Abstract
Key message
FKF1 dimerization is crucial for proper FT levels to fine-tune flowering time. Attenuating FKF1 homodimerization increased CO abundance by enhancing its COP1 binding, thereby accelerating flowering under long days.
Abstract
In Arabidopsis (Arabidopsis thaliana), the blue-light photoreceptor FKF1 (FLAVIN-BINDING, KELCH REPEAT, F-BOX 1) plays a key role in inducing the expression of FLOWERING LOCUS T (FT), encoding the main florigenic signal in plants, in the late afternoon under long-day conditions (LDs) by forming dimers with FT regulators. Although structural studies have unveiled a variant of FKF1 (FKF1 I160R) that disrupts homodimer formation in vitro, the mechanism by which disrupted FKF1 homodimer formation regulates flowering time remains elusive. In this study, we determined that the attenuation of FKF1 homodimer formation enhances FT expression in the evening by promoting the increased stability of CONSTANS (CO), a primary activator of FT, in the afternoon, thereby contributing to early flowering. In contrast to wild-type FKF1, introducing the FKF1 I160R variant into the fkf1 mutant led to increased FT expression under LDs. In addition, the FKF1 I160R variant exhibited diminished dimerization with FKF1, while its interaction with GIGANTEA (GI), a modulator of FKF1 function, was enhanced under LDs. Furthermore, the FKF1 I160R variant increased the level of CO in the afternoon under LDs by enhancing its binding to COP1, an E3 ubiquitin ligase responsible for CO degradation. These findings suggest that the regulation of FKF1 homodimerization and heterodimerization allows plants to finely adjust FT expression levels around dusk by modulating its interactions with GI and COP1.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The timely flowering of plants in the appropriate seasons is crucial for their reproductive fitness (Song et al. 2015; Exposito-Alonso 2020). In many plant species, seasonal flowering is regulated by the expression of FLOWERING LOCUS T (FT), which encodes a mobile florigenic signal (Corbesier et al. 2007; Jaeger and Wigge 2007; Mathieu et al. 2007). FT protein is produced in the vasculature of leaves and travels to the shoot apical meristem (Corbesier et al. 2007; Jaeger and Wigge 2007; Mathieu et al. 2007; Takagi et al. 2023). In Arabidopsis (Arabidopsis thaliana), a facultative long-day plant, FT is strongly expressed under long-day conditions (LDs), thereby promoting flowering (Suárez-López et al. 2001). Under LDs, FT expression largely relies on the function of the transcriptional activator CONSTANS (CO), whose stabilization during the daytime is controlled by photoreceptors: This regulation determines the timing of flowering (Samach et al. 2000; Valverde et al. 2004; Song et al. 2015; Takagi et al. 2023).
Under conventional laboratory LD conditions, FT mRNA abundance only peaks at the end of the day (Suárez-López et al. 2001). However, under natural LDs, in which a simulated red-to-far-red ratio mimicking that of sunlight and daily temperature cycles is superimposed onto LDs, the daily expression of FT peaks in both the morning and evening (Song et al. 2018; Kim et al. 2024). This bimodal expression pattern is determined by the regulation of CO stability and CO-independent regulatory pathways (Song et al. 2018; Kim et al. 2024; Lee et al. 2023; Takagi et al. 2023). Unlike FT expression, the daily profile of CO levels shows two peaks under both conventional laboratory LDs and natural LDs: a narrow peak early in the morning and a broad peak in the late afternoon (Valverde et al. 2004; Song et al. 2012, 2018). During the morning under natural LDs, CO is stabilized by the activity of the far-red-light-responsible photoreceptor phytochrome A (phyA) (Song et al. 2018). EARLY FLOWERING 3 (ELF3), an interaction partner of CONSTITUTIVE PHOTOMORPHOGENIC 1 (COP1), forms a protein complex with CO and mediates its degradation (Jang et al. 2008; Song et al. 2018). PhyA interacts with ELF3 in the morning (Song et al. 2018), which might interfere with the function of ELF3 in destabilizing CO. By contrast, the red-light-activated phyB, the major red-light photoreceptor in plants, facilitates the destabilization of CO by forming a complex with CO and HOS1 (HIGH EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 1), an E3 ubiquitin ligase that degrades its proteolytic targets such as CO (Lazaro et al. 2015). In addition, the F-box blue-light photoreceptor ZEITLUPE (ZTL), which possesses E3 ubiquitin ligase activity, binds to and degrades CO during the morning (Song et al. 2014).
In the afternoon under conventional laboratory LDs, FKF1 (FLAVIN-BINDING, KELCH REPEAT, F-BOX 1), a ZTL homologous protein, strongly promotes the stability of CO by interacting with CO through its LOV (Light, Oxygen, or Voltage) domain (Song et al. 2012). Since the LOV domain is responsible for blue-light absorption, the FKF1–CO interaction increases under blue light (Ito et al. 2012; Song et al. 2012). Moreover, FKF1 binds to COP1 and inhibits its homodimer formation, leading to a decrease in COP1-dependent CO degradation (Lee et al. 2017). In addition to FKF1, CRYPTOCHROME 2 (CRY2), another type of blue-light photoreceptor, also contributes to CO stabilization in the afternoon. CRY2 interacts with COP1 and SUPRESSOR OF PHYA-105 1 (SPA1) under blue light and suppresses the formation of a complex between COP1 and SPA1, resulting in the attenuation of CO degradation mediated by the E3 ubiquitin ligase activity of COP1 (Jang et al. 2008; Zuo et al. 2011).
Among photoreceptors, two homologous blue-light receptors, ZTL and FKF1, exhibit contrasting functions not only in regulating CO stability but also in regulating the seasonal expression of FT (Takagi et al. 2023). Under natural LDs, ZTL forms a protein complex with the FT inhibitors TARGET OF EAT1 (TOE1) and TOE2, which specifically inhibits FT expression in the morning (Zhang et al. 2015; Kim et al. 2024). In contrast to ZTL, FKF1 promotes FT transcription more prominently in the afternoon under natural LDs (Song et al. 2018). Besides stabilizing CO protein, FKF1 induces FT expression by removing CYCLING DOF FACTOR 1 (CDF1), a transcriptional repressor of CO and FT, from the FT promoter in the afternoon via its E3 ubiquitin ligase activity (Imaizumi et al. 2005; Sawa et al. 2007; Song et al. 2012). The diel accumulation patterns of FKF1 and ZTL are remarkably similar, with both peaking in the afternoon (Kim et al. 2024). However, ZTL is more abundant in the morning than FKF1 (Kim et al. 2024). The regulation of FT expression by the opposing functions of FKF1 and ZTL involves dimer formation between these proteins and the role of their partner protein GIGANTEA (GI). Higher abundance of ZTL in the morning inhibits the activity of FKF1 in promoting flowering through heterodimer formation (Takase et al. 2011; Hwang et al. 2019). Under LDs, FKF1 and ZTL interact with GI in a blue-light-dependent manner in the afternoon through their LOV domains. These interactions play crucial roles in inactivating ZTL during this time window and facilitating FKF1-induced CDF1 degradation (Kim et al. 2007; Sawa et al. 2007; Hwang et al. 2019). Consequently, the binding of GI to ZTL leads to the sequestration of FKF1, as well as CO, from ZTL, resulting in the stabilization of CO and the induction of FT expression by FKF1 in the afternoon (Hwang et al. 2019).
