Abstract
DNA extracted from faeces may be a valuable source of information about the animal itself, as well as its microflora. An isolated reindeer population from Svalbard (Rangifer tarandus platyrhynchus) was tested for the presence of Shiga toxin encoding genes in the collected faecal samples. Even though the reindeers were not interacting with any other ruminants, which are considered to be a major reservoir of Shiga toxin containing bacteria, the stx1 gene was detected in 9 out of 10 tested samples, and the stx2 subtype c was found in five tested samples. This is an exceptionally high proportion, especially in the case of stx1, when compared to those observed in semi-domesticated or wild cervid populations in less remote locations. Distribution of the investigated genes in a small, local population indicates a different pattern of transmission, which seems to favour bacteria carrying the stx1 genes over those carrying the stx2 genes. The overall high percentage of animals with microbiota containing the stx genes suggests an important role of these genes in such an extreme environment for either hosts or their gut bacteria. Additionally, male-specific DNA found in the faeces was isolated in order to establish a given animal’s sex. PCR based on two pairs of primers, DBY7 and DBY8 gave the expected length of product characteristic for Y chromosome. The results of molecular sexing were consistent with field observations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
DNA extracted from faeces can constitute a source of information about the presence of Shiga toxin produced by Escherichia coli (STEC). The main reservoir for Shiga toxin pathogens is ruminants, mainly cattle, which are their asymptomatic carriers, but they can be also found in other domestic and wild animals (Gyles 2007; Fernández et al. 2009; Franklin et al. 2013). Natural occurrence of the stx (Shiga) genes in deer faeces samples from different parts of the world has been reported (Asakura et al. 1998; Aschfalk et al. 2003; Lillehaug et al. 2005; Kistler et al. 2011; Hofer et al. 2012; Eggert et al. 2013). One of the virulence factors of STEC is Shiga toxins (Stx) encoded on lambdoid prophage genomes (Łoś et al. 2013). These proteins are divided into two major types: Stx1 and Stx2, based on amino acid sequence. The Stx1 protein, produced by STEC, is identical to the Shiga toxin derived from Shigella dysenteriae I. The Stx2 protein is homologous to Stx1 at the level of 55 % for the A subunit and 57 % for the B subunit (Nataro and Kaper 1998). There are several variants of the Stx2 toxin, differing in nucleotide sequences: Stx2a, Stx2b, Stx2c, Stx2d, Stx2e, Stx2f and Stx2g. Strains producing the Stx2a, Stx2c and Stx2d subtypes are often associated with development of haemolytic uremic syndrome (HUS), while other subtypes are less frequently associated with a human disease (Fuller et al. 2011). The STEC strains may encode either only one type of the Shiga toxin, or both types, or different variants of Stx2 (Nataro and Kaper 1998). Bacteria producing the Stx2 toxin are more pathogenic, and an infection caused by them more frequently leads to the development of HUS, contrary to those encoding the Stx1 toxin alone (Law 2000). Shiga toxin causes diseases in humans ranging from mild diarrhoea to HUS, with a mortality rate up to 10 % among young children and the elderly. This syndrome is characterized by acute renal failure, anaemia and thrombocytopenia (Razzaq 2006). Transmission occurs after eating contaminated food or drinking contaminated water, and less than 100 cells may be enough to cause a disease.
Svalbard reindeer (Rangifer tarandus platyrhynchus) is the most isolated subspecies of Rangifer, endemic to Svalbard archipelago in the High Arctic. However, contrary to other Rangifer subspecies, the Svalbard reindeers live individually or in small groups (Alendal et al. 1979). They do not undertake long seasonal migrations, nor are they nomadic within seasons like the mainland reindeer and caribou are (Tyler and Øritsland 1989). Their home ranges are relatively small; the females may remain within 1.5 km of the same spot for several months, or travel on average less than 0.7 km per day during the whole year (Tyler and Øritsland 1989). Some females are philopatric over periods of several years, so the home ranges of daughters could adjoin those of their mothers (Tyler and Øritsland 1990). Outside the mating season, the spatial sexual segregation occurs, calves stay with the females, while the males live separately (Loe et al. 2006).
The Svalbard reindeers show seasonal variation in selection of grazing areas and diet. During the summer, they spend most of their time accumulating fat, particularly feeding on a high-quality vascular plant species growing in the wet areas, in the valleys and lowland plains, often in the vicinity of large seabird colonies (Jakubas et al. 2008). In the winter, when vegetation is of a lower quality and access to it is limited, they forage on the dry and exposed ridges, where little snow accumulation occurs (Bjune 2000).
