-Garcia A, Alvarez-Garcia JA, Soria-Morillo LM (2018) Deep neural network for traffic sign recognition systems: an analysis of spatial transformers and stochastic optimisation methods. Neural Netw 99:158–165
Trebeschi S, van Griethuysen JJM, Lambregts DMJ et al (2017) Deep learning for fully-automated localization and segmentation of rectal cancer on multiparametric MR. Sci Rep 7:5301
Nai YH, Teo BW, Tan NL et al (2021) Comparison of metrics for the evaluation of medical segmentations using prostate MRI dataset. Comput Biol Med 134:104497
van Griethuysen JJM, Fedorov A, Parmar C et al (2017) Computational radiomics system to decode the radiographic phenotype. Cancer Res 77:e104–e107
Wong LM, Ai QYH, Mo FKF, Poon DMC, King AD (2021) Convolutional neural network in nasopharyngeal carcinoma: how good is automatic delineation for primary tumor on a non-contrast-enhanced fat-suppressed T2-weighted MRI? Jpn J Radiol 39:571–579
Hashemi-Beni L, Gebrehiwot A, Karimoddini A, Shahbazi A, Dorbu F (2022) Deep convolutional neural networks for weeds and crops discrimination from UAS imagery. Front Remote Sens 3:755939
Long Z, Zhang D, Guo H, Wang W (2021) Automated segmentation of knee menisci from magnetic resonance images by using ATTU-Net: a pilot study on small datasets. OSA Continuum 4:3096–3107
Suzuki K, Otsuka Y, Nomura Y, Kumamaru KK, Kuwatsuru R, Aoki S (2022) Development and validation of a modified three-dimensional U-Net deep-learning model for automated detection of lung nodules on chest CT images from the Lung Image Database Consortium and Japanese datasets. Acad Radiol 29(Suppl 2):S11–S17
Xue C, Yuan J, Zhou Y, Wong OL, Cheung KY, Yu SK (2022) Acquisition repeatability of MRI radiomics features in the head and neck: a dual-3D-sequence multi-scan study. Vis Comput Ind Biomed Art 5:10
Funding
This study was funded by the Taiwanese Ministry of Science and Technology (grant numbers: MOST 109–2314-B-182A-045, MOST 111–2314-B-182A-041, and MOST 110–2314-B-182A-090 (Taiwan)).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Shu-Hang Ng.
Conflict of interest
The authors declare no competing interests.
Statistics and biometry
No complex statistical methods were necessary for this paper.
Informed consent
Written informed consent was waived by the Institutional Review Board.
Ethical approval
Institutional Review Board approval was obtained.
Study subjects or cohorts overlap
None of the study subjects or cohorts has been previously reported.
Methodology
• retrospective
• diagnostic or prognostic study
• performed at one institution
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lin, YC., Lin, G., Pandey, S. et al. Fully automated segmentation and radiomics feature extraction of hypopharyngeal cancer on MRI using deep learning. Eur Radiol 33, 6548–6556 (2023). https://doi.org/10.1007/s00330-023-09827-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-023-09827-2