Structural studies have demonstrated that the LOV domains of FKF1 and ZTL intrinsically form homodimers (Nakasako et al. 2005; Pudasaini et al. 2017; Kwon et al. 2022). However, upon binding to GI, the LOV homodimers are converted to monomers (Kwon et al. 2022). Considering their roles in modulating CO stability, these observations suggest that the monomeric and dimeric states of FKF1 and ZTL (determined via their LOV domains) might be crucial for their functions in regulating flowering time (Song et al. 2012, 2014; Hwang et al. 2019). Although we previously demonstrated that the Ile160 residue in the FKF1 LOV domain is important for homodimer formation in vitro (Pudasaini et al. 2017), the biological relevance of conformational changes in FKF1 in regulating flowering time remains unclear.
In this study, to explore the effects of structural changes in FKF1 on flowering time in Arabidopsis at the molecular level, we investigated the properties associated with the regulation of CO stabilization and FT gene expression using the FKF1 I160R variant, which shows reduced homodimer formation. Our findings reveal the altered binding affinities between FKF1 and complex-forming proteins, confirming their potential to finely regulate flowering time in plants at the mechanistic level.
Materials and methods
Plant materials and growth conditions
All Arabidopsis thaliana plants [WT, fkf1 (Nelson et al. 2000), FKF1:HA-FKF1/fkf1 (Imaizumi et al. 2003), FKF1:HA-FKF1 GI:GI-TAP/fkf1 gi-2 (Sawa et al. 2007), GI:GI-TAP/gi-2 (David et al. 2006), CO:HA-CO (Song et al. 2012), FKF1:HA-FKF1 I160R/fkf1, FKF1:FKF1 I160R GI:GI-TAP/fkf1 gi-2, CO:HA-CO/cop1-4, FKF1:HA-FKF1 I160R CO:HA-CO/fkf1] used in this study are the Columbia (Col-0) ecotype.
The I160R mutation in the FKF1 gene was introduced using the FKF1 I160R reverse primer (5ʹ-GAAGACCTGTATCCCACGTACGTGTGTAATGGT-3ʹ, the underline indicates the sequences that encode arginine).
Plants were grown on soil in a plant growth room for flowering time measurement and tobacco transient expression. For Arabidopsis co-immunoprecipitation assays as well as protein and gene expression analyses, seeds were sterilized, stratified at 4 °C for 3 days, and sown onto 0.5X Murashige Skoog (MS) media (Duchefa Biochemie) with 1% (w/v) sucrose.
Seedlings were grown in LDs (16-h light/8-h dark) with constant 22 °C under full-spectrum white LED light (4,470 K) (Bissol LED) with a fluence rate of 80–100 μmol m−2 s−1. For light quality treatment, plants grown in LDs were transferred to LDs with monochromatic blue (450 nm) or red (660 nm) ranging from 40 to 50 μmol m−2 s−1 on day 8 and harvested at ZT12 on day 10.
To analyze the flowering time of Arabidopsis, the number of rosette and cauline leaves on the main stem were counted when the inflorescence reached 3 cm long after bolting. The analysis of flowering time was carried out independently in triplicate, involving 12 individual plants per trial. Each iteration of the experiment yielded consistent and comparable results.
RNA isolation and gene expression analysis
For diel gene expression analysis, seedlings grown on agar plates under LDs for 10 days were harvested at 3 h intervals from ZT1 to ZT22. Total RNA was extracted using the Higene Total RNA Prep Kit (BioFact). Subsequently, 2 μg of isolated RNA was employed to synthesize first-strand cDNA using the DiastarTM RT kit (SolGent) with an oligo dT primer. Following dilution with 40 μl of water, 2 μl of cDNA was utilized for quantitative polymerase chain reaction (qPCR) performed on a CFX96 thermal real-time cycler (Bio-Rad). Primers and PCR conditions for FKF1, CO, FT, and IPP2 were previously detailed (Song et al. 2012). Relative expression levels were determined by normalizing against IPP2 expression.
Immunoblot analysis and protein quantification
To analyze FKF1, FKF1 I160R, and CO protein abundance, FKF1:HA-FKF1/fkf1, FKF1:HA-FKF1 I160R/fkf1, CO:HA-CO, and FKF1:HA-FKF1 I160R CO:HA-CO/fkf1 transgenic plants were grown on agar plates under LDs. Seedlings were collected at 4-h intervals from ZT0 to ZT20 on day 10, with an additional harvest at ZT0.5 for CO. Protein extraction, nuclei isolation, and Western blot procedures were previously described (Song et al. 2012). In brief, total proteins were extracted using an extraction buffer [50 mM sodium-phosphate pH 7.4, 100 mM NaCl, 10% glycerol, 5 mM EDTA, 1% NP-40, 0.5% SDS, 0.5% sodium deoxycholate, 50 μM MG-132, and protease inhibitor tablets-EDTA free (Roche)]. Nuclei proteins were obtained using the CelLytic Plant Nuclei Isolation/Extraction Kit (Sigma) following the manufacturer's protocol. Detection of HA-FKF1 and HA-CO proteins was accomplished using the anti-HA antibody (3F10, Roche). Actin and Histone H3 served as internal loading controls for whole protein extract and nuclei proteins, respectively, and were detected using anti-actin (AC009, ABclonal) and anti-histone H3 (ab1791, Abcam) antibodies, respectively. For protein quantification, immunoreactive proteins were visualized with SuperSignal West Pico (Thermo Fisher) and/or ECL select Femto (Bio-Rad) mixture solution, followed by imaging using the ChemiDocTM Touch imaging system (Bio-Rad). The captured image was analyzed for quantification using the Image Lab software (Bio-Rad). Relative amounts of HA-fusion proteins were normalized based on the expression values of actin and Histone H3 for HA-FKF1 and HA-CO, respectively.