The Svalbard reindeers are mainly distributed throughout Spitsbergen, i.e. the largest island of the Svalbard archipelago. By 1925, they had almost gone extinct because of over-hunting in the late nineteenth and early twentieth centuries (Tyler 1987). Over recent decades, their population subsequently increased. In Hornsund, the southernmost fiord of Spitsbergen, the reindeer began occurring regularly only since the 1990s (Fossa et al. 2002); however, their local population is still small and does not exceed 20 individuals. The Hornsund area is uninhabited by humans, except for the Polish Polar Station, where nine to ten people spend the whole year. Additionally, during the summer months, small groups of scientists stay in Hornsund for several days or weeks. Visitors coming from cruise ships spend a few hours at this site, taking short walks in a limited area. Among domesticated animals, two outdoor and enchained dogs are kept in the station. The Svalbard reindeer are the only ruminants and representatives of only one of the three wild terrestrial mammal species (including the Arctic fox Vulpes lagopus and amphibious polar bear Ursus maritimus) inhabiting the archipelago (Aanes 2005). Interestingly, the presence of the Shiga toxin genes encoded by STEC has never been investigated in the Svalbard reindeer population.
The aim of this study was to investigate whether the stx gene reservoirs are present in such remote and isolated areas, where contact of indigenous fauna with humans and domesticated animals or other wild mammals is minimal; hence, probability of transmission of bacteria carrying the stx genes from any human-associated or wild mammal-associated reservoir is very limited. To accomplish that, we tested for the presence of the stx1 and stx2 genes in the reindeer faeces originating from the isolated population of the Spitsbergen Island. These results, although limited in scope, may provide useful information about prevalence of the stx-producing microbes in such isolated and severe areas. It is worth noting that presence of the stx genes does not necessarily conclude the presence of the Shiga toxin producing bacteria; however, it can indicate a natural source of these genes. In order to make our inference more complete, we also performed molecular sex identification among the tested reindeer population, targeting male-specific DNA. In case of extensive studies, especially when there is no possibility to determine the sex of individual reindeer by observation, additional knowledge about population parameters may indicate a potential pattern of a particular gene distribution.
Materials and methods
Faeces sample collection
Ten reindeer faecal samples (R1–R10) were collected between July 26th and August 5th of 2013, in the northern part of the Hornsund fiord (SW Spitsbergen; 77°00′N, 15°33′E). The sampling area covered less than 2 km2, between the seashore and the Ariekammen and Skoddefjellet mountains (Fig. 1). Reindeer individuals were being followed until they produced a fresh sample of faeces, which were collected, stored separately in plastic bags, and frozen at −20 °C within max. 30 min of collection time. In order to minimize human interference in the wild population and due to reindeer herd structure, we had assumed a sampling strategy that may result in multiple sample testing. Five samples (R3, R5, R6, R8 and R10) were collected from adult male reindeers living separately. The remaining samples (R1, R2, R4, R7, and R9) were collected in the area where three females and two young individuals (more than 1-year old) were present. However, it was not possible to link the faeces to the individual reindeer in this group due to the tight range of the herd. The individuals seemed to be in good condition; their appearance and behaviour did not reveal any abnormalities. Sex observation was conducted and documented for later comparison with sex identification using PCR methods.
Maps of the study area. Right location of the Hornsund fiord on Spitsbergen. Left close-up of the north part of the Hornsund fiord. Grey shape indicates the reindeer’s grazing area, where individuals were observed and faecal samples (R1–R10) were collected. Black spots indicate soil sampling sites (S1–S2). Map was prepared with the use of CorelDraw
Soil sample collection
Soil sample S1 was collected in the area, where reindeer were observed grazing. Lush vegetation occurring in the area is associated with a large breeding colony of planktivorous seabirds, little auks (Alle alle), which intensively fertilize the adjacent tundra with nutrients from their excrements (Stempniewicz 1990; Jakubas et al. 2008). Sample S1 was collected down the slope, at the seabird colony edge. A second soil sample (S2) was collected in a topographically similar location but not under a routine seabird flight route and hence was not supplied with the said nutrients. This area does not attract reindeer, due to sparse vegetation. In the study area, typical Arctic soils occur, mostly gleysols and regosols, ranging in depth from 15 to 20 cm (Fischet and Skiba 1993). Therefore, soil samples were collected from the upper 15 cm of the soil layer. Collected soil samples were kept separately in plastic bags and frozen at −20 °C within max. 30 min of collection time. Locations of soil sampling spots, adjacent to the coverage area of reindeer population, are marked in Fig. 1. Soil samples tested in this project were collected under a separate project, intended to determine the structure of bacterial communities in soil samples from Spitsbergen.
