Abstract
Objectives
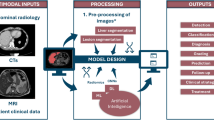
Machine learning (ML) for medical imaging is emerging for several organs and image modalities. Our objectives were to provide clinicians with an overview of this field by answering the following questions: (1) How is ML applied in liver computed tomography (CT) imaging? (2) How well do ML systems perform in liver CT imaging? (3) What are the clinical applications of ML in liver CT imaging?
Methods
A systematic review was carried out according to the guidelines from the PRISMA-P statement. The search string focused on studies containing content relating to artificial intelligence, liver, and computed tomography.
Results
One hundred ninety-one studies were included in the study. ML was applied to CT liver imaging by image analysis without clinicians’ intervention in majority of studies while in newer studies the fusion of ML method with clinical intervention have been identified. Several were documented to perform very accurately on reliable but small data. Most models identified were deep learning-based, mainly using convolutional neural networks. Potentially many clinical applications of ML to CT liver imaging have been identified through our review including liver and its lesion segmentation and classification, segmentation of vascular structure inside the liver, fibrosis and cirrhosis staging, metastasis prediction, and evaluation of chemotherapy.
Conclusion
Several studies attempted to provide transparent result of the model. To make the model convenient for a clinical application, prospective clinical validation studies are in urgent call. Computer scientists and engineers should seek to cooperate with health professionals to ensure this.
Key Points
• ML shows great potential for CT liver image tasks such as pixel-wise segmentation and classification of liver and liver lesions, fibrosis staging, metastasis prediction, and retrieval of relevant liver lesions from similar cases of other patients.
• Despite presenting the result is not standardized, many studies have attempted to provide transparent results to interpret the machine learning method performance in the literature.
• Prospective studies are in urgent call for clinical validation of ML method, preferably carried out by cooperation between clinicians and computer scientists.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
For several tasks related to medical imaging, ML is emerging as a new reliable tool due to its high performance and a superior capacity to build complex models for making predictions [1]. More than 220 medical devices using ML have been approved in the USA and Europe [2]. This development has increased steadily since 2014. Today, ML software can be considered a medical device [3].
Computer tomography (CT) imaging plays an essential role in diagnostics and post-treatment follow-up in liver diseases [4]. Applying ML-based tools to CT images has shown promising results [5]. It has been tested theoretically for tasks including identification and segmentation of the liver, lesions, blood vessels, and bile ducts in the liver [6], quantification of liver tissue characteristics [7], evaluation of cancer treatment, and prediction of liver disease [8, 9].
A recently published systematic review and meta-analysis demonstrated the diagnostic accuracy of deep learning (DL) in ophthalmology, respiratory medicine, and breast surgery [10]. In addition, a limited literature review has been published in the subfield of ML applied to liver imaging [11,12,13]. However, the performance and clinical applicability of ML in liver imaging are not comprehensively addressed in the literature.
A search in PROSPERO—a database of prospectively registered systematic reviews in health and social care [14]—did not reveal any forthcoming publication in this rapidly developing field. We, therefore, conducted a systematic review from a clinical perspective.
This review aims to answer the following questions: (1) How is ML applied in CT liver imaging? (2) How well do ML systems perform in liver CT imaging? (3) What are the clinical applications of ML in liver CT imaging?
Some important part of this article is given in the electronic supplementary material due to length of the article.
Methods
This systematic review was conducted in accordance with the guidelines for the “Preferred Reporting Items for Systematic Reviews and Meta-Analyses” extension for diagnostic accuracy studies statement [15]. A selection and retrieval of studies from the literature was done in accordance with Cochrane handbook for systematic review [16]. A search was conducted in Medline, EMBASE, and Web of Science and included studies published between January 1, 2011, and October 31, 2021. The search string consisted of exploded MeSH-terms, Emtree-terms, and free text to find all studies containing the terms “Artificial intelligence” AND “Computed tomography” AND “liver” (or containing all possible synonyms of all three) in the title, abstract, or keywords. The exact search string was given in the electronic supplementary material.
When considering study quality, we identified characteristics as important given in the electronic supplementary material. The suggested list is comprehensive, and studies might be quite informative with minimal risk of bias, without meeting all requirements [17]. Yet, if a study followed only few of the characteristics, it was not considered well-documented for clinical use.
Results
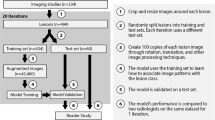
The search was conducted in two phases, one in October 2020 and one in October 2021. There were 191 studies included for review. The selection process is illustrated in the PRISMA flow diagram in Fig. 1 [18]. The selected studies are summarized in Table 1 and details given in the electronic supplementary material.
We encountered studies with 19 different aims. To make comparison and discussion more feasible, we divided these studies into five groups according to study aim: (1) liver segmentation; (2) lesion segmentation; (3) lesion detection; (4) classification of liver or liver lesions; (5) miscellaneous/other. Aims are illustrated in electronic supplementary material. There is some overlap in the groups due to several studies having multiple aims. Detailed characteristics of included studies are given in supplementary tables.
Liver segmentation
The aim of liver segmentation was the primary or secondary study aim in eighty-four of the included studies. Of those, fifty-one are journal articles [20, 24, 29,30,31,32,33,34,35, 38,39,40,41, 43,44,45,46,47, 49, 55,56,57,58, 62, 63, 65, 68, 70,71,72,73,74,75,76,77,78,79, 81, 84,85,86,87, 89, 91, 93,94,95, 97, 98, 196, 197], and 33 are proceeding papers [19, 21,22,23, 25, 26, 36, 37, 42, 48, 51, 53, 54, 59,60,61, 64, 66, 67, 69, 80, 82, 83, 88, 90, 92, 96, 99, 100, 103, 198]. The liver segmentation was done from the CT as a whole liver, not the clinical segmentation, e.g., Couinaud segments of the liver. Overall, this group of studies has contributed considerably with technically sound methods and experimented with various subdomains of ML, especially DL.
The quality of many recent studies has improved using external validation method to provide better generalizability. Though comparing directly with human experts is preferred, only eleven studies were found to do so.
The study group gives insinuation of obtaining labeled medical data which is challenging, as two-thirds of studies used datasets open for public use for training or testing their ML model. The dataset from LiTS 2017, which was the most frequently used, included 131 patients in their test set [199].
The attempt of transparency in reporting models’ performance was seen in many studies, though out of eighty-seven studies, only 11 reported their results with confidence interval or standard error; thus, further analyses of the result were not feasible in the group.
DICE score was used in most studies in this group to describe the model’s ability to predict which pixel contains the liver. The highest DICE reported was a score of 0.9851 [41], and the lowest score was 0.75 [94]. Other measures to describe the model’s performance were scattered, including AUC-ROC and accuracy (Table 2). Dong et al also reported a DICE of 0.92 and an accuracy of 0.9722 from their study, and the AUC of 0.96. References of studies in the group are in Table 3.
Lesion segmentation
This group of studies performed segmentation of liver lesions from CT images with ML. The model’s goal was the highest possible truthfulness of segmented lesions compared to ground truth. Sixty studies had lesion segmentation as a primary or secondary study aim. Thirty-six are journal articles [24, 29, 31, 32, 38, 46, 47, 55, 56, 62, 72, 78, 84, 91, 93, 94, 97, 98, 102, 111, 115, 117, 118, 122, 124, 125, 130, 133,134,135, 137, 138, 140, 201], and twenty-four [22, 37, 42, 64, 65, 68, 82, 88, 92, 96, 99, 103, 108, 121, 124, 126,127,128,129, 131, 132, 136, 139, 200] are proceedings papers.
Several models have shown remarkable segmenting ability for predicted lesions larger than 2 cm in diameter, while almost every model is still struggling to segment lesion size less than 1 cm in diameter. However, this is comparable with clinicians in the clinical setting. Another limitation for the model to predict the lesion was quality of CT images. Several more recent studies used voxel-wise (3D pixels) classification. This could use more available information and give output in 3D to improve performance.
Validation of the model with external validation and ML to humans is improving for this group, and twenty-six studies have used external validation. Only six studies have compared their model with human experts.
More than half of the studies have reported performance in a DICE score in this group. The score range was seen skewed in different studies with the range of 0.44–0.96; a selection of lesion size played a key role here for higher performance or higher DICE score. Another informative measure called Volume Overlap Error (VOE) gives the difference between predicted and ground truth in an area. Thus, 0 is the optimal score. Twenty-two studies reported VOE, with a 0.01–0.46 mm range. Other measures were dispersed in different studies, including accuracy, AUC, precision, or PPV. Few studies have reported their performance with confidence intervals or standard errors—references of studies in the group in Table 3.
