Abstract
Key message
111 PHD genes were newly identified in rye genome and ScPHD5’s role in regulating cold tolerance and flowering time was suggested.
Abstract
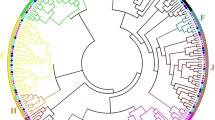
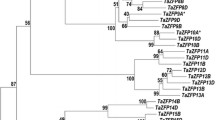
Plant homeodomain (PHD)-finger proteins regulate the physical properties of chromatin and control plant development and stress tolerance. Although rye (Secale cereale L.) is a major winter crop, PHD-finger proteins in rye have not been studied. Here, we identified 111 PHD genes in the rye genome that exhibited diverse gene and protein sequence structures. Phylogenetic tree analysis revealed that PHDs were genetically close in monocots and diverged from those in dicots. Duplication and synteny analyses demonstrated that ScPHDs have undergone several duplications during evolution and that high synteny is conserved among the Triticeae species. Tissue-specific and abiotic stress-responsive gene expression analyses indicated that ScPHDs were highly expressed in spikelets and developing seeds and were responsive to cold and drought stress. One of these genes, ScPHD5, was selected for further functional characterization. ScPHD5 was highly expressed in the spike tissues and was localized in the nuclei of rye protoplasts and tobacco leaves. ScPHD5-overexpressing Brachypodium was more tolerant to freezing stress than wild-type (WT), with increased CBF and COR gene expression. Additionally, these transgenic plants displayed an extremely early flowering phenotype that flowered more than two weeks earlier than the WT, and vernalization genes, rather than photoperiod genes, were increased in the WT. RNA-seq analysis revealed that diverse stress response genes, including HSPs, HSFs, LEAs, and MADS-box genes, were also upregulated in transgenic plants. Our study will help elucidate the roles of PHD genes in plant development and abiotic stress tolerance in rye.








Similar content being viewed by others
References
Aasland R, Gibson TJ, Stewart AF (1995) The PHD finger: implications for chromatin-mediated transcriptional regulation. Trends Biochem Sci 20:56–59
Alves SC, Worland B, Thole V, Snape JW, Bevan MW, Vain P (2009) A protocol for Agrobacterium-mediated transformation of Brachypodium distachyon community standard line Bd21. Nat Protoc 4:638–649
Andrews S (2010) FastQC: a quality control tool for high throughput sequence data. Babraham Bioinformatics. Cambridge, United Kingdom, Babraham Institute
Arora R, Agarwal P, Ray S, Singh AK, Singh VP, Tyagi AK, Kapoor S (2007) MADS-box gene family in rice: genome-wide identification, organization and expression profiling during reproductive development and stress. BMC Genomics 8:242
Bailey TL, Johnson J, Grant CE, Noble WS (2015) The MEME suite. Nucleic Acids Res 43:W39–W49
Bates LS, Waldren Ra, Teare I (1973) Rapid determination of free proline for water-stress studies. Plant Soil 39:205–207
Bienz M (2006) The PHD finger, a nuclear protein-interaction domain. Trends Biochem Sci 31:35–40
Bredow M, Vanderbeld B, Walker VK (2016) Knockdown of ice-binding proteins in brachypodium distachyon demonstrates their role in freeze protection. PLoS ONE 11:e0167941
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13:1194–1202
Chen H, Nelson R, Sherwood J (1994) Enhanced recovery of transformants of Agrobacterium tumefaciens after freeze-thaw transformation and drug selection. Biotechniques 16(664–668):670
Chen L, Zhao Y, Xu S, Zhang Z, Xu Y, Zhang J, Chong K (2018) OsMADS57 together with OsTB1 coordinates transcription of its target OsWRKY94 and D14 to switch its organogenesis to defense for cold adaptation in rice. New Phytol 218:219–231
Chou KC, Shen HB (2010) Plant-mPLoc: a top-down strategy to augment the power for predicting plant protein subcellular localization. PLoS ONE 5:e11335
Curtis MD, Grossniklaus U (2003) A gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant Physiol 133:462–469
