Abstract
Key message
Recently published high-quality reference genome assemblies indicate that, in addition to RDR1-deficiency, the loss of several key RNA silencing-associated genes may contribute to the hypersusceptibility of Nicotiana benthamiana to viruses.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Antiviral RNA silencing is the primary defense measure against viruses in plants (Lopez-Gomollon and Baulcombe 2022). Its main steps have been clarified in recent years and can be summarized as follows. The pathway is triggered by virus-derived double-stranded RNAs (dsRNAs) produced by virus-encoded RNA-dependent RNA polymerases (RDRs). These dsRNAs are processed by RNase III-like enzymes (DCLs), resulting in 21–24 nt long primary viral small interfering RNA duplexes (vsiRNAs). The process is assisted by double-stranded RNA-binding proteins such as DRB4 and possibly DRB3 and DRB2 (Qu et al. 2008; Raja et al. 2014; Barton et al. 2017; Fátyol et al. 2020; Incarbone et al. 2021). RNA silencing is typically accompanied by an amplification step catalyzed by host-encoded RDRs (mainly RDR1 and RDR6), which synthesize dsRNAs using aberrant single-stranded (ss) viral RNAs as templates. These dsRNAs are also processed by DCL-DRB complexes, yielding secondary vsiRNA duplexes. Eventually, one strand of the vsiRNA duplex is incorporated into an Argonaute (AGO) protein, resulting in an RNA-induced silencing complex (RISC). Antiviral RISCs are able to limit the replication of invading viruses in a sequence-specific manner by RNA cleavage or translational repression. As a highly effective countermeasure however, viruses produce proteins that, in addition to their canonical roles in the viral life cycle, are capable of suppressing antiviral RNA silencing in various steps (viral suppressors of RNA silencing, VSRs) (Ding 2023).
Studies based on the extensive genetic resources of the model plant Arabidopsis thaliana have played an instrumental role in clarifying the details of antiviral RNA silencing. Additionally, research using the LAB strain of Nicotiana benthamiana has contributed greatly to understanding the molecular arms race between plants and viruses (Bally et al. 2018). This strain has a 72 bp insertion in its RDR1 gene, that prematurely terminates the gene’s open reading frame. It is generally assumed that this dysfunction affects the plant’s response to viral infection, and most importantly, makes the plant susceptible to more than 500 different viruses. Curiously, however, Ying et al. found that supplementing the LAB strain with RDR1 from Nicotiana tabacum does not lead to increased resistance, but rather enhances virus infection (Ying et al. 2010). Thus, the authors suggested that RDR1 could be a negative regulator of RDR6 and that its inactivation in the LAB strain was likely due to high viral pressure to enhance RDR6's antiviral role. In the latter scenario, RDR1 deactivation is a consequence rather than a cause of the plant’s greater susceptibility to viruses. Either way, the findings of two recent studies reporting high-quality chromosome-level reference genome assemblies of N. benthamiana and N. tabacum are compatible with both hypotheses (Ranakawa et al. 2023; Wang et al. 2024). Comparative analysis of the two genomes revealed, that in addition to having a previously reported defect in RDR1, several additional key silencing genes were lost from one or both subgenomes of N. benthamiana, while both homeologs were retained in N. tabacum. The purpose of this Focus article is to link these findings to other recent studies on N. benthamiana’s RNA silencing-associated genes.
Secondary vsiRNAs are key to effective antiviral protection. Due to the aforementioned mutation of the RDR1 gene, their production in the LAB strain of N. benthamiana is RDR6-dependent. The function of RDR6 in N. benthamiana has been studied primarily in a plant line, in which the gene was silenced with a short-hairpin RNA (Schwach et al. 2005). Although the use of such a system provides invaluable information about gene function in many contexts, its application to study host-virus interactions can be problematic because viruses use various measures to compromise RNA silencing, which can also reduce the effectiveness of gene knock-down. To circumvent this potential problem, previously we created a bona fide rdr6 mutant of N. benthamiana, using CRISPR/Cas9 genome editing (Ludman and Fátyol 2019). In the plant’s genome two potential RDR6-like ORFs could be identified. However, our results demonstrated that one of these alone can provide the RDR6 activity necessary for secondary vsiRNA production. These findings are consistent with the two reports mentioned above, which show that the genome of N. benthamiana contains a single functional RDR6 gene and an RDR6 pseudogene (Ranakawa et al. 2023; Wang et al. 2024).