Lesion detection
Twenty studies had lesion detection as a primary or secondary study aim. This involves simply detecting if lesions are present in a CT image. Fifteen of them are proceedings papers [23, 26, 27, 87, 102, 104, 105, 107,108,109,110, 112,113,114, 119], and five are journal articles [101, 106, 111, 115, 202].
Several newer studies have detected lesions before segmentation of the lesions or diagnosis of the lesions with ML from CT liver images but have not reported performance of the lesion detection task of the model; thus, this group is smaller.
External validation was reported only in four studies. Most studies acquired their training data from local hospitals, and only eight studies have used data sets open for public use. DL was the choice of a subdomain of ML for this group.
Reporting of performance was seen as transparent and detailed in newer studies in all groups. In this group, performance was primarily reported in accuracy and precision, but five studies reported only false positives and true positive rate [26, 87, 101, 104, 115]. Two studies presented its result with a confidence interval or standard error. It is worth mentioning that the study reporting the best precision only performed internal validation on the relatively small, public dataset 3D-IRCADb—references of studies in the group in Table 3.
Classification of liver or lesions
Included studies in this group classifying the type and severity of lesions or tumors, grading hepatocellular carcinoma (HCC), and differentiating between HCC, hemangioma, and metastases. Most studies differed only between two categories, such as classifying tumors as either benign or malign. Forty-seven studies had the classification of liver or liver lesions as a study aim. Thirty-four of them journal articles [56, 71, 72, 74, 78, 141,142,143,144,145,146, 148,149,150,151,152, 154, 156,157,158,159,160,161, 164,165,166,167,168,169,170,171,172, 202, 203], and thirteen are proceedings papers [27, 64, 65, 68, 75, 82, 119, 147, 153, 155, 162, 163, 204]. For classification of liver or liver lesions, traditional machine learning, e.g., support vector machines and random forest models, and deep learning models were commonly used.
Nine studies compared their model performance directly to one or more clinicians in a competition-based comparison. Only 12 studies have used datasets open for public validation, and even fewer are needed for training purposes.
Accuracy was a method of choice to present the performance in this group; thirty-one studies reported accuracy, with a range of 0.76–0.99. Sixteen studies reported AUC, with a range of 0.68–0.97. Precision was reported in fourteen studies. The precision range was 0.82–1.00. Note that both Sreeja et al and Romero et al reported a perfect precision of 1.0, which Sreeja et al commented was possible due to the small size of their data set [153, 155]. Only three studies presented their result with a confidence interval—references of studies in the group are in Table 3.
Other/miscellaneous
The last and most diverse category we found eligible to compare was miscellaneous, including 29 journal article [6, 8, 9, 33, 50, 52, 56, 71, 161, 164, 173,174,175,176,177,178,179, 181,182,183,184, 186,187,188, 191, 194, 195, 205, 206] and 8 proceeding paper [27, 180, 185, 189, 190, 192, 193, 207] total thirty-seven studies. The aims of the studies are clinical-oriented.
Seven studies have performed liver fibrosis staging [33, 173,174,175,176,177,178] according to “Metavir” or “Fibrosis-4” classification [208, 209]. Four compared algorithms performance with human expert while two studies performed external validation. Only two studies used public dataset for liver segmenting purpose; however, private datasets were used for fibrosis staging training and validation purpose in all the seven studies. ML method like SVM, k-nearest neighbor were used traditionally but in the recent studies, CNN-based systems using different classifier to extract the feature from the liver image are gaining more attention. Jung et al used liver and spleen volumetric indices and perform the pathologic liver fibrosis staging with CNN [177]. Comparison of ML algorithm to 3 radiologists’ assessment of liver fibrosis staging was performed with more accurate result in ML group [33].
Six studies segmented blood vessels in the liver from CT images, including portal and liver veins [52, 179, 183,184,185, 191]. Twelve studies reported a DICE score with a range of 0.68–0.98. The four studies reported accuracy with a range of 0.91–0.98, with a mean of 0.96 and a median of 0.97. Five studies stated that they externally validated their model.
Five retrieved focal liver lesion images as a study aim [50, 186, 187, 192, 206]. These studies showed how models could improve clinical workflow by retrieving similar cases in medical records, including earlier expert opinions.
Two studies, published as journal articles, predicted liver metastases within colorectal cancer patients [8, 9]. They reported AUC equal to 0.86 ± 0.01(12) and 0.747 ± 0.036.
One study focused on the segmentation of bile ducts and stones in the intrahepatic bile duct—hepatolith and reported DICE of 0.90 and 0.71 for bile duct and hepatolith segmentation, respectively [6].
Three study focused on response evaluation after chemotherapy or radio-embolization of malignant liver lesions using texture analysis [161, 181, 182]. They compared texture analysis predictions with survival and serologic response and reported an accuracy of 0.97, sensitivity of 0.93, and specificity of 1.0. This was after training on sixty-two patients and testing using cross-validation.
Two recent studies have predicted liver reserve function using Child–Pugh classification [164, 189] and Thuring et al have compared the results from their ML model with results from clinicians. Prediction of Child–Pugh accuracy was 53%, classification of Child–Pugh A vs B: accuracy was 78%, sensitivity 81%, specificity 70%, and AUC 0.80. Wang et al had preoperatively predicted early recurrence in HCC. One study has predicted overall survival of patients with unresectable HCC treated by transarterial chemoembolization [176]. This study also presented fusion of clinical data with ML model. References of studies in the group in Table 3.
Discussion
We found that ML is applied to liver CT imaging for various clinical oriented aims and covering a broad spectrum of applications.
At least one-third of studies were documented to perform very accurately on reliable, but small data. Unfortunately, reporting of performance was seldom appropriate due to lack of details. To our knowledge, there exists no standardized form of presenting results for machine learning models applied to medical imaging.
Several studies reported models that were close to clinical application. However, clinical validation with thorough documentation of both model and data (training and validation) to assess quality and generalizability were lacking. Evaluation of the model by only analysis of a result parameters would be imperfect [210].
Almost all studies that performed segmentation of liver structures from the CT images of the abdomen used deep learning models, mainly the subtype CNN. Open-access datasets and competitions like LiTS 2017 contribute substantially to the development of ML applied to liver imaging, as more than 280 studies report their model performance in a standardized format, and the competition is still ongoing with cumulative comparison. U-Net a sub domain of CNN is used by many participants and have shown promising result. The distribution of sources of dataset used by studies included in this review is illustrated in Fig. 2. The use of complex models and targeting for complex aims like automatic liver fibrosis staging, treatment response evaluation, prediction of occurrence of liver metastases, and liver blood vessels segmentation for traditional anatomical landmarks, e.g., Coineaud classification, are getting more common and may herald a maturing process in the field.
ML systems showed promising results on retrospective data for several tasks related to CT imaging, as some segmentation studies reported models with more than 98% ability to predict which pixels or voxels contained liver in abdominal CT scans. Further, several studies reported 95% performance compared to ground truth for liver or liver lesions classification. In recent years, identified studies have used ML for prediction of occurrence or treatment effect of liver metastases, liver vessel segmentation, and evaluation of treatment effect on liver malignancy. These showed results around 70–80% of ground truth.
Other applications such as classification of liver fibrosis stage and prediction of benign or malign lesions showed promising results and potential for the high impact of ML in future routine clinical practice.
Reporting of model performance should give in the state-of-the-art visualization methods, e.g., confusion matrix. In the studies like segmentation task, measuring parameter like mean surface distance with standard error should be reported to get overall transparency of the model performance [116]. Sixty-two studies identified in this review have such breach in reporting of model performance. This makes it difficult to get a good overall understanding of the field, especially for clinicians. We encourage the readers to assess such results with caution.
Further, reporting of standard error and confidence intervals was often lacking. We recommend that it should be reported by default. This problem was also seen in other applications of ML to medical images, and we concur with the need for reporting standards for medical application as stated by Aggarwal et al [10].
There are potentially many applications of ML in liver CT imaging have been identified thorough this review, especially in the miscellaneous group aims are clinically derived, while segmenting of liver and its lesions could implement as diagnostic and treatment planning tool. Studies in classification group could serve diagnosis of different lesions, e.g., different types of malign and benign tumors, or severity of the liver cirrhosis. Despite the promising performance reported in many studies, clinical applications of ML in liver CT imaging have to pass through the corridor of clinical validation and clinical trials [210].