DNA extraction from faecal samples
After thawing, samples were homogenized using an MP FastPrep-24 Instrument (MP Biomedicals Inc.). Nucleic acids were extracted from 200 mg of faeces by the GeneMATRIX Faecal DNA Purification Kit (Eurx Ltd.) and stored at −20 °C until further use. To avoid cross-contamination of samples, this process was performed with sterile equipment. The quantity and quality of the extracted DNA were evaluated by using Nano Drop spectrophotometer and agarose gel electrophoresis.
DNA extraction from soil
DNA was isolated from soil samples using the FastDNA® SPIN Kit for Soil and the FastPrep® Instrument (MP Biomedicals, Santa Ana, CA) and stored at −20 °C until further use. To avoid cross-contamination of samples, this process was performed with sterile equipment. The quantity and quality of the extracted DNA were evaluated by using Nano Drop spectrophotometer and agarose gel electrophoresis.
Stx gene identification by PCR amplification and sequencing
A PCR reaction for amplification of the stx1 gene was performed with the extracted DNA samples using primers listed in Table 1. Due to low amount of the product obtained during amplification of the stx2 gene, a nested PCR reaction was performed, using two sets of primers presented in Table 1.
The primer sequences (synthesized by Sigma-Aldrich, Germany) and the expected product size for each pair of primers are shown in Table 1. Amplification reactions were performed in the final volume of 25 µl, with the use of SapphireAmp® Fast PCR Master Mix (Takara, Japan). For each reaction, 1 µl of the DNA template was used. All reactions were performed using a thermal cycler (Eppendorf) under the following conditions: 94 °C for 1 min for initial denaturation of the DNA, followed by 40 cycles of 98 °C for 5 s, 66 °C for 5 s and 72 °C for 10 s.
The amplified products were visualized by gel electrophoresis to confirm their specific size and then were sent out for sequencing (Genomed, Poland). Analyses of the obtained data were conducted using the MEGA (version 6) software and aligned using BLASTN (NCBI/BLAST). The Stx2 sequences were also aligned using BLASTP (NCBI/BLAST).
Sex identification by PCR amplification (reindeer male-specific DNA)
The PCR amplification for reindeer sex identification was performed according to a procedure reported by Hellborg and Ellegren (2003) with the use of DBY4, DBY7, DBY8, DBY9, UBE1Y6 and UTY5 primers, constructed for the reindeer (Table 1). All reactions were performed using a thermal cycler (Eppendorf) with touchdown PCR where temperatures from 60 to 50 °C or 55 to 45 °C were applied in the annealing step, according to the protocol described, in order to lower the specificity of primers. The amplification reactions were performed with the use of DreamTaq Green PCR Master Mix (Thermo Scientific), and amplification was performed at least three times for each individual reindeer with each primer pair and each pair of primers.
Results
Faecal samples
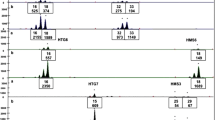
Ten faecal samples from a Svalbard reindeer population collected in the Hornsund fiord were analysed for the presence of the stx1 and stx2 genes. DNA obtained by extraction demonstrated high purity and low degradation for nine samples. Sample R8 was characterized by lower DNA amount and lower purity when compared to other samples. Results for detection of the stx genes in each individual are summarized in Table 2. Comparison of stx1 and stx2 sequences found in reindeer faecal samples is available as supplementary data in Online Resource 1 and Online Resource 2, respectively. Nucleotide sequence analysis of the stx1 gene revealed no differences in sequences among all individuals. Also, the nucleotide sequence analysis of the stx2 gene indicated no differences among individuals. Furthermore, based on protein alignment of the stx2 gene, the sequences obtained are consistent with the c subtype (GenBank accession number ABB36584.1; Online Resource 3). Some of the stx2 gene nucleotides isolated from reindeer faeces differ from the reference sequence of the c subtype, but mutations are located in the third position of the affected codons and consequently do not change the protein sequence.