Decena MA, Gálvez-Rojas S, Agostini F, Sancho R, Contreras-Moreira B, Des Marais DL, Hernandez P, Catalán P (2021) Comparative genomics, evolution, and drought-induced expression of dehydrin genes in model Brachypodium grasses. Plants 10:2664
Draper J, Mur LA, Jenkins G, Ghosh-Biswas GC, Bablak P, Hasterok R, Routledge AP (2001) Brachypodium distachyon. A new model system for functional genomics in grasses. Plant Physiol 127:1539–1555
Feng Y, Liu QP, Xue QZ (2004) Comparative phylogenetic analysis of the rice and Arabidopsis PHD-finger proteins. Yi Chuan Xue Bao 31:1284–1293
Fernández Gómez J, Wilson ZA (2014) A barley PHD finger transcription factor that confers male sterility by affecting tapetal development. Plant Biotechnol J 12:765–777
Fernández-Calleja M, Casas AM, Igartua E (2021) Major flowering time genes of barley: allelic diversity, effects, and comparison with wheat. Theor Appl Genet 134:1867–1897
Finn RD, Clements J, Eddy SR (2011) HMMER web server: interactive sequence similarity searching. Nucleic Acids Res 39(S2):W29–W37
Fornara F, Parenicová L, Falasca G, Pelucchi N, Masiero S, Ciannamea S, Lopez-Dee Z, Altamura MM, Colombo L, Kater MM (2004) Functional characterization of OsMADS18, a member of the AP1/SQUA subfamily of MADS box genes. Plant Physiol 135:2207–2219
Gaut BS, Morton BR, McCaig BC, Clegg MT (1996) Substitution rate comparisons between grasses and palms: synonymous rate differences at the nuclear gene Adh parallel rate differences at the plastid gene rbcL. Proc Natl Acad Sci 93:10274–10279
Geiger H, Miedaner T (2009) Rye (Secale cereale L.). Cereals. https://doi.org/10.1007/978-0-387-72297-9
Guo M, Liu JH, Ma X, Luo DX, Gong ZH, Lu MH (2016) The plant heat stress transcription factors (HSFs): structure, regulation, and function in response to abiotic stresses. Front Plant Sci 7:114
Hu B, Jin J, Guo AY, Zhang H, Luo J, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31:1296–1297
Hwarari D, Guan Y, Ahmad B, Movahedi A, Min T, Hao Z, Lu Y, Chen J, Yang L (2022) ICE-CBF-COR signaling cascade and its regulation in plants responding to cold stress. Int J Mol Sci 23:1549
Jeong HJ, Yang J, Cho LH, An G (2016) OsVIL1 controls flowering time in rice by suppressing OsLF under short days and by inducing Ghd7 under long days. Plant Cell Rep 35:905–920
Jia X, Zhang X, Qu J, Han R (2016) Optimization conditions of wheat mesophyll protoplast isolation. Agric Sci 07:850–858
Jones P, Binns D, Chang HY, Fraser M, Li W, McAnulla C, McWilliam H, Maslen J, Mitchell A, Nuka G (2014) InterProScan 5: genome-scale protein function classification. Bioinformatics 30:1236–1240
Joo JH, Wang S, Chen J, Jones A, Fedoroff NV (2005) Different signaling and cell death roles of heterotrimeric G protein α and β subunits in the Arabidopsis oxidative stress response to ozone. Plant Cell 17:957–970
Jung WJ, Seo YW (2019) Identification of novel C-repeat binding factor (CBF) genes in rye (Secale cereale L.) and expression studies. Gene 5:82–94
Kang H, Zhang M, Zhou S, Guo Q, Chen F, Wu J, Wang W (2016) Overexpression of wheat ubiquitin gene, Ta-Ub2, improves abiotic stress tolerance of Brachypodium distachyon. Plant Sci 248:102–115
Khong GN, Pati PK, Richaud F, Parizot B, Bidzinski P, Mai CD, Bes M, Bourrie I, Meynard D, Beeckman T, Selvaraj MG, Manabu I, Genga AM, Brugidou C, Do VN, Guiderdoni E, Morel JB, Gantet P (2015) OsMADS26 negatively regulates resistance to pathogens and drought tolerance in rice. Plant Physiol 169:2935–2949
Kim D, Paggi JM, Park C, Bennett C, Salzberg SL (2019) Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat Biotechnol 37:907–915
Kim DY, Lee YJ, Hong MJ, Kim JH, Seo YW (2021) Genome wide analysis of U-box E3 ubiquitin ligases in wheat (Triticum aestivum L.). Int J Mol Sc 22:2699
Kitagawa S, Shimada S, Murai K (2012) Effect of Ppd-1 on the expression of flowering-time genes in vegetative and reproductive growth stages of wheat. Genes Genet Syst 87:161–168
Kolde R, Kolde MR, (2018). Package ‘pheatmap’. R Package. 1.