The main issues identified in the literature were limited access to high-quality data and lack of clinical validation. External validation is becoming more popular among developers, illustrated in Fig. 3, but it is insufficient to qualify for medical application. There is an urgent need for a shift in focus towards clinical validation in this field. Scholars should perform feasibility studies in clinical routine, and design and carry out prospective studies to validate the performance of ML tools in realistic clinical settings. Developers should seek to collaborate with clinicians in this process. Strength and weakness of the study as well future perspective is given in the supplementary material.
Conclusion
We found reports of many ML applications to liver CT images in the literature, including automatic liver and lesion segmentation, lesion detection, liver or lesion classification, liver vessel segmentation including bile ducts, fibrosis staging, metastasis prediction, and evaluation of chemotherapy as treatment of hepatocellular carcinoma and retrieval of relevant liver lesions from other similar cases. Several were documented to perform very accurately on reliable but small data. Deep learning models and classification models of ML were commonly used. However, presenting the result of studies is not standardized in the literature. Some studies were close to reporting sufficient details on clinical relevance, data characteristics and quality, algorithm characteristics and bias, and performance measures on external data to be considered ready for clinical use. Further prospective, clinical studies are recommended, and the need for a more interactive technological and medical research is evident to achieve a secure clinical use of ML methodology in this field.
Code availability
Custom code or mathematical algorithms were not used and do not play any role in our conclusion.
Abbreviations
- 3D RA U-Net:
-
3D hybrid residual attention U-shaped neural network
- A:
-
Article
- ACM:
-
Auto-context model
- AHC Blocks:
-
Attention hybrid connection blocks
- ANN:
-
Artificial neural network
- ASM:
-
Active shape model
- BPSO:
-
Binary particle swarm optimization
- CDNN:
-
Convolutional—deconvolutional neural network
- CEDCNN:
-
Cascade encoder-decoder CNN
- CENet:
-
Contour embedded neural network
- CNN:
-
Convolutional neural network
- CRF:
-
Conditional random field
- CT:
-
Computed tomography
- DBN-DNN:
-
Deep belief network-deep neural network
- DCT:
-
Discrete cosine transforms
- DL:
-
Deep learning
- DLA:
-
Deep learning algorithm
- DLO:
-
Dice loss
- DResU-Net:
-
Deep residual U-net
- DRL:
-
Deep reinforcement learning
- ELM:
-
Extreme learning machine
- FCMC:
-
Fuzzy C-means clustering
- FCN:
-
Fully convolutional neural network
- FCNN:
-
Fully convolutional neural network
- GAN:
-
Generative adversarial network
- GDL:
-
Generalized dice loss
- GLC U-Net:
-
Global and local contexts composition U-shaped neural network
- GTL:
-
Generalized Teverskry loss
- GWO:
-
Grey wolf optimization
- HCC:
-
Hepatocellular carcinom
- HCC:
-
Hepatocellular carcinoma
- HDCNN:
-
Hybridized fully convolutional neural network
- k-NN:
-
k-nearest neighbor
- ML:
-
Machine learning
- MOGA:
-
Multi objective genetic algorithm
- MPNet:
-
Message passing neural network
- MRF:
-
Markov random field
- MSCA:
-
Mean-shift clustering algorithm
- MW-U-Net:
-
Modality weighted U-net
- PCA:
-
Principal component analysis
- PNN:
-
Probabilistic neural network
- PP:
-
Proceeding paper from conference
- R-CNN:
-
Region based convolutional neural network
- RES-U-Net:
-
Residual U-net
- RFC:
-
Random forest classifier
- RL:
-
Reinforcement learning
- RPN:
-
Region proposal network
- SSD:
-
Single-shot multibox detector
- SSD:
-
Support vector machine
- TDP:
-
Three-dimensional dual path multiscale convolutional neural network
- TL:
-
Teverskry loss
- U-NET:
-
U-shaped neural network (referes to the model architecture)
- U-RES-Net:
-
U-shaped residual neural network
- VGG 16:
-
Visual Geometry group 16 (Personal name of model named after a research group)
- VOE:
-
Volume overlap error
References
Baştanlar Y, Özuysal M (2014) Introduction to machine learning. In: Yousef M, Allmer J (eds) miRNomics: MicroRNA Biology and Computational Analysis. Humana Press, Totowa, NJ, pp 105–128
Muehlematter UJ, Daniore P, Vokinger KN (2021) Approval of artificial intelligence and machine learning-based medical devices in the USA and Europe (2015–20): a comparative analysis. Lancet Digit Health 3:e195–e203
FDA (2021) Artificial Intelligence and Machine Learning (AI/ML) Software as a Medical Device Action Plan 2021. FDA. Available via https://www.fda.gov/medical-devices/software-medical-device-samd/artificial-intelligence-and-machine-learning-software-medical-device. Accessed 24.01.2023
Rubin GD (2014) Computed tomography: revolutionizing the practice of medicine for 40 years. Radiology 273:S45–S74
Cardobi N, Dal Palu A, Pedrini F et al (2021) An overview of artificial intelligence applications in liver and pancreatic imaging. Cancers 13:11
Fu X, Cai N, Huang K et al (2019) M-Net: a novel U-Net with multi-stream feature fusion and multi-scale dilated convolutions for bile ducts and hepatolith segmentation. IEEE Access 7:148645–148657
Decharatanachart P, Chaiteerakij R, Tiyarattanachai T, Treeprasertsuk S (2021) Application of artificial intelligence in chronic liver diseases: a systematic review and meta-analysis. BMC Gastroenterol 21:10
Lee S, Choe EK, Kim SY, Kim HS, Park KJ, Kim D (2020) Liver imaging features by convolutional neural network to predict the metachronous liver metastasis in stage I-III colorectal cancer patients based on preoperative abdominal CT scan. BMC Bioinf 21:382
Taghavi M, Trebeschi S, Simões R et al (2021) Machine learning-based analysis of CT radiomics model for prediction of colorectal metachronous liver metastases. Abdom Radiol (NY) 46:249–256
Aggarwal R, Sounderajah V, Martin G et al (2021) Diagnostic accuracy of deep learning in medical imaging: a systematic review and meta-analysis. NPJ Digital Med 4:65
Zhou LQ, Wang JY, Yu SY et al (2019) Artificial intelligence in medical imaging of the liver. World J Gastroenterol 25:672–682
Park HJ, Park B, Lee SS (2020) Radiomics and deep learning: hepatic applications. Korean J Radiol 21:387–401
Azer SA (2019) Deep learning with convolutional neural networks for identification of liver masses and hepatocellular carcinoma: a systematic review. World J Gastrointest Oncol 11:1218–1230
Schiavo JH (2019) PROSPERO: an international register of systematic review protocols. Med Ref Serv Q 38:171–180
McInnes MDF, Moher D, Thombs BD et al (2018) Preferred Reporting Items for a Systematic Review and Meta-analysis of diagnostic test accuracy studies: the PRISMA-DTA statement. JAMA 319:388–396
Cumpston M, Li T, Page MJ et al (2019) Updated guidance for trusted systematic reviews: a new edition of the Cochrane Handbook for Systematic Reviews of Interventions. Cochrane Database Syst Rev 10:Ed000142
de Hond AAH, Leeuwenberg AM, Hooft L et al (2022) Guidelines and quality criteria for artificial intelligence-based prediction models in healthcare: a scoping review. NPJ Digit Med 5:2
Haddaway NR, Page MJ, Pritchard CC, McGuinness LA (2022) PRISMA2020: an R package and Shiny app for producing PRISMA 2020-compliant flow diagrams, with interactivity for optimised digital transparency and Open Synthesis. Campbell Syst Rev 18:e1230
Mubashir A, Yuan D, Syed Furqan Q, Jian Y (2019) Convolutional-neural-network-based feature extraction for liver segmentation from CT imagesProcSPIE, pp 1117934
Ahn Y, Yoon JS, Lee SS et al (2020) Deep learning algorithm for automated segmentation and volume measurement of the liver and spleen using portal venous phase computed tomography images. Korean J Radiol 21:987–997
Bhavya A, Aditya B, Karthik K (2018) Automatic and fast CT liver segmentation using sparse ensemble with machine learned contextsProcSPIE, pp 105740L