For stx1, 9 out of 10 samples tested positive. For stx2, 5 out of 10 samples tested positive. For each stx2 positive sample, the stx1 gene was also detected. For four samples, only the stx1 gene was detected, and for one sample neither the stx1 nor stx2 gene was detected. The pattern of positive stx1 detection is not related to the animal’s sex, as 9 out of 10 individuals were positive for stx1. Analogously, there is no clear pattern for occurrence of stx2, as it was detected in 3 out of 5 males. In the R9 sample, no stx gene was detected and this individual is a part of a group of three females and two young individuals.
Molecular sex identification was performed for all reindeer faecal samples tested. In case of all samples, amplification with the DBY4, DBY9, UBE1Y6 and UTY5 forward and revers primers did not yield the expected product. The failure of Y-specific amplification was due to the occurrence of non-specific amplification, and these were excluded from further analysis. For the DBY7 primers, we obtained specific products only for five of all the tested individuals (Table 2). For the R8 individual, no amplification product was visible except for one reproducible faint product for the DBY7 amplification. For the DBY8 primer, in 7 samples, we obtained a product of about 200 bp in length, as expected. In the R4, R8 and R9 samples, we did not obtain any product (Table 2) for the DBY8 amplification. The reindeer sex assessment based on molecular technics, relying on the results for two pairs of primers, DBY7 and DBY8 (which gave the expected product size), was consistent with field observations for the R3, R5, R6 and R10 individuals. Our data also indicated that the R2 individual (assigned as a young reindeer of unknown sex or adult female) was in fact a male. DBY8 primer pairs also indicated that the R1 and R7 individuals were males as well (Table 2).
Soil samples
PCR reactions for soil DNA samples were performed, and the size of the obtained amplicons was positively verified on an agarose gel. The S1 sample was negative for both, the stx1 and stx2 genes; however, sample S2 was positive for stx1, but not stx2. Sequencing confirmed the presence of the stx1 gene in the S2 soil sample. The nucleotide sequence analysis of this stx1 gene revealed no differences between DNA obtained from the reindeer faecal samples and the sequence of DNA extracted from the soil sample (Online Resource 1).
Discussion
To our knowledge, this is the first study denoting the presence of Shiga toxin genes in Svalbard reindeer. In this study, we detected the stx1 gene in 90 % of the tested faecal samples and in 50 % of the tested samples the stx2 gene was determined to be present, which is a considerably high percentage of detection frequency when compared to the Norwegian reindeer (Rangifer tarandus tarandus; Aschfalk et al. 2003) or a white-tailed deer (Odocoileus virginianus) from Louisiana (Dunn et al. 2004) and Pennsylvania (Kistler et al. 2011). Lillehaug et al. (2005) reported that no E. coli O157 isolates, both Shiga toxins producing as well as not Shiga toxins producing, were observed in four different wild cervid species: red deer (Cervus elaphus), roe deer (Capreolus capreolus), moose (Alces alces) and reindeer from the central Norway. In Pennsylvanian white-tailed deer, the presence of stx2 was detected at a similar percentage (46 %) as in the study presented here; however, the number of the tested samples was five times higher and was based on a different sampling method. On the other hand, stx1, which was detected in majority of the tested faecal samples, was detected only in 10 % of the Pennsylvanian deer samples, but every sample positive for stx1 was also positive for stx2 (Kistler et al. 2011). In an extensive study of four different species of free-living ruminants, including red deer, roe deer, fallow deer Dama dama and mouflon Ovis musimon in the south-western Spain, both stx genes were detected in 15.4 % of samples (Sánchez et al. 2009). In that study, the presence of both genes was observed in half of the tested samples. As already mentioned, bacteria producing the Stx2 toxin are more commonly associated with infections and more frequently lead to the HUS development in humans, contrary to those individuals infected only with the strains producing Stx1 toxin only (Law 2000). Therefore, a broad distribution of the stx1 gene in the tested Svalbard reindeer population and the presence of the stx2 gene in only half of the tested samples result in a lower pathogenic potential than was found in the other studies, where the presence of the stx2 gene was much higher than that of the stx1 gene.
