Abstract
Objectives
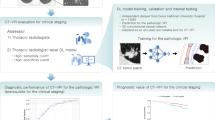
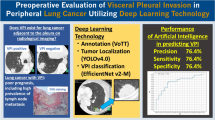
To develop and validate a preoperative CT-based deep learning model for the prediction of visceral pleural invasion (VPI) in early-stage lung cancer.
Methods
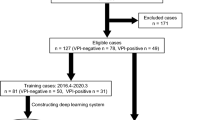
In this retrospective study, dataset 1 (for training, tuning, and internal validation) included 676 patients with clinical stage IA lung adenocarcinomas resected between 2009 and 2015. Dataset 2 (for temporal validation) included 141 patients with clinical stage I adenocarcinomas resected between 2017 and 2018. A CT-based deep learning model was developed for the prediction of VPI and validated in terms of discrimination and calibration. An observer performance study and a multivariable regression analysis were performed.
Results
The area under the receiver operating characteristic curve (AUC) of the model was 0.75 (95% CI, 0.67–0.84), which was comparable to those of board-certified radiologists (AUC, 0.73–0.79; all p > 0.05). The model had a higher standardized partial AUC for a specificity range of 90 to 100% than the radiologists (all p < 0.05). The high sensitivity cutoff (0.245) yielded a sensitivity of 93.8% and a specificity of 31.2%, and the high specificity cutoff (0.448) resulted in a sensitivity of 47.9% and a specificity of 86.0%. Two of the three radiologists provided highly sensitive (93.8% and 97.9%) but not specific (48.4% and 40.9%) diagnoses. The model showed good calibration (p > 0.05), and its output was an independent predictor for VPI (adjusted odds ratio, 1.07; 95% CI, 1.03–1.11; p < 0.001).
Conclusions
The deep learning model demonstrated a radiologist-level performance. The model could achieve either highly sensitive or highly specific diagnoses depending on clinical needs.
Key Points
• The preoperative CT-based deep learning model demonstrated an expert-level diagnostic performance for the presence of visceral pleural invasion in early-stage lung cancer.
• Radiologists had a tendency toward highly sensitive, but not specific diagnoses for the visceral pleural invasion.




Similar content being viewed by others
Abbreviations
- AUC:
-
Area under the receiver operating characteristic curve
- CI:
-
Confidence interval
- IQR:
-
Interquartile range
- NPV:
-
Negative predictive value
- OR:
-
Odds ratio
- PPV:
-
Positive predictive value
- VPI:
-
Visceral pleural invasion
References
Rami-Porta R, Bolejack V, Crowley J et al (2015) The IASLC Lung Cancer Staging Project: proposals for the revisions of the T descriptors in the forthcoming eighth edition of the TNM classification for lung cancer. J Thorac Oncol 10:990–1003
Kamigaichi A, Tsutani Y, Kagimoto A et al (2020) Comparing segmentectomy and lobectomy for clinical stage IA solid-dominant lung cancer measuring 2.1-3 cm. Clin Lung Cancer. https://doi.org/10.1016/j.cllc.2020.04.015
Smith CB, Swanson SJ, Mhango G, Wisnivesky JP (2013) Survival after segmentectomy and wedge resection in stage I non-small-cell lung cancer. J Thorac Oncol 8:73–78
Tsutani Y, Miyata Y, Nakayama H et al (2014) Segmentectomy for clinical stage IA lung adenocarcinoma showing solid dominance on radiology. Eur J Cardiothorac Surg 46:637–642
Zhong C, Fang W, Mao T, Yao F, Chen W, Hu D (2012) Comparison of thoracoscopic segmentectomy and thoracoscopic lobectomy for small-sized stage IA lung cancer. Ann Thorac Surg 94:362–367
Lopez Guerra JL, Gomez DR, Lin SH et al (2013) Risk factors for local and regional recurrence in patients with resected N0-N1 non-small-cell lung cancer, with implications for patient selection for adjuvant radiation therapy. Ann Oncol 24:67–74
Takizawa H, Kondo K, Kawakita N et al (2018) Autofluorescence for the diagnosis of visceral pleural invasion in non-small-cell lung cancer. Eur J Cardiothorac Surg 53:987–992
Mizuno T, Arimura T, Kuroda H, Sakakura N, Yatabe Y, Sakao Y (2018) P2.16-34 visceral pleural invasion is closely associated with nodal spread in cStage IA lung adenocarcinoma. J Thorac Oncol 13:S845
Ahn SY, Park CM, Jeon YK et al (2017) Predictive CT features of visceral pleural invasion by T1-sized peripheral pulmonary adenocarcinomas manifesting as subsolid nodules. AJR Am J Roentgenol 209:561–566
Hsu JS, Han IT, Tsai TH et al (2016) Pleural tags on CT scans to predict visceral pleural invasion of non-small cell lung cancer that does not abut the pleura. Radiology 279:590–596
Imai K, Minamiya Y, Ishiyama K et al (2013) Use of CT to evaluate pleural invasion in non-small cell lung cancer: measurement of the ratio of the interface between tumor and neighboring structures to maximum tumor diameter. Radiology 267:619–626
Yang S, Yang L, Teng L et al (2018) Visceral pleural invasion by pulmonary adenocarcinoma ≤3 cm: the pathological correlation with pleural signs on computed tomography. J Thorac Dis 10:3992–3999
Ehteshami Bejnordi B, Veta M, Johannes van Diest P et al (2017) Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women with breast cancer. JAMA 318:2199–2210