Albishri AA, Shah SJH, Lee Y (2019) CU-Net: cascaded U-Net model for automated liver and lesion segmentation and summarization. In: IEEE International Conference on Bioinformatics and Biomedicine (BIBM), San Diego, pp 1416–1423
Ali L, Khelil K, Wajid SK et al (2017) Machine learning based computer-aided diagnosis of liver tumour. In: IEEE 16th International Conference on Cognitive Informatics and Cognitive Computing (ICCI*CC), Oxford, pp 139–114
Alirr OI (2020) Deep learning and level set approach for liver and tumor segmentation from CT scans. J Appl Clin Med Phys 21:200–209
Astono I, Welsh JS, Chalup S (2018) Adjacent network for semantic segmentation of liver CT scans. In: 18th IEEE International Conference on Bioinformatics and Bioengineering, Taichung, pp 35–40
Ben-Cohen A, Diamant I, Klang E, Amitai M, Greenspan H (2016) Fully convolutional network for liver segmentation and lesions detection. In: 2nd International Workshop on Deep Learning in Medical Image Analysis (DLMIA) / 1st International Workshop on Large-Scale Annotation of Biomedical Data and Expert Label Synthesis (LABELS). Springer International Publishing Ag, Athens, pp 77–85
Bevilacqua V, Brunetti A, Trotta GF et al (2017) A novel approach for hepatocellular carcinoma detection and classification based on triphasic CT Protocol2017 IEEE Congress on Evolutionary Computation (CEC), pp 1856–1863
Bhole C, Morsillo N, Pal C (2011) 3D segmentation in CT imagery with conditional random fields and histograms of oriented gradients. In: Suzuki K, Wang F, Shen D, Yan P (eds) Machine Learning in Medical Imaging. Springer, Berlin Heidelberg, Berlin, Heidelberg, pp 326–334
Budak U, Guo Y, Tanyildizi E, Sengur A (2020) Cascaded deep convolutional encoder-decoder neural networks for efficient liver tumor segmentation. Med Hypotheses 134:8
Cai J (2019) Segmentation and diagnosis of liver carcinoma based on adaptive scale-kernel fuzzy clustering model for CT images. J Med Syst 43:322
Chen Y, Wang K, Liao X et al (2019) Channel-Unet: a spatial channel-wise convolutional neural network for liver and tumors segmentation. Front Gen 10
Chlebus G, Schenk A, Moltz JH, van Ginneken B, Hahn HK, Meine H (2018) Automatic liver tumor segmentation in CT with fully convolutional neural networks and object-based postprocessing. Sci Rep 8:15497
Choi KJ, Jang JK, Lee SS et al (2018) Development and validation of a deep learning system for staging liver fibrosis by using contrast agent-enhanced CT images in the liver. Radiol 289:688–697
Chung M, Lee J, Lee M, Lee J, Shin Y-G (2020) Deeply self-supervised contour embedded neural network applied to liver segmentation. Comput Methods Programs Biomed 192:105447
Danciu M, Gordan M, Florea C, Orghidan R, Sorantin E, Vlaicu A (2013) A hybrid 3D learning-and-interaction-based segmentation approach applied on CT liver volumes. Radioeng 22:100–113
Danciu M, Gordan M, Florea C, Vlaicu A (2012) 3D DCT supervised segmentation applied on liver volumes 2012. 35th International Conference on Telecommunications and Signal Processing (TSP), pp 779–783
Delmoral JC, Costa DC, Borges D, Tavares JMRS (2019) Segmentation of pathological liver tissue with dilated fully convolutional networks: a preliminary study2019 IEEE 6th Portuguese Meeting on Bioengineering (ENBENG), pp 1–4
Dong X, Zhou Y, Wang L, Peng J, Lou Y, Fan Y (2020) Liver cancer detection using hybridized fully convolutional neural network based on deep learning framework. IEEE Access 8:129889–129898
Dou Q, Yu LQ, Chen H et al (2017) 3D deeply supervised network for automated segmentation of volumetric medical images. Med Image Anal 41:40–54
Guo X, Schwartz LH, Zhao B (2019) Automatic liver segmentation by integrating fully convolutional networks into active contour models. Med Phys 46:4455–4469
He B, Huang C, Sharp G et al (2016) Fast automatic 3D liver segmentation based on a three-level AdaBoost-guided active shape model. Med Phys 43:2421
Heker M, Ben-Cohen A, Greenspan H (2019) Hierarchical fine-tuning for joint liver lesion segmentation and lesion classification in CT2019 41st Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), pp 895–898
Hu P, Wu F, Peng J, Liang P, Kong D (2016) Automatic 3D liver segmentation based on deep learning and globally optimized surface evolution. Phys Med Biol 61:8676–8698
Huang W, Tan ZM, Lin Z et al (2012) A semi-automatic approach to the segmentation of liver parenchyma from 3D CT images with extreme learning machine. In: 34th Annual International Conference of the IEEE Engineering-in-Medicine-and-Biology-Society (EMBS). IEEE, San Diego, pp 3752–3755
Ji H, He J, Yang X, Deklerck R, Cornelis J (2013) ACM-based automatic liver segmentation from 3-D CT images by combining multiple atlases and improved mean-shift techniques. IEEE J Biomed Health Inform 17:690–698
Jiang H, Li S, Li S (2018) Registration-based organ positioning and joint segmentation method for liver and tumor segmentation. Biomed Res Int 2018:8536854
Jiang H, Shi T, Bai Z, Huang L (2019) AHCNet: an application of attention mechanism and hybrid connection for liver tumor segmentation in CT volumes. IEEE Access 7:24898–24909
Jin X, Ye H, Li L, Xia Q (2017) Image segmentation of liver CT based on fully convolutional network2017 10th International Symposium on Computational Intelligence and Design (ISCID), pp 210–213
Kavur AE, Gezer NS, Barış M et al (2020) Comparison of semi-automatic and deep learning-based automatic methods for liver segmentation in living liver transplant donors. Diagn Interv Radiol 26:11–21
Kumar A, Dyer S, Kim J et al (2016) Adapting content-based image retrieval techniques for the semantic annotation of medical images. Comput Med Imaging Graph 49:37–45
Zheng H, Lin L, Hu H et al (2019) Semi-supervised segmentation of liver using adversarial learning with deep atlas prior. In: Shen D, Liu T, Peters TM et al (eds) Medical Image Computing and Computer Assisted Intervention – MICCAI 2019. Springer International Publishing, Cham, pp 148–156
Zhang R, Zhou Z, Wu W, Lin CC, Tsui PH, Wu S (2018) An improved fuzzy connectedness method for automatic three-dimensional liver vessel segmentation in CT images. J Healthc Eng 2018:2376317
Zhang L, Xu L (2018) An automatic liver segmentation algorithm for CT images U-net with separated paths of feature extraction2018 IEEE 3rd International Conference on Image, Vision and Computing (ICIVC), pp 294–298
Xu W, Liu H, Wang X, Qian Y (2019) Liver segmentation in CT based on ResUNet with 3D probabilistic and geometric post process2019 IEEE 4th International Conference on Signal and Image Processing (ICSIP), pp 685–689
Xi XF, Wang L, Sheng VS, Cui Z, Fu B, Hu F (2020) Cascade U-ResNets for simultaneous liver and lesion segmentation. IEEE Access 8:68944–68952
Xin S, Shi H, Jide A, Zhu M, Ma C, Liao H (2020) Automatic lesion segmentation and classification of hepatic echinococcosis using a multiscale-feature convolutional neural network. Med Biol Eng Comput 58:659–668
Xia K, Yin H, Qian P, Jiang Y, Wang S (2019) Liver semantic segmentation algorithm based on improved deep adversarial networks in combination of weighted loss function on abdominal CT images. IEEE Access 7:96349–96358
Winkel DJ, Weikert TJ, Breit H-C et al (2020) Validation of a fully automated liver segmentation algorithm using multi-scale deep reinforcement learning and comparison versus manual segmentation. Eur J Radiol 126:108918
Wang C, Song H, Chen L et al (2018) Automatic liver segmentation using multi-plane integrated fully convolutional neural networks2018 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), pp 1–6
Tian J, Liu L, Shi Z, Xu F (2019) Automatic Couinaud segmentation from CT volumes on liver using GLC-UNet. In: Suk H-I, Liu M, Yan P, Lian C (eds) Machine Learning in Medical Imaging. Springer International Publishing, Cham, pp 274–282