A high frequency of occurrence of the stx genes in the studied population might be explained by conditions facilitating the transmission of faecal bacterial flora between the individuals remaining in close proximity, when utilizing small pasture area during the summer (during one week of sample collection, the reindeer were observed grazing within less than 2 km2). Also, during the winter, reindeer aggregate in small ice-free areas, where they may find food (Alendal and Byrkjedal 1974; Tyler and Øritsland 1989). Moreover, the philopatry in females (a tendency of females to stay in the mothers’ area, Tyler and Øritsland 1990), as well as the fact that the reindeer is a polygynous species (males mate with many females), increases the possibility of stx transmission. Ogden et al. (2004) showed that among seasonally housed cattle, transmission of E. coli O157 with virulence markers is increasing due to crowding during the winter months. Similarly in the Svalbard reindeer, mating season and gathering in snow-free areas during the winter, may favour the stx gene horizontal transfer. Fernández et al. (2009) are suggesting that the type of the stx gene is not only influenced by the age of the cattle, but also by the season in which samples were collected. In this study, no stx sequence was detected in sample 9, which originated from either a young individual with unknown sex or an adult female. This lack of the stx genes could be a result of a young age of the individual, which had not yet gained a particular gut flora, as the other individuals from the tested herd. Unfortunately, we could not indicate with certainty which samples originated from the young individuals and which are from adult females, as during sampling we could not relate faecal samples to individuals of that herd with satisfactory probability.
In amplification targeting the male-specific DNA, we were not able to confirm with certainty if the lack of amplification product for DBY8 and multiple products obtained with the DBY7 primers for the R4, R8, and R9 samples were due to a lower quality of the extracted DNA, an amplification error or simply due to lack of the Y chromosome. As amplification reactions were repeated at least three times for each individual, we strongly suggest that individuals R4 and R9 were females, which is consistent with the observations carried out in the field while sampling. In the case of the R8 sample, also because of a direct identification during sampling and due to one positive reaction with the DBY7 primers, we rather suggest that lack of the amplification product, not only for the DBY8 primers, but for all tested primers, is due to a lower quality and lower quantity of the obtained DNA. Also, as we typed the R9 individual as a female, we cannot indicate any reasonable argument of lack of any of the stx genes in that particular sample, and additionally we cannot confirm nor exclude the young age of that individual. Moreover, this particular individual was characterized by a bit different bacterial species structure than the rest of the individuals, although no statistically significant differences were found at the high taxonomic ranks in the faecal bacterial composition between particular reindeer individuals (Zielińska et al. 2016), which may be the result of a young age of that individual. Additionally, these analyses indicate that the primers for DBY7 and DBY8 can be used in the future studies as effective molecular markers in sex identification for the Svalbard reindeer. Barbosa et al. (2009) published a report that after amplification of DNA from the red deer with the use of the DBY8 primers, one should obtain a 200 bp product which is characteristic for males, and a 350 bp product characteristic for females. In this study, after amplification with the DBY8 primers, we obtained only a 200 bp length product (in seven tested samples, Table 2). Taken together, it indicates that although the reported marker is allowing identification of both females and males of the red deer, it is not suitable for reindeer sample assessment. However, due to a small size of the tested samples, such an analysis should be performed on a larger set of samples, with both male and female origin. In this study, based on molecular sex identification targeting male-specific DNA, we were able to confirm with two different pairs of primers that there were two males among a tight reindeer group, and with the use of just one pair of primers, we confirmed one male, which can suggest that samples from one individual may have been tested multiple times. Therefore, due to one season sample collection, the small sample size and uncertainty with regard to the age and sex of some individuals, as well as possible multiple sampling from the same individual, we cannot predict any pattern of the stx gene occurrence related to the sex or age of the animals. Thus, based on this study any well-documented sex or age distribution pattern that could imply any stx gene distribution cannot be indicated.