Lee JH, Yoo SJ, Park SH, Hwang I, Lee JS, Ahn JH (2007) Role of SVP in the control of flowering time by ambient temperature in Arabidopsis. Genes Dev 21:397–402
Lee S, Woo YM, Ryu SI, Shin YD, Kim WT, Park KY, Lee IJ, An G (2008) Further characterization of a rice AGL12 group MADS-box gene, OsMADS26. Plant Physiol 147:156–168
Letunic I, Bork P (2019) Interactive tree of life (iTOL) v4: recent updates and new developments. Nucleic Acids Res 47:W256–W259
Li G, Wang L, Yang J, He H, Jin H, Li X, Ren T, Ren Z, Li F, Han X (2021) A high-quality genome assembly highlights rye genomic characteristics and agronomically important genes. Nat Genet 53:574–584
Li Q, Wang Y, Wang F, Guo Y, Duan X, Sun J, An H (2016) Functional conservation and diversification of APETALA1/FRUITFULL genes in Brachypodium distachyon. Physiol Plant 157:507–518
Liao Y, Smyth GK, Shi W (2014) featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30:923–930
Liu J, Shi Y, Yang S (2018) Insights into the regulation of C-repeat binding factors in plant cold signaling. J Integr Plant Biol 60:780–795
Liu Y, Liu C, Li Z, Xia M, Jiang H, Cheng B, Zhou J, Zhu S (2011) Overexpression of a plant homeodomain (PHD)-finger transcription factor, OsPHD1, can enhance stress tolerance in rice. J Agric Biotechnol 19:462–469
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2- ΔΔCT method. Methods 25:402–408
López-González L, Mouriz A, Narro-Diego L, Bustos R, Martínez-Zapater JM, Jarillo JA, Piñeiro M (2014) Chromatin-dependent repression of the Arabidopsis floral integrator genes involves plant specific PhD-containing proteins. Plant Cell 26:3922–3938
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:1–21
Ma L, Zhu T, Wang H, Zhou H, Shao L, Ding Q, Zhang D, Ma L (2020) Genome-wide identification, phylogenetic analysis and expression profiling of the late embryogenesis-abundant (LEA) gene family in Brachypodium distachyon. Funct Plant Biol 48:386–401
Ma S, Wang M, Wu J, Guo W, Chen Y, Li G, Wang Y, Shi W, Xia G, Fu D (2021) WheatOmics: a platform combining multiple omics data to accelerate functional genomics studies in wheat. Mol Plant 14:1965–1968
Martis MM, Zhou R, Haseneyer G, Schmutzer T, Vrána J, Kubaláková M, König S, Kugler KG, Scholz U, Hackauf B (2013) Reticulate evolution of the rye genome. Plant Cell 25:3685–3698
Matsubara K, Yamanouchi U, Nonoue Y, Sugimoto K, Wang ZX, Minobe Y, Yano M (2011) Ehd3, encoding a plant homeodomain finger-containing protein, is a critical promoter of rice flowering. Plant J 66:603–612
Michaels SD, Amasino RM (1999) FLOWERING LOCUS C encodes a novel MADS domain protein that acts as a repressor of flowering. Plant Cell 11:949–956
Miura K, Renhu N, Suzaki T (2020) The PHD finger of Arabidopsis SIZ1 recognizes trimethylated histone H3K4 mediating SIZ1 function and abiotic stress response. Commun Biol 3:23
Molitor AM, Bu Z, Yu Y, Shen WH (2014) Arabidopsis AL PHD-PRC1 complexes promote seed germination through H3K4me3-to-H3K27me3 chromatin state switch in repression of seed developmental genes. PLOS Genet 10:e1004091
Moon J, Suh SS, Lee H, Choi KR, Hong CB, Paek NC, Kim SG, Lee I (2003) The SOC1 MADS-box gene integrates vernalization and gibberellin signals for flowering in Arabidopsis. Plant J 35:613–623
Musselman CA, Kutateladze TG (2011) Handpicking epigenetic marks with PHD fingers. Nucleic Acids Res 39:9061–9071
Oliveros JC, (2007). VENNY. An interactive tool for comparing lists with venn diagrams. http://bioinfogp.cnb.csic.es/tools/venny/index.html