Co-immunoprecipitation experiments
The co-immunoprecipitation assays were conducted following previously established procedures (Song et al. 2012). For the examination of FKF1 homodimerization and interactions involving FKF1 with ZTL, as well as FKF1 with CO in tobacco cells, agrobacteria strain GV3101 carrying 35S:FKF1-TAP, 35S:HA-FKF1, 35S:HA-FKF1 I160R, 35S:ZTL-TAP, or 35S:CO-TAP constructs were infiltrated into 3- to 4-week-old Nicotiana benthamiana leaves. Subsequently, the tobacco plants were grown under LDs for 3 days, and the infiltrated leaves were harvested to obtain whole protein extracts. An equivalent amount of ground tissues was utilized for the co-immunoprecipitation experiment. To investigate in vivo interactions, GI:GI-TAP, FKF1:HA-FKF1/fkf1, FKF1:HA-FKF1 I160R/fkf1 #2–3, FKF1:HA-FKF1 GI:GI-TAP/fkf1 gi-2, FKF1:HA-FKF1 I160R GI:GI-TAP/fkf1 gi-2, and CO:HA-CO/cop1-4 plants were grown under the specified conditions. All co-immunoprecipitation experiments were conducted using 10-day-old seedlings and were repeated three times with independently harvested samples. The proteins GI-TAP, HA-FKF1, and COP1 were detected through Western blotting using anti-protein A (Sigma), anti-HA (3F10, Roche), and anti-COP1 (Lian et al. 2011) antibodies, respectively.
Expression of the FKF1 I160R protein
An FKF1 I160R variant composed of residues 28–174 was previously cloned into a pGST-Parallel vector using the NcoI and XhoI cut sites (Pudasaini et al. 2017). 24 L of FKF1 I160R proteins were expressed in E. coli JM109DE3. JM109DE3 cells transformed with pGST-FKF1 I160R were grown at 37 °C until an OD600 of 0.4–0.5, and the temperature was reduced to 18 °C for 40 min prior to induction with 0.3 mM IPTG (RPI). Cells were harvested after about 20 h and pelleted in a buffer containing 50 mM Hepes (pH 8.0), 100 mM NaCl, and 10% glycerol and stored at – 80 °C.
Purification of the FKF1 I160R protein
The pellet from a 24-l expression of FKF1 I160R proteins was thawed and lysed via sonication. The lysate was clarified via centrifugation at 22,000 RPM for 1 h. The supernatant was applied to glutathione affinity resin (Qiagen) and purified in the light to enhance protein stability. After binding, GST-tagged FKF1 I160R was treated on a column with 4 mg of TEV protease at 22 °C for 2 h. Proteins were then eluted by the addition of buffer containing 50 mM Hepes (pH 8.0), 100 mM NaCl, and 10% glycerol. Eluted protein was concentrated to 5 ml and further purified by Size Exclusion Chromatography using a Superdex S200 column equilibrated in the same buffer. Fractions containing FKF1 I160R proteins were pooled together, and concentrated to 500 μl, resulting in a final overall yield of about 0.3 mg of FKF1 I160R proteins.
Kinetic characterization of the FKF1 I160R variant
For kinetic characterization, the purified FKF1 I160R protein was diluted in a buffer containing 50 mM Hepes (pH 8.0), 100 mM NaCl, 10% glycerol, and 1.0 M imidazole (pH 8.0) to a final imidazole concentration of 300 mM to accelerate the recovery kinetics. A 100 μl sample was placed into a 1.0 cm pathlength quartz cuvette, and spectra were recorded every 1800 s for 24 h. Kinetics were extracted from the absorbance trace at 450 nm through a single exponential fit.
Statistical analysis
Statistical analyses, including calculations of mean, standard error of the mean (SEM), and t tests, were conducted using Microsoft Excel software. The significance of differences between two experimental groups was determined using a two-tailed Student’s t test. Significance levels were set at P values of < 0.05, < 0.01, and < 0.001 for assessing the significance of differences between data sets.
Results
The FKF1 I160R variant shows enhanced activity in regulating flowering time
The blue-light photoreceptor FKF1 is pivotal in the regulation of photoperiod-dependent flowering (Song et al. 2015). The LOV domain within FKF1 is indispensable for its functionality, where the light-dependent reversible formation of a Flavin-C4a within the LOV domain regulates function, and the LOV domain mediates dimer formation. Mutations within the LOV domain compromise its interactions with GI and CO in planta under blue-light conditions (Sawa et al. 2007; Song et al. 2012; Pudasaini et al. 2017). Moreover, the LOV domain forms a homodimer through equivalent anti-parallel contacts following the β-scaffold (Pudasaini et al. 2017). The I160R variant of the FKF1 LOV domain, characterized by a mutation at position Ile160 within one of the β-sheets, disrupts FKF1 LOV homodimer formation in vitro (Pudasaini et al. 2017), suggesting that this amino acid residue may be crucial for determining the homodimeric and heterodimeric states of FKF1 in Arabidopsis. Notably, the I160R variant demonstrates photochemistry consistent with WT FKF1 (Supplementary Fig. S1), thereby allowing one to isolate the biological effects of homo/hetero dimerization independent of photochemical function. Therefore, we explored the effect of changes in dimer formation on the role of FKF1 in regulating flowering time, focusing on the I160R mutation. Considering that FKF1 predominantly accumulates in the afternoon and that its function is specifically linked to the evening peak levels of FT rather than the morning FT peak (Imaizumi et al. 2003; Song et al. 2018), we opted to employ conventional laboratory LDs, hereafter referred to as LDs, instead of natural LDs. We generated the FKF1:HA-FKF1 I160R expression cassette, designed to express the full-length FKF1 I160R with an N-terminal hemagglutinin (HA) tag, driven by the FKF1 promoter, and introduced this construct into the Arabidopsis fkf1 mutant (Imaizumi et al. 2003).
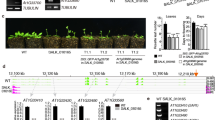
We compared the effects of the wild-type version of FKF1 and the FKF1 I160R variant on complementing the late-flowering phenotype of the fkf1 mutant by measuring total leaf number at bolting. Under LDs, wild-type and fkf1 plants bolted with an average leaf number of 19.7 and 44.0, respectively (Fig. 1A, B). Wild-type FKF1 nearly complemented the late flowering of the fkf1 mutant, with 23.2 leaves at bolting (Fig. 1A, B). Notably, two transgenic lines of FKF1:HA-FKF1 I160R/fkf1 flowered significantly earlier than wild-type plants (Fig. 1A, B). These results indicate that the I160R mutation in FKF1 enhances its function in regulating flowering time.