For reindeer faecal samples tested in this study, the stx1 gene was detected more frequently in comparison with stx2, which may suggest that microbes that carry stx1 are dominant. However, in many studies regarding ruminants, stx2 was shown to be detected more often than stx1 (Sánchez et al. 2009; Kistler et al. 2011). In contrast, only the stx1 gene was detected in samples taken from a reindeer cadaver found in Norway, but no association with mortality was established (Aschfalk et al. 2003). Additionally, the mechanisms of action of both stx genes are believed to be alike, but cytotoxicity of stx2 may be responsible for more serious response in humans. It has been shown previously that the stx2c subtype detected in this study, as well as stx2a or stx2d, is more often associated with HUS and display a subtle difference in receptor preference (Persson et al. 2007; Fuller et al. 2011). There is no information available about any symptoms of the STEC infections among people visiting or living in the Polish Polar Station. Despite that the polar environment characterizes with a minimal human interference, the natural occurrence and distribution of the Stx2c subtype are not incontestable. In a polar environment, almost untouched by humans, could be a reservoir potentially dangerous to humans visiting this area. The food brought by people, dogs living in the Polar Station, livestock or simply contamination carried, e.g. on shoe soles, could be some of the potential sources of the stx genes in the tested area. Also, other wild mammals or birds breeding in Svalbard may constitute a natural reservoir of the stx genes. Reindeer may utilize droppings of barnacle geese (Branta leucopsis) as an alternative food source (Van der Wal and Loonen 1998) and also ingest soil near the Arctic fox dens, probably to supplement their diets with microelements from prey remains (Stempniewicz and Iliszko 2010). It is also possible that the stx genes were transferred to Svalbard with cattle and pigs, which were bred until the 90s in Barentsburg, the second (after Longyearbyen) largest settlement on Svalbard, 150–200 km away from Hornsund (Umbreit 2005).
Limited soil sample size analysis in this study does not allow for presenting any pattern of the stx gene distribution in Arctic environment. This additional stx gene analysis in already acquired soil samples collected in the course of another study was performed in order to enrich inference or partially determine the distribution pattern in reindeer environment. However, the presence of the stx1 gene in sample S2, localized away from the routine flight route of the seabirds, may suggest a natural presence of the stx genes in the Arctic environment without a clear pattern attributed to a zoonotic transmission. Moreover, the extraction method applied might be insufficient for some types of bacteria, and additionally soil texture may induce underrepresentation of some types of bacteria. In the future, a more complex analysis should be performed with a larger number of samples and utilizing several DNA extraction methods in order to improve the analysis and facilitate inference about the stx gene distribution in the polar environment.
In case of DNA extracted from faeces, it can be a very valuable source of information about endemic population of the Svalbard reindeer, not only about microbiological aspects, like bacterial flora occurring in the digestive tract of reindeer (Zielińska et al. 2016), but also about the stx genes as potential pathogenic factors or establishing the gender of reindeer through male-specific DNA amplification. This is, to our knowledge, the first report showing the presence of the stx genes in faecal samples of this High Arctic cervid species. Although we studied limited number of samples, it was enough to prove the occurrence of the stx genes in wild reindeer population inhabiting this remote area almost untouched by humans. However, our study is limited to the Hornsund area, and thus the distribution of the stx genes occurrence in Svalbard requires further studies. The fact that a small, isolated and located so far north cervid population carries the stx genes in their faecal flora may suggest a very broad distribution of bacteria bearing these genes among various wild vertebrate populations of the remote polar regions.
References