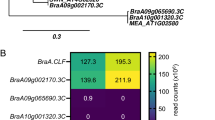
In addition to regulating host gene expression AGO1 has been consistently implicated in antiviral protection (Silva-Martins et al. 2020). In the new reports four AGO1 homeologues—two AGO1A (Nbe10g25940 and Nbe02g25990) and two AGO1B (Nbe06g21200 and Nbe05g23300)—were annotated in the N. benthamiana genome, a number lower than that in N. tabacum, which has six of them (Ranakawa et al. 2023; Wang et al. 2024). The functions of the N. benthamiana AGO1 homeologues were previously studied in plants, where they were selectively inactivated using CRISPR/Cas9 (Ludman and Fátyol 2021). Interestingly, inactivation of only one of the AGO1A or AGO1B homeologues was sufficient to produce distinct phenotypes in the plants. The ago1a mutants developed normally but were hypersusceptible to TCV infection. AGO1B deficiency, however, was incompatible with normal development as homozygous ago1b plants could not be obtained, indicating embryonic lethality. Even heterozygotes exhibited severe leaf malformations and were almost completely sterile. Nevertheless, these heterozygotes were hypersusceptible to TCV infection, indicating that AGO1B is also involved in antiviral protection. The two newly annotated AGO1 homeologues bear high overall similarity to those previously inactivated by genome editing, but both carry deletions in their N-terminal domains (Fig. S1). Although less conserved than the rest of the protein, the N terminal region of AGOs is implicated in a number of important functions (Martín-Merchán et al. 2023). Whether the aforementioned deletions affect the activities of the two newly annotated AGO1 homeologues remains to be determined.
AGO2 is involved in various stress responses in plants, including antiviral protection. This was conclusively proven more than 10 years ago by demonstrating that the ago2 mutant A. thaliana is hypersusceptible to TCV and CMV infections (Harvey et al. 2011). Later, the critical role of the N. benthamiana AGO2 orthologue in limiting virus infections was also corroborated using genome-edited ago2 mutant plants (Ludman et al. 2017). Inactivating two alleles of a single AGO2 gene increased the susceptibility of the plant to a number of different viruses, providing evidence for the existence of one functional AGO2 gene, despite the plant's allotetraploid genome. This finding has now been also confirmed by the papers of Wang et al. (2024) and Ranakawa et al. (2023).
The investigation of the antiviral role of N. benthamiana AGO5 has been the subject of two recent reports, both of which employed CRISPR/Cas9 genome editing to inactivate the gene (Ludman et al. 2023; Tu et al. 2023). The AGO5 gene (Nbe10g12070 in Wang et al. 2024) encoding the 978 aa long AGO5 protein was inactivated in both studies, resulting in increased susceptibility to several viruses. Consistent with the allotetraploid nature of N. benthamiana, a second related AGO5-like ORF (Nbe02g30110 in Wang et al. 2024) can also be retrieved from the plant’s genome. This ORF encodes a protein of 919 aa, which overall exhibits 86.6% identity and 88% similarity to the AGO5 protein encoded by Nbe10g12070. Importantly however, this AGO5 variant carries a deletion that removes the N-terminal 56 aa segment of its PIWI domain, including the first catalytically essential aspartic acid (Fig. S2). Consequently, this protein is most likely dysfunctional, which is consistent with the finding that a mutation in Nbe10g12070 alone is sufficient to increase the plant’s susceptibility to viral infections.