Tang M, Valipour S, Zhang Z, Cobzas D, Jagersand M (2017) A deep level set method for image segmentation. In: Cardoso MJ, Arbel T, Carneiro G et al (eds) Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support. Springer International Publishing, Cham, pp 126–134
Seo H, Huang C, Bassenne M, Xiao R, Xing L (2020) Modified U-Net (mU-Net) with incorporation of object-dependent high level features for improved liver and liver-tumor segmentation in CT images. IEEE Trans Med Imaging 39:1316–1325
Selvi E, Selver MA, Güzeliş C, Dicle O (2014) A higher-order neural network design for improving segmentation performance in medical image series. J Phys: Conf Ser 490:012079
Selvathi D, Malini C, Shanmugavalli P (2013) Automatic segmentation and classification of liver tumor in CT images using adaptive hybrid technique and Contourlet based ELM classifier2013 International Conference on Recent Trends in Information Technology (ICRTIT), pp 250–256
Sayed GI, Hassanien AE, Schaefer G (2016) An automated computer-aided diagnosis system for abdominal CT liver images 20th conference on medical image understanding and analysis (MIUA 2016), Loughborough Univ, Loughborough, England, pp 68–73
Sakboonyara B, Taeprasartsit P (2019) U-Net and mean-shift histogram for efficient liver segmentation from CT images2019 11th International Conference on Knowledge and Smart Technology (KST), pp 51–56
K S, H LU, H KIM, S K, M T (2018) ROI-based fully automated liver registration in multi-phase CT Images2018 18th International Conference on Control, Automation and Systems (ICCAS), pp 645–649
Raj A, Jayasree M (2016) Automated liver tumor detection using Markov random field segmentation International conference on emerging trends in engineering, science and technology (ICETEST - 2015), Trichur, India, pp 1305–1310
Rafiei S, Nasr-Esfahani E, Najarian K, Karimi N, Samavi S, Soroushmehr SMR (2018) Liver segmentation in CT images using three dimensional to two dimensional fully convolutional network2018 25th IEEE International Conference on Image Processing (ICIP), pp 2067–2071
Qin W, Wu J, Han F et al (2018) Superpixel-based and boundary-sensitive convolutional neural network for automated liver segmentation. Phys Med Biol 63:095017
Ponnoprat D, Inkeaw P, Chaijaruwanich J et al (2020) Classification of hepatocellular carcinoma and intrahepatic cholangiocarcinoma based on multi-phase CT scans. Med Biol Eng Comput 58:2497–2515
Ouhmich F, Agnus V, Noblet V, Heitz F, Pessaux P (2019) Liver tissue segmentation in multiphase CT scans using cascaded convolutional neural networks. Int J Comput Assist Radiol Surg 14:1275–1284
Ng YS, Xi Y, Qian Y et al (2020) Use of spectral detector computed tomography to improve liver segmentation and volumetry. J Comput Assist Tomogr 44:197–203
Nayak A, Baidya Kayal E, Arya M et al (2019) Computer-aided diagnosis of cirrhosis and hepatocellular carcinoma using multi-phase abdomen CT. Int J Comput Assist Radiol Surg 14:1341–1352
Mukherjee DP, Higashiura K, Okada T et al (2013) Utilizing disease-specific organ shape components for disease discrimination: application to discrimination of chronic liver disease from CT data16th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI) pp 235-242. Nagoya Univ, Nagoya, Japan
Morshid A, Elsayes KM, Khalaf AM et al (2019) A machine learning model to predict hepatocellular carcinoma response to transcatheter arterial chemoembolization. Radiol Artif Intell 1
Mohagheghi S, Foruzan AH (2020) Incorporating prior shape knowledge via data-driven loss model to improve 3D liver segmentation in deep CNNs. Int J Comput Assist Radiol Surg 15:249–257
Mofrad FB, Zoroofi RA, Tehrani-Fard AA, Akhlaghpoor S, Sato Y (2014) Classification of normal and diseased liver shapes based on Spherical Harmonics coefficients. J Med Syst 38:20
Meng L, Tian Y, Bu S (2020) Liver tumor segmentation based on 3D convolutional neural network with dual scale. J Appl Clin Med Phys 21:144–157
Luo S, Li J (2014) Accurate object segmentation using novel active shape and appearance models based on support vector machine learning2014 International Conference on Audio, Language and Image Processing, pp 347–351
Lu F, Wu F, Hu P, Peng Z, Kong D (2017) Automatic 3D liver location and segmentation via convolutional neural network and graph cut. Int J Comput Assist Radiol Surg 12:171–182
Selvaraj G, Janakiraman S (2013) Improved feature selection based on particle swarm optimization for liver disease diagnosis. In: 4th International Conference on Swarm, Evolutionary, and Memetic Computing (SEMCCO). Springer-Verlag Berlin, SRM University, Chennai, pp 214–225
Li XH, Huang C, Jia FC, Li ZM, Fang CH, Fan YF (2014) Automatic liver segmentation using statistical prior models and free-form deformation. In: International Workshop on Medical Computer Vision - Algorithms for Big Data (MICCAI-bigMCV), Cambridge, pp 181–188
Li X, Chen H, Qi X, Dou Q, Fu CW, Heng PA (2018) H-DenseUNet: hybrid densely connected unet for liver and tumor segmentation from CT volumes. IEEE Trans Med Imaging 37:2663–2674
Liu Z, Song YQ, Sheng VS et al (2019) Liver CT sequence segmentation based with improved U-Net and graph cut. Expert Systems with Applications 126:54–63
Linguraru MG, Richbourg WJ, Liu J et al (2012) Tumor burden analysis on computed tomography by automated liver and tumor segmentation. IEEE Trans Med Imaging 31:1965–1976
Afifi A, Nakaguchi T (2015) Unsupervised detection of liver lesions in CT images. In: 37th Annual International Conference of the IEEE-Engineering-in-Medicine-and-Biology-Society (EMBC). IEEE, Milan, pp 2411–2414
Roth K, Hesser J, Konopczynski T (2020) Mask mining for improved liver lesion segmentation. In: IEEE 17th International Symposium on Biomedical Imaging (ISBI). IEEE, Iowa, pp 943–947
Tran ST, Cheng CH, Liu DG (2021) A multiple layer U-Net, U-n-Net, for liver and liver tumor segmentation in CT. IEEE Access 9:3752–3764
Xu HL, Wang BH, Xue WG et al (2019) Automatic segmentation of liver CT image based on dense pyramid network. In: 1st International Workshop on Multiscale Multimodal Medical Imaging (MMMI). Springer International Publishing, Shenzhen, pp 10–16
Yu AH, Liu Z, Sheng VS et al (2021) CT segmentation of liver and tumors fused multi-scale features. Intell Autom Soft Comput 30:589–599
Zhang Y, Tian J, Zhong C et al (2021) DARN: Deep attentive refinement network for liver tumor segmentation from 3D CT volume. In: 25th International Conference on Pattern Recognition (ICPR). IEEE Computer Society, Electrical Network, pp 7796–7803
Ayalew YA, Fante KA, Mohammed MA (2021) Modified U-Net for liver cancer segmentation from computed tomography images with a new class balancing method. BMC Biomedical Engineering 3:4
Chen WF, Ou HY, Liu KH et al (2021) In-series U-Net network to 3D tumor image reconstruction for liver hepatocellular carcinoma recognition. Diagnostics 11:18
Chung M, Lee J, Park S, Lee CE, Lee J, Shin YG (2021) Liver segmentation in abdominal CT images via auto-context neural network and self-supervised contour attention*. Artif Intell Med 113:12
Elmenabawy NA, Elnakib A, Moustafa HED (2020) Deep joint segmentation of liver and cancerous nodules from Ct images2020 37th National Radio Science Conference (NRSC), pp 296–301
Fan TL, Wang GL, Li Y, Wang HR (2020) MA-Net: a multi-scale attention network for liver and tumor segmentation. IEEE Access 8:179656–179665
He K, Liu XM, Shahzad R et al (2021) Advanced deep learning approach to automatically segment malignant tumors and ablation zone in the liver with contrast-enhanced CT. Front Oncol 11:10
Kwon J, Choi K (2020) Trainable multi-contrast windowing for liver CT segmentation. In: IEEE International Conference on Big Data and Smart Computing (BigComp). IEEE, Busan, pp 169–172
Lei T, Zhou WZ, Zhang YX et al (2020) Lightweight v-net for liver segmentation. In: 2020 IEEE International Conference on Acoustics, Speech, and Signal Processing. IEEE, Barcelona, pp 1379–1383
Ben-Cohen A, Klang E, Kerpel A, Konen E, Amitai MM, Greenspan H (2018) Fully convolutional network and sparsity-based dictionary learning for liver lesion detection in CT examinations. Neurocomputing 275:1585–1594