Aanes R (2005) Svalbard reindeer. In: Kovacs KM (ed) Bird and mammals of Svalbard. Norvegian Polar Institute, Tromso, pp 74–76
Alendal E, Byrkjedal I (1974) Population size and reproduction of the reindeer (Rangifer tarandus platyrhynchus) on Nordenskiold Land, Svalbard. Norsk Polarinst Årbok 2:139–152
Alendal E, de Bie S, van Wieren SE (1979) Size and composition of the wild reindeer (Rangifer tarandus platyrhynchus) population in the Southeast Svalbard Nature Reserve. Holarct Ecol 2:101–107. doi:10.1111/j.1600-0587.1979.tb00687.x
Asakura H, Makino S, Shirahata T, Tsukamoto T, Kurazono H, Ikeda T, Takeshi K (1998) Detection and genetical characterization of Shiga toxin-producing Escherichia coli from wild deer. Microbiol Immunol 42:815–822
Aschfalk A, Kemper N, Höller C (2003) Bacteria of pathogenic importance in faeces from cadavers of free-ranging or corralled semi-domesticated reindeer in northern Norway. Vet Res Commun 27:93–100
Barbosa AM, Fernandez-Garcia JL, Carranza J (2009) A new marker for rapid sex identification of red deer (Cervus elaphus). Hystrix Ital J Mammal 20:169–172
Bjune AE (2000) Pollen analysis of faeces as a method of demonstrating seasonal variations in the diet of Svalbard reindeer (Rangifer tarandus platyrhynchus). Polar Res 19:192. doi:10.1111/j.1751-8369.2000.tb00342.x
Dunn JR, Keen JE, Moreland D, Thompson RA (2004) Prevalence of Escherichia coli O157: H7 in White-tailed Deer from Louisiana. J Wildl Dis 40:361–365. doi:10.7589/0090-3558-40.2.361
Eggert M, Stüber E, Heurich M, Fredriksson-Ahomaa M, Burgos Y, Beutin L, Märtlbauer E (2013) Detection and characterization of Shiga toxin-producing Escherichia coli in faeces and lymphatic tissue of free-ranging deer. Epidemiol Infect 14:251–259. doi:10.1017/S0950268812000246
Fernández D, Rodríguez EM, Arroyo GH, Padola NL, Parma AE (2009) Seasonal variation of Shiga toxin-encoding genes (stx) and detection of E. coli O157 in dairy cattle from Argentina. J Appl Microbiol 106:1260–1267. doi:10.1111/j.1365-2672.2008.04088.x
Fischet Z, Skiba S (1993) Some remarks about bioenergetic aspects of tundra soil. Pol Polar Res 14:345–355
Fossa AM, Nybakken L, Oht M (2002) Vegetation and environmental relations along the western coast of Spitsbergen, Svalbard. In: Jónsdóttir IS (ed) Biodiversity in arctic plant communities. UNIS Publication Series. AB-306 reports 8-21
Franklin AB, Vercauteren KC, Maguire H, Cichon MK, Fischer JW, Lavelle MJ et al (2013) Wild ungulates as disseminators of Shiga toxin-producing Escherichia coli in urban areas. PLoS One. doi:10.1371/journal.pone.0081512
Fuller CA, Pellino CA, Flagler MJ, Strasser JE, Weiss AA (2011) Shiga toxin subtypes display dramatic differences in potency. Infect Immun 79:1329–1337. doi:10.1128/IAI.01182-10
Gyles CL (2007) Shiga toxin-producing Escherichia coli: an overview. J Anim Sci 85:45–62. doi:10.2527/jas.2006-508
Hellborg L, Ellegren YH (2003) Chromosome conserved anchored tagged sequences (YCATS) for the analysis of mammalian male-specific DNA. Mol Ecol. doi:10.1046/j.1365-294X.2003.01702.x
Hofer E, Cernela N, Stephan R (2012) Shiga toxin subtypes associated with Shiga toxin-producing Escherichia coli strains isolated from red deer, roe deer, chamois, and ibex. Foodborne Pathog Dis 9:792–795. doi:10.1089/fpd.2012.1156
Jakubas D, Zmudczynska K, Wojczulanis-Jakubas K, Stempniewicz L (2008) Faeces deposition and numbers of vertebrate herbivores in the vicinity of planktivorous and piscivorous seabird colonies in Hornsund, Spitsbergen. Pol Polar Res 29:45–58
Kistler WA, Mulugeta S, Mauro SA (2011) Detection of stx1 and stx2 genes in Pennsylvanian white-tailed deer. Toxins 3:640–646. doi:10.3390/toxins3060640
Law D (2000) Virulence factors of Escherichia coli O157 and other Shiga toxin-producing E. coli. J Appl Microbiol 88:729–745
Lillehaug A, Bergsjø B, Schau J, Bruheim T, Vikøren T, Handeland K (2005) Campylobacter spp., Salmonella spp., Verocytotoxic Escherichia coli, and Antibiotic Resistance in Indicator Organisms in Wild Cervids. Acta Vet Scand 46:23–32
Loe LE, Irvine RJ, Bonenfant C, Stien A, Langvatn R, Albon SD, Mysterud A (2006) Testing five hypotheses of sexual segregation in an arctic ungulate. J Anim Ecol. doi:10.1111/j.1365-2656.2006.01069.x
Łoś M, Łoś JM, Węgrzyn G (2007) Rapid identification of Shiga toxin-producing Escherichia coli (STEC) using electric biochips. Diag Mol Pathol. doi:10.1097/PDM.0b013e31815a5127