In addition to the silencing-related genes discussed thus far, one copy of each of DCL2 DCL3 and DRB4 genes present in N. tabacum has been lost in N. benthamiana, and the number of AGO4-like genes is also lower in this species than in N. tabacum (Ranakawa et al. 2023; Wang et al. 2024). Because the hierarchical actions of multiple AGOs relying on both primary and secondary vsiRNAs, are needed to establish potent antiviral resistance (Ludman et al. 2023), all of the above deficiencies conceivably contribute to N. benthamiana’s hypersusceptibility to viruses (Fig. 1). In this context, an interesting question arises as to why during the evolution of N. benthamiana, genes associated with RNA silencing appeared to have been preferentially lost. Although there is no definitive answer yet, based on existing data, two non-exclusive hypotheses can be formulated as possible explanations. In the Suaveolentes section of the genus Nicotiana, to which N. benthamiana belongs, allopolyploidization was likely key to adapting to Australia's harshest climatic and ecological regions (Ranakawa et al. 2023). RNA silencing appears to be particularly important in relation to the genomic instability that often accompanies allopolyploid hybridization, both as cause and effect (Comai 2000). Alternatively, or perhaps complementarily to the above, it has also been suggested that impairing the plant’s viral defenses leads to metabolic changes that promotes early vigor and, consequently, more successful adaptation to adverse environmental conditions (Bally et al. 2015). Although the details are obviously sketchy, both of the above could have significant but difficult to predict consequences for the evolution of the plant’s RNA silencing apparatus.
Annotated homeologues of key antiviral RNA silencing genes in the N. tabacum and N. benthamiana genomes. The genes were placed within the framework of the canonical antiviral RNA silencing pathway. Known pseudogenes were omitted. The gene identifiers used are from Wang et al. 2024. The RDR1 gene of the widely used LAB strain of N. benthamiana carries an inactivating 72 bp insertion and is therefore considered pseudogene in this strain (highlighted in red). N. benthamiana’s antiviral silencing genes, for which mutants have already been generated through genome editing, are highlighted in green (color figure online)
In any case, the impaired silencing is at least one of the factors that underlies N. benthamiana’s unparalleled susceptibility to viruses, thereby making it an exceptional model system for studying their interactions with the host. Additionally, in recent years, N. benthamiana has increasingly become the manufacturing platform of choice for molecular pharming. Among other factors, weak RNA silencing, which allows for efficient protein production, also contributes to the plant’s popularity. Further improvement of this and other beneficial properties of N. benthamiana can now rely not only on rapidly evolving genome editing technologies but also on the genomic resources reported in the studies of Wang et al. (2024) and Ranakawa et al. (2023).
Data availability
All data supporting the findings of this study are available within the paper and within its supplementary materials published online.
References
Bally J, Nakasugi K, Jia F, Jung H, Ho SY, Wong M, Paul CM, Naim F, Wood CC, Crowhurst RN, Hellens RP, Dale JL, Waterhouse PM (2015) The extremophile Nicotiana benthamiana has traded viral defence for early vigour. Nat Plants 1:15165
Bally J, Jung H, Mortimer C, Naim F, Philips JG, Hellens R, Bombarely A, Goodin MM, Waterhouse PM (2018) The rise and rise of Nicotiana benthamiana: a plant for all reasons. Annu Rev Phytopathol 56:405–426
Barton DA, Roovers EF, Gouil Q, da Fonseca GC, Reis RS, Jackson C, Overall RL, Fusaro AF, Waterhouse PM (2017) Live cell imaging reveals the relocation of dsRNA binding proteins upon viral infection. Mol Plant Microbe Interact 30(6):435–443
Comai L (2000) Genetic and epigenetic interactions in allopolyploid plants. Plant Mol Biol 43(2–3):387–399
Ding SW (2023) Transgene silencing, RNA interference, and the antiviral defense mechanism directed by small interfering RNAs. Phytopathology 113(4):616–625
Fátyol K, Fekete KA, Ludman M (2020) Double-stranded-RNA-binding protein 2 participates in antiviral defense. J Virol 94(11):e00017-20