Bevilacqua V, Carnimeo L, Brunetti A et al (2016) Synthesis of a neural network classifier for hepatocellular carcinoma grading based on triphasic CT images. In: 1st International Conference on Recent Trends in Image Processing and Pattern Recognition (RTIP2R). Springer-Verlag Berlin, Karnatak Arts Sci & Commerce Coll, Bidar, pp 356–368
Chen L, Song H, Li Q, Cui YT, Yang J, Hu XHT (2019) Liver segmentation in CT images using a non-local fully convolutional neural network. In: IEEE International Conference on Bioinformatics and Biomedicine (BIBM). IEEE, San Diego, pp 639–642
Frid-Adar M, Diamant I, Klang E, Amitai M, Goldberger J, Greenspan H (2017) Modeling the intra-class variability for liver lesion detection using a multi-class patch-based CNN. In: 3rd International Workshop on Patch-Based Techniques in Medical Images (Patch-MI). Springer International Publishing Ag, Quebec City, pp 129–137
Furuzuki M, Lu HM, Kim H et al (2019) A detection method for liver cancer region based on faster R-CNN. In: 19th International Conference on Control, Automation and Systems (ICCAS). IEEE, Jeju, pp 808–811
Gong H, Yu LF, Leng S et al (2019) A deep learning- and partial least square regression-based model observer for a low-contrast lesion detection task in CT. Med Phys 46:2052–2063
Huang WM, Li N, Lin ZP et al (2013) Liver tumor detection and segmentation using kernel-based extreme learning machine. In: 35th Annual International Conference of the IEEE-Engineering-in-Medicine-and-Biology-Society (EMBC). IEEE, Osaka, pp 3662–3665
Jin XY, Du ZH, Zhang T, Li LJ (2017) A disease detection method of liver based on improved convolutional neural network. In: 10th International Symposium on Computational Intelligence and Design (ISCID). IEEE, Hangzhou, pp 96–98
Jin XY, Jin QL, Yang X (2015) A disease detection method of liver based on improved back propagation neural network. In: 8th International Symposium on Computational Intelligence and design (ISCID). IEEE, Hangzhou, pp 111–113
Kim B, Kim J, Lee J-G, Kim DH, Park SH, Ye JC (2019) Unsupervised deformable image registration using cycle-consistent CNN. In: Shen D, Liu T, Peters TM et al (eds) Medical Image Computing and Computer Assisted Intervention – MICCAI 2019. Springer International Publishing, Cham, pp 166–174
Vivanti R, Szeskin A, Lev-Cohain N, Sosna J, Joskowicz L (2017) Automatic detection of new tumors and tumor burden evaluation in longitudinal liver CT scan studies. Int J Comput Assist Radiol Surg 12:1945–1957
Tao QY, Ge ZY, Cai JF, Yin JX, See S (2019) Improving deep lesion detection using 3D contextual and spatial attention. In: 10th International Workshop on Machine Learning in Medical Imaging (MLMI) / 22nd International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI). Springer International Publishing Ag, Shenzhen, pp 185–193
Liang D, Lin L, Chen X et al (2019) Multi-stream scale-insensitive convolutional and recurrent neural networks for liver tumor detection in dynamic Ct Images2019 IEEE International Conference on Image Processing (ICIP), pp 794–798
Lee S-g, Bae JS, Kim H, Kim JH, Yoon S (2018) Liver lesion detection from weakly-labeled multi-phase CT volumes with a grouped single shot MultiBox detector. In: Frangi AF, Schnabel JA, Davatzikos C, Alberola-López C, Fichtinger G (eds) Medical Image Computing and Computer Assisted Intervention – MICCAI 2018. Springer International Publishing, Cham, pp 693–701
Yang CJ, Wang CK, Fang YD et al (2021) Clinical application of mask region-based convolutional neural network for the automatic detection and segmentation of abnormal liver density based on hepatocellular carcinoma computed tomography datasets. PLoS ONE 16:e0255605
Zhou J, Gandomi AH, Chen F, Holzinger A (2021) Evaluating the quality of machine learning explanations: a survey on methods and metrics. Electronics 10:593
Almotairi S, Kareem G, Aouf M, Almutairi B, Salem MA (2020) Liver tumor segmentation in CT scans using modified SegNet. Sensors (Basel) 20
Anter AM, Hassenian AE (2019) CT liver tumor segmentation hybrid approach using neutrosophic sets, fast fuzzy c-means and adaptive watershed algorithm. Artif Intell Med 97:105–117
Chen X, Lin LF, Liang D et al (2019) A dual-attention dilated residual network for liver lesion classification and localization on CT images. In: 26th IEEE International Conference on Image Processing (ICIP). IEEE, Taipei, pp 235–239
Deng ZF, Guo QZ, Zhu ZL (2019) Dynamic regulation of level set parameters using 3D convolutional neural network for liver tumor segmentation. J Healthc Eng 2019:17
Huang W, Yang Y, Lin Z et al (2014) Random feature subspace ensemble based extreme learning machine for liver tumor detection and segmentation2014 36th Annual International Conference of the IEEE Engineering in Medicine and Biology Society, pp 4675–4678
Kadoury S, Vorontsov E, Tang A (2015) Metastatic liver tumour segmentation from discriminant Grassmannian manifolds. Phys Med Biol 60:6459
Zhou JY, Huang WM, Xiong W, Chen WY, Venkatesh SK (2013) Segmentation of hepatic tumor from abdominal CT data using an improved support vector machine framework. In: 35th Annual International Conference of the IEEE-Engineering-in-Medicine-and-Biology-Society (EMBC). IEEE, Osaka, pp 3347–3350
Zhang Y, Pan X, Li C, Wu T (2020) 3D liver and tumor segmentation with CNNs based on region and distance metrics. Appl Sci. https://doi.org/10.3390/app10113794
Zhang Y, Jiang B, Wu J et al (2020) Deep learning initialized and gradient enhanced level-set based segmentation for liver tumor from CT images. IEEE Access 8:76056–76068
Zhang X, Tian J, Xiang DH, Li XL, Deng KX (2011) Interactive liver tumor segmentation from CT scans using support vector classification with watershed. In: 33rd Annual International Conference of the IEEE Engineering-in-Medicine-and-Biology-Society (EMBS). IEEE, Boston, pp 6005–6008
Wu Y, Zhou Q, Hu H, Rong G, Li Y, Wang S (2019) Hepatic lesion segmentation by combining plain and contrast-enhanced CT images with modality weighted U-Net2019 IEEE International Conference on Image Processing (ICIP), pp 255–259
Wei Y, Jiang X, Liu K et al (2019) A hybrid multi-atrous and multi-scale network for liver lesion detection. In: Suk H-I, Liu M, Yan P, Lian C (eds) Machine Learning in Medical Imaging. Springer International Publishing, Cham, pp 364–372
Vorontsov E, Tang A, Pal C, Kadoury S (2018) Liver lesion segmentation informed by joint liver segmentation2018 IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018), pp 1332–1335
Vorontsov E, Tang A, Roy D, Pal CJ, Kadoury S (2017) Metastatic liver tumour segmentation with a neural network-guided 3D deformable model. Med Biol Eng Comput 55:127–139
Todoroki Y, Iwamoto Y, Lin L, Hu H, Chen YW (2019) Automatic detection of focal liver lesions in multi-phase CT images using a multi-channel & multi-scale CNN2019 41st Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), pp 872–875
Sun C, Guo S, Zhang H, Li J, Ma S, Li X (2017) Liver lesion segmentation in CT images with MK-FCN2017 IEEE 2nd Advanced Information Technology, Electronic and Automation Control Conference (IAEAC), pp 1794–1798
Shimizu A, Narihira T, Kobatake H, Furukawa D, Nawano S, Shinozaki K (2013) Ensemble learning based segmentation of metastatic liver tumours in contrast-enhanced computed tomography. IEICE Trans Inf Syst 96-D:864–868
Moawad AW, Fuentes D, Khalaf AM et al (2020) Feasibility of automated volumetric assessment of large hepatocellular carcinomas’ responses to transarterial chemoembolization. Front Oncol 10:572
Radu C, Fisher P, Mitrea D et al (2020) Integration of real-time image fusion in the robotic-assisted treatment of hepatocellular carcinoma. Biol (Basel) 9
Haq MNU, Irtaza A, Nida N, Shah MA, Zubair L (2021) Liver tumor segmentation using resnet based mask-R-CNN2021 International Bhurban Conference on Applied Sciences and Technologies (IBCAST), pp 276–281