Complementation of the fkf1 mutant with the FKF1 I160R variant promotes flowering. A Flowering phenotypes of representative 5- to 6-week-old Arabidopsis thaliana plants (Col-0 ecotype) grown under long-day conditions (LDs). Scale bar is 1 cm. B Total leaf number at bolting of wild-type (WT, Col-0 ecotype), fkf1 mutant, FKF1:HA-FKF1/fkf1, and FKF1:HA-FKF1 I160R/fkf1 #2-3, and FKF1:HA-FKF1 I160R/fkf1 #11-3 plants. The data underwent statistical analysis using a two-tailed Student's t test. **p < 0.01 and ***p ≤ 0.001. Data represent means ± SEM
The FKF1 I160R variant exhibits reduced protein stability but enhanced efficacy in inducing FT expression
We investigated the molecular characteristics of the FKF1 I160R variant associated with the promotion of flowering. FKF1 mRNA levels and FKF1 protein abundance exhibit daily rhythms, with peak levels in the afternoon under LDs (Imaizumi et al. 2003). In two FKF1:HA-FKF1 I160R/fkf1 transgenic lines, the diel profile of FKF1 I160R mRNA under LDs was either similar to or higher than that of FKF1 mRNA in FKF1:HA-FKF1/fkf1 plants, with peak levels at Zeitgeber Time 10 (ZT10), resembling that of wild-type plants (Fig. 2A, B). We previously observed that introducing the I160R mutation into a truncated version of FKF1 comprising amino acid residues 28–174 of the LOV domain led to a transition from homodimer to monomer formation and increased susceptibility to protein degradation in vitro (Pudasaini et al. 2017). Consistent with this observation in vitro, FKF1 I160R protein levels in FKF1:HA-FKF1 I160R/fkf1 plants were much lower than FKF1 protein levels in FKF1:HA-FKF1/fkf1 plants while exhibiting the same daily rhythm (Fig. 2C, D), confirming the finding that the FKF1 I160R variant is more susceptible to proteolysis than the wild-type form (Pudasaini et al. 2017). These results suggest that changes in the abundance of FKF1 protein may not be the primary factor contributing to the early flowering of FKF1:HA-FKF1I160R/fkf1 plants.
Introducing the FKF1 I160R variant into the fkf1 mutant increases FT mRNA levels. Comparative analysis of the levels of FKF1 mRNA (A, B), FKF1 protein (C, D), CO mRNA (E, F), and FT mRNA (G, H) in WT, FKF1:HA-FKF1/fkf1, and FKF1:HA-FKF1I160R/fkf1 #2-3 (A, C, E, G), and FKF1:HA-FKF1 I160R/fkf1 #11-3 (B, D, F, H) plants grown under LDs. For gene expression analysis, plants were harvested every 3 h from ZT1 to ZT22 on day 10. FKF1, CO, and FT transcript levels were normalized to IPP2 levels. To compare protein abundance, plants were collected every 4 h from ZT0 to ZT20 on day 10. HA-FKF1 proteins were detected by immunoblotting with anti-HA antibody. Relative protein abundance was normalized to Actin levels. Data represent means ± SEM from at least three independent biological replicates
Under LDs, FKF1 induces FT mRNA expression in the evening through multiple feedforward mechanisms that promote flowering (Song et al. 2012). In addition to its role in CO stabilization, FKF1 promotes the expression of CO and FT in the afternoon by directly alleviating their transcriptional repression imposed by CDF proteins via its E3 ubiquitin ligase activity (Imaizumi et al. 2005; Sawa et al. 2007; Fornara et al. 2009; Song et al. 2012). Therefore, we examined CO and FT transcript levels under LDs. In both FKF1:HA-FKF1 I160R/fkf1 lines, CO mRNA levels were comparable to those in wild-type and FKF1:HA-FKF1/fkf1 plants (Fig. 2E, F). In contrast to CO, there was a notable increase in the abundance of FT mRNA in FKF1:HA-FKF1 I160R/fkf1 lines (Fig. 2G, H). This increase appeared to correlate with the protein levels of the FKF1 I160R variant (Fig. 2C, D). These findings suggest that the FKF1 I160R variant selectively regulates FT over CO expression. Considering the simultaneous inhibition of CO and FT expression by CDF1 (Song et al. 2012), these findings further support the notion that this regulatory mechanism is likely unrelated to the degradation of CDF by FKF1.
The FKF1 I160R variant displays increased interaction with GI but decreased binding to ZTL
The transition between monomeric and dimeric states appears to be a critical factor for the role of FKF1 in regulating flowering time. FKF1 has an inherent capacity to form a homodimer but is converted into a monomer upon binding with GI (Kwon et al. 2022). Under LDs, FKF1 forms a complex with GI in the afternoon. This complex formation enhances the E3 ubiquitin ligase activity of FKF1 by facilitating the recognition of its proteolytic targets, CDF proteins, resulting in the induction of CO and FT expression during this time period (Sawa et al. 2007). Conversely, under short-day conditions, the interaction between FKF1 and GI is impaired during the daytime, contributing to very low expression levels of CO and FT (Sawa et al. 2007). The I160R mutation abolished the homodimerization of FKF1 via its LOV domain in vitro (Pudasaini et al. 2017). However, the I160 site would not disrupt the FKF1-GI interface predicted from the structure of GI bound to LKP2 (Kwon et al. 2022 and Supplementary Fig. S2). These observations suggest that the heightened functionality of FKF1 in elevating FT mRNA levels due to the I160R mutation could be attributed to changes in the dimerization state of FKF1. Consequently, we investigated the effect of the I160R mutation on FKF1 dimerization in plant cells.
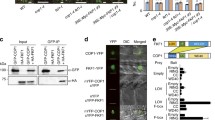
To substantiate the reduced FKF1 homodimerization observed in vitro, we transiently overexpressed TAP-tagged FKF1 (FKF1-TAP) with either HA-FKF1 or the HA-FKF1 I160R variant in Nicotiana benthamiana leaves under LDs. Co-immunoprecipitation (co-IP) revealed a decrease in the binding affinity of FKF1-TAP to the HA-FKF1 I160R variant compared to HA-FKF1 (Fig. 3A, B). Next, we asked whether the I160R mutation influences the complex formation of FKF1 with GI. We generated FKF1:HA-FKF1 I160R GI:GI-TAP/fkf1 transgenic plants and subjected them to co-IP assays. Under LDs, the interaction between the FKF1 I160R variant and GI significantly increased compared to that between FKF1 and GI (Fig. 3C, D). As the LOV domain of FKF1 exists as a monomer when bound to GI (Kwon et al. 2022), these results suggest that the I160R mutation might facilitate the conversion of FKF1 homodimers to monomers, promoting subsequent GI binding. As FKF1 interacts with GI in a blue-light-dependent manner, we evaluated the impact of light quality on this interaction. Although some variations were observed, a similar trend in complex formation between FKF1–GI and FKF1 I160R–GI was observed under white-, red-, and blue-light conditions (Supplementary Fig. S3). These results suggest that the I160R mutation has minimal effects on the blue-light-mediated interaction.