Harvey JJ, Lewsey MG, Patel K, Westwood J, Heimstädt S, Carr JP, Baulcombe DC (2011) An antiviral defense role of AGO2 in plants. PLoS ONE 6(1):e14639
Incarbone M, Clavel M, Monsion B, Kuhn L, Scheer H, Vantard É, Poignavent V, Dunoyer P, Genschik P, Ritzenthaler C (2021) Immunocapture of dsRNA-bound proteins provides insight into tobacco rattle virus replication complexes and reveals arabidopsis DRB2 to be a wide-spectrum antiviral effector. Plant Cell 33(11):3402–3420
Lopez-Gomollon S, Baulcombe DC (2022) Roles of RNA silencing in viral and non-viral plant immunity and in the crosstalk between disease resistance systems. Nat Rev Mol Cell Biol 23:645–662
Ludman M, Burgyán J, Fátyol K (2017) Crispr/Cas9 mediated inactivation of argonaute 2 reveals its differential involvement in antiviral responses. Sci Rep 7(1):1010
Ludman M, Fátyol K (2019) The virological model plant, Nicotiana benthamiana expresses a single functional RDR6 homeolog. Virology 537:143–148
Ludman M, Fátyol K (2021) Targeted inactivation of the AGO1 homeologues of Nicotiana benthamiana reveals their distinct roles in development and antiviral defence. New Phytol 229(3):1289–1297
Ludman M, Szalai G, Janda T, Fátyol K (2023) Hierarchical contribution of Argonaute proteins to antiviral protection. J Exp Bot 74(21):6760–6772
Martín-Merchán A, Moro B, Bouet A, Bologna NG (2023) Domain organization, expression, subcellular localization, and biological roles of Argonaute proteins in Arabidopsis. J Exp Bot 74(7):2374–2388
Qu F, Ye X, Morris TJ (2008) Arabidopsis DRB4, AGO1, AGO7, and RDR6 participate in a DCL4-initiated antiviral RNA silencing pathway negatively regulated by DCL1. Proc Natl Acad Sci U S A 105(38):14732–14737
Raja P, Jackel JN, Li S, Heard IM, Bisaro DM (2014) Arabidopsis double-stranded RNA binding protein DRB3 participates in methylation-mediated defense against geminiviruses. J Virol 88(5):2611–2622
Ranawaka B, An J, Lorenc MT, Jung H, Sulli M, Aprea G, Roden S, Llaca V, Hayashi S, Asadyar L, LeBlanc Z, Ahmed Z, Naim F, de Campos SB, Cooper T, de Felippes FF, Dong P, Zhong S, Garcia-Carpintero V, Orzaez D, Dudley KJ, Bombarely A, Bally J, Winefield C, Giuliano G, Waterhouse PM (2023) A multi-omic Nicotiana benthamiana resource for fundamental research and biotechnology. Nat Plants 9(9):1558–1571
Schwach F, Vaistij FE, Jones L, Baulcombe DC (2005) An RNA-dependent RNA polymerase prevents meristem invasion by potato virus X and is required for the activity but not the production of a systemic silencing signal. Plant Physiol 138(4):1842–1852
Silva-Martins G, Bolaji A, Moffett P (2020) What does it take to be antiviral? An Argonaute-centered perspective on plant antiviral defense. J Exp Bot 71(20):6197–6210
Tu CW, Huang YW, Lee CW, Kuo SY, Lin NS, Hsu YH, Hu CC (2023) Argonaute 5-mediated antiviral defense and viral counter-defense in Nicotiana benthamiana. Virus Res 334:199179
Ying XB, Dong L, Zhu H, Duan CG, Du QS, Lv DQ, Fang YY, Garcia JA, Fang RX, Guo HS (2010) RNA-dependent RNA polymerase 1 from Nicotiana tabacum suppresses RNA silencing and enhances viral infection in Nicotiana benthamiana. Plant Cell 22(4):1358–1372
Wang J, Zhang Q, Tung J, Zhang X, Liu D, Deng Y, Tian Z, Chen H, Wang T, Yin W, Li B, Lai Z, Dinesh-Kumar SP, Baker B, Li F (2024) High-quality assembled and annotated genomes of Nicotiana tabacum and Nicotiana benthamiana reveal chromosome evolution and changes in defense arsenals. Mol Plant 17(3):423–437
Funding
Open access funding provided by Hungarian University of Agriculture and Life Sciences. This work was supported by grants from the National Research Development and Innovation Office, Hungary (K124705, K142626, PD121287 and FK124661).
Author information
Authors and Affiliations
Contributions
KF planned and designed the research; KF, ML and AS performed experiments and analyzed data; KF and AS obtained grant support; KF wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Communicated by Günther Hahne.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ludman, M., Anita, S. & Fátyol, K. Deficiency of multiple RNA silencing-associated genes may contribute to the increased susceptibility of Nicotiana benthamiana to viruses. Plant Cell Rep 43, 177 (2024). https://doi.org/10.1007/s00299-024-03262-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00299-024-03262-3