Anil BC, Dayananda P (2021) Automatic liver tumor segmentation based on multi-level deep convolutional networks and fractal residual network. IETE J Res. https://doi.org/10.1080/03772063.2021.1878066:1-9
Aslam MS, Younas M, Sarwar MU et al (2021) Liver-tumor detection using CNN ResUNet. Comput Mater Continua 67
Dey R, Hong Y (2020) Hybrid cascaded neural network for liver lesion segmentation. In: IEEE 17th International Symposium on Biomedical Imaging (ISBI). IEEE, Iowa, pp 1173–1177
Hamard A, Frandon J, Larbi A et al (2020) Impact of ultra-low dose CT acquisition on semi-automated RECIST tool in the evaluation of malignant focal liver lesions. Diagn Interv Imaging 101:473–479
AmirHosseini B, Hosseini R (2019) An improved fuzzy-differential evolution approach applied to classification of tumors in liver CT scan images. Med Biol Eng Comput 57:2277–2287
Balagourouchetty L, Pragatheeswaran JK, Pottakkat B, G R, (2020) GoogLeNet-based ensemble FCNet classifier for focal liver lesion diagnosis. IEEE J Biomed Health Inform 24:1686–1694
Cao SE, Zhang LQ, Kuang SC et al (2020) Multiphase convolutional dense network for the classification of focal liver lesions on dynamic contrast-enhanced computed tomography. World J Gastroenterol 26:3660–3672
Das A, Acharya UR, Panda SS, Sabut S (2019) Deep learning based liver cancer detection using watershed transform and Gaussian mixture model techniques. Cogn Syst Res 54:165–175
Devi RM, Seenivasagam V (2020) Automatic segmentation and classification of liver tumor from CT image using feature difference and SVM based classifier-soft computing technique. Soft Comput 24:18591–18598
Jiang HY, Zheng RP, Yi DH, Zhao D (2013) A novel multiinstance learning approach for liver cancer recognition on abdominal CT images based on CPSO-SVM and IO. Comput Math Methods Med 2013:10
Jin XY, Zhang T, Li LJ, Wu HT, Sun B (2016) Lesion recognition method of liver CT images based on random forest. In: 8th International Conference on Information Technology in Medicine and Education (ITME). IEEE, Fuzhou, pp 227–230
Kabe GK, Song YQ, Liu Z (2020) Optimization of FireNet for liver lesion classification. Electronics 9:16
Khalili K, Lawlor RL, Pourafkari M et al (2020) Convolutional neural networks versus radiologists in characterization of small hypoattenuating hepatic nodules on CT: a critical diagnostic challenge in staging of colorectal carcinoma. Sci Rep 10:10
Kumar SS, Moni RS, Rajeesh J (2013) An automatic computer-aided diagnosis system for liver tumours on computed tomography images. Comput Electr Eng 39:1516–1526
Kutlu H, Avci E (2019) A novel method for classifying liver and brain tumors using convolutional neural networks, discrete wavelet transform and long short-term memory networks. Sensors 19:16
Yasaka K, Akai H, Abe O, Kiryu S (2018) Deep learning with convolutional neural network for differentiation of liver masses at dynamic contrast-enhanced CT: a preliminary study. Radiology 286:887–896
Sreeja P, Hariharan S (2017) Image analysis for the detection and diagnosis of hepatocellular carcinoma from abdominal CT images. In: International Conference on Internet of Things for Technological Development (IoT4TD). Springer-Verlag Singapore Pte Ltd, Gandhinagar, pp 107–117
Shi WQ, Kuang SC, Cao S et al (2020) Deep learning assisted differentiation of hepatocellular carcinoma from focal liver lesions: choice of four-phase and three-phase CT imaging protocol. Abdom Radiol (NY) 45:2688–2697
Romero FP, Diler A, Bisson-Gregoire G et al (2019) End-to-end discriminative deep network for liver lesion classification. In: 16th IEEE International Symposium on Biomedical Imaging (ISBI). IEEE, Venice, pp 1243–1246
Renukadevi T, Karunakaran S (2020) Optimizing deep belief network parameters using grasshopper algorithm for liver disease classification. Int J Imaging Syst Technol 30:168–184
Rajathi GI, Jiji GW (2019) Chronic liver disease classification using hybrid whale optimization with simulated annealing and ensemble classifier. Symmetry-Basel 11:21
Peng J, Kang S, Ning Z et al (2020) Residual convolutional neural network for predicting response of transarterial chemoembolization in hepatocellular carcinoma from CT imaging. Eur Radiol 30:413–424
Özyurt F, Tuncer T, Avci E, Koç M, Serhatlioğlu İ (2019) A novel liver image classification method using perceptual hash-based convolutional neural network. Arab J Sci Eng 44:3173–3182
Mala K, Sadasivam V, Alagappan S (2015) Neural network based texture analysis of CT images for fatty and cirrhosis liver classification. Appl Soft Comput 32:80–86
Maaref A, Romero FP, Montagnon E et al (2020) Predicting the response to FOLFOX-based chemotherapy regimen from untreated liver metastases on baseline CT: a deep neural network approach. J Digit Imaging 33:937–945
Li J, Sun J, Shen NY, Chen EL, Zhang YC (2019) A CAD system for liver cancer diagnosis based on multi-phase CT images features with BP network. In: 11th International Conference on Intelligent Human-Machine Systems and Cybernetics (IHMSC). IEEE, Zhejiang University, Hangzhou, pp 67–70
Liang D, Lin L, Hu H et al (2018) Combining convolutional and recurrent neural networks for classification of focal liver lesions in multi-phase CT images. In: Frangi AF, Schnabel JA, Davatzikos C, Alberola-López C, Fichtinger G (eds) Medical Image Computing and Computer Assisted Intervention – MICCAI 2018. Springer International Publishing, Cham, pp 666–675
Thuring J, Rippel O, Haarburger C et al (2020) Multiphase CT-based prediction of Child-Pugh classification: a machine learning approach. Eur Radiol Exp 4:9
Wang MY, Fu FF, Zheng BJ et al (2021) Development of an AI system for accurately diagnose hepatocellular carcinoma from computed tomography imaging data. Br J Cancer 125:1111–1121
Wang Q, Wang Z, Sun Y et al (2020) SCCNN: a diagnosis method for hepatocellular carcinoma and intrahepatic cholangiocarcinoma based on Siamese cross contrast neural network. IEEE Access 8:85271–85283
Xu HY, Zou XH, Zhao YN et al (2021) Differentiation of intrahepatic cholangiocarcinoma and hepatic lymphoma based on radiomics and machine learning in contrast-enhanced computer tomography. Technol Cancer Res Treat 20:7
Zhang J, Huang Z, Cao L et al (2020) Differentiation combined hepatocellular and cholangiocarcinoma from intrahepatic cholangiocarcinoma based on radiomics machine learning. Ann Transl Med 8:119
Giannini V, Rosati S, Defeudis A et al (2020) Radiomics predicts response of individual HER2-amplified colorectal cancer liver metastases in patients treated with HER2-targeted therapy. Int J Cancer 147:3215–3223
Homayounieh F, Singh R, Nitiwarangkul C et al (2020) Semiautomatic segmentation and radiomics for dual-energy CT: A pilot study to differentiate benign and malignant hepatic lesions. AJR Am J Roentgenol 215:398–405
Mao B, Zhang LZ, Ning PG et al (2020) Preoperative prediction for pathological grade of hepatocellular carcinoma via machine learning-based radiomics. Eur Radiol 30:6924–6932
Mokrane FZ, Lu L, Vavasseur A et al (2020) Radiomics machine-learning signature for diagnosis of hepatocellular carcinoma in cirrhotic patients with indeterminate liver nodules. Eur Radiol 30:558–570
Budai BK, Tóth A, Borsos P et al (2020) Three-dimensional CT texture analysis of anatomic liver segments can differentiate between low-grade and high-grade fibrosis. BMC Med Imaging 20:108
Huo Y, Terry JG, Wang J et al (2019) Fully automatic liver attenuation estimation combing CNN segmentation and morphological operations. Med Phys 46:3508–3519
Kayaaltı Ö, Aksebzeci BH, Karahan İÖ et al (2014) Liver fibrosis staging using CT image texture analysis and soft computing. Appl Soft Comput 25:399–413
Yasaka K, Akai H, Kunimatsu A, Abe O, Kiryu S (2018) Deep learning for staging liver fibrosis on CT: a pilot study. Eur Radiol 28:4578–4585
Son JH, Lee SS, Lee Y et al (2020) Assessment of liver fibrosis severity using computed tomography-based liver and spleen volumetric indices in patients with chronic liver disease. Eur Radiol 30:3486–3496