The I160R mutation influences FKF1 dimerization in planta. Co-immunoprecipitation (co-IP) assays using proteins expressed in leaves of Nicotiana benthamiana (A, B, E, F) and Arabidopsis transgenic lines (C, D). TAP-tagged proteins were immunoprecipitated with anti-Protein A antibody. Relative amounts of complex formation were quantified. A, C, E Representative images of the results of co-IP assays. B, D, F Bar graphs representing the amounts of co-precipitated HA-tagged proteins calculated as (HA-tagged proteinIP/TAP-tagged proteinIP)/(HA-tagged proteinInput/TAP-tagged proteinInput). A, B FKF1-TAP proteins were coexpressed with either HA-FKF1 or HA-FKF1 I160R protein in 3-week-old tobacco plants. C, D Arabidopsis plants were grown on agar medium under LDs and harvested at ZT12 on day 10. E, F ZTL-TAP proteins were transiently coexpressed with either HA-FKF1 or HA-FKF1 I160R in 3-week-old tobacco plants. The data underwent statistical analysis using a two-tailed Student's t test. *p < 0.05 and **p ≤ 0.01. ns not significant. Means ± SEM were calculated from three biological replicates
In contrast to GI, ZTL, a homolog of FKF1, inhibits the function of FKF1 in regulating flowering by directly binding to this protein (Takase et al. 2011; Hwang et al. 2019). Thus, we examined whether the increased levels of the monomeric form of FKF1 affect its interaction with ZTL. In tobacco leaf cells, the amount of the FKF1 I160R variant co-immunoprecipitated with ZTL-TAP was significantly reduced compared to FKF1 (Fig. 3E, F), implying that ZTL may interact with the FKF1 homodimer. Collectively, these findings suggest that the conversion of FKF1 homodimers to monomers could play a pivotal role in promoting the levels of FT mRNA by modulating complex formation with FT regulators such as GI and ZTL.
The FKF1 I160R variant enhances CO stability through increased binding to COP1
CO serves as the primary FT activator, and FKF1 stabilizes CO in the afternoon through a direct interaction via its LOV domain (Samach et al. 2000; Song et al. 2012). Gene expression analysis showed that the FKF1 I160R variant selectively enhances FT expression over CO (Fig. 2E–H). This finding prompted us to hypothesize that the increased levels of FKF1 monomers resulting from the I160R mutation might increase the interaction of FKF1 with CO and/or enhance CO stability. To explore these possibilities, we carried out co-IP assays using proteins transiently overexpressed in tobacco leaves. Unexpectedly, the binding of CO to the FKF1 I160R variant was significantly reduced compared to its binding to FKF1 (Fig. 4A, B). We then analyzed CO protein levels using CO:HA-CO and CO:HA-CO FKF1:FKF1-I160R/fkf1 lines. Under LDs, the daily expression profile of CO in the CO:HA-CO FKF1:HA-FKF1 I160R/fkf1 line showed enhanced protein abundance in the afternoon, particularly at ZT12 (Fig. 4C, D). These results suggest that the FKF1 I160R variant indirectly upregulates CO stability rather than complex formation. COP1 is the main E3 ubiquitin ligase that is responsible for CO degradation throughout the day (Jang et al. 2008; Liu et al. 2008; Sarid-Krebs et al. 2015). Under LDs, FKF1 suppresses COP1 homodimerization by directly binding to it in the presence of light, thereby potentially diminishing COP1 activity and increasing the stabilization of CO in the afternoon (Lee et al. 2017). Therefore, we reasoned that the I160R mutation in FKF1 may increase its binding to COP1. We performed co-IP analysis using FKF1:HA-FKF1/fkf1 and FKF1:HA-FKF1 I160R/fkf1 #2–3 plants grown under LDs. The level of COP1 that co-precipitated with HA-FKF1 I160R notably increased compared to FKF1 in the late afternoon (Fig. 4E, F). These findings suggest that the FKF1 monomer preferentially interacts with COP1, leading to increased CO abundance.
The FKF1 I160R variant shows enhanced binding affinity to COP1. A, C, E Representative images of the results of co-IP assays (A, C) and protein expression analysis (E). B, D, F Relative amounts of complex formation (B, F) and protein abundance (D) were quantified. A, B CO-TAP proteins were coexpressed with either HA-FKF1 or HA-FKF1 I160R in leaf cells of 3-week-old tobacco plants. The data underwent statistical analysis using a two-tailed Student's t test. **p ≤ 0.01. C, D CO:HA-CO and CO:HA-CO FKF1:FKF1 I160R/fkf1 plants were grown under LDs and harvested at the indicated time points on day 10. Relative protein abundance was normalized to Histone H3 levels. Histone H3 protein was detected by immunoblotting with anti-Histone H3 antibody. E, F Seedlings were grown under LDs and sampled at ZT12 on day 10. E HA-tagged proteins were immunoprecipitated with anti-HA antibody. Endogenous COP1 proteins were detected by immunoblotting with anti-COP1 antibody. F Bar graphs representing the amounts of co-immunoprecipitated COP1 protein calculated as (COP1IP/HA-tagged proteinIP)/(COP1Input/HA-tagged proteinInput). The data underwent statistical analysis using a two-tailed Student's t test. ns not significant. The means ± SEM of normalized values calculated from three biological replicates are shown
Discussion
Many plants constantly monitor changes in daylength and thus coordinate the timing of the transition to the reproductive phase with favorable seasons, thereby maximizing seed production (Andrés and Coupland 2012; Song et al. 2015). In Arabidopsis, the blue-light photoreceptor FKF1 acts as the main photoperiod sensor involved in the photoperiod-dependent accumulation of FT transcript (Sawa et al. 2007; Song et al. 2012, 2015). In this study, using the FKF1 I160R variant, which shows attenuated homodimerization, we demonstrated that the regulation of FKF1 dimerization is important for optimizing the levels of FT induction in the afternoon under LDs (Fig. 5).