Łoś JM, Łoś M, Węgrzyn A, Węgrzyn G (2013) Altruism of Shiga toxin-producing Escherichia coli: recent hypothesis versus experimental results. Front Cell Infect Microbiol. doi:10.3389/fcimb.2012.00166
Nataro JP, Kaper JB (1998) Diarrheagenic Escherichia coli. Clin Microbiol Res 11:142–403
Ogden ID, Mac Rae M, Strachan NJC (2004) Is the prevalence and shedding concentrations of E. coli O157 in beef cattle in Scotland seasonal? FEMS Microbiol Lett 233:297–300
Persson S, Olsen KEP, Ethelberg S, Scheutz F (2007) Subtyping method for Escherichia coli Shiga Toxin (Verocytotoxin) 2 variants and correlations to clinical manifestations. J Clin Microbiol. doi:10.1128/JCM.02591-06
Razzaq S (2006) Hemolytic uremic syndrome: an emerging health risk. Am Fam Physician 74:991–996
Sánchez S, García-Sánchez A, Martínez R, Blanco J, Blanco JE, Blanco M et al (2009) Detection and characterisation of Shiga toxin-producing Escherichia coli other than Escherichia coli O157:H7 in wild ruminants. Vet J. doi:10.1016/j.tvjl.2008.01.011
Stempniewicz L (1990) Biomass of Dovekie excreta in the vicinity of a breeding colony. Colon Waterbird. doi:10.2307/1521421
Stempniewicz L, Iliszko L (2010) Glaucous gulls kleptoparasiting arctic foxes in Magdalenefiorden, NW Spitsbergen. Arctic. doi:10.14430/arctic651
Tyler NJC (1987) Natural limitation of the abundance of the High Arctic Svalbard reindeer. PhD Thesis, University of Cambridge, Cambridge
Tyler NJC, Øritsland NA (1989) Why don’t Svalbard reindeer migrate? Holarct Ecol 12:369–376. doi:10.1111/j.1600-0587.1989.tb00911.x
Tyler NJC, Øritsland NA (1990) Home ranges in Svalbard reindeer. Rangifer. doi:10.7557/2.10.3.846
Umbreit A (2005) Spitsbergen: Svalbard, Franz Josef Land, Jan Mayen. Chalfont St PeterBradt Travel Guides, 246
Van der Wal R, Loonen MJ (1998) Goose droppings as food for reindeer. Can J Zool. doi:10.1139/z98-033
Zielińska S, Kidawa D, Stempniewicz L, Łoś M, Łoś JM (2016) New insights into the microbiota of the Svalbard reindeer Rangifer tarandus platyrhynchus. Front Microbiol. doi:10.3389/fmicb.2016.00170
Acknowledgments
We are grateful to Mateusz Barcikowski for participation in the sample collection in the field. Collection of the reindeer faecal samples was possible due to a financial support from the National Science Centre (Poland) based on the decision (DEC-2011/01/N/NZ8/04569). All fieldwork has been performed under permission of the Governor of Svalbard. This work was in a large part supported by the National Science Centre, Poland (Grant No. 2011/01/D/NZ2/04817) and Ministry of Science and Higher Education, Poland (Grant No. 0312/IP1/2011/71). We are also grateful to Michalina Filipiak for providing primers. We thank Klaudia Milewska and Tomasz Gąsior for their invaluable help with sequence analysis.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Online Resource 1
Comparison of the stx1 gene sequence from reindeer samples (R1–R8, R10) and soil sample (S2) to the reference sequence AE005174.2. There are no differences in nucleotide sequences among the reindeer samples, the soil sample and the reference sequence. (PDF 402 kb)
Online Resource 2
Comparison of the stx2 gene sequences from reindeer samples (R1, R3, R6, R7, R10) to reference sequences CP008957.1 and DQ235774.1/ABB36584.1 (characteristic for the c subtype). There are no differences in nucleotide sequences among the reindeer samples and the reference sequences CP008957.1. Nucleotide differences between the reindeer samples and the reference sequence of the c subtype are shown in colour. (PDF 694 kb)
Online Resource 3
Comparison of the protein sequence of Stx2 from reindeer samples (R1, R3, R6, R7, R10) to the reference sequence DQ235774.1/ABB36584.1 (characteristic for the c subtype). There are no differences in the protein sequence among the reindeer samples and the reference sequences. (PDF 346 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Zielińska, S., Kidawa, D., Stempniewicz, L. et al. DNA extracted from faeces as a source of information about endemic reindeer from the High Arctic: detection of Shiga toxin genes and the analysis of reindeer male-specific DNA. Polar Biol 40, 659–666 (2017). https://doi.org/10.1007/s00300-016-1990-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00300-016-1990-2