Pang F, Niu J, Solanki MK, Nosheen S, Liu Z, Wang Z (2022) PHD-finger family genes in wheat (Triticum aestivum L.): evolutionary conservatism, functional diversification, and active expression in abiotic stress. Front Plant Sci 13:1016831
Qin M, Luo W, Zheng Y, Guan H, Xie X (2019a) Genome-wide identification and expression analysis of the PHD-finger gene family in Solanum tuberosum. PLoS ONE 14:e0226964
Qin Z, Bai Y, Muhammad S, Wu X, Deng P, Wu J, An H, Wu L (2019b) Divergent roles of FT-like 9 in flowering transition under different day lengths in Brachypodium distachyon. Nat Commun 10:812
R Core Team (2023) R: a language and environment for statistical computing. Vienna, Austria, R Foundation for Statistical Computing (https://www.R-project.org/)
Rabanus-Wallace MT, Hackauf B, Mascher M, Lux T, Wicker T, Gundlach H, Baez M, Houben A, Mayer KFX, Guo L (2021) Chromosome-scale genome assembly provides insights into rye biology, evolution and agronomic potential. Nat Genet 53:564–573
Ream TS, Woods DP, Schwartz CJ, Sanabria CP, Mahoy JA, Walters EM, Kaeppler HF, Amasino RM (2014) Interaction of photoperiod and vernalization determines flowering time of Brachypodium distachyon. Plant Physiol 164:694–709
Schindler U, Beckmann H, Cashmore AR (1993) HAT3.1, a novel Arabidopsis homeodomain protein containing a conserved cysteine-rich region. Plant J 4:137–150
Scholthof KBG, Irigoyen S, Catalan P, Mandadi KK (2018) Brachypodium: a monocot grass model genus for plant biology. Plant Cell 30:1673–1694
Stothard P (2000) The sequence manipulation suite: JavaScript programs for analyzing and formatting protein and DNA sequences. Biotechniques 28:1102–1104
Sun M, Jia B, Yang J, Cui N, Zhu Y, Sun X (2017) Genome-wide identification of the PHD-finger family genes and their responses to environmental stresses in Oryza sativa L. Int J Mol Sci 18:2005
Sung S, Amasino RM (2004) Vernalization in Arabidopsis thaliana is mediated by the PHD finger protein VIN3. Nature 427:159–164
Tamura K, Stecher G, Kumar S (2021) MEGA11: molecular evolutionary genetics analysis version 11. Mol Biol Evol 38(7):3022–3027
Teo ZWN, Zhou W, Shen L (2019) Dissecting the function of MADS-box transcription factors in orchid reproductive development. Front Plant Sci 10:1474
Thumuluri V, Almagro Armenteros JJ, Johansen AR, Nielsen H, Winther O (2022) DeepLoc 2.0: multi-label subcellular localization prediction using protein language models. Nucleic Acids Res 50:W228–W234
Tian T, Liu Y, Yan H, You Q, Yi X, Du Z, Xu W, Su Z (2017) agriGO v2.0: a GO analysis toolkit for the agricultural community, 2017 update. Nucleic Acids Res 45:W122–W129
Wang M, Zou Z, Li Q, Xin H, Zhu X, Chen X, Li X (2017) Heterologous expression of three camellia sinensis small heat shock protein genes confers temperature stress tolerance in yeast and Arabidopsis thaliana. Plant Cell Rep 36:1125–1135
Wang Q, Liu J, Wang Y, Zhao Y, Jiang H, Cheng B (2015) Systematic analysis of the maize PHD-finger gene family reveals a subfamily involved in abiotic stress response. Int J Mol Sci 16:23517–23544
Wang W, Wu Y, Shi R, Sun M, Li Q, Zhang G, Wu J, Wang Y, Wang W (2020) Overexpression of wheat α-mannosidase gene TaMP impairs salt tolerance in transgenic Brachypodium distachyon. Plant Cell Rep 39:653–667
Wei B, Zhang RZ, Guo JJ, Liu DM, Li AL, Fan RC, Mao L, Zhang XQ (2014) Genome-wide analysis of the MADS-box gene family in Brachypodium distachyon. PLoS ONE 9:e84781
Wei W, Zhang YQ, Tao JJ, Chen HW, Li QT, Zhang WK, Ma B, Lin Q, Zhang JS, Chen SY (2015) The Alfin-like homeodomain finger protein AL5 suppresses multiple negative factors to confer abiotic stress tolerance in arabidopsis. Plant J 81:871–883