Hwang EJ, Park S, Jin KN et al (2019) Development and validation of a deep learning-based automated detection algorithm for major thoracic diseases on chest radiographs. JAMA Netw Open 2:e191095
Ding Y, Sohn JH, Kawczynski MG et al (2019) A deep learning model to predict a diagnosis of Alzheimer disease by using (18)F-FDG PET of the brain. Radiology 290:456–464
Kim H, Goo JM, Kim YT, Park CM (2019) CT-defined visceral pleural invasion in T1 lung adenocarcinoma: lack of relationship to disease-free survival. Radiology 292:741–749
Moons KG, Altman DG, Reitsma JB et al (2015) Transparent Reporting of a multivariable prediction model for Individual Prognosis or Diagnosis (TRIPOD): explanation and elaboration. Ann Intern Med 162:W1–W73
Goldstraw P, Chansky K, Crowley J et al (2016) The IASLC Lung Cancer Staging Project: proposals for revision of the TNM stage groupings in the forthcoming (eighth) edition of the TNM classification for lung cancer. J Thorac Oncol 11:39–51
Travis WD, Brambilla E, Rami-Porta R et al (2008) Visceral pleural invasion: pathologic criteria and use of elastic stains: proposal for the 7th edition of the TNM classification for lung cancer. J Thorac Oncol 3:1384–1390
Venkatraman ES (2000) A permutation test to compare receiver operating characteristic curves. Biometrics 56:1134–1138
DeLong ER, DeLong DM, Clarke-Pearson DL (1988) Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 44:837–845
McClish DK (1989) Analyzing a portion of the ROC curve. Med Decis Making 9:190–195
McNemar Q (1947) Note on the sampling error of the difference between correlated proportions or percentages. Psychometrika 12:153–157
Leisenring W, Alonzo T, Pepe MS (2000) Comparisons of predictive values of binary medical diagnostic tests for paired designs. Biometrics 56:345–351
Tammemagi MC, Ten Haaf K, Toumazis I et al (2019) Development and validation of a multivariable lung cancer risk prediction model that includes low-dose computed tomography screening results: a secondary analysis of data from the National Lung Screening Trial. JAMA Netw Open 2:e190204
Walsh CG, Sharman K, Hripcsak G (2017) Beyond discrimination: a comparison of calibration methods and clinical usefulness of predictive models of readmission risk. J Biomed Inform 76:9–18
Iizuka S, Kawase A, Oiwa H, Ema T, Shiiya N, Funai K (2019) A risk scoring system for predicting visceral pleural invasion in non-small lung cancer patients. Gen Thorac Cardiovasc Surg 67:876–879
Hsu JS, Jaw TS, Yang CJ et al (2017) Convex border of peripheral non-small cell lung cancer on CT images as a potential indicator of pleural invasion. Medicine (Baltimore) 96:e7323
Brown LM, Louie BE, Jackson N, Farivar AS, Aye RW, Vallieres E (2016) Recurrence and survival after segmentectomy in patients with prior lung resection for early-stage non-small cell lung cancer. Ann Thorac Surg 102:1110–1118
Karimi D, Dou H, Warfield SK, Gholipour A (2019) Deep learning with noisy labels: exploring techniques and remedies in medical image analysis. arXiv preprint arXiv:1912.02911
Johnson JM, Khoshgoftaar TM (2019) Survey on deep learning with class imbalance. J Big Data 6:27. https://doi.org/10.1186/s40537-019-0192-5
Acknowledgments
We sincerely thank Myunghee Lee and Ju Young Jeong for their assistance in the data acquisition.
Funding
This study was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF), funded by the Ministry of Science and ICT (grant number: NRF-2020R1C1C1003684), Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Hyungjin Kim.
Conflict of interest
Activities related to the present article: none.
Activities not related to the present article: H.K. received a research grant from Lunit. E.J.H. and C.M.P. received research grants from Lunit and Coreline Soft. J.M.G. received research grants from Lunit, INFINITT Healthcare, and DONGKOOK Pharmaceutical.
Statistics and biometry
No complex statistical methods were necessary for this paper.
Informed consent
Written informed consent was waived by the Institutional Review Board.
Ethical approval
Institutional Review Board approval was obtained.
Study subjects or cohorts overlap
Some study subjects or cohorts have been previously reported (Radiology. 2019 Sep;292(3):741–749.; Radiology. 2019 Mar;290(3):807–813.; Eur Radiol. 2019 Nov;29(11):6069–6079.; Lung Cancer. 2019 Aug;134:151–157.; J Thorac Oncol. 2018 Dec;13(12):1864–1872.).
Methodology
• retrospective
• diagnostic or prognostic study
• performed at one institution
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 29 kb)
Rights and permissions
About this article
Cite this article
Choi, H., Kim, H., Hong, W. et al. Prediction of visceral pleural invasion in lung cancer on CT: deep learning model achieves a radiologist-level performance with adaptive sensitivity and specificity to clinical needs. Eur Radiol 31, 2866–2876 (2021). https://doi.org/10.1007/s00330-020-07431-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-020-07431-2