A hypothetical model describing the role of FKF1 dimerization in fine-tuning FT expression under long-day conditions. Under a long photoperiod, the abundance of CO protein exhibits dual peaks in the morning and late afternoon. In the morning, the COP1 dimer binds to CO and initiates its degradation via E3 ubiquitin ligase activity, resulting in reduced CO abundance. In the afternoon, the LOV-domain-containing blue-light photoreceptor FKF1 is expressed and naturally forms homodimers. The FKF1 homodimer interacts with and stabilizes CO, promoting its accumulation. Concurrently, the blue-light-activated FKF1 homodimer binds to GI through its LOV domain in the late afternoon. Upon GI binding, the FKF1 homodimer is converted to the monomeric form, causing a conformational change in its Kelch domain and adopting an open structure. Subsequently, the GI-bound FKF1 monomeric form interacts with COP1 through its Kelch domain, disrupting the COP1 homodimer. As a result, the COP1 monomer is unable to form a complex with SPA1 for CO degradation, leading to the significant accumulation of CO protein in the late afternoon. Consequently, the increased activity of CO induces peak levels of FT expression around dusk, thereby promoting flowering. During the night, due to impaired FKF1–GI complex formation, COP1 dimers accumulate and actively degrade CO
The interaction of FKF1 with GI is important for the ubiquitin ligase activity and stability of FKF1 (Sawa et al. 2007; Hwang et al. 2019). Under LDs, GI binding facilitates the function of FKF1 in the degradation of CDF1, allowing CO to be expressed in the afternoon (Sawa et al. 2007). However, the involvement of FKF1–GI complex formation in the induction of FT transcription remains poorly understood. In the current study, reducing the formation of FKF1 homodimers led to increased levels of FT mRNA but not CO mRNA (Figs. 2G, H and 3A, B), indicating that the conversion of FKF1 homodimers to monomers is crucial for the selective role of FKF1 in inducing FT expression. During the daytime under LDs, the FKF1 I160R variant demonstrated enhanced interactions with GI and COP1, resulting in elevated CO protein abundance (Figs. 3C, D and 4C–F). These findings may, at least in part, explain the heightened levels of FT mRNA observed in FKF1:HA-FKF1 I160R/fkf1 plants around dusk under LDs (Fig. 2E–H). Notably, FKF1 protein accumulation appears to rely on the function of GI under LDs (Fornara et al. 2009; Hwang et al. 2019). Further investigation is needed to identify the factors contributing to the increased susceptibility of the monomeric form of FKF1 to degradation, despite its enhanced binding to GI (Figs. 2C, D and 4C, D).
The perception of blue light through the LOV domain is essential for the function of FKF1 in accelerating flowering time through interactions with not only GI but also CO (Imaizumi et al. 2003; Sawa et al. 2007; Song et al. 2012). While FKF1 interacts with GI in a blue-light-dependent manner using its LOV domain, the presence of blue light may not affect the conversion between the FKF1 homodimer and monomer (Sawa et al. 2007; Kwon et al. 2022). Conversely, GI binding appears to disrupt LOV domain homodimerization after photoactivation (Kwon et al. 2022). Our co-IP analysis revealed that even with the I160R mutation, FKF1 maintained its ability to bind to GI in a blue-light-dependent manner (Supplementary Fig. S3), supporting the previous observation (Kwon et al. 2022). Under LDs, FKF1 also binds to CO through the LOV domain, and this binding is enhanced by blue light (Song et al. 2012). However, in contrast to GI, the binding of FKF1 to CO was significantly reduced by the I160R mutation (Fig. 4A, B), indicating that FKF1 may interact with CO as a homodimer or the I160 site may lie within the FKF1-CO interface. Considering the promotion of flowering via elevated FT induction in FKF1:HA-FKF1 I160R/fkf1 plants (Fig. 2G, H), the conversion of FKF1 to the monomeric form appears to offer several advantages in its function, which counteract the adverse effects of this conversion on protein stability and CO binding. The interaction between FKF1 I160R and ZTL significantly decreased compared to that between FKF1 and ZTL (Fig. 3E, F). Since ZTL inhibits FKF1 activity and competes with it for GI binding (Takase et al. 2011; Hwang et al. 2019), the reduced complex formation may contribute to the increased interaction of the FKF1 monomers with GI (Fig. 3C, D).
The interaction between ZTL and GI through the LOV domain results in diminished affinity towards its proteolytic targets, PRR5 and TOC1 (Kim et al. 2007). The formation of the ZTL–GI complex induces a conformational change in the Kelch domain, leading to a closed configuration (Kwon et al. 2022). This change might contribute to the increased abundance of PRR5 and TOC1 during the night (Kim et al. 2007). By contrast, when FKF1 interacts with GI in a blue-light-specific manner, the Kelch domain appears to adopt an open configuration, allowing it to recognize CDF1, a substrate for the E3 ubiquitin ligase activity of FKF1 (Imaizumi et al. 2005; Sawa et al. 2007; Fornara et al. 2009; Kwon et al. 2022). Under LDs, FKF1 binds to the COP1 monomer under blue light through the Kelch domain (Lee et al. 2017). However, unlike CDF1, COP1 protein abundance is unaffected by the mutation or overexpression of FKF1 (Lee et al. 2017). Instead, FKF1 binding inhibits COP1 homodimerization, which mediates the degradation of CO by forming a tetramer complex with SPA1 (Laubinger et al. 2006; Jang et al. 2008; Zhu et al. 2008; Lee et al. 2017). Consistent with this finding, our data show that the enhanced interaction of FKF1 with COP1 due to the I160R mutation led to increased CO protein abundance (Fig. 4C–F). Considering the enhanced binding affinity of FKF1 to GI due to the I160R mutation (Fig. 3C, D), these findings support the notion that GI binding may facilitate the dissociation of FKF1 homodimers and the complex formation of FKF1 monomers with COP1, allowing CO to accumulate in the afternoon under LDs by decreasing COP1–SPA1 complex formation (Fig. 5). Furthermore, given that COP1 destabilizes GI in conjunction with ELF3 (Yu et al. 2008), the heightened formation of the FKF1 and COP1 complex due to the I160R mutation could potentially enhance GI accumulation. This, in turn, might enhance the abundance of FT mRNA via the GI-mediated microRNA172 pathway, independently of CO transcription (Jung et al. 2007).