Yin Y, Yakar D, Dierckx R, Mouridsen KB, Kwee TC, de Haas RJ (2021) Liver fibrosis staging by deep learning: a visual-based explanation of diagnostic decisions of the model. Eur Radiol 31:9620–9627
Ahmadi K, Karimi A, Fouladi Nia B (2016) New technique for automatic segmentation of blood vessels in CT scan images of liver based on optimized fuzzy c-means method. Comput Math Methods Med 2016:5237191
Ben-Cohen A, Klang E, Amitai MM, Goldberger J, Greenspan H (2018) Anatomical data augmentation for CNN based pixel-wise classification2018 IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018), pp 1096–1099
Conze PH, Noblet V, Rousseau F et al (2017) Scale-adaptive supervoxel-based random forests for liver tumor segmentation in dynamic contrast-enhanced CT scans. Int J Comput Assist Radiol Surg 12:223–233
Gensure RH, Foran DJ, Lee VM et al (2012) Evaluation of hepatic tumor response to yttrium-90 radioembolization therapy using texture signatures generated from contrast-enhanced CT images. Acad Radiol 19:1201–1207
Huang Q, Sun J, Ding H, Wang X, Wang G (2018) Robust liver vessel extraction using 3D U-Net with variant dice loss function. Comput Biol Med 101:153–162
Zeng YZ, Zhao YQ, Liao M, Zou BJ, Wang XF, Wang W (2016) Liver vessel segmentation based on extreme learning machine. Phys Med 32:709–716
Yu W, Fang B, Liu Y, Gao M, Zheng S, Wang Y (2019) Liver vessels segmentation based on 3d residual U-NET2019 IEEE International Conference on Image Processing (ICIP), pp 250–254
Yang W, Lu Z, Yu M, Huang M, Feng Q, Chen W (2012) Content-based retrieval of focal liver lesions using bag-of-visual-words representations of single- and multiphase contrast-enhanced CT images. J Digit Imaging 25:708–719
Wang J, Han XH, Xu Y et al (2017) Sparse codebook model of local structures for retrieval of focal liver lesions using multiphase medical images. Int J Biomed Imaging 2017:1413297
Li Q, Yu B, Tian X, Cui X, Zhang R, Guo Q (2020) Deep residual nets model for staging liver fibrosis on plain CT images. Int J Comput Assist Radiol Surg 15:1399–1406
Sun W, Qin N, Huang D, Liu Z, Ni S (2020) QN-S3VM method for evaluation of liver functional reserve2020 Chinese Automation Congress (CAC), pp 5629–5634
Xu M, Wang Y, Chi Y, Hua X (2020) Training liver vessel segmentation deep neural networks on noisy labels from contrast CT imaging2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI), pp 1552–1555
Yang JZ, Fu MH, Hu Y (2021) Liver vessel segmentation based on inter-scale V-Net. Math Biosci Eng 18:4327–4340
Yoshinobu Y, Iwamoto Y, Han XH et al (2020) Deep learning method for content-based retrieval of focal liver lesions using multiphase contrast-enhanced computer tomography images. In: IEEE International Conference on Consumer Electronics (ICCE). IEEE, Las Vegas, pp 598–601
Gu J, Zhao Z, Zeng Z et al (2020) Multi-phase cross-modal learning for noninvasive gene mutation prediction in hepatocellular carcinoma42nd Annual International Conference of the IEEE-Engineering-in-Medicine-and-Biology-Society (EMBC), Montreal, Canada, pp 5814–5817
Kobe A, Zgraggen J, Messmer F et al (2021) Prediction of treatment response to transarterial radioembolization of liver metastases: radiomics analysis of pre-treatment cone-beam CT: a proof of concept study. Eur J Radiol Open 8:100375
Li X, Qi Z, Du H et al (2022) Deep convolutional neural network for preoperative prediction of microvascular invasion and clinical outcomes in patients with HCCs. Eur Radiol 32:771–782
Ahmad M, Ai DN, Xie GW et al (2019) Deep belief network modeling for automatic liver segmentation. IEEE Access 7:20585–20595
Zhang Y, Peng C, Peng L et al (2022) DeepRecS: from RECIST diameters to precise liver tumor segmentation. IEEE J Biomed Health Inform 26:614–625
Zhang Y, He Z, Zhong C, Zhang Y, Shi Z (2017) Fully convolutional neural network with post-processing methods for automatic liver segmentation from CT2017 Chinese Automation Congress (CAC), pp 3864–3869
Bilic P, Christ P, Li HB et al (2023) The liver tumor segmentation benchmark (LiTS). Med Image Anal 84:102680
Chen XY, Zhang R, Yang PK (2019) Feature fusion encoder decoder network for automatic liver lesion segmentation. In: 16th IEEE International Symposium on Biomedical Imaging (ISBI). IEEE, Venice, pp 430–433
Vivanti R, Joskowicz L, Lev-Cohain N, Ephrat A, Sosna J (2018) Patient-specific and global convolutional neural networks for robust automatic liver tumor delineation in follow-up CT studies. Med Biol Eng Comput 56:1699–1713
Zhou J, Wang W, Lei B et al (2020) Automatic detection and classification of focal liver lesions based on deep convolutional neural networks: a preliminary study. Front Oncol 10:581210
Adcock A, Rubin D, Carlsson G (2014) Classification of hepatic lesions using the matching metric. Comput Vis Image Underst 121:36–42
Liang D, Lin LF, Hu HJ et al (2018) Residual convolutional neural networks with global and local pathways for classification of focal liver lesions. In: 15th Pacific Rim International Conference on Artificial Intelligence (PRICAI) / 15th Pacific Rim Knowledge Acquisition Workshop (PKAW). Springer International Publishing Ag, Nanjing, pp 617–628
Wang W, Chen Q, Iwamoto Y et al (2020) Deep fusion models of multi-phase CT and selected clinical data for preoperative prediction of early recurrence in hepatocellular carcinoma. IEEE Access 8:139212–139220
Zhang L, Xia W, Yan ZP et al (2020) Deep learning predicts overall survival of patients with unresectable hepatocellular carcinoma treated by transarterial chemoembolization plus sorafenib. Front Oncol 10:593292
Wang J, Han XH, Xu Y et al (2017) Tensor sparse representation of temporal features for content-based retrieval of focal liver lesions using multi-phase medical images2017 IEEE International Symposium on Multimedia (ISM), pp 507–510
Group TFMCS, Bedossa P (1994) Intraobserver and interobserver variations in liver biopsy interpretation in patients with chronic hepatitis C. Hepatology 20:15–20
Sterling RK, Lissen E, Clumeck N et al (2006) Development of a simple noninvasive index to predict significant fibrosis in patients with HIV/HCV coinfection. Hepatology 43:1317–1325
WHO (2021) Generating evidence for artificial intelligence-based medical devices: a framework for training, validation and evaluation. World Health Organization WHO.int. Available via https://www.who.int/publications/i/item/9789240038462. Accessed 24.01.2023
Acknowledgements
Infrastructure support for this research was provided by the University Hospital of North Norway and The Arctic University of Norway (UiT).
Guidance and support while writing this manuscript from Professor Arthur Revhaug MD PhD at the Arctic University of Norway (UiT). Arthur.revhaug@uit.no .
Funding
Open access funding provided by UiT The Arctic University of Norway (incl University Hospital of North Norway) The authors state that this work has not received any funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Keyur Radiya.
Conflict of interest
The authors of this manuscript declare no relationships with any companies, whose products or services may be related to the subject matter of the article.
Statistics and biometry
No complex statistical methods were necessary for this paper.
Ethical approval
Institutional Review Board approval was not required because systematic review article and not an experiment.
Methodology
• Systematic review
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Keyur Radiya and Henrik Lykke Joakimsen share first authorship of this review.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Radiya, K., Joakimsen, H.L., Mikalsen, K.Ø. et al. Performance and clinical applicability of machine learning in liver computed tomography imaging: a systematic review. Eur Radiol 33, 6689–6717 (2023). https://doi.org/10.1007/s00330-023-09609-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-023-09609-w