Wen F, Wu X, Li T, Jia M, Liu X, Li P, Zhou X, Ji X, Yue X (2017) Genome-wide survey of heat shock factors and heat shock protein 70s and their regulatory network under abiotic stresses in brachypodium distachyon. PLoS ONE 12:e0180352
Winicov I, Valliyodan B, Xue L, Hoober JK (2004) The MsPRP2 promoter enables strong heterologous gene expression in a root specific manner and is enhanced by overexpression of Alfin1. Planta 219:925–935
Woods DP, Ream TS, Bouché F, Lee J, Thrower N, Wilkerson C, Amasino RM (2017) Establishment of a vernalization requirement in Brachypodium distachyon requires REPRESSOR OF VERNALIZATION1. Proc Nat Aca Sci 114:6623–6628
Wu S, Wu M, Dong Q, Jiang H, Cai R, Xiang Y (2016a) Genome-wide identification, classification and expression analysis of the PHD-finger protein family in populus trichocarpa. Gene 575:75–89
Wu ZG, Jiang W, Chen SL, Mantri N, Tao ZM, Jiang CX (2016b) Insights from the cold transcriptome and metabolome of dendrobium officinale: global reprogramming of metabolic and gene regulation networks during cold acclimation. Front Plant Sci 7:1653
Yang R, Hong Y, Ren Z, Tang K, Zhang H, Zhu JK, Zhao C (2019) A role for PICKLE in the regulation of cold and salt stress tolerance in Arabidopsis. Front Plant Sci 10:900
Yin X, Liu X, Xu B, Lu P, Dong T, Yang D, Ye T, Feng YQ, Wu Y (2019) OsMADS18, a membrane-bound MADS-box transcription factor, modulates plant architecture and the abscisic acid response in rice. J Exp Bot 70:3895–3909
Yoon JS, Kim JY, Kim DY, Seo YW (2021) A novel wheat ASR gene, TaASR2D, enhances drought tolerance in Brachypodium distachyon. Plant Physiol Biochem 159:400–414
Yoon JS, Kim JY, Lee MB, Seo YW (2019) Over-expression of the Brachypodium ASR gene, BdASR4, enhances drought tolerance in Brachypodium distachyon. Plant Cell Rep 38:1109–1125
Yu CS, Chen YC, Lu CH, Hwang JK (2006) Prediction of protein subcellular localization. Proteins: Struct Funct Bioinf 64:643–651
Zhai M, Sun Y, Jia C, Peng S, Liu Z, Yang G (2016) Over-expression of JrsHSP17.3 gene from Juglans regia confer the tolerance to abnormal temperature and NaCl stresses. J Plant Biol 59:549–558
Zhang Z (2022) KaKs_calculator 3.0: calculating selective pressure on coding and non-coding sequences. Genomics Proteomics Bioinformatics 20:536–540
Zhang Z, Huang R (2013) Analysis of malondialdehyde, chlorophyll proline, soluble sugar, and glutathione content in arabidopsis seedling. Bio Protoc 3:e817–e817
Funding
This work was supported by the Basic Science Research Program through the National Research Foundation of Korea, funded by the Ministry of Education (2021R1I1A1A01048945) and by a grant from Korea University.
Author information
Authors and Affiliations
Contributions
WJJ and YWS conceived and planned the experiments. WJJ, JHJ, and JSY performed the experiments. WJJ wrote the draft manuscript. All authors provided critical feedback and helped shape the research, analysis, and manuscript.
Corresponding author
Ethics declarations
Conflict interest
The authors declare that they have no competing interests.
Additional information
Communicated by Sheng Ying.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jung, W.J., Jeong, J.H., Yoon, J.S. et al. Genome-wide identification of the plant homeodomain-finger family in rye and ScPHD5 functions in cold tolerance and flowering time. Plant Cell Rep 43, 142 (2024). https://doi.org/10.1007/s00299-024-03226-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00299-024-03226-7