In addition to regulating CO stability, FKF1 may directly induce FT expression through GI binding. Both FKF1 and GI associate with the FT promoter, and FKF1 relieves FT repression by CDF1 (Sawa et al. 2007; Song et al. 2012), indicating that FKF1 likely forms a complex with GI at the FT locus. Moreover, GI interacts with FT repressors, such as SHORT VEGETATIVE PHASE (SVP) and TEMPRANILLOs (TEMs) (Sawa et al. 2007). As FKF1 and GI bind to the FT promoter regions near its SVP-binding sites (Lee et al. 2007; Sawa et al. 2007; Song et al. 2012), perhaps FKF1 is involved in reducing FT repression by SVP and TEMs by forming a complex with GI. Together, our findings suggest that the regulation of FKF1 dimerization provides a new layer that determines the optimal levels of FT expression during the afternoon. This regulatory mechanism might enable plants to finely adjust their flowering time in response to dynamic environmental fluctuations.
Data availability
The data supporting the findings of this study are provided in the main text and supplementary materials.
References
Andrés F, Coupland G (2012) The genetic basis of flowering responses to seasonal cues. Nat Rev Genet 13:627–639
Corbesier L, Vincent C, Jang S, Fornara F, Fan Q, Searle I, Giakountis A, Farrona S, Gissot L, Turnbull C, Coupland G (2007) FT protein movement contributes to long-distance signaling in floral induction of Arabidopsis. Science 316:1030–1033
David KM, Armbruster U, Tama N, Putterill J (2006) Arabidopsis GIGANTEA protein is post-transcriptionally regulated by light and dark. FEBS Lett 580:1193–1197
Emsley P, Cowtan K (2004) Coot: model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr 60:2126–2132
Exposito-Alonso M (2020) Seasonal timing adaptation across the geographic range of Arabidopsis thaliana. Proc Natl Acad Sci USA 117:9665–9667
Fornara F, Panigrahi KCS, Gissot L, Sauerbrunn N, Rühl M, Jarillo JA, Coupland G (2009) Arabidopsis DOF transcription factors act redundantly to reduce CONSTANS expression and are essential for a photoperiodic flowering response. Dev Cell 17:75–86
Hwang DY, Park S, Lee S, Lee SS, Imaizumi T, Song YH (2019) GIGANTEA regulates the timing stabilization of CONSTANS by altering the interaction between FKF1 and ZEITLUPE. Mol Cells 42:693–701
Imaizumi T, Tran HG, Swartz TE, Briggs WR, Kay SA (2003) FKF1 is essential for photoperiodic-specific light signalling in Arabidopsis. Nature 426:302–306
Imaizumi T, Schultz TF, Harmon FG, Ho LA, Kay SA (2005) FKF1 F-box protein mediates cyclic degradation of a repressor of CONSTANS in Arabidopsis. Science 309:293–297
Ito S, Song YH, Imaizumi T (2012) LOV domain-containing F-box proteins: light-dependent protein degradation modules in Arabidopsis. Mol Plant 5:573–582
Jaeger KE, Wigge PA (2007) FT protein acts as a long-range signal in Arabidopsis. Curr Biol 17:1050–1054
Jang S, Marchal V, Panigrahi KCS, Wenkel S, Soppe W, Deng XW, Velverde F, Coupland G (2008) Arabidopsis COP1 shapes the temporal pattern of CO accumulation conferring a photoperiodic flowering response. Embo J 27:1277–1288
Jung JH, Seo YH, Seo PJ, Reyes JL, Yun J, Chua NH, Park CM (2007) The GIGANTEA-regulated microRNA172 mediates photoperiodic flowering independent of CONSTANS in Arabidopsis. Plant Cell 19:2736–2748
Kim WY, Fujiwara S, Suh SS, Kim J, Kim Y, Han L, David K, Putterill J, Nam HG, Somers DE (2007) ZEITLUPE is a circadian photoreceptor stabilized by GIGANTEA in blue light. Nature 449:356–360
Kim H, Kang HW, Hwang DY, Lee N, Kubota A, Imaizumi T, Song YH (2024) Low temperature-mediated repression and far-red light-mediated induction determine morning FLOWERING LOCUS T expression levels. J Integr Plant Biol 66:103–120
Krissinel E, Henrick K (2007) Inference of macromolecular assemblies from crystalline state. J Mol Biol 372:774–797
Kwon E, Patha D, Dahal P, Tandukar S, Jung HS, Kim WY, Kim DY (2022) Structural analysis of the regulation of blue-light receptors by GIGANTEA. Cell Rep 39:110700
Laubinger S, Marchal V, Gourrierec JL, Wenkel S, Adrian J, Jang S, Kulajta C, Barun H, Coupland G, Hoecker U (2006) Arabidopsis SPA proteins regulate photoperiodic flowering and interact with the floral inducer CONSTANS to regulate its stability. Development 133:3213–3222
Lazaro A, Mouriz A, Piñeiro M, Jarillo JA (2015) Red light-mediated degradation of CONSTANS by the E3 ubiquitin ligase HOS1 regulates photoperiodic flowering in Arabidopsis. Plant Cell 27:2437–2454
Lee JH, Yoo SJ, Park SH, Hwang I, Lee JS, Ahn JH (2007) Role of SVP in the control of flowering time by ambient temperature in Arabidopsis. Genes Dev 21:397–402
Lee BD, Kim MR, Kang MY, Cha JY, Han SH, Nawkar GM, Sakuraba Y, Lee SY, Imaizumi T, McClung CR, Kim WY, Paek NC (2017) The F-box protein FKF1 inhibits dimerization of COP1 in the control of photoperiodic flowering. Nat Commun 8:2259
Lee N, Ozaki Y, Hempton AK, Takagi H, Purusuwashi S, Song YH, Endo M, Kubota A, Imaizumi T (2023) The FLOWERING LOCUS T gene expression is controlled by high-irradiance response and external coincidence mechanism in long days in Arabidopsis. New Phytol 239:208–221
Lian HL, He SB, Zhang YC, Zhu DM, Zhang JY, Jia KP, Sun SX, Li L, Yang HQ (2011) Blue-light-dependent interaction of cryptochrome 1 with SPA1 defines a dynamics singaling mechanism. Genes Dev 25:1023–1028
Liu LJ, Zhang YC, Li QL, Sang Y, Mao J, Lian HL, Wang L, Yang HQ (2008) COP1-mediated ubiquitination of CONSTANS is implicated in cryptochrome regulation of flowering in Arabidopsis. Plant Cell 20:292–306
Mathieu J, Warthmann N, Küttner F, Schmid M (2007) Export of FT protein from phloem companion cells is sufficient for floral induction in Arabidopsis. Curr Biol 17:1055–1060
Nakasako M, Natsuoka D, Zikihara K, Tokutomi S (2005) Quaternary structure of LOV-domain containing polypeptide of Arabidopsis FKF1 protein. FEBS Lett 579:1067–1071
Nelson DC, Lasswell J, Rogg LE, Cohen MA, Bartel B (2000) FKF1, a clock-controlled gene that regulates the transition to flowering in Arabidopsis. Cell 101:331–340
Pudasaini A, Zoltowski BD (2013) Zeitlupe senses blue-light fluence to mediate circadian timing in Arabidopsis thaliana. Biochemistry 52:7150–7158
Pudasaini A, Shim JS, Song YH, Shi H, Kiba T, Somers DE, Imaizumi T, Zoltowski BD (2017) Kinetics of the LOV domain of ZEITLUPE determine its circadian function in Arabidopsis. Elife 6:e21646
Samach A, Onouchi H, Gold SE, Ditta GS, Schwarz-Sommer Z, Yanofsky MF, Coupland G (2000) Distinct roles of CONSTANS target genes in reproductive development of Arabidopsis. Science 288:1613–1616
Sarid-Krebs L, Panigrahi KCS, Fornara F, Takahashi Y, Hayama R, Jang S, Tilmes V, Valverde F, Coupland G (2015) Phosphorylation of CONSTANS and its COP1-dependent degradation during photoperiodic flowering of Arabidopsis. Plant J 84:451–463
Sawa M, Nusinow DA, Kay SA, Imaizumi T (2007) FKF1 and GIGANTEA complex formation is required for day-length measurement in Arabidopsis. Science 318:261–265
Song YH, Smith RW, To BJ, Millar AJ, Imaizumi T (2012) FKF1 conveys timing information for CONSTANS stabilization in photoperiodic flowering. Science 336:1045–1049
Song YH, Estrada DA, Johnson RS, Kim SK, Lee SY, MacCoss MJ, Imaizumi T (2014) Distinct roles of FKF1, GIGANTEA, and ZEITLUPE proteins in the regulation of CONSTANS stability in Arabidopsis photoperiodic flowering. Proc Natl Acad Sci USA 111:17672–17677
Song YH, Shim JS, Kinmonth-Schultz HA, Imaizumi T (2015) Photoperiodic flowering: time measurement mechanisms in leaves. Annu Rev Plant Biol 66:441–464
Song YH, Kubota A, Kwon MS, Covington MF, Lee N, Taagen ER, LaboyCintrón D, Hwang DY, Akiyama R, Hodge SK, Huang H, Nguyen NH, Nusinow DA, Millar AJ, Shimizu KK, Imaizumi T (2018) Molecular basis of flowering under natural long-day conditions in Arabidopsis. Nat Plants 4:824–835
Suárez-López P, Wheatley K, Robson F, Onouchi H, Valverde F, Coupland G (2001) CONSTANS mediates between the circadian clock and the control of flowering in Arabidopsis. Nat 410:1116–1120. https://doi.org/10.1038/35074138
Takagi H, Hempton AK, Imaizumi T (2023) Photoperiodic flowering in Arabidopsis: Multilayered regulatory mechanisms of CONSTANS and the florigen FLOWERING LOCUS T. Plant Commun 4:100552
Takase T, Nishiyama Y, Tanihigashi H, Ogura Y, Miyazaki Y, Yamada Y, Kiyosue T (2011) LOV KELCH PROTEIN2 and ZEITLUPE repress Arabidopsis photoperiodic flowering under non-inductive conditions, dependent on FLAVIN-BINDING KELCH REPEAT F-BOX1. Plant J 67:608–621
Valverde F, Mouradov A, Soppe W, Ravenscroft D, Samach A, Coupland G (2004) Photoreceptor regulation of CONSTANS protein in photoperiodic flowering. Science 303:1003–1006
Yu JW, Rubio V, Lee NY, Bai S, Lee SY, Kim SS, Liu L, Zhang Y, Irigoyen ML, Sullivan JA, Zhang Y, Lee I, Xie Q, Paek NC, Deng XW (2008) COP1 and ELF3 control circadian function and photoperiodic flowering by regulating GI stability. Mol Cell 32:617–630
Zhang B, Wang L, Zeng L, Zhang C, Ma H (2015) Arabidopsis TOE proteins convey a photoperiodic signal to antagonize CONSTANS and regulate flowering time. Genes Dev 29:975–987
Zhu D, Maier A, Lee JH, Laubinger S, Saijo Y, Wang H, Qu LJ, Hoecker U, Deng XW (2008) Biochemical characterization of Arabidopsis complexes containing CONSTITUTIVELY PHOTOMORPHOGENIC1 and SUPPRESSOR OF PHYA proteins in light control of plant development. Plant Cell 20:2307–2323
Zuo Z, Liu H, Liu B, Liu X, Lin C (2011) Blue light-dependent interaction of CRY2 with SPA1 regulates COP1 activity and floral initiation in Arabidopsis. Curr Biol 21:841–847
Acknowledgements
This work was supported by National Institutes of Health (NIH) (No. R01GM079712 to T.I. and No. 2R15GM109282-03 to B.D.Z) and the National Research Foundation (NRF) of Korea grant funded by the Korean government (MSIT) (No. NRF-2020R1A2C1014655 and No. NRF-2021R1A4A1032888 to Y.H.S.).
Funding
Open Access funding enabled and organized by Seoul National University. Foundation for the National Institutes of Health, R01GM079712, Takato Imaizumi, 2R15GM109282-03, Brian D Zoltowski, the National Research Foundation (NRF) of Korea, NRF-2020R1A2C1014655, Young Hun Song, National Research Foundation (NRF) of Korea, NRF-2021R1A4A1032888, Young Hun Song.
Author information
Authors and Affiliations
Contributions
S.W.C performed most of the research. J.L. carried out the photochemistry experiments. J.S.P. conducted the analysis of CO protein abundance. H.K. analyzed and interpreted the data. M.C. performed a flowering time experiment. H.Q.Y. and T.I. reviewed the manuscript. B.Z. contributed to the experimental design and wrote the manuscript. Y.H.S. conceptualized the experiments, supervised the study, and wrote the manuscript. All the authors read and approved of its content.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Communicated by Woe-Yeon Kim.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cho, S.W., Lokhandwala, J., Park, J.S. et al. Disrupting FKF1 homodimerization increases FT transcript levels in the evening by enhancing CO stabilization. Plant Cell Rep 43, 121 (2024). https://doi.org/10.1007/s00299-024-03207-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00299-024-03207-w